| Gene symbol |
Time course plot |
Accession |
Description |
Treatment |
15min |
30min |
1hr |
2hr |
4hr |
8hr |
20hr |
AK045366
| 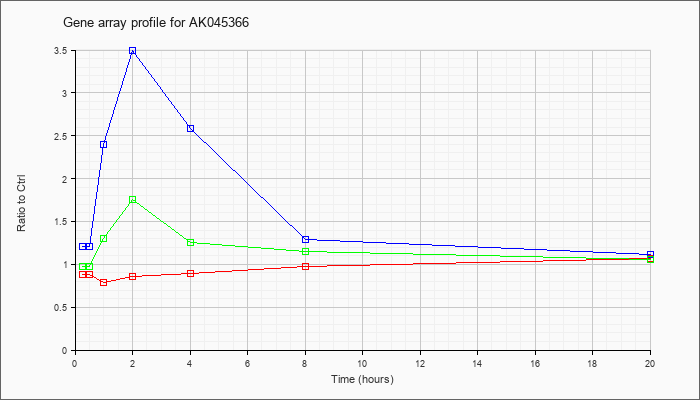 |
AK045366 |
adult male corpora quadrigemina cDNA, RIKEN full-length enriched library, clone:B230107A03 product:unclassifiable, full insert sequence [AK045366] |
KLA | .88 |
.80 |
.79 |
.86 |
.89 |
.97 |
1.07 |
| ATP | 1.21 |
1.67 |
2.40 |
3.50 |
2.58 |
1.29 |
1.12 |
| KLA/ATP | .97 |
.98 |
1.30 |
1.76 |
1.26 |
1.15 |
1.06 |
|
AK169742
| 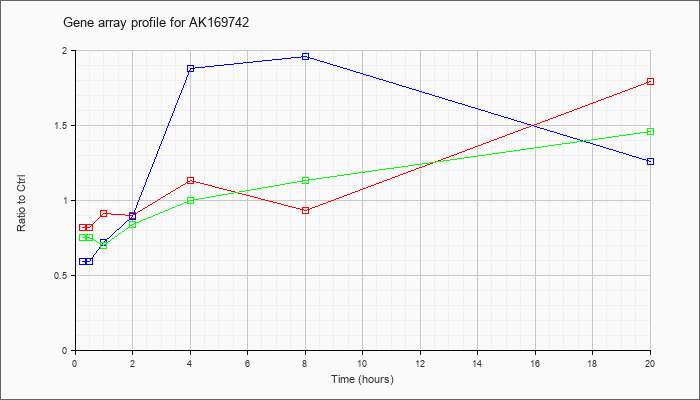 |
AK169742 |
2 days neonate thymus thymic cells cDNA, RIKEN full-length enriched library, clone:E430016M08 product:glyceraldehyde-3-phosphate dehydrogenase, full insert sequence. [AK169742] |
KLA | .82 |
.94 |
.91 |
.90 |
1.13 |
.93 |
1.79 |
| ATP | .59 |
.66 |
.72 |
.89 |
1.88 |
1.96 |
1.26 |
| KLA/ATP | .75 |
.69 |
.70 |
.84 |
1.00 |
1.13 |
1.46 |
|
| Akt1 | 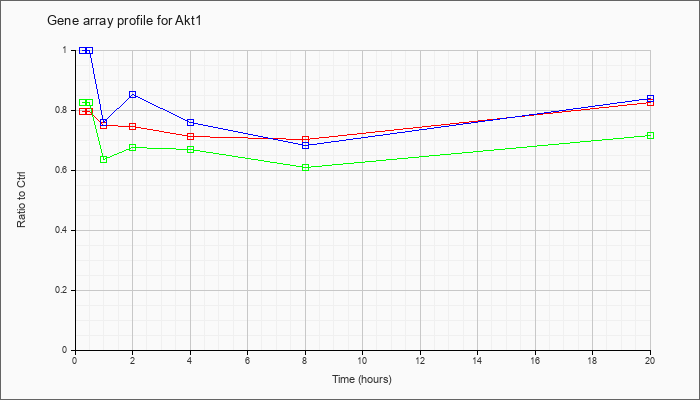 |
NM_009652 |
thymoma viral proto-oncogene 1 (Akt1), mRNA [NM_009652] |
KLA | .80 |
.80 |
.75 |
.75 |
.71 |
.70 |
.82 |
| ATP | 1.00 |
1.02 |
.76 |
.85 |
.76 |
.68 |
.84 |
| KLA/ATP | .82 |
.80 |
.64 |
.68 |
.67 |
.61 |
.72 |
|
| Akt2 | 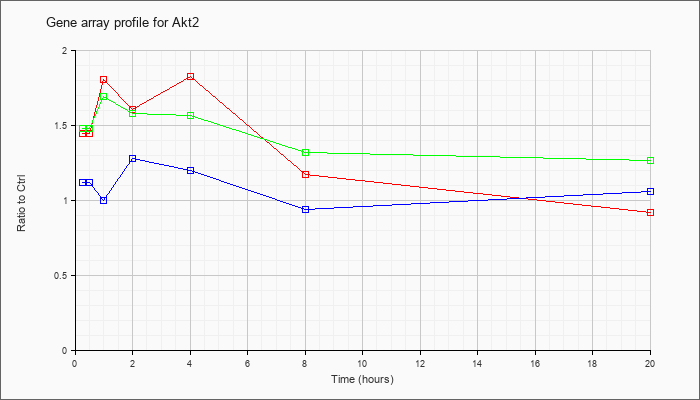 |
NM_007434 |
thymoma viral proto-oncogene 2 (Akt2), transcript variant 2, mRNA [NM_007434] |
KLA | 1.45 |
1.49 |
1.81 |
1.61 |
1.83 |
1.17 |
.92 |
| ATP | 1.12 |
1.12 |
1.00 |
1.28 |
1.20 |
.94 |
1.06 |
| KLA/ATP | 1.48 |
1.70 |
1.69 |
1.58 |
1.57 |
1.32 |
1.27 |
|
