| Gene symbol |
Time course plot |
Accession |
Description |
Treatment |
15min |
30min |
1hr |
2hr |
4hr |
8hr |
20hr |
1700055N
04Rik | 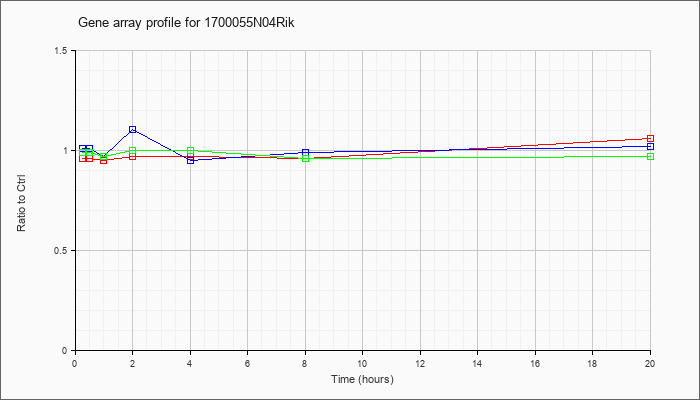 |
AK081788 |
16 days embryo head cDNA, RIKEN full-length enriched library, clone:C130077J09 product:weakly similar to ALDEHYDE DEHYDROGENASE 7 (EC 1.2.1.5) [Homo sapiens], full insert sequence [AK081788] |
KLA | .96 |
.98 |
.95 |
.97 |
.97 |
.96 |
1.06 |
| ATP | 1.01 |
1.00 |
.97 |
1.10 |
.95 |
.99 |
1.02 |
| KLA/ATP | .99 |
1.00 |
.97 |
1.00 |
1.00 |
.96 |
.97 |
|
4930438A
08Rik | 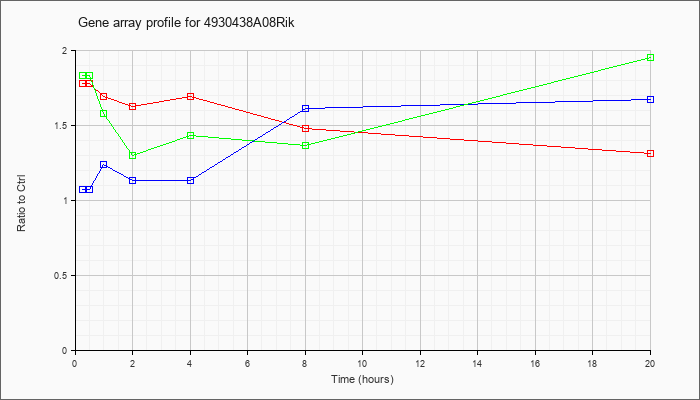 |
XM_001477470 |
gb|PREDICTED: Mus musculus RIKEN cDNA 4930438A08 gene (4930438A08Rik), mRNA [XM_001477470] |
KLA | 1.78 |
1.45 |
1.69 |
1.62 |
1.69 |
1.48 |
1.31 |
| ATP | 1.07 |
1.04 |
1.24 |
1.13 |
1.13 |
1.61 |
1.67 |
| KLA/ATP | 1.83 |
1.46 |
1.58 |
1.30 |
1.43 |
1.36 |
1.95 |
|
| Adh1 | 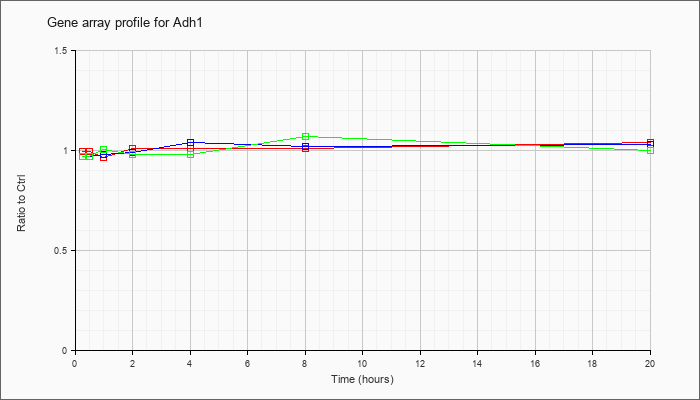 |
AK082149 |
0 day neonate cerebellum cDNA, RIKEN full-length enriched library, clone:C230014D21 product:unclassifiable, full insert sequence [AK082149] |
KLA | .94 |
.96 |
.97 |
.95 |
1.01 |
.95 |
.96 |
| ATP | .99 |
1.05 |
.93 |
.91 |
1.02 |
1.04 |
1.00 |
| KLA/ATP | .87 |
.88 |
.94 |
.92 |
.86 |
1.04 |
.93 |
|
| Adh1 |  |
NM_007409 |
alcohol dehydrogenase 1 (class I) (Adh1), mRNA [NM_007409] |
KLA | 1.02 |
.98 |
.96 |
1.04 |
1.01 |
1.04 |
1.08 |
| ATP | .96 |
1.03 |
1.00 |
1.03 |
1.05 |
1.01 |
1.04 |
| KLA/ATP | 1.02 |
1.03 |
1.04 |
1.01 |
1.04 |
1.09 |
1.04 |
|
| Adh4 | 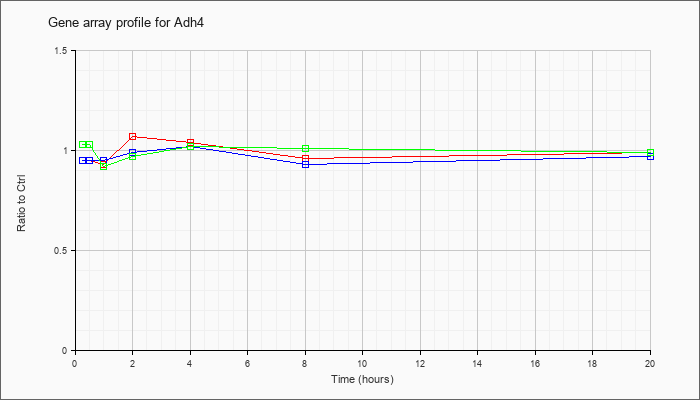 |
NM_011996 |
alcohol dehydrogenase 4 (class II), pi polypeptide (Adh4), mRNA [NM_011996] |
KLA | .95 |
.94 |
.93 |
1.07 |
1.04 |
.96 |
.99 |
| ATP | .95 |
.95 |
.95 |
.99 |
1.02 |
.93 |
.97 |
| KLA/ATP | 1.03 |
1.02 |
.92 |
.97 |
1.02 |
1.01 |
.99 |
