| Gene symbol |
Time course plot |
Accession |
Description |
Treatment |
15min |
30min |
1hr |
2hr |
4hr |
8hr |
20hr |
| Aldh1b1 | 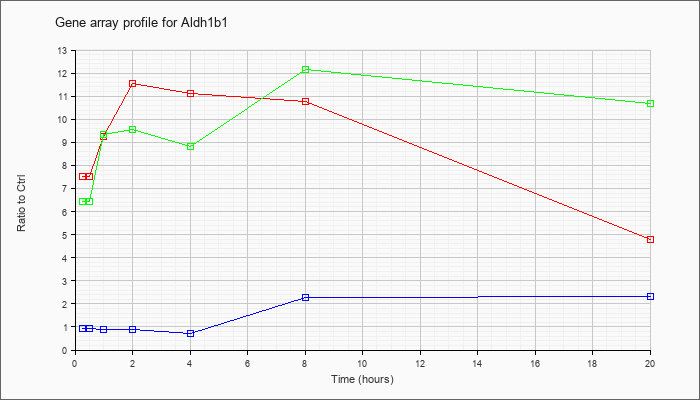 |
NM_028270 |
aldehyde dehydrogenase 1 family, member B1 (Aldh1b1), nuclear gene encoding mitochondrial protein, mRNA [NM_028270] |
KLA | 7.51 |
7.83 |
9.27 |
11.57 |
11.10 |
10.78 |
4.79 |
| ATP | .93 |
.91 |
.90 |
.89 |
.73 |
2.27 |
2.30 |
| KLA/ATP | 6.45 |
7.40 |
9.33 |
9.54 |
8.82 |
12.15 |
10.67 |
|
| Aldh2 | 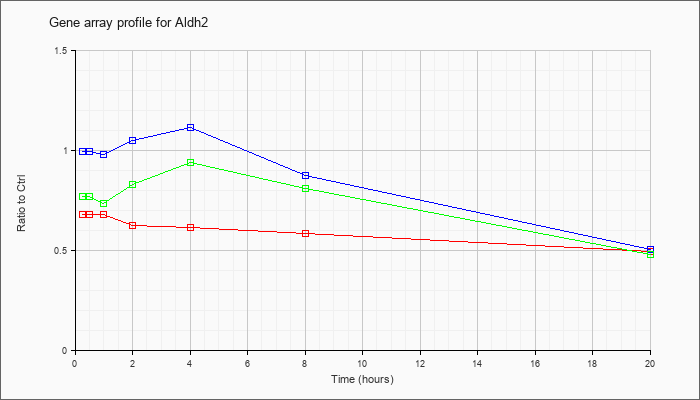 |
AK163452 |
adult male corpora quadrigemina cDNA, RIKEN full-length enriched library, clone:B230105J09 product:aldehyde dehydrogenase 2, mitochondrial, full insert sequence [AK163452] |
KLA | .62 |
.65 |
.59 |
.60 |
.63 |
.64 |
.49 |
| ATP | 1.01 |
.91 |
1.06 |
.93 |
1.18 |
.93 |
.41 |
| KLA/ATP | .70 |
.68 |
.70 |
.84 |
1.16 |
.96 |
.45 |
|
| Aldh2 |  |
NM_009656 |
