| Gene symbol |
Time course plot |
Accession |
Description |
Treatment |
15min |
30min |
1hr |
2hr |
4hr |
8hr |
20hr |
4933405O
20Rik | 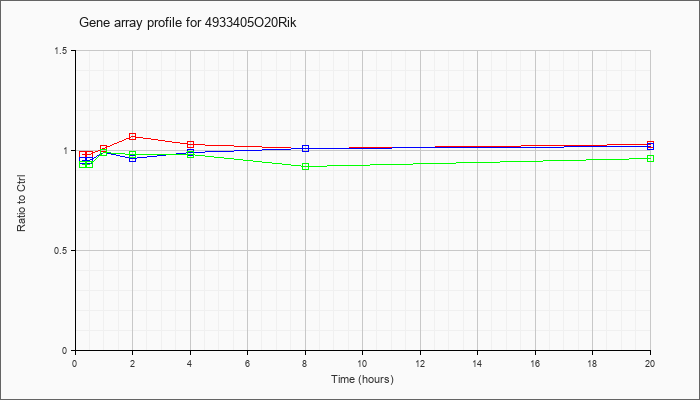 |
NM_172901 |
RIKEN cDNA 4933405O20 gene (4933405O20Rik), mRNA [NM_172901] |
KLA | .98 |
.99 |
1.01 |
1.07 |
1.03 |
1.01 |
1.03 |
| ATP | .95 |
.97 |
.99 |
.96 |
.99 |
1.01 |
1.02 |
| KLA/ATP | .93 |
1.01 |
.99 |
.98 |
.98 |
.92 |
.96 |
|
| Aadat | 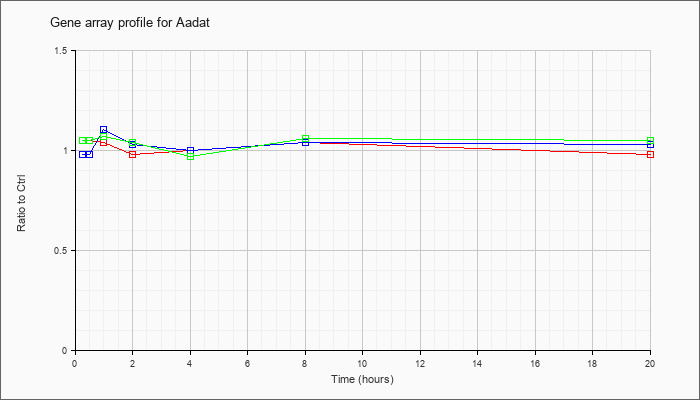 |
NM_011834 |
aminoadipate aminotransferase (Aadat), mRNA [NM_011834] |
KLA | 1.05 |
1.06 |
1.04 |
.98 |
1.00 |
1.04 |
.98 |
| ATP | .98 |
.99 |
1.10 |
1.03 |
1.00 |
1.04 |
1.03 |
| KLA/ATP | 1.05 |
1.09 |
1.07 |
1.04 |
.97 |
1.06 |
1.05 |
|
| Aco1 | 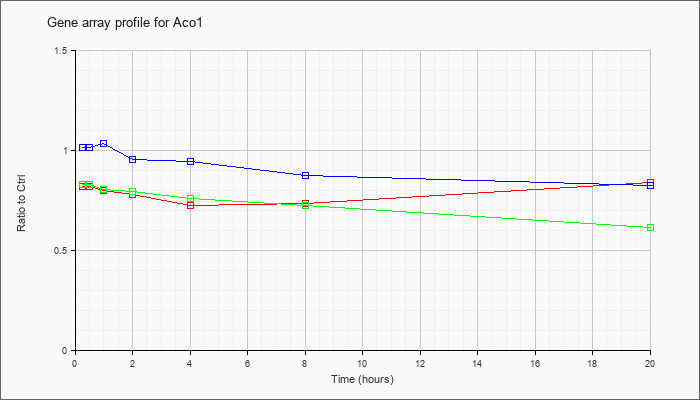 |
NM_007386 |
aconitase 1 (Aco1), mRNA [NM_007386] |
KLA | .82 |
.85 |
.80 |
.78 |
.73 |
.74 |
.84 |
| ATP | 1.02 |
1.07 |
1.04 |
.96 |
.95 |
.88 |
.83 |
| KLA/ATP | .83 |
.86 |
.81 |
.80 |
.76 |