| Akt3 | 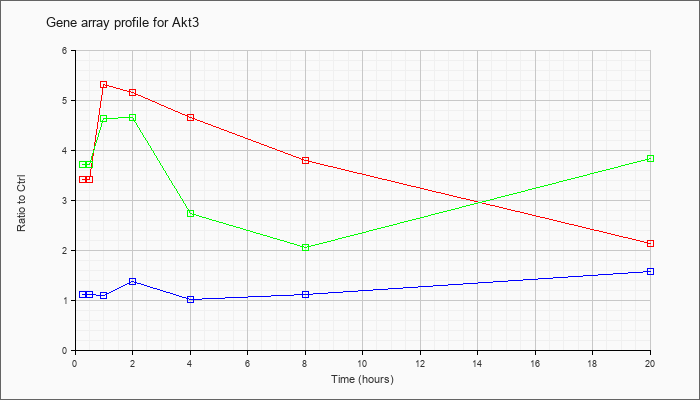 |
NM_011785 |
thymoma viral proto-oncogene 3 (Akt3), mRNA [NM_011785] |
KLA | 3.42 |
3.45 |
5.31 |
5.16 |
4.65 |
3.80 |
2.12 |
| ATP | 1.12 |
1.22 |
1.10 |
1.37 |
1.02 |
1.11 |
1.56 |
| KLA/ATP | 3.72 |
4.17 |
4.62 |
4.64 |
2.74 |
2.05 |
3.82 |
|
| Aldoa | 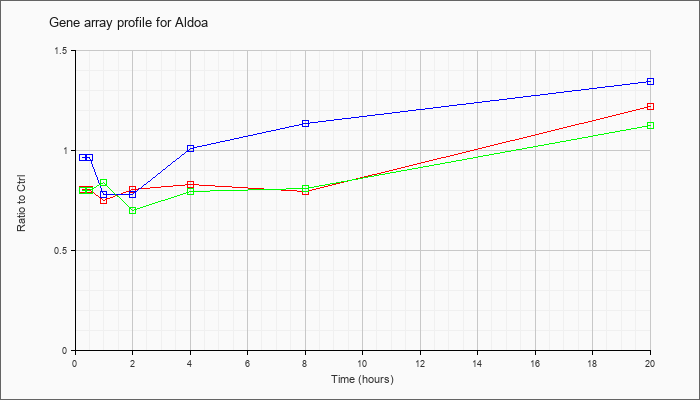 |
NM_007438 |
aldolase 1, A isoform (Aldoa), mRNA [NM_007438] |
KLA | .81 |
.83 |
.75 |
.81 |
.83 |
.80 |
1.22 |
| ATP | .97 |
.93 |
.78 |
.78 |
1.01 |
1.14 |
1.35 |
| KLA/ATP | .80 |
.86 |
.84 |
.70 |
.80 |
.81 |
1.12 |
|
| Angpt1 | 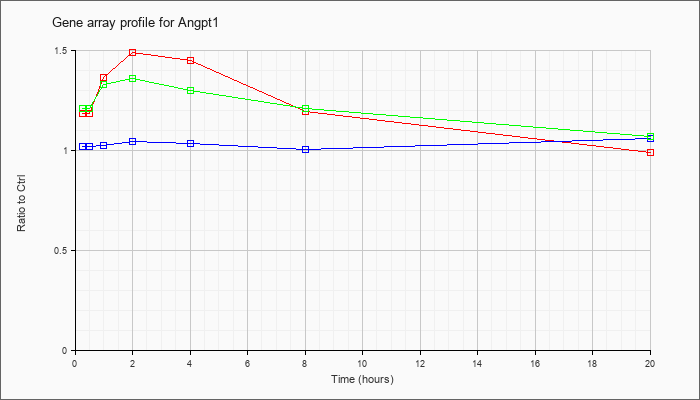 |
NM_009640 |
angiopoietin 1 (Angpt1), mRNA [NM_009640] |
KLA | 1.19 |
1.24 |
1.37 |
1.49 |
1.45 |
1.20 |
.99 |
| ATP | 1.02 |
1.05 |
1.03 |
1.05 |
1.04 |
1.01 |
1.06 |
| KLA/ATP | 1.21 |
1.23 |
1.33 |
1.36 |
1.30 |
1.21 |
1.07 |
|
| Angpt2 | 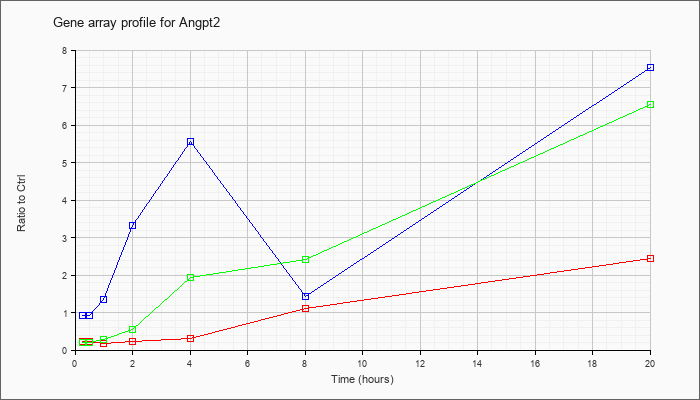 |
NM_007426 |
angiopoietin 2 (Angpt2), mRNA [NM_007426] |
KLA | .22 |
.22 |
.18 |
.22 |
.31 |
1.11 |
2.43 |
| ATP | .93 |
.92 |
1.36 |
3.33 |
5.55 |
1.44 |
7.53 |
| KLA/ATP | .20 |
.20 |
.28 |
.54 |
1.94 |
2.41 |
6.55 |
|
| Angpt4 | 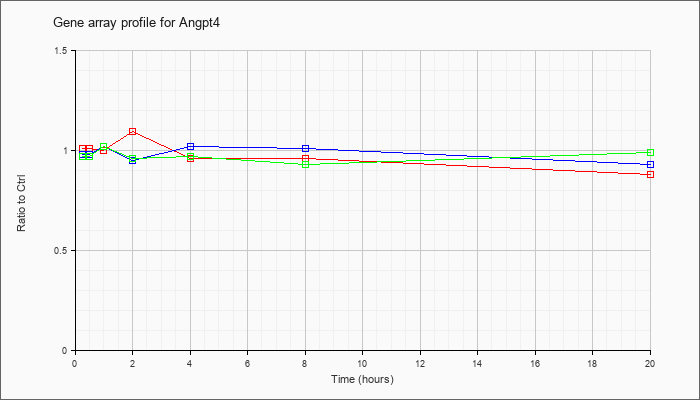 |
NM_009641 |
angiopoietin 4 (Angpt4), mRNA [NM_009641] |
KLA | 1.01 |
.99 |
1.00 |
1.09 |
.96 |
.96 |
.88 |
| ATP | .98 |
.97 |
1.02 |
.95 |
1.02 |
1.01 |
.93 |
| KLA/ATP | .97 |
1.01 |
1.02 |
.96 |
.97 |
.93 |
.99 |
|
| Arnt | 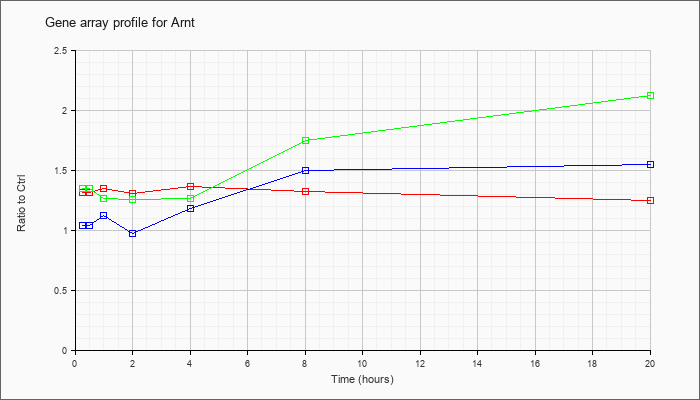 |
AK037762 |