aldehyde dehydrogenase 2, mitochondrial (Aldh2), nuclear gene encoding mitochondrial protein, mRNA [NM_009656] |
KLA | .74 |
.81 |
.77 |
.65 |
.60 |
.53 |
.50 |
| ATP | .98 |
1.10 |
.90 |
1.17 |
1.04 |
.82 |
.60 |
| KLA/ATP | .84 |
.86 |
.77 |
.82 |
.72 |
.66 |
.51 |
|
| Aldh3a2 | 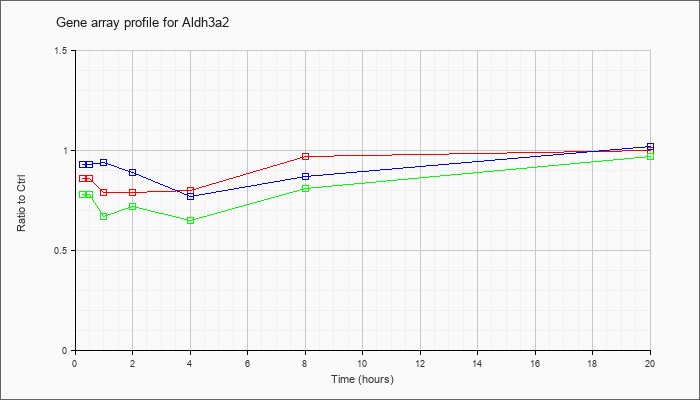 |
NM_007437 |
aldehyde dehydrogenase family 3, subfamily A2 (Aldh3a2), mRNA [NM_007437] |
KLA | .86 |
.83 |
.79 |
.79 |
.80 |
.97 |
1.00 |
| ATP | .93 |
.98 |
.94 |
.89 |
.77 |
.87 |
1.02 |
| KLA/ATP | .78 |
.81 |
.67 |
.72 |
.65 |
.81 |
.97 |
|
| Aldh7a1 | 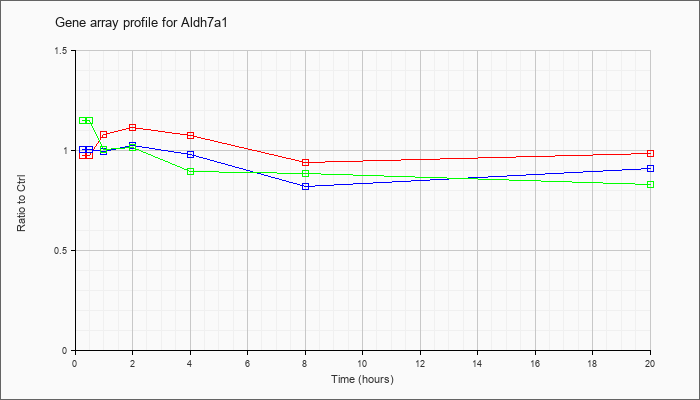 |
NM_138600 |
aldehyde dehydrogenase family 7, member A1 (Aldh7a1), mRNA [NM_138600] |
KLA | .98 |
1.03 |
1.08 |
1.12 |
1.08 |
.94 |
.99 |
| ATP | 1.01 |
1.03 |
1.00 |
1.03 |
.98 |
.82 |
.91 |
| KLA/ATP | 1.15 |
1.12 |
1.01 |
1.02 |
.90 |
.89 |
.83 |
|
| Aldh9a1 | 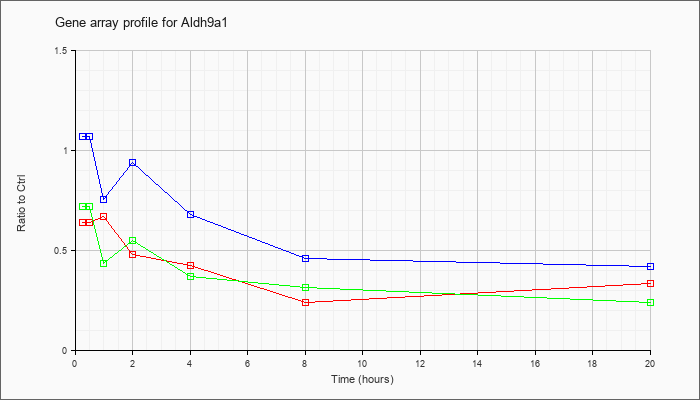 |
NM_019993 |