|
| Adh5 | 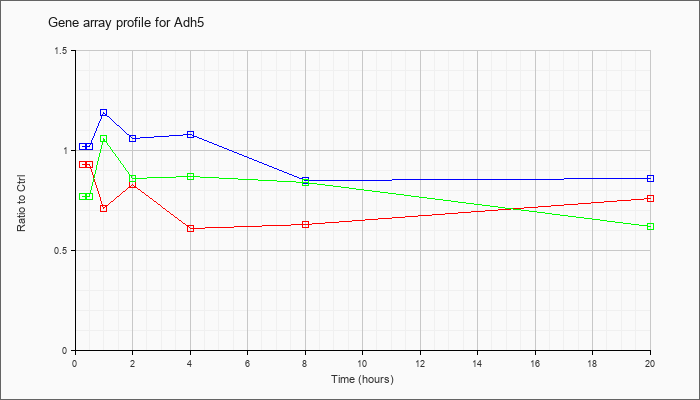 |
NM_007410 |
alcohol dehydrogenase 5 (class III), chi polypeptide (Adh5), mRNA [NM_007410] |
KLA | .93 |
.76 |
.71 |
.83 |
.61 |
.63 |
.76 |
| ATP | 1.02 |
.98 |
1.19 |
1.06 |
1.08 |
.85 |
.86 |
| KLA/ATP | .77 |
.76 |
1.06 |
.86 |
.87 |
.84 |
.62 |
|
| Adh7 | 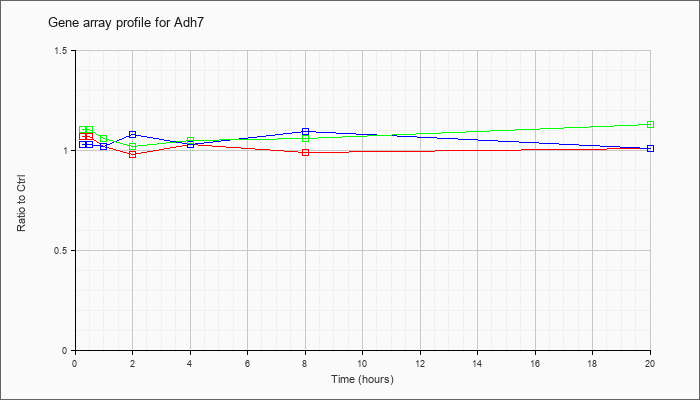 |
NM_009626 |
alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide (Adh7), mRNA [NM_009626] |
KLA | 1.07 |
1.01 |
1.02 |
.98 |
1.03 |
.99 |
1.01 |
| ATP | 1.03 |
1.09 |
1.02 |
1.08 |
1.03 |
1.09 |
1.01 |
| KLA/ATP | 1.10 |
1.03 |
1.06 |
1.02 |
1.05 |
1.06 |
1.13 |
|
| Aldh1a3 | 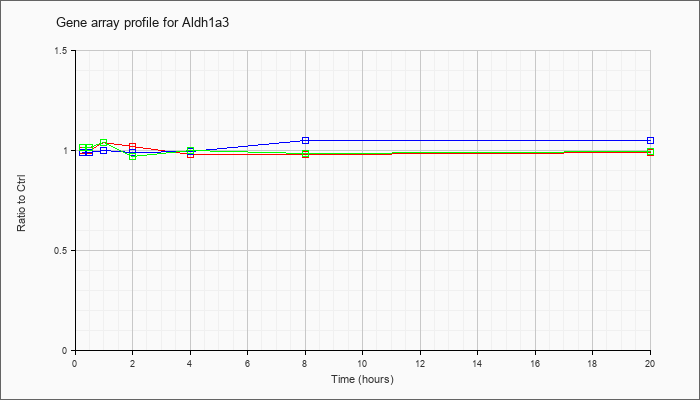 |
AK086764 |
15 days embryo head cDNA, RIKEN full-length enriched library, clone:D930050B08 product:aldehyde dehydrogenase family 1, subfamily A3, full insert sequence. [AK086764] |
KLA | .97 |
1.05 |
1.05 |
1.01 |
1.00 |
.98 |
.97 |
| ATP | 1.00 |
1.03 |
.99 |
.92 |
1.01 |
1.09 |
1.02 |
| KLA/ATP | 1.01 |
.99 |
1.06 |
1.00 |
.97 |
.98 |
1.00 |
|
| Aldh1a3 |  |
NM_053080 |
aldehyde dehydrogenase family 1, subfamily A3 (Aldh1a3), mRNA [NM_053080] |
KLA | 1.03 |
.96 |
1.03 |
1.03 |
.96 |
.98 |
1.01 |
| ATP | .98 |
1.04 |
1.01 |
1.06 |
.98 |
1.01 |
1.08 |
| KLA/ATP | 1.02 |
1.00 |
1.02 |
.94 |
1.03 |
.99 |
.99 |
|
| Aldh3a1 | 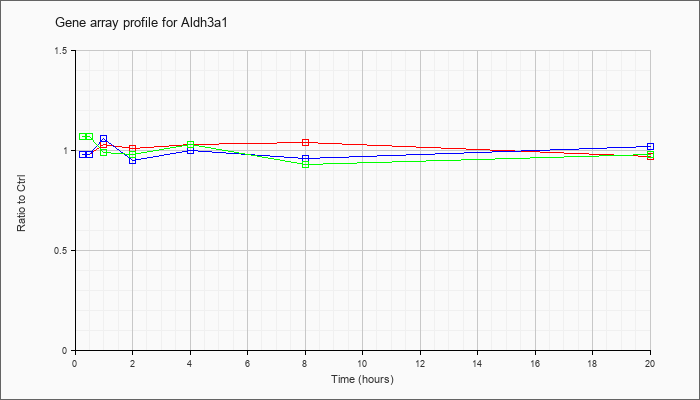 |
NM_007436 |
aldehyde dehydrogenase family 3, subfamily A1 (Aldh3a1), transcript variant 1, mRNA [NM_007436] |
KLA | .98 |
.98 |
1.03 |
1.01 |
1.03 |
1.04 |
.97 |
| ATP | .98 |
1.01 |
1.06 |
.95 |
1.00 |
.96 |
1.02 |
| KLA/ATP | 1.07 |
1.08 |
.99 |
.98 |
1.03 |
.93 |
.98 |
|
| Aldh3b1 | 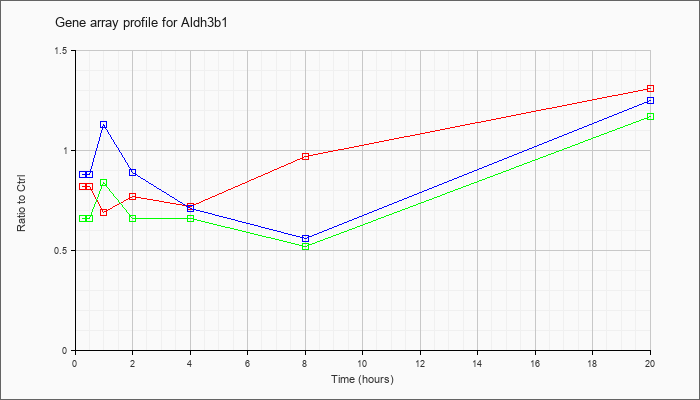 |
NM_026316 |
aldehyde dehydrogenase 3 family, member B1 (Aldh3b1), mRNA [NM_026316] |
KLA | .82 |
.77 |
.69 |
.77 |
.72 |