16 days neonate thymus cDNA, RIKEN full-length enriched library, clone:A130047M16 product:aryl hydrocarbon receptor nuclear translocator, full insert sequence. [AK037762] |
KLA | 1.01 |
.87 |
.97 |
.92 |
1.13 |
.91 |
1.00 |
| ATP | 1.16 |
1.22 |
1.22 |
.84 |
1.57 |
1.19 |
1.05 |
| KLA/ATP | 1.06 |
1.06 |
1.08 |
.95 |
1.40 |
1.18 |
1.06 |
|
| Arnt |  |
NM_001037737 |
aryl hydrocarbon receptor nuclear translocator (Arnt), transcript variant 1, mRNA [NM_001037737] |
KLA | 1.23 |
1.27 |
1.32 |
1.23 |
1.34 |
1.33 |
1.30 |
| ATP | 1.02 |
1.09 |
.96 |
.90 |
.99 |
1.33 |
1.38 |
| KLA/ATP | 1.30 |
1.28 |
1.05 |
1.12 |
1.07 |
1.64 |
1.89 |
|
| Arnt |  |
U14333 |
aromatic hydrocarbon receptor nuclear translocator Arnt (ARNT) mRNA, complete cds. [U14333] |
KLA | 1.78 |
1.78 |
1.78 |
1.85 |
1.64 |
1.71 |
1.40 |
| ATP | .96 |
1.05 |
1.37 |
1.24 |
1.19 |
2.13 |
2.37 |
| KLA/ATP | 1.72 |
1.87 |
1.87 |
1.83 |
1.53 |
2.53 |
3.65 |
|
| Bcl2 | 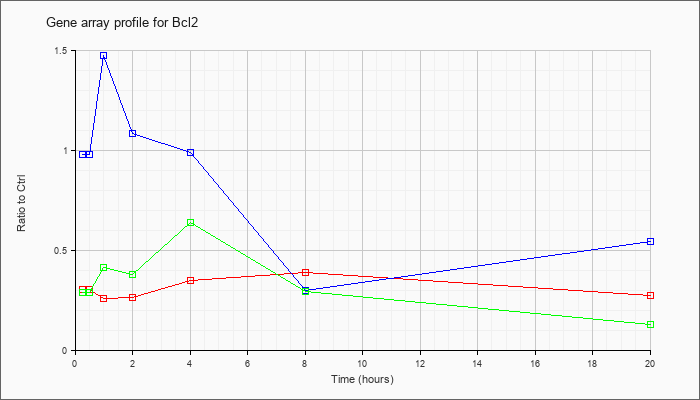 |
NM_009741 |
B-cell leukemia/lymphoma 2 (Bcl2), transcript variant 1, mRNA [NM_009741] |
KLA | .30 |
.26 |
.25 |
.25 |
.34 |
.38 |
.26 |
| ATP | .97 |
.89 |
1.50 |
1.04 |
.99 |
.29 |
.52 |
| KLA/ATP | .28 |
.26 |
.40 |
.33 |
.62 |
.29 |
.12 |
|
| Bcl2 |  |
NM_177410 |
B-cell leukemia/lymphoma 2 (Bcl2), transcript variant 2, mRNA [NM_177410] |
KLA | .33 |
.33 |
.37 |
.42 |
.49 |
.46 |
.39 |
| ATP | 1.07 |
1.12 |
1.22 |
1.56 |
1.00 |
.38 |
.76 |
| KLA/ATP | .35 |
.34 |
.54 |
.89 |
.76 |
.36 |
.23 |
|
| Camk2a | 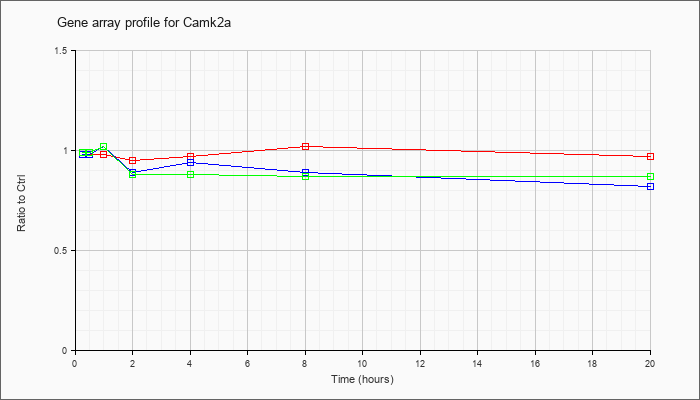 |
NM_177407 |