aldehyde dehydrogenase 9, subfamily A1 (Aldh9a1), mRNA [NM_019993] |
KLA | .64 |
.64 |
.67 |
.48 |
.42 |
.24 |
.33 |
| ATP | 1.07 |
1.02 |
.75 |
.94 |
.68 |
.46 |
.42 |
| KLA/ATP | .72 |
.65 |
.43 |
.55 |
.37 |
.31 |
.24 |
|
| Gulo | 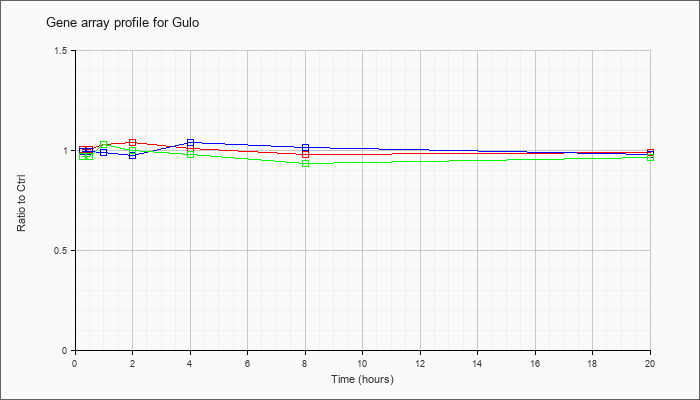 |
NM_178747 |
gulonolactone (L-) oxidase (Gulo), mRNA [NM_178747] |
KLA | 1.01 |
.97 |
1.03 |
1.04 |
1.01 |
.98 |
.99 |
| ATP | 1.00 |
.99 |
.99 |
.98 |
1.04 |
1.02 |
.98 |
| KLA/ATP | .97 |
.98 |
1.03 |
1.00 |
.98 |
.94 |
.97 |
|
| Miox | 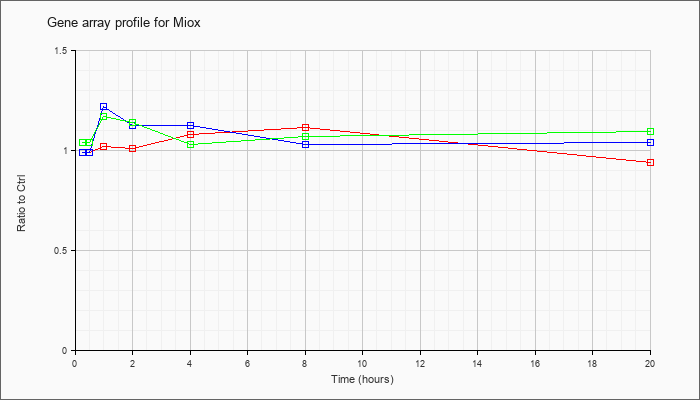 |
AF197127 |
unknown mRNA. [AF197127] |
KLA | .99 |
1.00 |
1.02 |
1.01 |
1.08 |
1.11 |
.94 |
| ATP | .99 |
1.04 |
1.22 |
1.12 |
1.12 |
1.03 |
1.04 |
| KLA/ATP | 1.04 |
1.01 |
1.17 |
1.14 |
1.03 |
1.07 |
1.09 |
|
Mm.30009
5 | 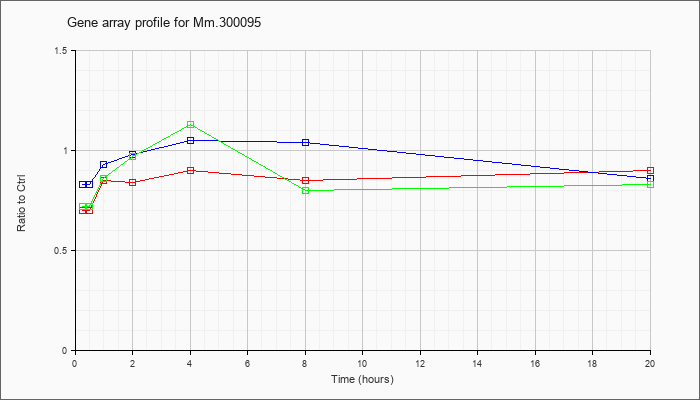 |
145699117 |
Unknown |
KLA | .70 |
.74 |
.85 |
.84 |
.90 |
.85 |
.90 |
| ATP | .83 |
.77 |
.93 |
.98 |
1.05 |
1.04 |