.97 |
1.31 |
| ATP | .88 |
.85 |
1.13 |
.89 |
.71 |
.56 |
1.25 |
| KLA/ATP | .66 |
.72 |
.84 |
.66 |
.66 |
.52 |
1.17 |
|
| Aldh3b2 | 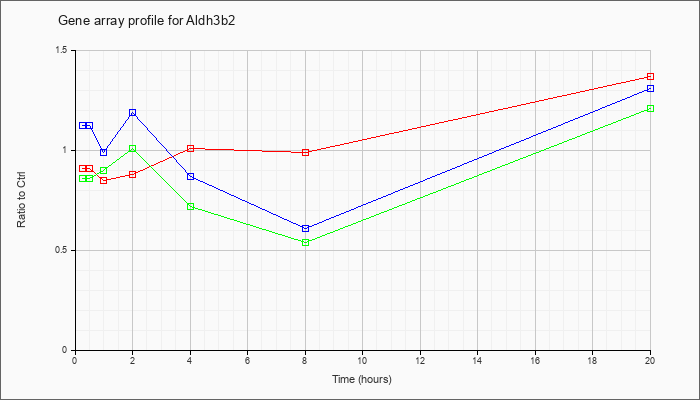 |
XM_129134 |
gb|PREDICTED: Mus musculus aldehyde dehydrogenase 3 family, member B2 (Aldh3b2), mRNA [XM_129134] |
KLA | .91 |
.93 |
.85 |
.88 |
1.01 |
.99 |
1.37 |
| ATP | 1.12 |
1.11 |
.99 |
1.19 |
.87 |
.61 |
1.31 |
| KLA/ATP | .86 |
1.02 |
.90 |
1.01 |
.72 |
.54 |
1.21 |
|
| Aoc2 | 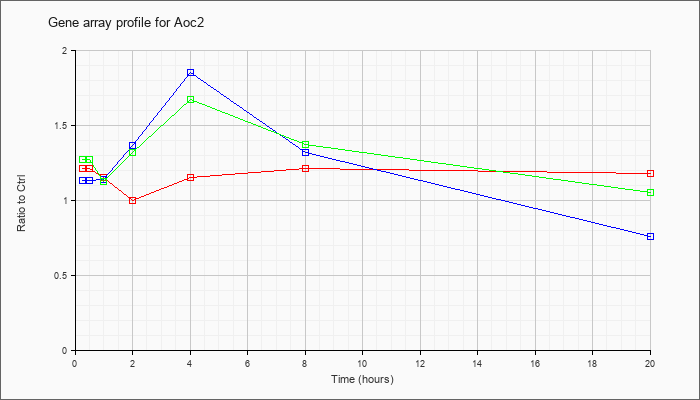 |
NM_178932 |
amine oxidase, copper containing 2 (retina-specific) (Aoc2), mRNA [NM_178932] |
KLA | 1.21 |
1.19 |
1.15 |
1.00 |
1.15 |
1.21 |
1.18 |
| ATP | 1.13 |
1.13 |
1.14 |
1.36 |
1.85 |
1.32 |
.76 |
| KLA/ATP | 1.27 |
1.34 |
1.12 |
1.32 |
1.67 |
1.37 |
1.05 |
|
| Aoc3 | 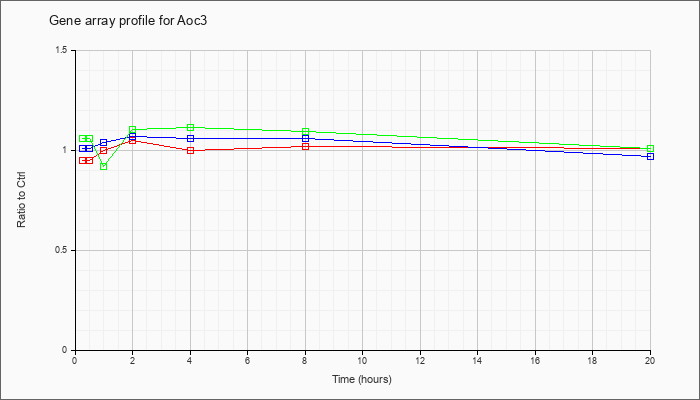 |
NM_009675 |
amine oxidase, copper containing 3 (Aoc3), mRNA [NM_009675] |
KLA | .95 |
1.06 |
1.00 |
1.05 |
1.00 |
1.02 |
1.01 |
| ATP | 1.01 |
.90 |
1.04 |
1.07 |
1.06 |
1.06 |
.97 |
| KLA/ATP | 1.06 |
.98 |
.92 |
1.10 |
1.11 |
1.09 |
1.01 |
|
| Aox1 | 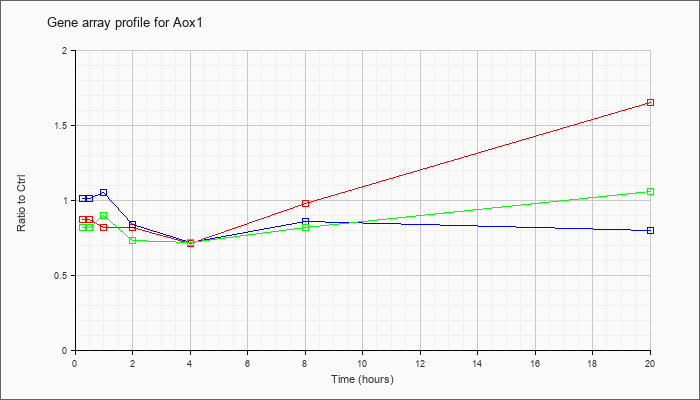 |
NM_009676 |
aldehyde oxidase 1 (Aox1), mRNA [NM_009676] |
KLA | .87 |
.83 |
.82 |
.82 |
.71 |
.98 |
1.65 |
| ATP | 1.01 |
1.02 |
1.05 |
.84 |
.72 |
.86 |
.80 |
| KLA/ATP | .82 |
.84 |
.90 |
.73 |
.72 |
.82 |
1.06 |
|
| Aox3 | 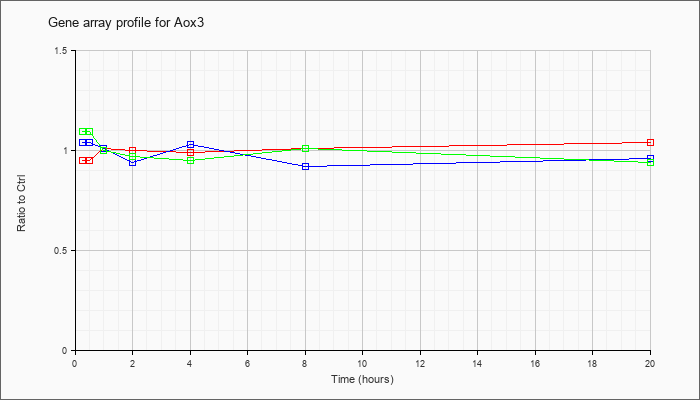 |
NM_023617 |
aldehyde oxidase 3 (Aox3), mRNA [NM_023617] |
KLA | .95 |
1.02 |
1.01 |
1.00 |
.99 |
1.01 |
1.04 |
| ATP | 1.04 |
1.03 |
1.01 |
.94 |
1.03 |
.92 |
.96 |
| KLA/ATP | 1.09 |
1.11 |
1.00 |
.97 |
.95 |
1.01 |
.94 |
|
| Aox4 | 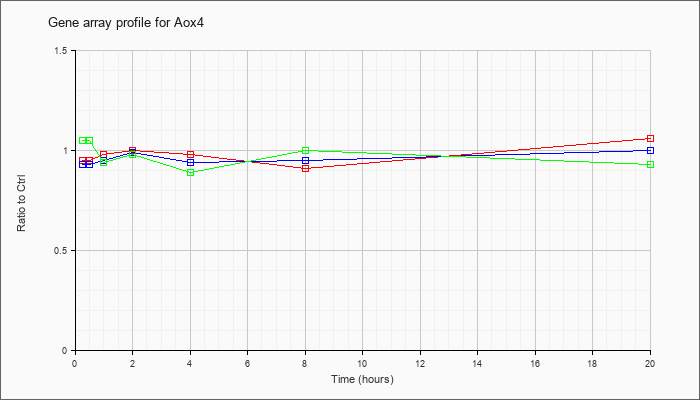 |
NM_023631 |
aldehyde oxidase 4 (Aox4), mRNA [NM_023631] |
KLA | .95 |
.97 |
.98 |
1.00 |
.98 |
.91 |
1.06 |
| ATP | .93 |
.99 |
.95 |
.99 |
.94 |