.73 |
.62 |
|
| Aco2 | 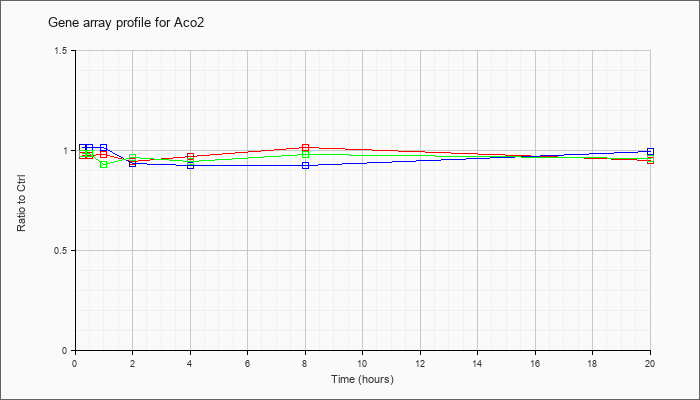 |
AK029122 |
10 days neonate skin cDNA, RIKEN full-length enriched library, clone:4732494K09 product:hypothetical protein, full insert sequence [AK029122] |
KLA | 1.02 |
1.02 |
.99 |
.98 |
.97 |
1.04 |
.83 |
| ATP | .98 |
1.04 |
1.05 |
.95 |
.95 |
1.02 |
1.00 |
| KLA/ATP | .96 |
.95 |
1.00 |
1.04 |
1.06 |
1.03 |
.99 |
|
| Aco2 |  |
NM_080633 |
aconitase 2, mitochondrial (Aco2), nuclear gene encoding mitochondrial protein, mRNA [NM_080633] |
KLA | .93 |
.96 |
.97 |
.91 |
.97 |
.99 |
1.07 |
| ATP | 1.05 |
1.04 |
.98 |
.92 |
.90 |
.83 |
.99 |
| KLA/ATP | 1.01 |
1.06 |
.86 |
.89 |
.83 |
.93 |
.93 |
|
| Acy1 | 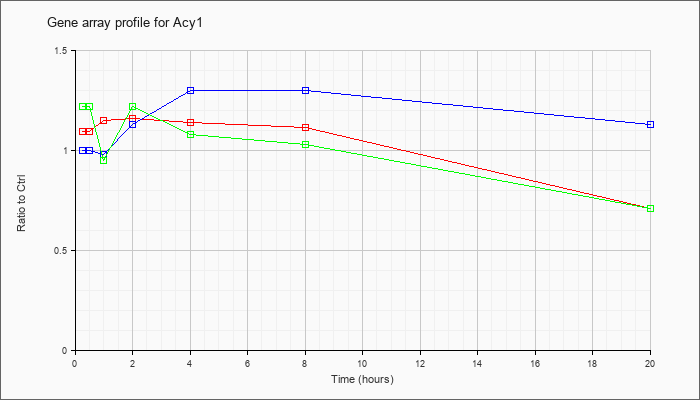 |
NM_025371 |
aminoacylase 1 (Acy1), mRNA [NM_025371] |
KLA | 1.09 |
1.23 |
1.15 |
1.16 |
1.14 |
1.11 |
.71 |
| ATP | 1.00 |
1.03 |
.98 |
1.13 |
1.30 |
1.30 |
1.13 |
| KLA/ATP | 1.22 |
1.18 |
.95 |
1.22 |
1.08 |
1.03 |
.71 |
|