calcium/calmodulin-dependent protein kinase II alpha (Camk2a), transcript variant 2, mRNA [NM_177407] |
KLA | .98 |
.97 |
.98 |
.95 |
.97 |
1.02 |
.97 |
| ATP | .98 |
1.11 |
1.02 |
.89 |
.94 |
.89 |
.82 |
| KLA/ATP | .99 |
.95 |
1.02 |
.88 |
.88 |
.87 |
.87 |
|
| Camk2b | 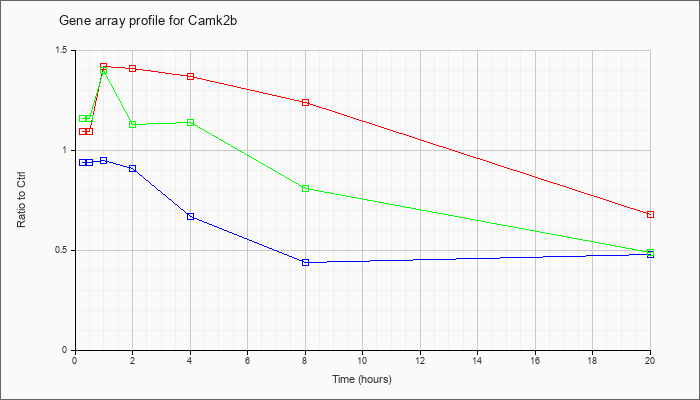 |
NM_007595 |
calcium/calmodulin-dependent protein kinase II, beta (Camk2b), mRNA [NM_007595] |
KLA | 1.09 |
1.28 |
1.42 |
1.41 |
1.37 |
1.24 |
.68 |
| ATP | .94 |
.87 |
.95 |
.91 |
.67 |
.44 |
.48 |
| KLA/ATP | 1.16 |
1.24 |
1.40 |
1.13 |
1.14 |
.81 |
.49 |
|
| Camk2d | 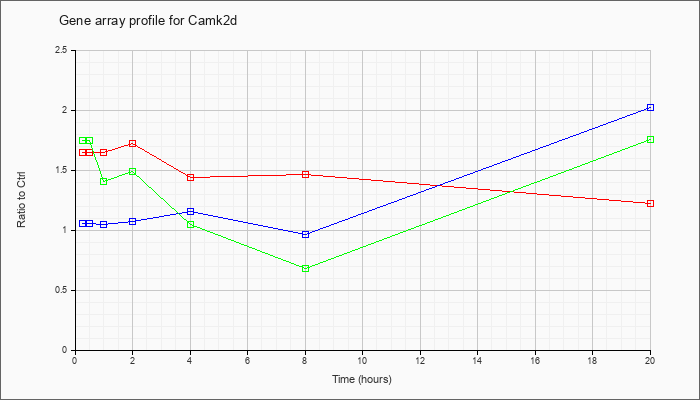 |
AK032524 |
adult male olfactory brain cDNA, RIKEN full-length enriched library, clone:6430579P05 product:calcium/calmodulin-dependent protein kinase II, delta, full insert sequence. [AK032524] |
KLA | 1.31 |
1.45 |
1.42 |
1.21 |
1.34 |
1.18 |
1.16 |
| ATP | 1.04 |
1.29 |
1.39 |
1.24 |
.94 |
.91 |
1.22 |
| KLA/ATP | 1.46 |
1.64 |
1.42 |
1.24 |
1.00 |
.93 |
1.10 |
|
| Camk2d |  |
NM_001025439 |
calcium/calmodulin-dependent protein kinase II, delta (Camk2d), transcript variant 1, mRNA [NM_001025439] |
KLA | 1.76 |
1.75 |
1.72 |
1.89 |
1.47 |
1.55 |
1.24 |
| ATP | 1.06 |
1.06 |
.93 |
1.01 |
1.22 |
.98 |
2.28 |
| KLA/ATP | 1.84 |
1.78 |
1.40 |
1.57 |
1.07 |
.60 |
1.97 |
|
| Camk2g | 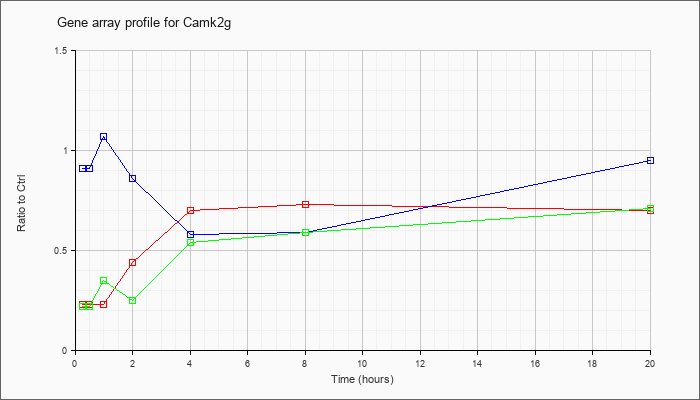 |
NM_178597 |
calcium/calmodulin-dependent protein kinase II gamma (Camk2g), transcript variant 1, mRNA [NM_178597] |
KLA | .23 |
.24 |
.23 |
.44 |
.70 |
.73 |
.70 |
| ATP | .91 |
.85 |