.95 |
1.00 |
| KLA/ATP | 1.05 |
1.00 |
.94 |
.98 |
.89 |
1.00 |
.93 |
|
| Comt | 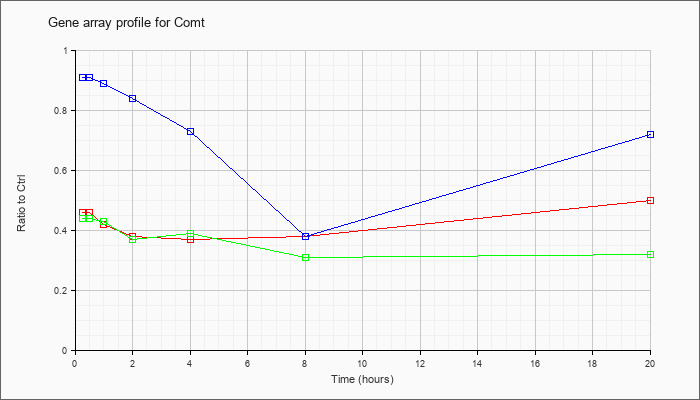 |
NM_007744 |
catechol-O-methyltransferase (Comt), transcript variant 2, mRNA [NM_007744] |
KLA | .46 |
.45 |
.42 |
.38 |
.37 |
.38 |
.50 |
| ATP | .91 |
.90 |
.89 |
.84 |
.73 |
.38 |
.72 |
| KLA/ATP | .44 |
.44 |
.43 |
.37 |
.39 |
.31 |
.32 |
|
| Dbh | 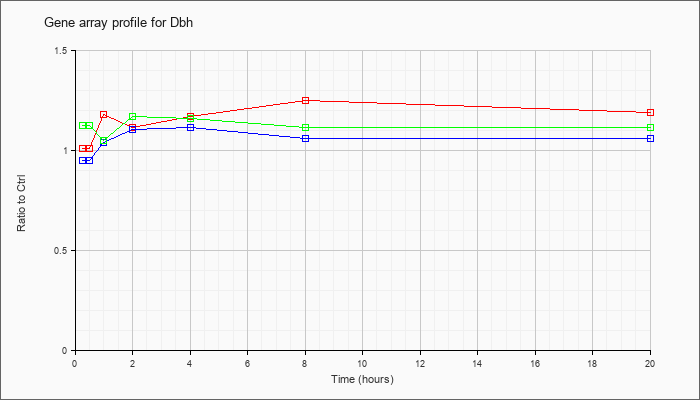 |
NM_138942 |
dopamine beta hydroxylase (Dbh), mRNA [NM_138942] |
KLA | 1.01 |
1.08 |
1.18 |
1.11 |
1.17 |
1.25 |
1.19 |
| ATP | .95 |
1.03 |
1.04 |
1.10 |
1.11 |
1.06 |
1.06 |
| KLA/ATP | 1.12 |
1.04 |
1.05 |
1.17 |
1.16 |
1.11 |
1.11 |
|
| Dct | 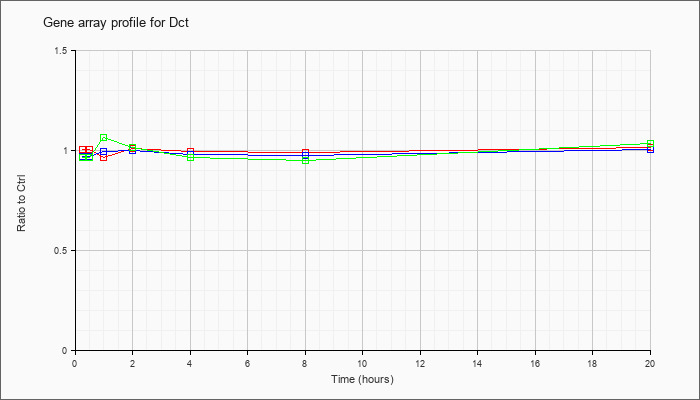 |
NM_010024 |
dopachrome tautomerase (Dct), mRNA [NM_010024] |
KLA | 1.01 |
.97 |
.97 |
1.01 |
1.00 |
.99 |
1.02 |
| ATP | .97 |
1.00 |
1.00 |
1.00 |
.98 |
.98 |
1.01 |
| KLA/ATP | .97 |
1.01 |
1.07 |
1.02 |
.97 |
.95 |
1.04 |
|
| Ddc | 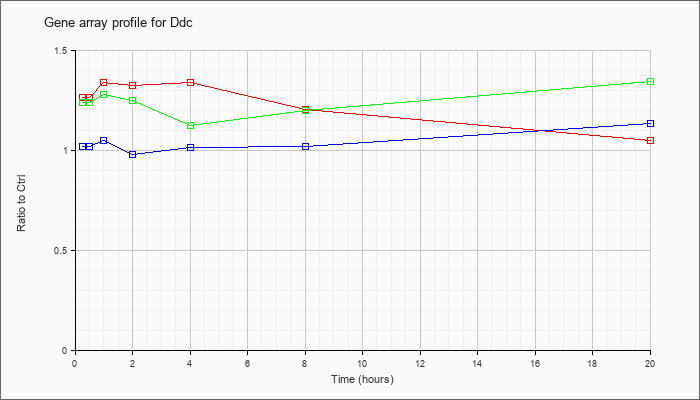 |
NM_016672 |
dopa decarboxylase (Ddc), mRNA [NM_016672] |
KLA | 1.27 |
1.28 |
1.34 |
1.33 |
1.34 |
1.21 |
1.05 |
| ATP | 1.02 |
.99 |
1.05 |
.98 |
1.02 |
1.02 |
1.14 |
| KLA/ATP | 1.24 |
1.22 |
1.28 |
1.25 |
1.12 |
1.20 |
1.35 |
|
| Fah | 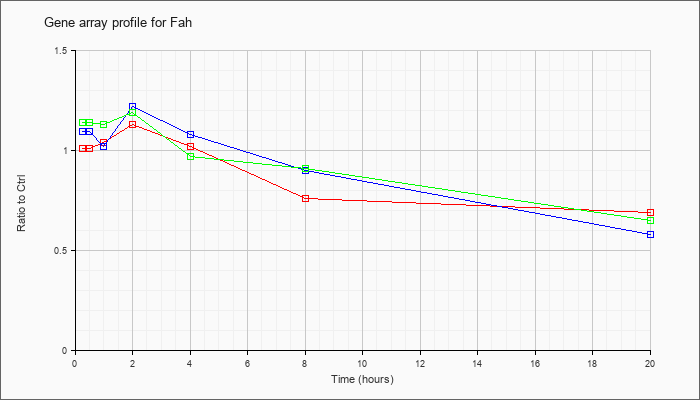 |
NM_010176 |
fumarylacetoacetate hydrolase (Fah), mRNA [NM_010176] |
KLA | 1.01 |
1.09 |
1.04 |
1.13 |
1.02 |
.76 |
.69 |
| ATP | 1.09 |
1.09 |
1.02 |
1.22 |
1.08 |
.90 |
.58 |
| KLA/ATP | 1.14 |
1.09 |
1.13 |
1.19 |
.97 |
.91 |
.65 |
|
| Fahd1 | 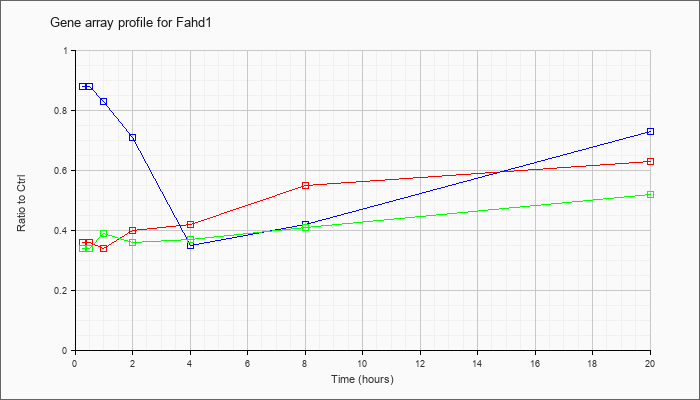 |
NM_023480 |
fumarylacetoacetate hydrolase domain containing 1 (Fahd1), nuclear gene encoding mitochondrial protein, mRNA [NM_023480] |
KLA | .36 |
.34 |
.34 |
.40 |
.42 |
.55 |
.63 |
| ATP | .88 |
.86 |
.83 |
.71 |
.35 |
.42 |
.73 |
| KLA/ATP | .34 |