| Bcat1 | 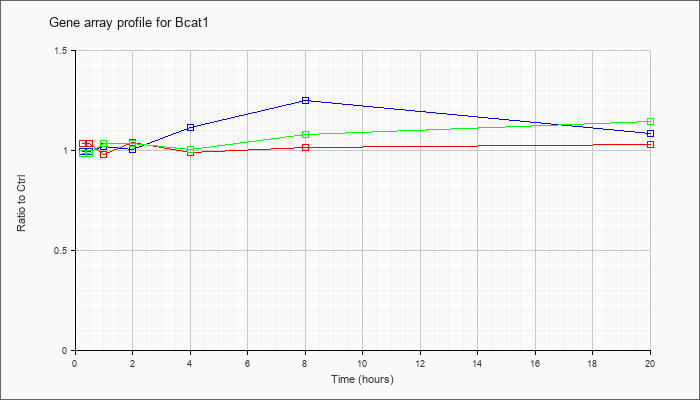 |
NM_001024468 |
branched chain aminotransferase 1, cytosolic (Bcat1), transcript variant 1, mRNA [NM_001024468] |
KLA | 1.04 |
1.03 |
.98 |
1.04 |
.99 |
1.02 |
1.03 |
| ATP | 1.00 |
1.01 |
1.02 |
1.01 |
1.12 |
1.25 |
1.09 |
| KLA/ATP | .99 |
1.00 |
1.04 |
1.04 |
1.01 |
1.08 |
1.15 |
|
| Bcat2 | 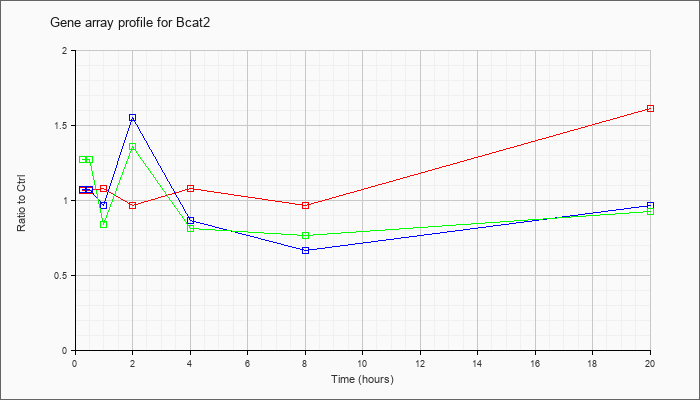 |
NM_009737 |
branched chain aminotransferase 2, mitochondrial (Bcat2), nuclear gene encoding mitochondrial protein, mRNA [NM_009737] |
KLA | 1.06 |
1.05 |
1.08 |
.97 |
1.08 |
.96 |
1.61 |
| ATP | 1.07 |
1.42 |
.96 |
1.55 |
.87 |
.66 |
.96 |
| KLA/ATP | 1.27 |
1.29 |
.84 |
1.36 |
.81 |
.76 |
.92 |
|
| Cs | 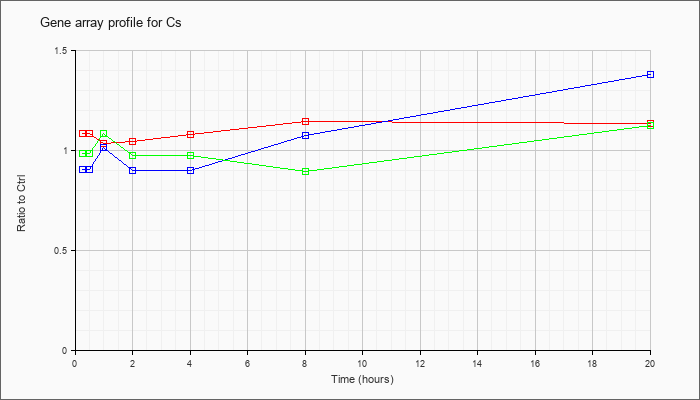 |
NM_026444 |
citrate synthase (Cs), nuclear gene encoding mitochondrial protein, mRNA [NM_026444] |
KLA | 1.09 |
1.07 |
1.04 |
1.05 |
1.08 |
1.15 |
1.14 |
| ATP | .91 |
.93 |
1.02 |
.90 |
.90 |
1.08 |
1.38 |
| KLA/ATP | .99 |
1.04 |