.86 |
| KLA/ATP | .72 |
.70 |
.86 |
.97 |
1.13 |
.80 |
.83 |
|
Mm.46685
9 | 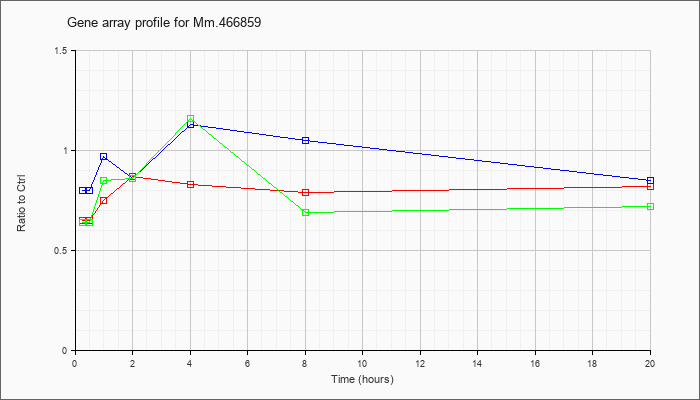 |
145699102 |
Unknown |
KLA | .65 |
.64 |
.75 |
.87 |
.83 |
.79 |
.82 |
| ATP | .80 |
.77 |
.97 |
.86 |
1.13 |
1.05 |
.85 |
| KLA/ATP | .64 |
.65 |
.85 |
.86 |
1.16 |
.69 |
.72 |
|
| Rgn | 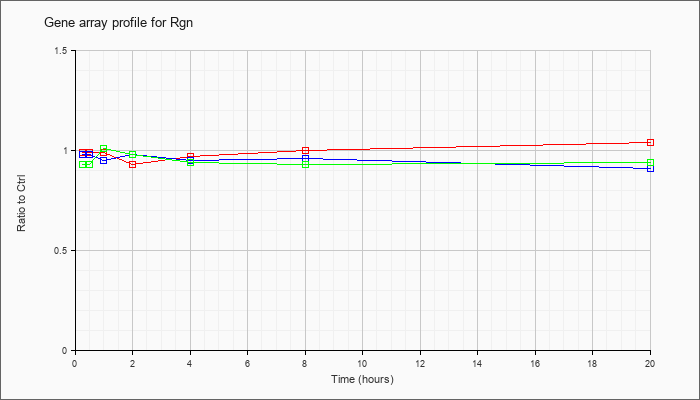 |
NM_009060 |
regucalcin (Rgn), mRNA [NM_009060] |
KLA | .99 |
.94 |
.99 |
.93 |
.97 |
1.00 |
1.04 |
| ATP | .98 |
1.03 |
.95 |
.98 |
.95 |
.96 |
.91 |
| KLA/ATP | .93 |
1.03 |
1.01 |
.98 |
.94 |
.93 |
.94 |
|
| Ugdh | 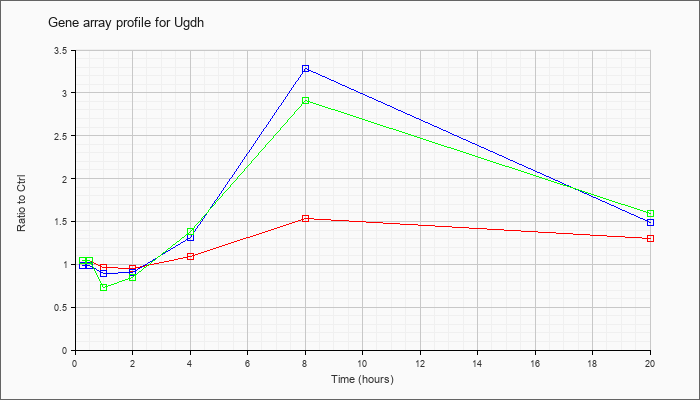 |
NM_009466 |
UDP-glucose dehydrogenase (Ugdh), mRNA [NM_009466] |
KLA | 1.04 |
1.06 |
.96 |
.95 |
1.09 |
1.54 |
1.30 |
| ATP | .99 |
1.06 |
.89 |
.91 |
1.31 |
3.29 |
1.49 |
| KLA/ATP | 1.04 |
.98 |
.73 |
.85 |
1.38 |
2.91 |
1.59 |
|
| Ugt1a1 | 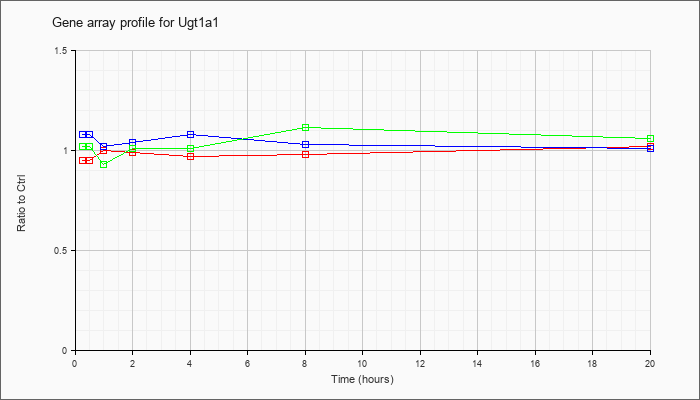 |
NM_201645 |