.32 |
.39 |
.36 |
.37 |
.41 |
.52 |
|
| Got1 | 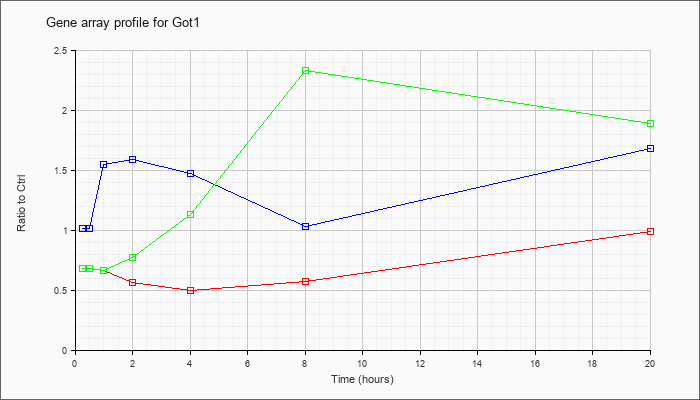 |
NM_010324 |
glutamate oxaloacetate transaminase 1, soluble (Got1), mRNA [NM_010324] |
KLA | .68 |
.70 |
.66 |
.56 |
.50 |
.57 |
.99 |
| ATP | 1.01 |
1.24 |
1.55 |
1.59 |
1.47 |
1.03 |
1.68 |
| KLA/ATP | .68 |
.71 |
.66 |
.77 |
1.13 |
2.33 |
1.89 |
|
| Got2 | 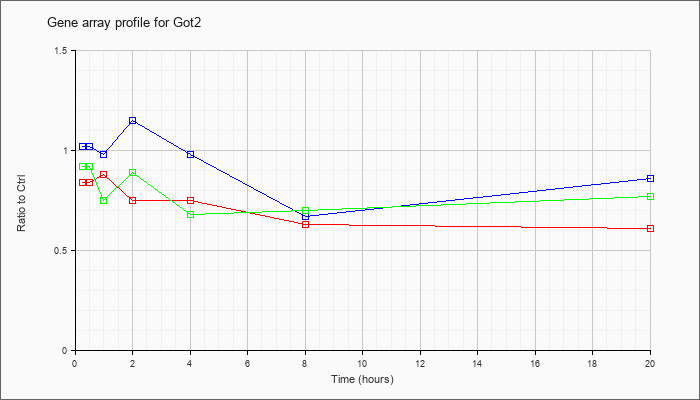 |
NM_010325 |
glutamate oxaloacetate transaminase 2, mitochondrial (Got2), nuclear gene encoding mitochondrial protein, mRNA [NM_010325] |
KLA | .84 |
.86 |
.88 |
.75 |
.75 |
.63 |
.61 |
| ATP | 1.02 |
1.12 |
.98 |
1.15 |
.98 |
.67 |
.86 |
| KLA/ATP | .92 |
.89 |
.75 |
.89 |
.68 |
.70 |
.77 |
|
| Gstz1 | 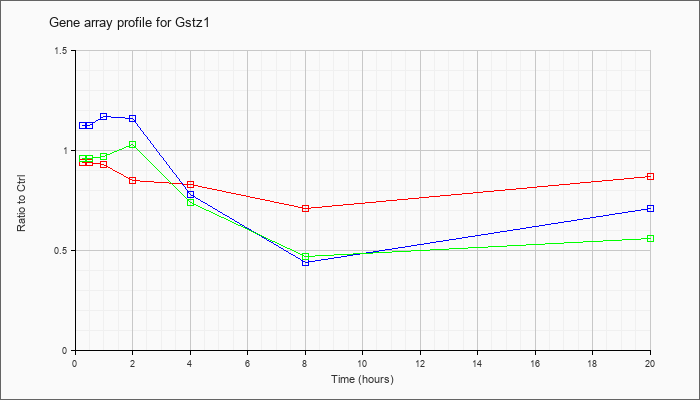 |
NM_010363 |
glutathione transferase zeta 1 (maleylacetoacetate isomerase) (Gstz1), mRNA [NM_010363] |
KLA | .94 |
.85 |
.93 |
.85 |
.83 |
.71 |
.87 |
| ATP | 1.12 |
1.08 |
1.17 |
1.16 |
.78 |
.44 |
.71 |
| KLA/ATP | .96 |
.98 |
.97 |
1.03 |
.74 |
.47 |
.56 |
|
| Hemk1 | 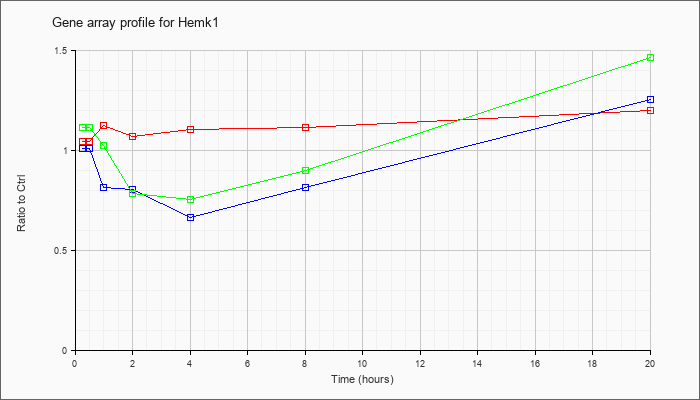 |
NM_133984 |
HemK methyltransferase family member 1 (Hemk1), mRNA [NM_133984] |
KLA | 1.05 |
1.10 |
1.13 |
1.07 |
1.10 |
1.12 |
1.20 |
| ATP | 1.01 |
1.00 |
.82 |
.81 |
.67 |
.82 |
1.26 |
| KLA/ATP | 1.11 |
1.15 |
1.03 |
.79 |
.76 |
.90 |
1.47 |
|
| Hgd | 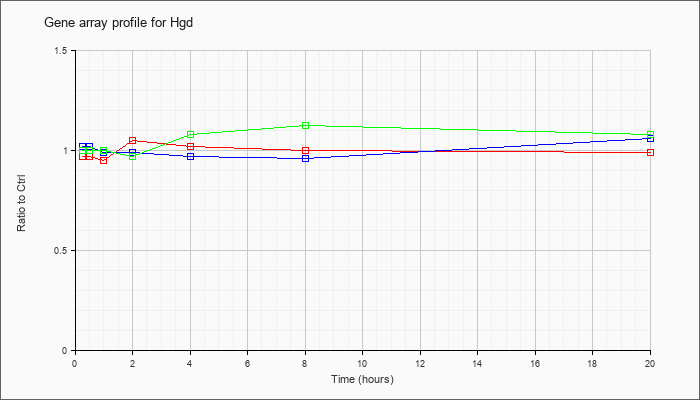 |
NM_013547 |
homogentisate 1, 2-dioxygenase (Hgd), mRNA [NM_013547] |
KLA | .97 |
1.03 |
.95 |
1.05 |
1.02 |
1.00 |
.99 |
| ATP | 1.02 |
1.00 |
.99 |
.99 |
.97 |
.96 |
1.06 |
| KLA/ATP | 1.00 |
.96 |
1.00 |
.97 |
1.08 |
1.12 |
1.08 |
|
| Hpd | 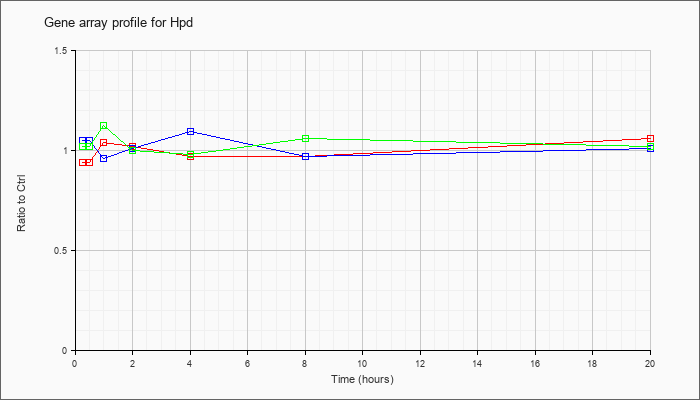 |
NM_008277 |
4-hydroxyphenylpyruvic acid dioxygenase (Hpd), mRNA [NM_008277] |
KLA | .94 |
1.10 |
1.04 |
1.02 |
.97 |
.97 |
1.06 |
| ATP | 1.05 |
1.07 |
.96 |
1.01 |
1.09 |
.97 |