1.07 |
.86 |
.58 |
.59 |
.95 |
| KLA/ATP | .22 |
.20 |
.35 |
.25 |
.54 |
.59 |
.71 |
|
| Cdkn1a | 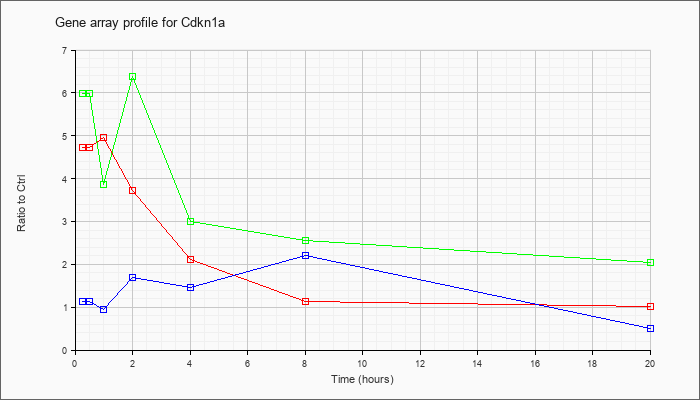 |
NM_007669 |
cyclin-dependent kinase inhibitor 1A (P21) (Cdkn1a), transcript variant 1, mRNA [NM_007669] |
KLA | 4.72 |
4.90 |
4.96 |
3.73 |
2.11 |
1.14 |
1.02 |
| ATP | 1.14 |
1.52 |
.95 |
1.69 |
1.47 |
2.21 |
.50 |
| KLA/ATP | 5.99 |
7.08 |
3.86 |
6.37 |
3.00 |
2.55 |
2.05 |
|
| Cdkn1b | 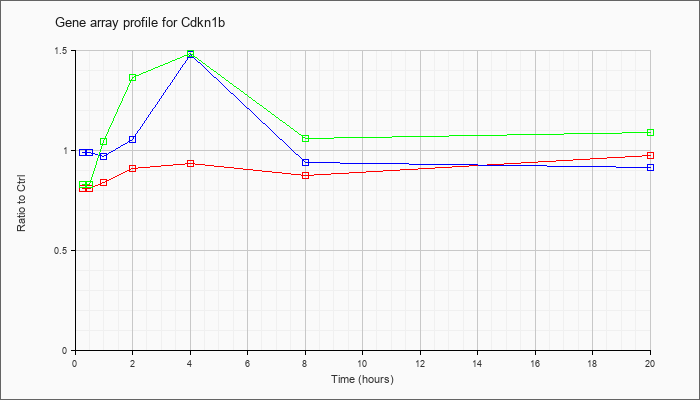 |
NM_009875 |
cyclin-dependent kinase inhibitor 1B (Cdkn1b), mRNA [NM_009875] |
KLA | .81 |
.84 |
.84 |
.91 |
.93 |
.88 |
.97 |
| ATP | .99 |
1.09 |
.97 |
1.06 |
1.48 |
.94 |
.91 |
| KLA/ATP | .83 |
.91 |
1.04 |
1.36 |
1.48 |
1.06 |
1.09 |
|
| Crebbp | 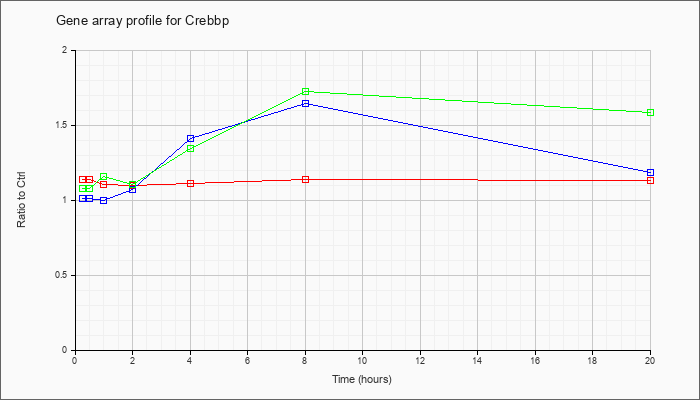 |
AK048818 |
0 day neonate cerebellum cDNA, RIKEN full-length enriched library, clone:C230072N21 product:hypothetical protein, full insert sequence [AK048818] |
KLA | 1.44 |
1.19 |
1.26 |
1.25 |
1.25 |
1.36 |
1.24 |
| ATP | .98 |
.90 |
1.04 |
1.11 |
1.95 |
2.72 |
1.51 |
| KLA/ATP | 1.17 |
1.27 |
1.45 |
1.28 |
1.98 |
2.80 |
2.52 |
|
| Crebbp |  |
NM_001025432 |
CREB binding protein (Crebbp), mRNA [NM_001025432] |
KLA | .97 |
1.01 |
.98 |
1.06 |
1.05 |
.99 |
1.07 |
| ATP | 1.07 |
1.05 |
.96 |
1.01 |
1.15 |
1.14 |
1.00 |
| KLA/ATP | 1.01 |
.97 |
.97 |
1.00 |
1.03 |
1.23 |
1.13 |
|
| Crebbp |  |
S66385 |
gb|CREB-binding protein [mice, brain, mRNA Partial, 7326 nt]. [S66385] |
KLA | 1.00 |
.98 |
1.08 |
.98 |
1.04 |
1.07 |
1.08 |
| ATP | .99 |