UDP glucuronosyltransferase 1 family, polypeptide A1 (Ugt1a1), mRNA [NM_201645] |
KLA | .95 |
1.07 |
1.00 |
.99 |
.97 |
.98 |
1.02 |
| ATP | 1.08 |
.95 |
1.02 |
1.04 |
1.08 |
1.03 |
1.01 |
| KLA/ATP | 1.02 |
.95 |
.93 |
1.01 |
1.01 |
1.11 |
1.06 |
|
| Ugt1a6b | 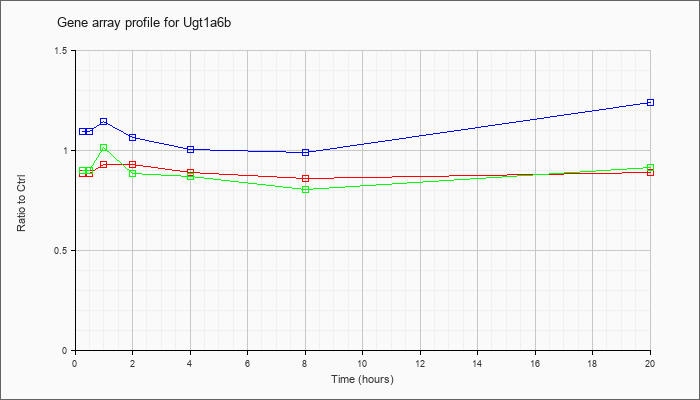 |
NM_201410 |
UDP glucuronosyltransferase 1 family, polypeptide A6B (Ugt1a6b), mRNA [NM_201410] |
KLA | .89 |
.86 |
.93 |
.93 |
.89 |
.86 |
.89 |
| ATP | 1.10 |
1.05 |
1.15 |
1.07 |
1.01 |
.99 |
1.24 |
| KLA/ATP | .90 |
.94 |
1.02 |
.89 |
.87 |
.81 |
.92 |
|
| Ugt1a9 | 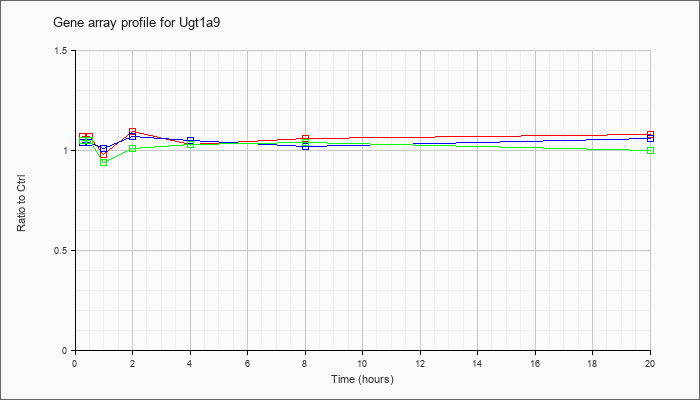 |
NM_201644 |
UDP glucuronosyltransferase 1 family, polypeptide A9 (Ugt1a9), mRNA [NM_201644] |
KLA | 1.07 |
.96 |
.98 |
1.09 |
1.03 |
1.06 |
1.08 |
| ATP | 1.04 |
.94 |
1.01 |
1.07 |
1.05 |
1.02 |
1.06 |
| KLA/ATP | 1.05 |
1.00 |
.94 |
1.01 |
1.03 |
1.04 |
1.00 |
|
| Ugt2a1 | 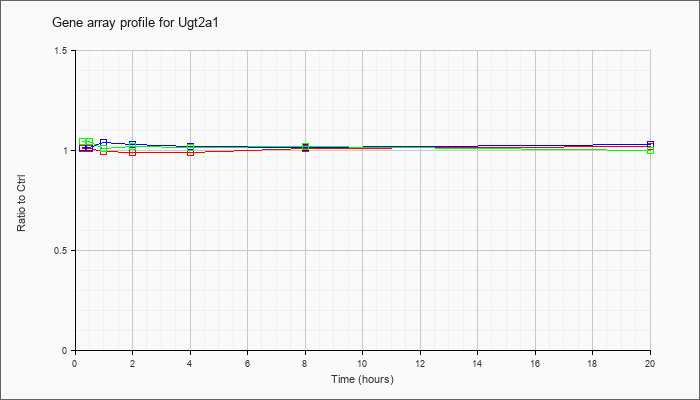 |
NM_053184 |