1.01 |
| KLA/ATP | 1.02 |
1.03 |
1.12 |
1.00 |
.98 |
1.06 |
1.02 |
|
| Il4i1 | 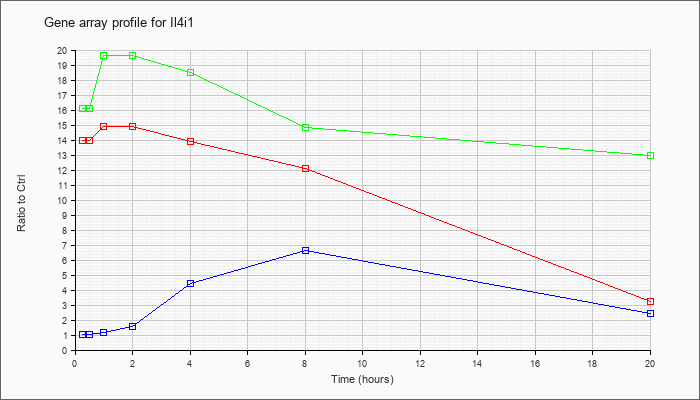 |
NM_010215 |
interleukin 4 induced 1 (Il4i1), mRNA [NM_010215] |
KLA | 13.98 |
13.74 |
14.90 |
14.92 |
13.89 |
12.12 |
3.25 |
| ATP | 1.02 |
.98 |
1.14 |
1.59 |
4.45 |
6.63 |
2.41 |
| KLA/ATP | 16.09 |
15.56 |
19.65 |
19.66 |
18.47 |
14.84 |
12.99 |
|
| Lao1 | 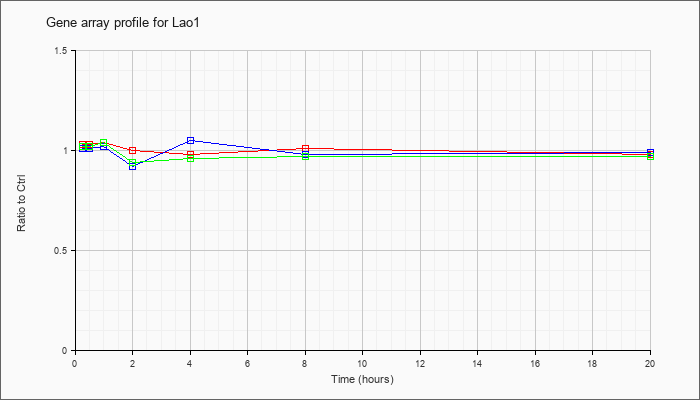 |
NM_133892 |
L-amino acid oxidase 1 (Lao1), mRNA [NM_133892] |
KLA | 1.03 |
1.03 |
1.04 |
1.00 |
.98 |
1.01 |
.98 |
| ATP | 1.01 |
1.03 |
1.02 |
.92 |
1.05 |
.98 |
.99 |
| KLA/ATP | 1.02 |
.96 |
1.04 |
.94 |
.96 |
.97 |
.97 |
|
| Maoa | 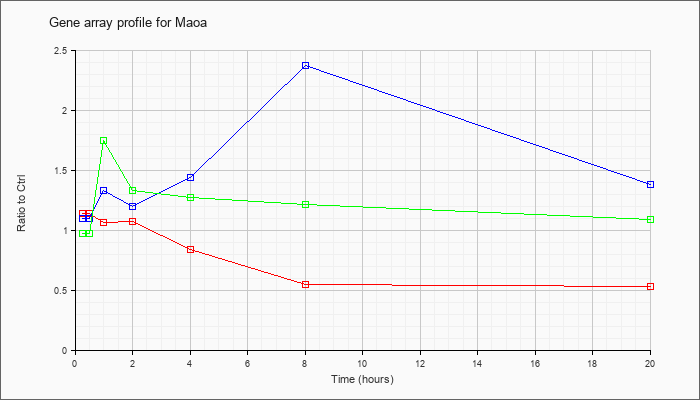 |
NM_173740 |
monoamine oxidase A (Maoa), nuclear gene encoding mitochondrial protein, mRNA [NM_173740] |
KLA | 1.14 |
1.11 |
1.06 |
1.07 |
.84 |
.55 |
.53 |
| ATP | 1.10 |
1.07 |
1.33 |
1.20 |
1.44 |
2.37 |
1.38 |
| KLA/ATP | .97 |
1.11 |
1.75 |
1.33 |
1.27 |
1.21 |
1.09 |
|
| Maob | 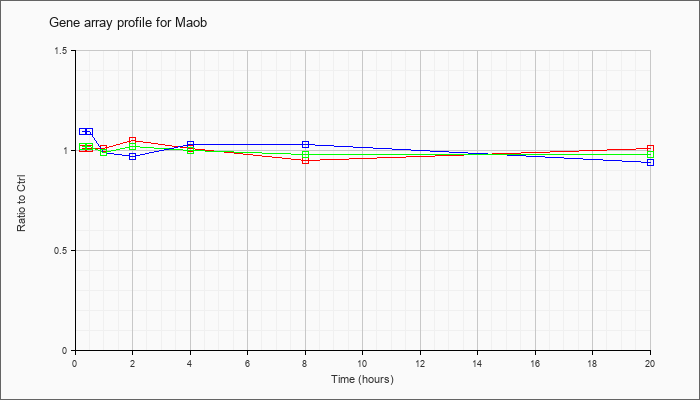 |
NM_172778 |
monoamine oxidase B (Maob), nuclear gene encoding mitochondrial protein, mRNA [NM_172778] |
KLA | 1.01 |
.97 |
1.01 |
1.05 |
1.01 |
.95 |
1.01 |
| ATP | 1.09 |
.98 |
.99 |
.97 |
1.03 |
1.03 |
.94 |
| KLA/ATP | 1.02 |
1.05 |
.99 |
1.02 |
1.00 |
.98 |
.98 |
|
| Mettl2 | 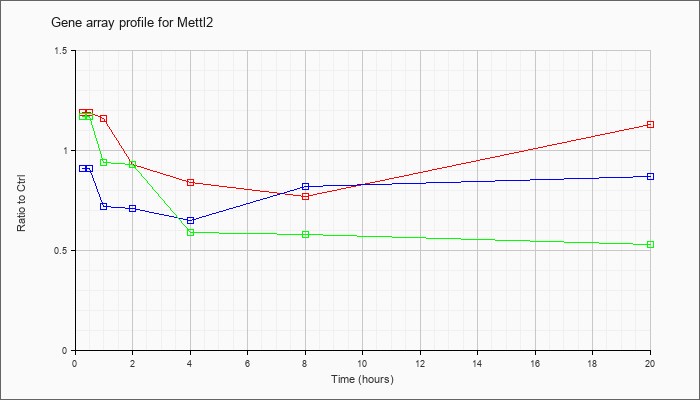 |
NM_172567 |
methyltransferase like 2 (Mettl2), mRNA [NM_172567] |
KLA | 1.19 |
1.15 |
1.16 |
.93 |
.84 |
.77 |
1.13 |
| ATP | .91 |
.87 |
.72 |
.71 |
.65 |
.82 |
.87 |
| KLA/ATP | 1.17 |
1.15 |
.94 |
.93 |
.59 |
.58 |
.53 |
|
| Mettl6 | 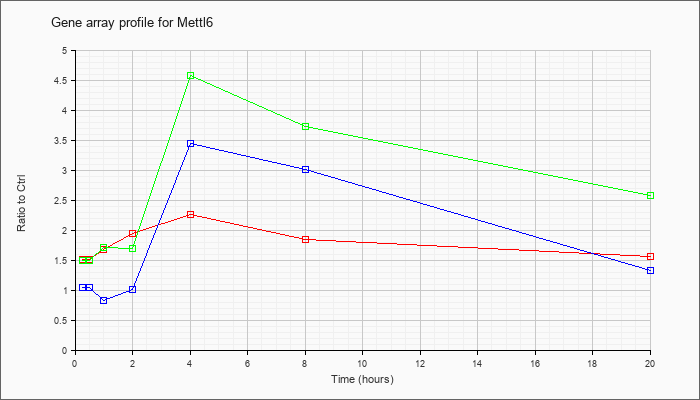 |
NM_025907 |
methyltransferase like 6 (Mettl6), mRNA [NM_025907] |
KLA | 1.52 |
1.55 |
1.68 |
1.94 |
2.26 |
1.84 |
1.56 |
| ATP | 1.05 |
.96 |
.83 |
1.01 |
3.44 |
3.01 |