.98 |
.99 |
1.09 |
1.13 |
1.08 |
1.05 |
| KLA/ATP | 1.06 |
1.00 |
1.05 |
1.03 |
1.02 |
1.14 |
1.11 |
|
| Cul2 | 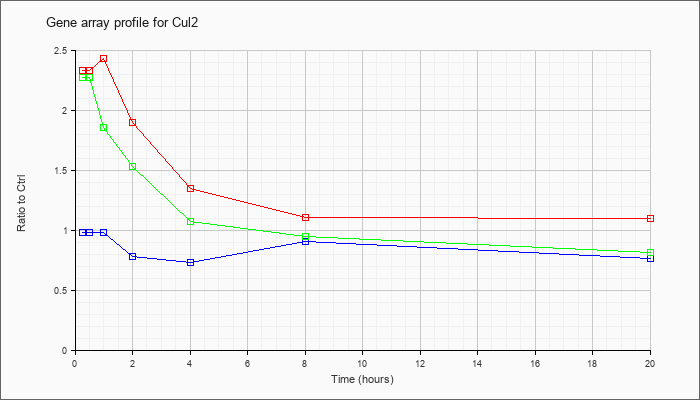 |
NM_029402 |
cullin 2 (Cul2), mRNA [NM_029402] |
KLA | 2.33 |
2.37 |
2.43 |
1.90 |
1.34 |
1.10 |
1.09 |
| ATP | .98 |
.92 |
.98 |
.78 |
.73 |
.90 |
.76 |
| KLA/ATP | 2.27 |
2.19 |
1.86 |
1.53 |
1.07 |
.95 |
.81 |
|
| Cybb | 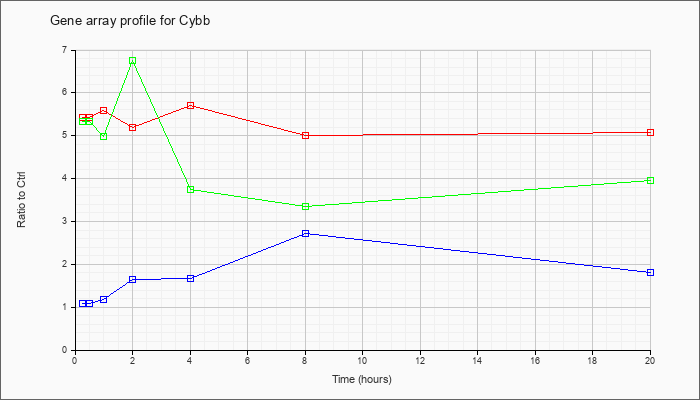 |
NM_007807 |
cytochrome b-245, beta polypeptide (Cybb), mRNA [NM_007807] |
KLA | 5.43 |
5.43 |
5.59 |
5.19 |
5.71 |
5.00 |
5.08 |
| ATP | 1.08 |
1.18 |
1.19 |
1.64 |
1.68 |
2.73 |
1.82 |
| KLA/ATP | 5.34 |
5.62 |
4.98 |
6.75 |
3.74 |
3.36 |
3.96 |
|
| Edn1 | 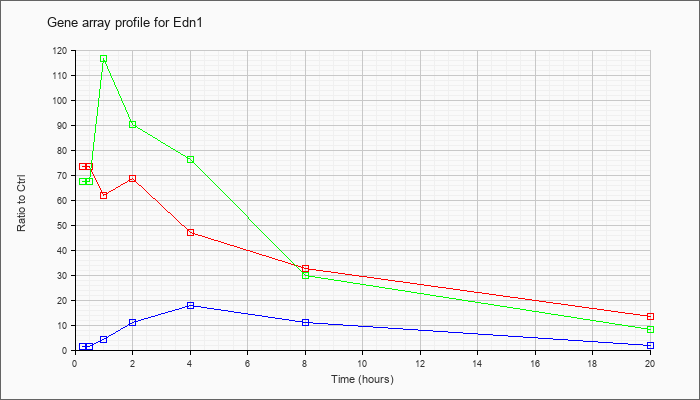 |
NM_010104 |
endothelin 1 (Edn1), mRNA [NM_010104] |
KLA | 73.36 |
71.33 |
61.92 |
68.69 |
46.94 |
32.60 |
13.54 |
| ATP | 1.23 |
1.89 |
4.18 |
10.86 |
17.85 |
11.03 |
1.91 |
| KLA/ATP | 67.27 |
76.78 |
116.70 |
90.29 |
76.38 |
29.71 |
8.04 |
|
| Egf | 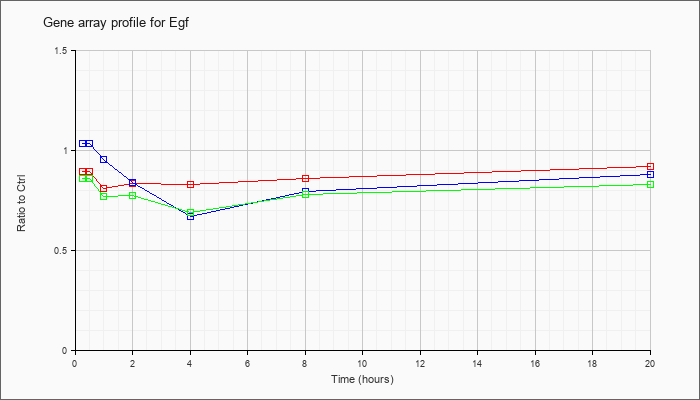 |
NM_010113 |
epidermal growth factor (Egf), mRNA [NM_010113] |
KLA | .89 |
.90 |
.81 |
.83 |
.83 |
.86 |
.92 |
| ATP | 1.03 |
1.03 |
.95 |
.84 |
.67 |
.79 |
.88 |
| KLA/ATP | .86 |
.86 |
.77 |
.77 |
.69 |
.78 |
.83 |
|
| Egfr | 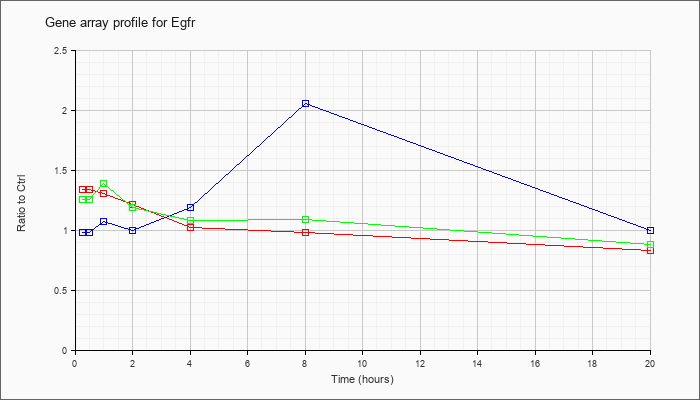 |
NM_007912 |
epidermal growth factor receptor (Egfr), transcript variant 2, mRNA [NM_007912] |
KLA | 1.10 |
1.07 |
1.14 |
1.11 |
1.08 |
1.03 |
.90 |
| ATP | .99 |
1.00 |
1.04 |
.98 |
.97 |
1.15 |
1.14 |
| KLA/ATP | 1.07 |
1.10 |