UDP glucuronosyltransferase 2 family, polypeptide A1 (Ugt2a1), mRNA [NM_053184] |
KLA | 1.01 |
1.03 |
.99 |
.99 |
.99 |
1.01 |
1.02 |
| ATP | 1.01 |
.93 |
1.04 |
1.03 |
1.02 |
1.01 |
1.03 |
| KLA/ATP | 1.04 |
1.01 |
1.01 |
1.02 |
1.01 |
1.02 |
1.00 |
|
| Ugt2a3 | 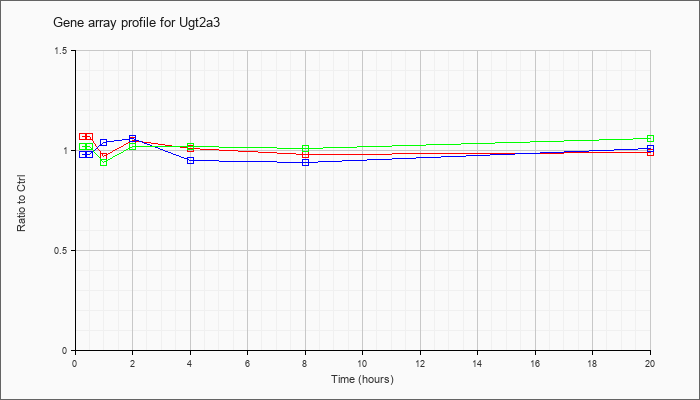 |
NM_028094 |
UDP glucuronosyltransferase 2 family, polypeptide A3 (Ugt2a3), mRNA [NM_028094] |
KLA | 1.07 |
1.01 |
.97 |
1.05 |
1.01 |
.98 |
.99 |
| ATP | .98 |
.96 |
1.04 |
1.06 |
.95 |
.94 |
1.01 |
| KLA/ATP | 1.02 |
1.00 |
.94 |
1.02 |
1.02 |
1.01 |
1.06 |
|
| Ugt2b1 | 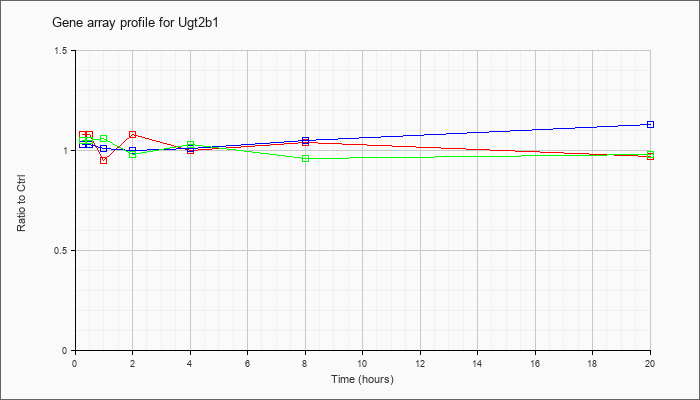 |
NM_152811 |
UDP glucuronosyltransferase 2 family, polypeptide B1 (Ugt2b1), mRNA [NM_152811] |
KLA | 1.08 |
1.01 |
.95 |
1.08 |
1.00 |
1.04 |
.97 |
| ATP | 1.03 |
1.14 |
1.01 |
1.00 |
1.01 |
1.05 |
1.13 |
| KLA/ATP | 1.05 |
.99 |
1.06 |
.98 |
1.03 |
.96 |
.98 |
|
| Ugt2b34 | 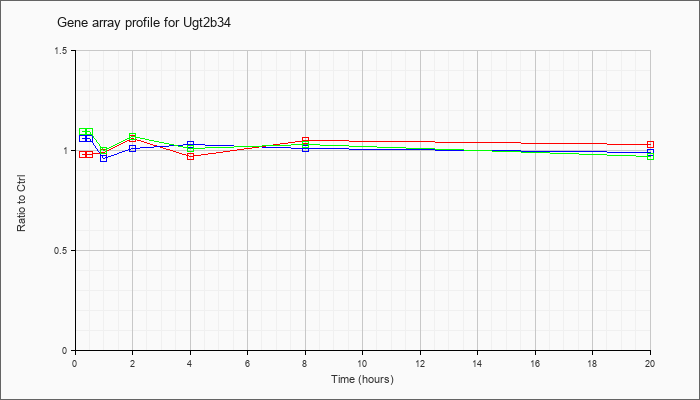 |