1.09 |
.98 |
.98 |
.90 |
1.13 |
|
| Csl | 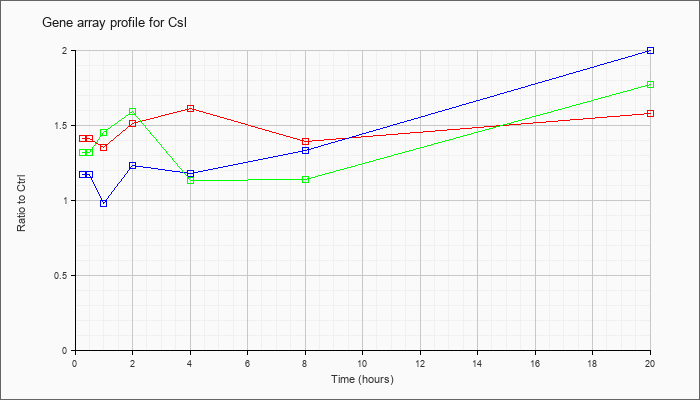 |
NM_027945 |
citrate synthase like (Csl), mRNA [NM_027945] |
KLA | 1.41 |
1.32 |
1.35 |
1.51 |
1.61 |
1.39 |
1.58 |
| ATP | 1.17 |
1.15 |
.98 |
1.23 |
1.18 |
1.33 |
2.00 |
| KLA/ATP | 1.32 |
1.53 |
1.45 |
1.59 |
1.13 |
1.14 |
1.77 |
|
| Got1 | 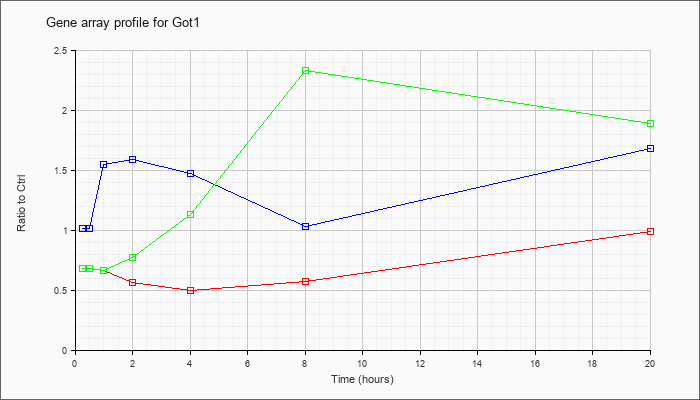 |
NM_010324 |
glutamate oxaloacetate transaminase 1, soluble (Got1), mRNA [NM_010324] |
KLA | .68 |
.70 |
.66 |
.56 |
.50 |
.57 |
.99 |
| ATP | 1.01 |
1.24 |
1.55 |
1.59 |
1.47 |
1.03 |
1.68 |
| KLA/ATP | .68 |
.71 |
.66 |
.77 |
1.13 |
2.33 |
1.89 |
|
| Got2 | 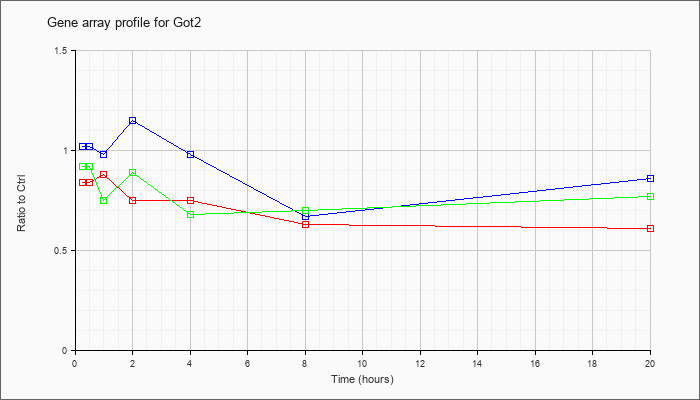 |
NM_010325 |
glutamate oxaloacetate transaminase 2, mitochondrial (Got2), nuclear gene encoding mitochondrial protein, mRNA [NM_010325] |
KLA | .84 |
.86 |
.88 |
.75 |
.75 |
.63 |
.61 |
| ATP | 1.02 |
1.12 |
.98 |
1.15 |
.98 |
.67 |
.86 |
| KLA/ATP | .92 |
.89 |
.75 |
.89 |
.68 |