1.16 |
1.08 |
1.01 |
.97 |
.94 |
|
| Egfr |  |
NM_207655 |
epidermal growth factor receptor (Egfr), transcript variant 1, mRNA [NM_207655] |
KLA | 1.57 |
1.44 |
1.47 |
1.31 |
.96 |
.92 |
.76 |
| ATP | .97 |
.93 |
1.10 |
1.00 |
1.41 |
2.97 |
.86 |
| KLA/ATP | 1.44 |
1.50 |
1.61 |
1.30 |
1.14 |
1.21 |
.82 |
|
| Egln1 | 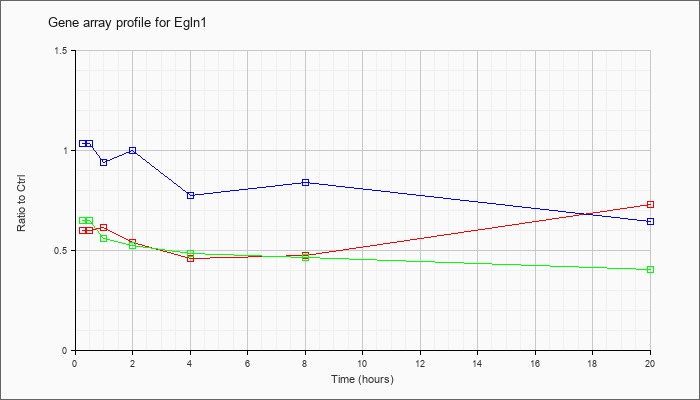 |
NM_053207 |
EGL nine homolog 1 (C. elegans) (Egln1), mRNA [NM_053207] |
KLA | .60 |
.60 |
.62 |
.54 |
.46 |
.48 |
.73 |
| ATP | 1.04 |
1.10 |
.94 |
1.00 |
.78 |
.84 |
.65 |
| KLA/ATP | .65 |
.64 |
.56 |
.53 |
.49 |
.47 |
.41 |
|
| Egln2 | 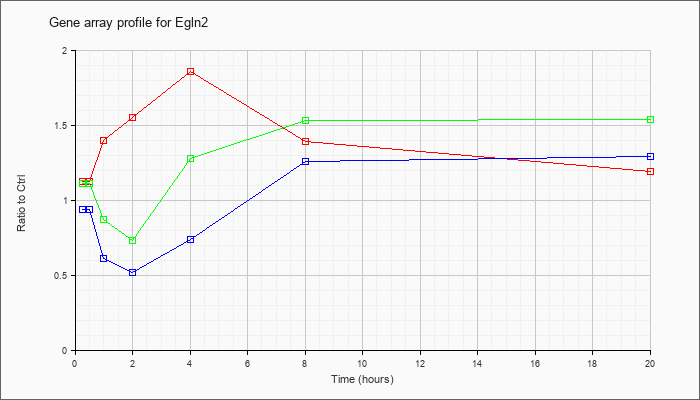 |
NM_053208 |
EGL nine homolog 2 (C. elegans) (Egln2), mRNA [NM_053208] |
KLA | 1.12 |
1.22 |
1.40 |
1.55 |
1.86 |
1.39 |
1.19 |
| ATP | .94 |
.82 |
.61 |
.52 |
.74 |
1.26 |
1.29 |
| KLA/ATP | 1.11 |
1.04 |
.87 |
.73 |
1.28 |
1.53 |
1.54 |
|
| Egln3 | 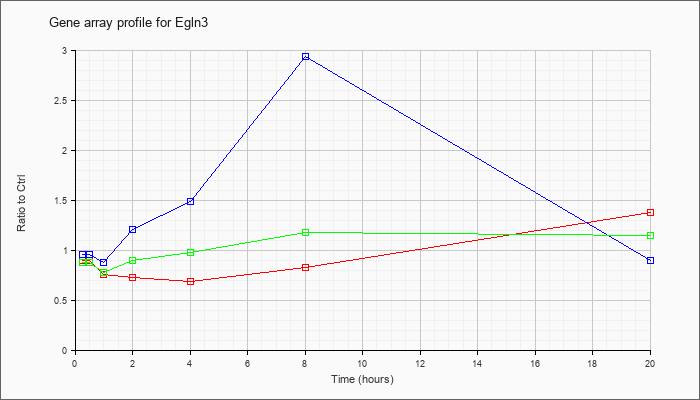 |
NM_028133 |
EGL nine homolog 3 (C. elegans) (Egln3), mRNA [NM_028133] |
KLA | .90 |
.90 |
.76 |
.73 |
.69 |
.83 |
1.38 |
| ATP | .96 |
1.03 |
.88 |
1.21 |
1.49 |
2.94 |
.90 |
| KLA/ATP | .88 |
.96 |
.78 |
.90 |
.98 |
1.18 |
1.15 |
|
| Eif4e | 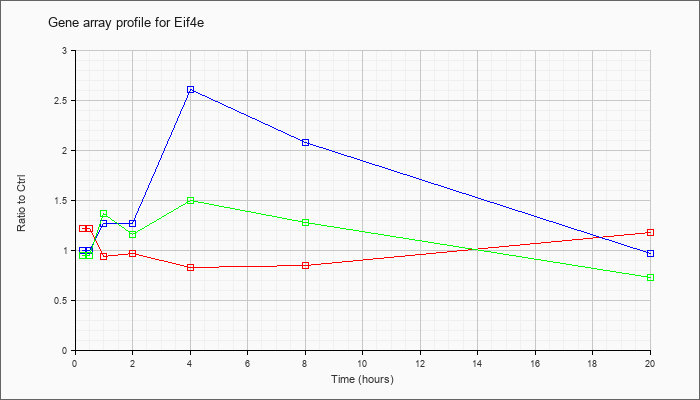 |
NM_007917 |
eukaryotic translation initiation factor 4E (Eif4e), mRNA [NM_007917] |
KLA | 1.22 |
1.06 |
.94 |
.97 |
.83 |
.85 |
1.18 |
| ATP | 1.00 |
1.01 |
1.27 |
1.27 |
2.61 |
2.08 |
.97 |
| KLA/ATP | .95 |
1.04 |
1.37 |
1.16 |
1.50 |
1.28 |
.73 |
|
| Eif4e2 | 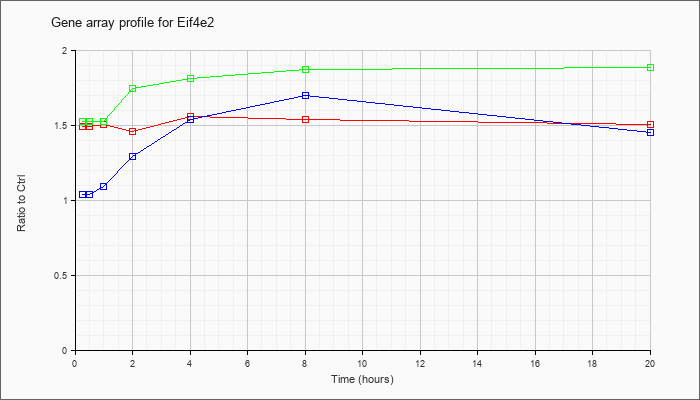 |