NM_153598 |
UDP glucuronosyltransferase 2 family, polypeptide B34 (Ugt2b34), mRNA [NM_153598] |
KLA | .98 |
1.08 |
.99 |
1.06 |
.97 |
1.05 |
1.03 |
| ATP | 1.06 |
1.04 |
.96 |
1.01 |
1.03 |
1.01 |
.99 |
| KLA/ATP | 1.09 |
.94 |
1.00 |
1.07 |
1.01 |
1.03 |
.97 |
|
| Ugt2b35 | 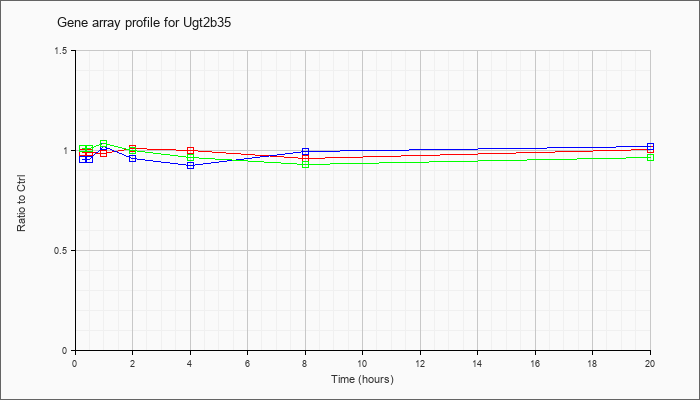 |
NM_172881 |
UDP glucuronosyltransferase 2 family, polypeptide B35 (Ugt2b35), mRNA [NM_172881] |
KLA | .99 |
1.02 |
.99 |
1.01 |
1.00 |
.96 |
1.01 |
| ATP | .96 |
1.02 |
1.02 |
.96 |
.93 |
1.00 |
1.02 |
| KLA/ATP | 1.01 |
.99 |
1.04 |
1.00 |
.97 |
.93 |
.97 |
|
| Ugt2b36 | 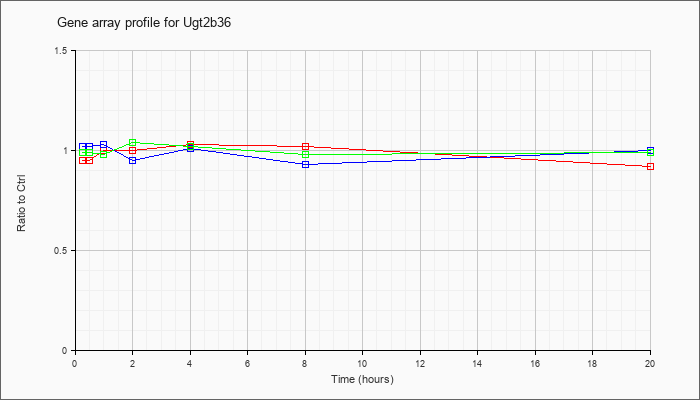 |
NM_001029867 |
UDP glucuronosyltransferase 2 family, polypeptide B36 (Ugt2b36), mRNA [NM_001029867] |
KLA | .95 |
.95 |
1.00 |
1.00 |
1.03 |
1.02 |
.92 |
| ATP | 1.02 |
1.07 |
1.03 |
.95 |
1.01 |
.93 |
1.00 |
| KLA/ATP | .99 |
1.05 |
.98 |
1.04 |
1.02 |
.98 |
.99 |
|