1.33 |
| KLA/ATP | 1.49 |
1.59 |
1.71 |
1.69 |
4.57 |
3.72 |
2.58 |
|
| Mif | 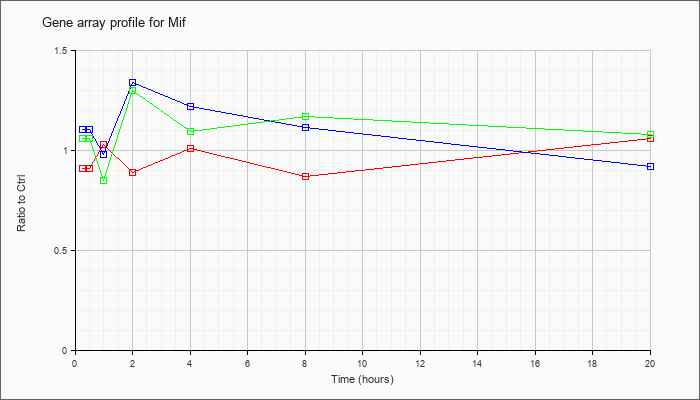 |
NM_010798 |
macrophage migration inhibitory factor (Mif), mRNA [NM_010798] |
KLA | .91 |
1.04 |
1.03 |
.89 |
1.01 |
.87 |
1.06 |
| ATP | 1.10 |
1.26 |
.98 |
1.34 |
1.22 |
1.11 |
.92 |
| KLA/ATP | 1.06 |
1.15 |
.85 |
1.30 |
1.09 |
1.17 |
1.08 |
|
Mm.15744
2 | 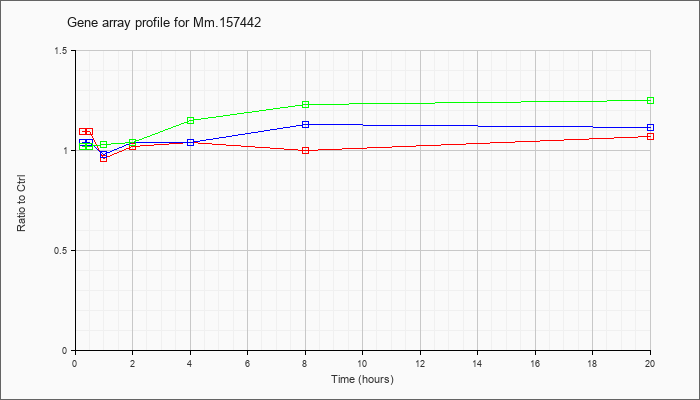 |
41152515 |
Unknown |
KLA | 1.09 |
1.07 |
.96 |
1.02 |
1.04 |
1.00 |
1.07 |
| ATP | 1.04 |
.96 |
.98 |
1.04 |
1.04 |
1.13 |
1.11 |
| KLA/ATP | 1.02 |
.97 |
1.03 |
1.04 |
1.15 |
1.23 |
1.25 |
|
Mm.23812
7 | 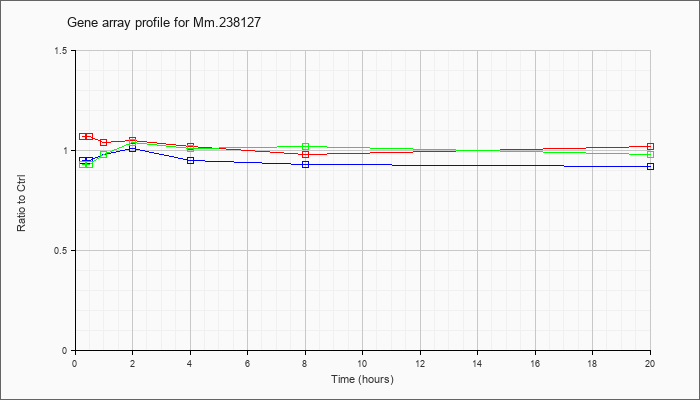 |
168823498 |
Unknown |
KLA | 1.07 |
.97 |
1.04 |
1.05 |
1.02 |
.98 |
1.02 |
| ATP | .95 |
1.00 |
.98 |
1.01 |
.95 |
.93 |
.92 |
| KLA/ATP | .93 |
.98 |
.98 |
1.04 |
1.01 |
1.02 |
.98 |
|
Mm.26684
3 | 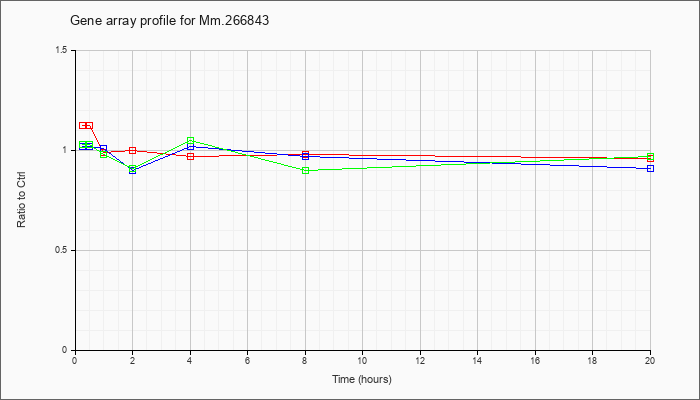 |
94405693 |
Unknown |
KLA | 1.12 |
.99 |
.99 |
1.00 |
.97 |
.98 |
.96 |
| ATP | 1.02 |
1.08 |
1.01 |
.90 |
1.02 |
.97 |
.91 |
| KLA/ATP | 1.03 |
.96 |
.98 |
.91 |
1.05 |
.90 |
.97 |
|
Mm.34137
| 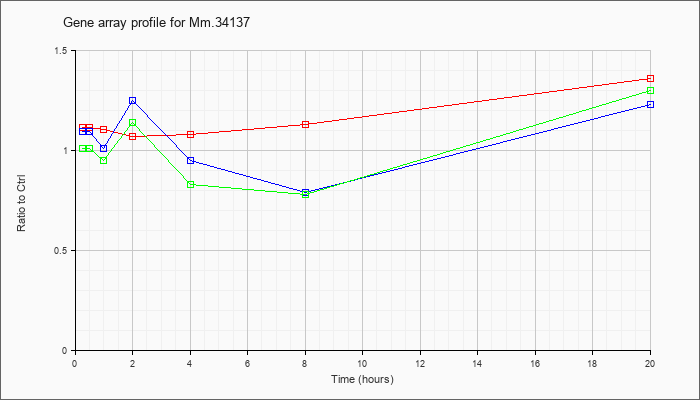 |
94405716 |
Unknown |
KLA | 1.11 |
1.12 |
1.10 |
1.07 |
1.08 |
1.13 |
1.36 |
| ATP | 1.09 |
1.14 |
1.01 |
1.25 |
.95 |
.79 |
1.23 |
| KLA/ATP | 1.01 |
1.24 |
.95 |
1.14 |
.83 |
.78 |
1.30 |
|
Mm.39052
9 | 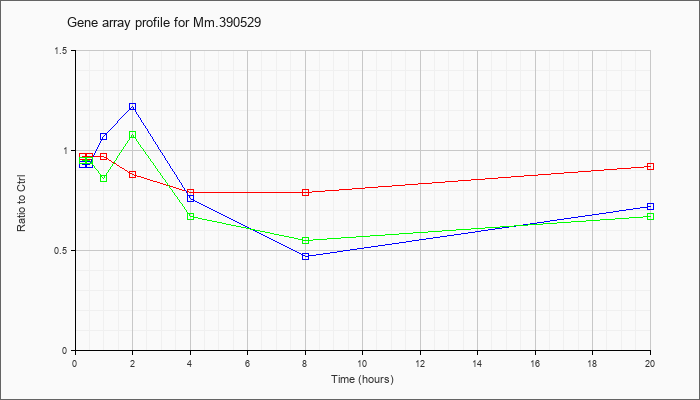 |
133892597 |
Unknown |
KLA | .97 |
1.02 |
.97 |
.88 |
.79 |
.79 |
.92 |
| ATP | .93 |
1.10 |
1.07 |
1.22 |
.76 |
.47 |
.72 |
| KLA/ATP | .95 |
.99 |
.86 |
1.08 |
.67 |
.55 |
.67 |
|
| Pnmt | 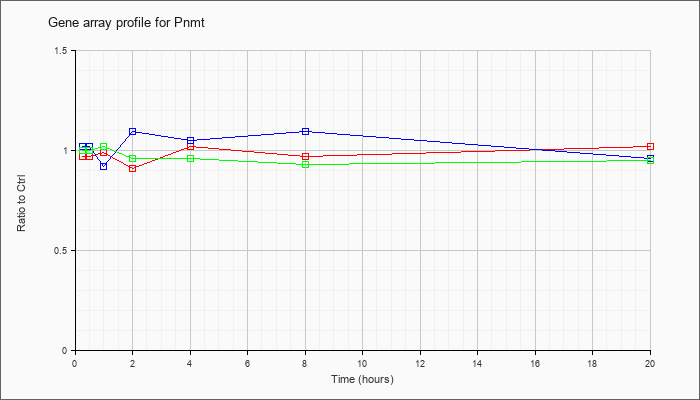 |
NM_008890 |
phenylethanolamine-N-methyltransferase (Pnmt), mRNA [NM_008890] |
KLA | .97 |
.99 |
.99 |
.91 |
1.02 |
.97 |
1.02 |
| ATP | 1.02 |
1.03 |
.92 |
1.09 |
1.05 |
1.09 |
.96 |