.70 |
.77 |
|
| Gpt | 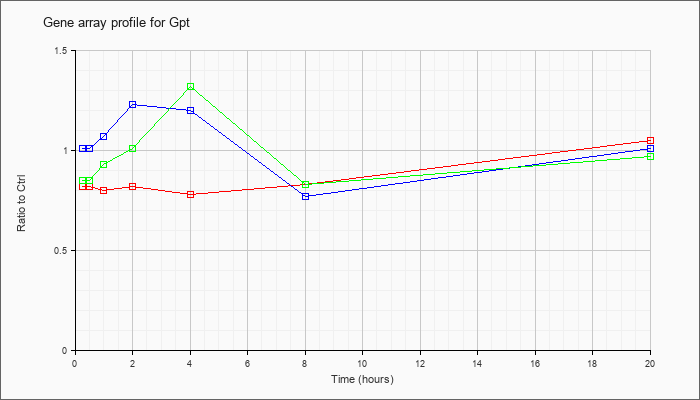 |
NM_182805 |
glutamic pyruvic transaminase, soluble (Gpt), mRNA [NM_182805] |
KLA | .82 |
.81 |
.80 |
.82 |
.78 |
.83 |
1.05 |
| ATP | 1.01 |
1.00 |
1.07 |
1.23 |
1.20 |
.77 |
1.01 |
| KLA/ATP | .85 |
.86 |
.93 |
1.01 |
1.32 |
.83 |
.97 |
|
| Gpt2 | 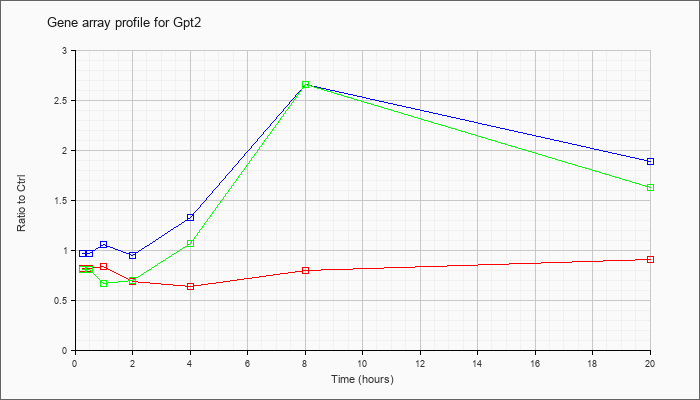 |
NM_173866 |
glutamic pyruvate transaminase (alanine aminotransferase) 2 (Gpt2), mRNA [NM_173866] |
KLA | .82 |
.87 |
.84 |
.69 |
.64 |
.80 |
.91 |
| ATP | .97 |
1.08 |
1.06 |
.95 |
1.33 |
2.66 |
1.89 |
| KLA/ATP | .81 |
.83 |
.67 |
.70 |
1.07 |
2.66 |
1.63 |
|
| Idh1 | 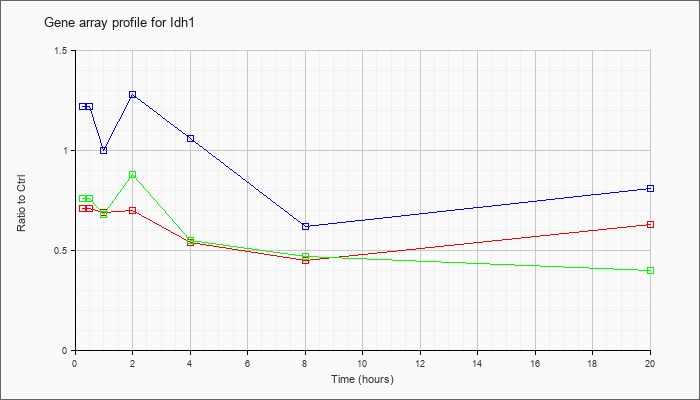 |
NM_010497 |
isocitrate dehydrogenase 1 (NADP+), soluble (Idh1), transcript variant 2, mRNA [NM_010497] |
KLA | .71 |
.74 |
.69 |
.70 |
.54 |
.45 |
.63 |
| ATP | 1.22 |
1.28 |
1.00 |
1.28 |
1.06 |
.62 |
.81 |
| KLA/ATP | .76 |
.81 |
.68 |
.88 |
.55 |
.47 |
.40 |
|
