Structure Database (LMSD)
Common Name
PIP3[3',4',5'] 17:0/20:4(5Z,8Z,11Z,14Z)
Systematic Name
1-heptadecanoyl-2-(5Z,8Z,11Z,14Z-eicosatetraenoyl)-sn-glycero-3-phospho-(1'-myo-inositol-3',4',5'-trisphosphate)
Synonyms
- PIP3[3',4',5'](17:0/20:4)
- PIP3(17:0/20:4)
LM ID
LMGP09010001
Formula
Exact Mass
Calculate m/z
1112.440474
Sum Composition
Abbrev Chains
PIP3 17:0_20:4
Status
Curated
3D model of PIP3[3',4',5'] 17:0/20:4(5Z,8Z,11Z,14Z)
Please note: Where there are chiral atoms but the stereochemistry is undefined, the 3D model takes an arbitrary conformation
Classification
Category
Main Class
Sub Class
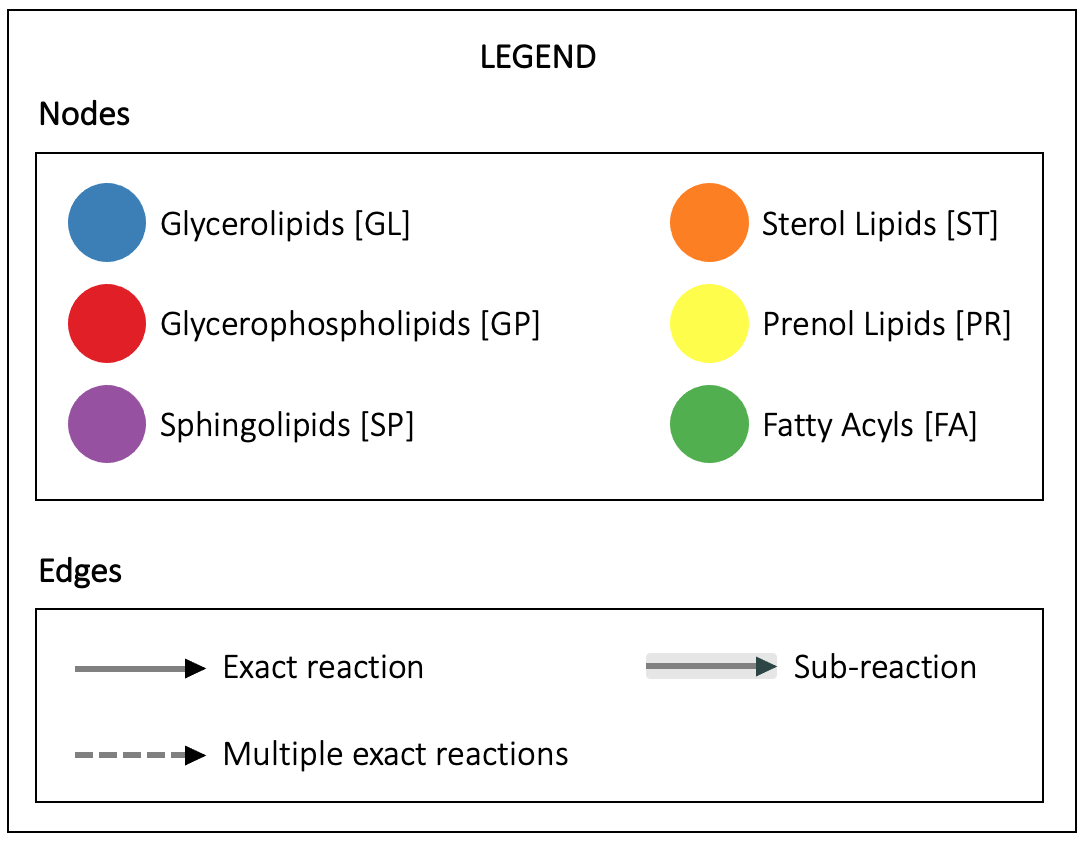
Reactions
Filter by species:
ⓘ
Reactions are shown if the E.C. number of the enzyme catalysing it is annotated in the UniProt database for a species belonging to the selected taxonomic class.
Click on an edge to display the reaction(s).