| Ugt2b37 | 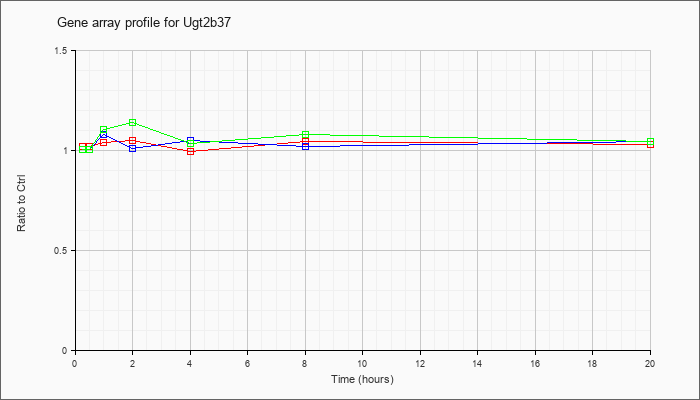 |
NM_053215 |
UDP glucuronosyltransferase 2 family, polypeptide B37 (Ugt2b37), mRNA [NM_053215] |
KLA | 1.02 |
1.06 |
1.04 |
1.05 |
1.00 |
1.05 |
1.03 |
| ATP | 1.01 |
1.03 |
1.08 |
1.01 |
1.05 |
1.02 |
1.05 |
| KLA/ATP | 1.01 |
.99 |
1.11 |
1.14 |
1.04 |
1.08 |
1.05 |
|
| Ugt2b38 | 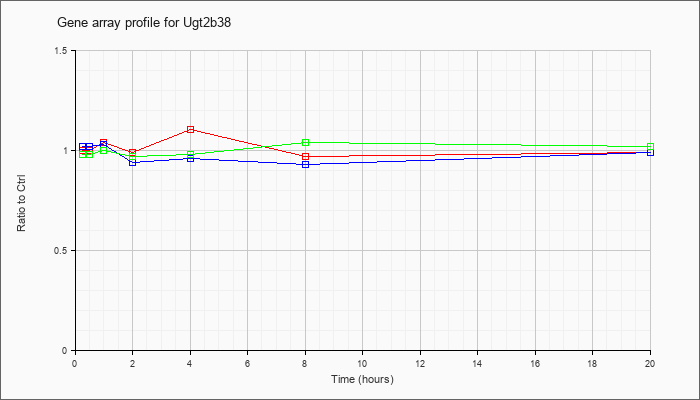 |
NM_133894 |
UDP glucuronosyltransferase 2 family, polypeptide B38 (Ugt2b38), mRNA [NM_133894] |
KLA | 1.00 |
.96 |
1.04 |
.99 |
1.10 |
.97 |
.99 |
| ATP | 1.02 |
1.04 |
1.03 |
.94 |
.96 |
.93 |
.99 |
| KLA/ATP | .98 |
.97 |
1.00 |
.97 |
.98 |
1.04 |
1.02 |
|
| Ugt2b5 | 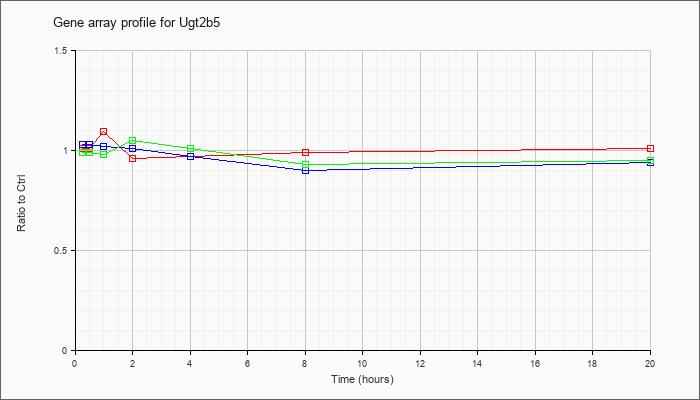 |
NM_009467 |
UDP glucuronosyltransferase 2 family, polypeptide B5 (Ugt2b5), mRNA [NM_009467] |
KLA | 1.01 |
.99 |
1.09 |
.96 |
.97 |
.99 |
1.01 |
| ATP | 1.03 |
1.02 |
1.02 |
1.01 |
.97 |
.90 |
.94 |
| KLA/ATP | .99 |
.99 |
.98 |
1.05 |
1.01 |
.93 |
.95 |
|