| Idh2 | 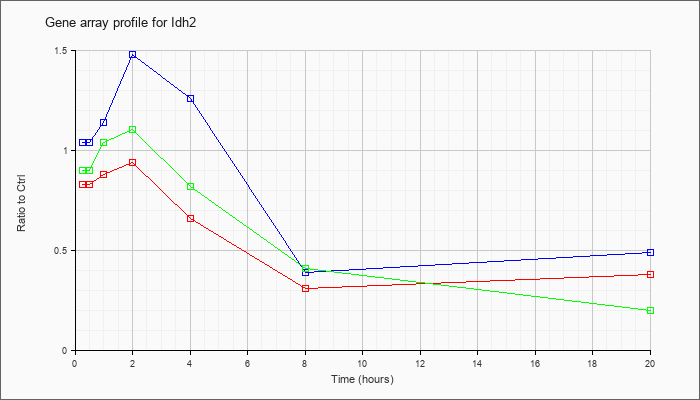 |
NM_173011 |
isocitrate dehydrogenase 2 (NADP+), mitochondrial (Idh2), nuclear gene encoding mitochondrial protein, mRNA [NM_173011] |
KLA | .83 |
.83 |
.88 |
.94 |
.66 |
.31 |
.38 |
| ATP | 1.04 |
1.09 |
1.14 |
1.48 |
1.26 |
.39 |
.49 |
| KLA/ATP | .90 |
.92 |
1.04 |
1.10 |
.82 |
.41 |
.20 |
|
| Idh3a | 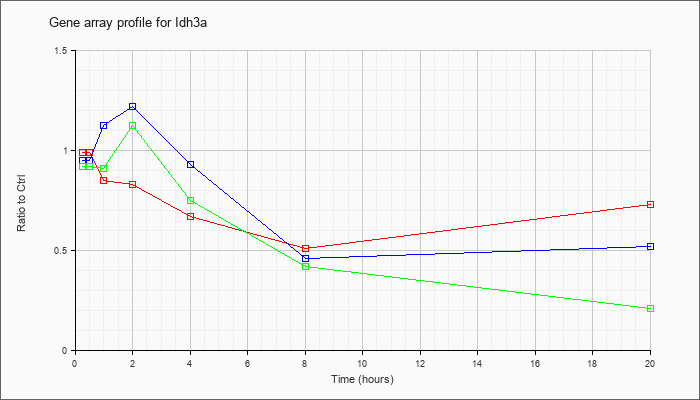 |
NM_029573 |
isocitrate dehydrogenase 3 (NAD+) alpha (Idh3a), nuclear gene encoding mitochondrial protein, mRNA [NM_029573] |
KLA | .99 |
1.06 |
.85 |
.83 |
.67 |
.51 |
.73 |
| ATP | .95 |
1.07 |
1.12 |
1.22 |
.93 |
.46 |
.52 |
| KLA/ATP | .92 |
1.00 |
.91 |
1.12 |
.75 |
.42 |
.21 |
|
| Idh3b | 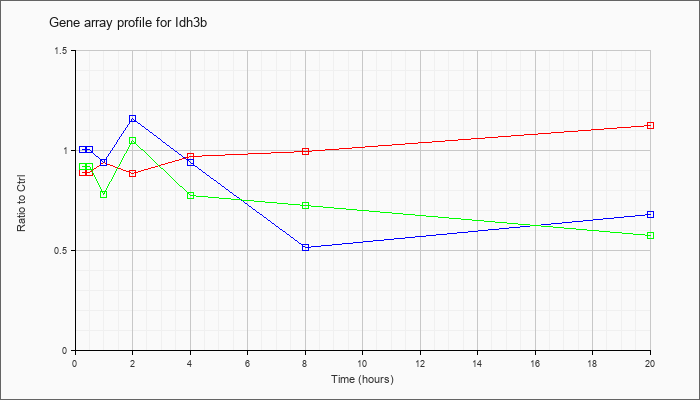 |
NM_130884 |
isocitrate dehydrogenase 3 (NAD+) beta (Idh3b), nuclear gene encoding mitochondrial protein, mRNA [NM_130884] |
KLA | .89 |
.94 |
.94 |
.89 |
.97 |
1.00 |
1.12 |
| ATP | 1.01 |
1.07 |
.94 |
1.16 |
.94 |
.52 |
.68 |
| KLA/ATP | .92 |