String Representations
InChiKey (Click to copy)
BKVMCHXQLWSEHU-TXXVHSBZSA-N
InChi (Click to copy)
InChI=1S/C46H84O22P4/c1-3-5-7-9-11-13-15-17-19-20-21-23-25-27-29-31-33-35-40(48)64-38(36-62-39(47)34-32-30-28-26-24-22-18-16-14-12-10-8-6-4-2)37-63-72(60,61)68-43-41(49)44(65-69(51,52)53)46(67-71(57,58)59)45(42(43)50)66-70(54,55)56/h11,13,17,19,21,23,27,29,38,41-46,49-50H,3-10,12,14-16,18,20,22,24-26,28,30-37H2,1-2H3,(H,60,61)(H2,51,52,53)(H2,54,55,56)(H2,57,58,59)/b13-11-,19-17-,23-21-,29-27-/t38-,41+,42+,43-,44+,45-,46-/m1/s1
SMILES (Click to copy)
[C@]([H])(OC(CCC/C=C\C/C=C\C/C=C\C/C=C\CCCCC)=O)(COP(=O)(O)O[C@H]1[C@H](O)[C@@H](OP(=O)(O)O)[C@H](OP(=O)(O)O)[C@@H](OP(O)(=O)O)[C@H]1O)COC(CCCCCCCCCCCCCCCC)=O
Other Databases
Calculated Physicochemical Properties
Heavy Atoms
72
Rings
1
Aromatic Rings
0
Rotatable Bonds
45
Van der Waals Molecular Volume
1048.86
Topological Polar Surface Area
349.10
Hydrogen Bond Donors
9
Hydrogen Bond Acceptors
22
logP
12.90
Molar Refractivity
271.19
Admin
Created at
-
Updated at
11th Nov 2025
LIPID MAPS® abbreviations for glycerophospholipids (GP)
The LIPID MAPS® glycerophospholipid abbreviations (PC, PE, etc.) are used here to refer to species with one or two radyl side-chains where the structures of the side chains are indicated within parentheses in the 'Headgroup(sn1/sn2)' format (e.g. PC(16:0/18:1(9Z)). By default, R stereochemistry at the C-2 carbon of glycerol and attachment of the headgroup at the sn3 position is assumed. Also, acyl chains are assumed by default. The 'O-' prefix is used to indicate the presence of an alkyl ether substituent e.g. PC(O-16:0/18:1(9Z)), whereas the 'P-' prefix is used for the 1Z-alkenyl ether (Plasmalogen) substituent e.g. PC(P-16:0/18:1(9Z)).
For molecules with opposite (S) stereochemistry at C2 of the glycerol group and attachment of the headgroup at the sn1 position, the stereochemistry specification of [S] is appended to the abbreviation. The 'Headgroup(sn3/sn2)' abbreviation format is used.
For molecules with unknown stereochemistry at the C-2 carbon of the glycerol group, the stereochemistry specification of [U] is appended to the abbreviation and the structure is drawn with C-2 stereochemistry unspecified.
The LIPID MAPS® glycerophospholipid abbreviations (PC, PE, etc.) are used here to refer to species with one or two radyl side-chains where the structures of the side chains are indicated within parentheses in the 'Headgroup(sn1/sn2)' format (e.g. PC(16:0/18:1(9Z)). By default, R stereochemistry at the C-2 carbon of glycerol and attachment of the headgroup at the sn3 position is assumed. Also, acyl chains are assumed by default. The 'O-' prefix is used to indicate the presence of an alkyl ether substituent e.g. PC(O-16:0/18:1(9Z)), whereas the 'P-' prefix is used for the 1Z-alkenyl ether (Plasmalogen) substituent e.g. PC(P-16:0/18:1(9Z)).
For molecules with opposite (S) stereochemistry at C2 of the glycerol group and attachment of the headgroup at the sn1 position, the stereochemistry specification of [S] is appended to the abbreviation. The 'Headgroup(sn3/sn2)' abbreviation format is used.
For molecules with unknown stereochemistry at the C-2 carbon of the glycerol group, the stereochemistry specification of [U] is appended to the abbreviation and the structure is drawn with C-2 stereochemistry unspecified.