| KLA/ATP | 1.00 |
1.02 |
1.02 |
.96 |
.96 |
.93 |
.95 |
|
| Tat | 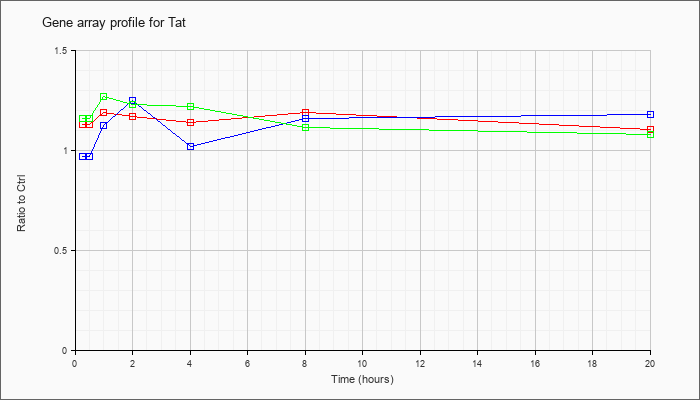 |
NM_146214 |
tyrosine aminotransferase (Tat), nuclear gene encoding mitochondrial protein, mRNA [NM_146214] |
KLA | 1.13 |
1.09 |
1.19 |
1.17 |
1.14 |
1.19 |
1.10 |
| ATP | .97 |
1.08 |
1.12 |
1.25 |
1.02 |
1.16 |
1.18 |
| KLA/ATP | 1.16 |
1.21 |
1.27 |
1.23 |
1.22 |
1.11 |
1.08 |
|
| Th | 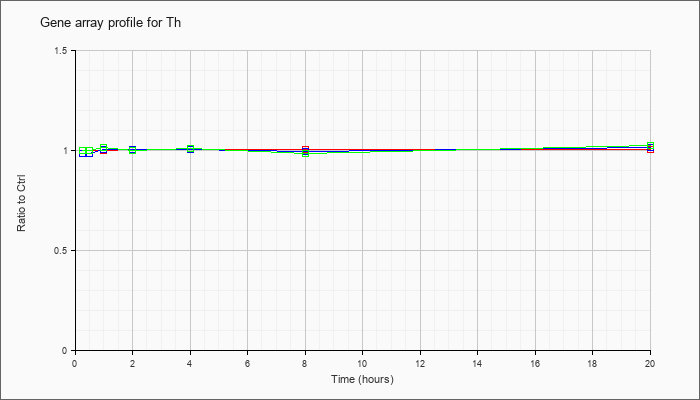 |
NM_009377 |
tyrosine hydroxylase (Th), mRNA [NM_009377] |
KLA | 1.00 |
1.01 |
1.00 |
1.00 |
1.00 |
1.00 |
1.00 |
| ATP | .98 |
1.01 |
1.00 |
1.00 |
1.00 |
.99 |
1.01 |
| KLA/ATP | 1.00 |
1.01 |
1.01 |
1.00 |
1.01 |
.98 |
1.02 |
|
| Tpo | 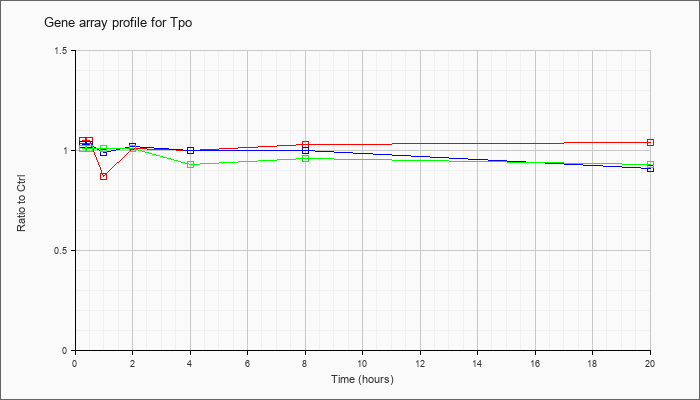 |
NM_009417 |
thyroid peroxidase (Tpo), mRNA [NM_009417] |
KLA | 1.05 |
1.05 |
.87 |
1.01 |
1.00 |
1.03 |
1.04 |
| ATP | 1.03 |
.95 |
.99 |
1.02 |
1.00 |
1.00 |
.91 |
| KLA/ATP | 1.01 |
.99 |
1.01 |
1.01 |
.93 |
.96 |
.93 |
|
| Tyr | 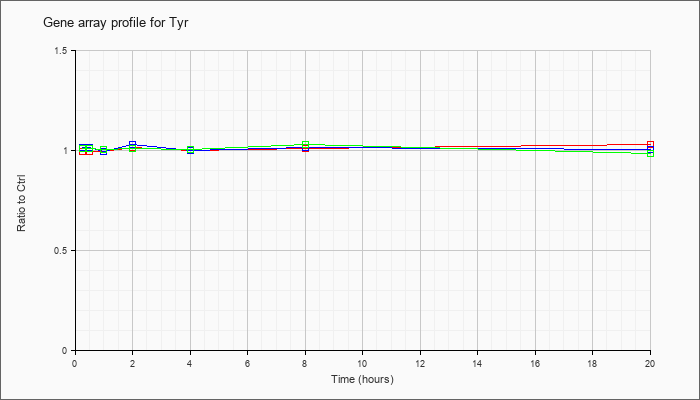 |
NM_011661 |
tyrosinase (Tyr), mRNA [NM_011661] |
KLA | .99 |
1.02 |
.99 |
1.01 |
1.00 |
1.01 |
1.03 |
| ATP | 1.01 |
.98 |
.99 |
1.03 |
1.00 |
1.01 |
1.00 |
| KLA/ATP | 1.01 |
1.00 |
1.00 |
1.01 |
1.00 |
1.03 |
.98 |
|
| Tyrp1 | 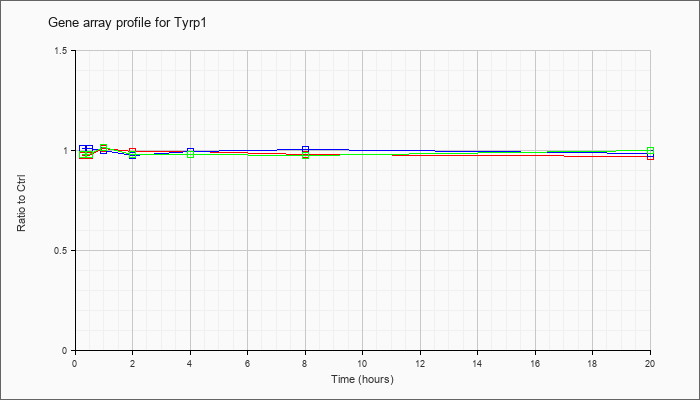 |
NM_031202 |
tyrosinase-related protein 1 (Tyrp1), mRNA [NM_031202] |
KLA | .97 |
.98 |
1.01 |
.99 |
.99 |
.98 |
.97 |
| ATP | 1.01 |
.99 |
1.00 |
.97 |
.99 |
1.00 |
.98 |
| KLA/ATP | .98 |
.99 |
1.01 |
.98 |
.98 |
.97 |
1.00 |
|
| Wbscr22 | 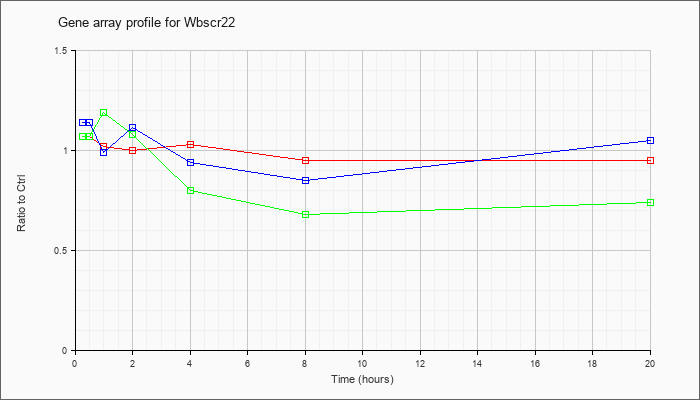 |
NM_025375 |
Williams Beuren syndrome chromosome region 22 (Wbscr22), mRNA [NM_025375] |
KLA | 1.07 |
1.03 |
1.02 |
1.00 |
1.03 |
.95 |
.95 |
| ATP | 1.14 |
1.21 |
.99 |
1.11 |
.94 |
.85 |
1.05 |
| KLA/ATP | 1.07 |
1.08 |
1.19 |
1.08 |
.80 |
.68 |
.74 |
|