.94 |
.78 |
1.05 |
.78 |
.73 |
.58 |
|
| Idh3g | 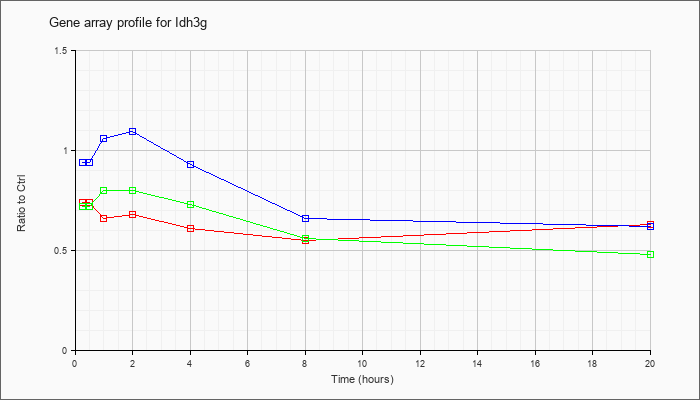 |
NM_008323 |
isocitrate dehydrogenase 3 (NAD+), gamma (Idh3g), nuclear gene encoding mitochondrial protein, mRNA [NM_008323] |
KLA | .74 |
.69 |
.66 |
.68 |
.61 |
.55 |
.63 |
| ATP | .94 |
.91 |
1.06 |
1.09 |
.93 |
.66 |
.62 |
| KLA/ATP | .72 |
.74 |
.80 |
.80 |
.73 |
.56 |
.48 |
|
| Nags | 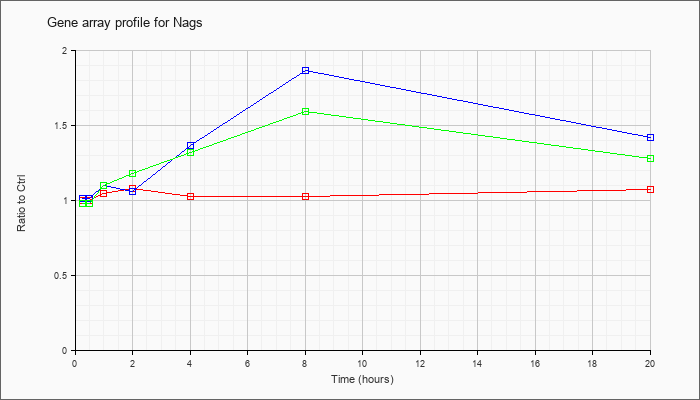 |
NM_145829 |
N-acetylglutamate synthase (Nags), transcript variant 1, mRNA [NM_145829] |
KLA | .99 |
.97 |
1.02 |
1.06 |
1.03 |
1.03 |
1.08 |
| ATP | .95 |
.98 |
1.07 |
1.12 |
1.17 |
1.38 |
1.26 |
| KLA/ATP | .97 |
1.05 |
1.10 |
1.42 |
1.31 |
1.52 |
1.27 |
|
| Nags |  |
NM_178053 |
N-acetylglutamate synthase (Nags), transcript variant 2, mRNA [NM_178053] |
KLA | 1.01 |
.98 |
1.07 |
1.10 |
1.02 |
1.02 |
1.06 |
| ATP | 1.07 |
1.06 |
1.12 |
1.00 |
1.56 |
2.35 |
1.57 |
| KLA/ATP | .98 |
1.02 |
1.09 |
.94 |
1.32 |
1.66 |
1.29 |
|