NM_001039169 |
eukaryotic translation initiation factor 4E member 2 (Eif4e2), transcript variant 2, mRNA [NM_001039169] |
KLA | 1.53 |
1.51 |
1.54 |
1.52 |
1.55 |
1.49 |
1.55 |
| ATP | 1.02 |
1.08 |
1.12 |
1.45 |
1.61 |
1.51 |
1.15 |
| KLA/ATP | 1.56 |
1.60 |
1.59 |
1.90 |
1.90 |
1.75 |
1.68 |
|
| Eif4e2 |  |
NM_023314 |
eukaryotic translation initiation factor 4E member 2 (Eif4e2), transcript variant 3, mRNA [NM_023314] |
KLA | 1.42 |
1.40 |
1.43 |
1.35 |
1.58 |
1.65 |
1.42 |
| ATP | 1.06 |
1.06 |
1.04 |
.98 |
1.41 |
2.07 |
2.04 |
| KLA/ATP | 1.46 |
1.48 |
1.40 |
1.45 |
1.64 |
2.12 |
2.30 |
|
Eif4ebp1
| 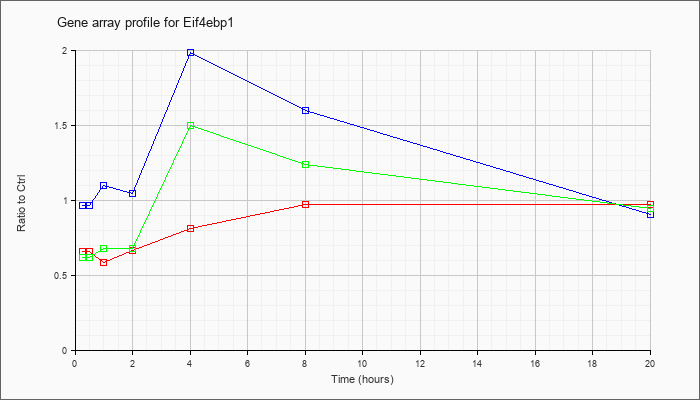 |
NM_007918 |
eukaryotic translation initiation factor 4E binding protein 1 (Eif4ebp1), mRNA [NM_007918] |
KLA | .66 |
.64 |
.59 |
.67 |
.81 |
.97 |
.97 |
| ATP | .97 |
.94 |
1.10 |
1.05 |
1.99 |
1.60 |
.91 |
| KLA/ATP | .62 |
.61 |
.68 |
.68 |
1.50 |
1.24 |
.95 |
|
| Eno1 | 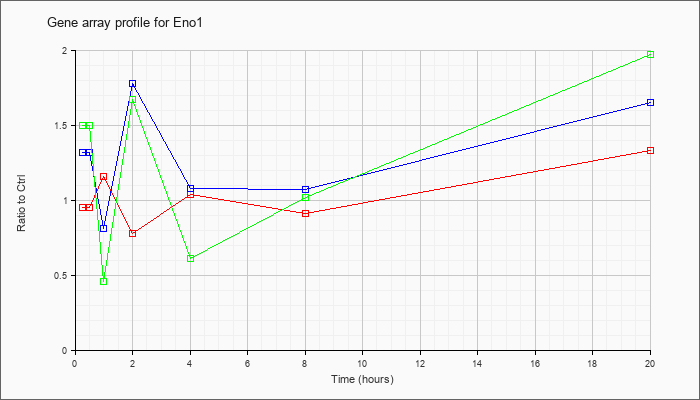 |
NM_023119 |
enolase 1, alpha non-neuron (Eno1), mRNA [NM_023119] |
KLA | .95 |
1.06 |
1.16 |
.78 |
1.04 |
.91 |
1.33 |
| ATP | 1.32 |
1.63 |
.81 |
1.78 |
1.08 |
1.07 |
1.65 |
| KLA/ATP | 1.50 |
1.43 |
.46 |
1.67 |
.61 |
1.02 |
1.97 |
|
| Eno2 | 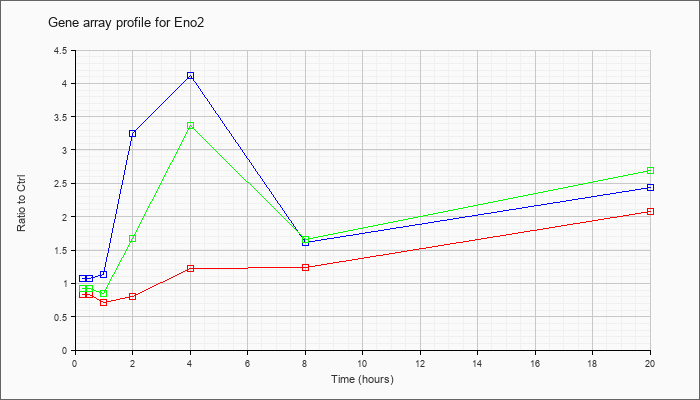 |
NM_013509 |
enolase 2, gamma neuronal (Eno2), mRNA [NM_013509] |
KLA | .84 |
.85 |
.72 |
.81 |
1.23 |
1.24 |
2.08 |
| ATP | 1.07 |
1.13 |
1.14 |
3.24 |
4.11 |
1.61 |
2.43 |
| KLA/ATP | .93 |
.89 |
.85 |
1.68 |
3.37 |
1.66 |
2.69 |
|
| Eno3 | 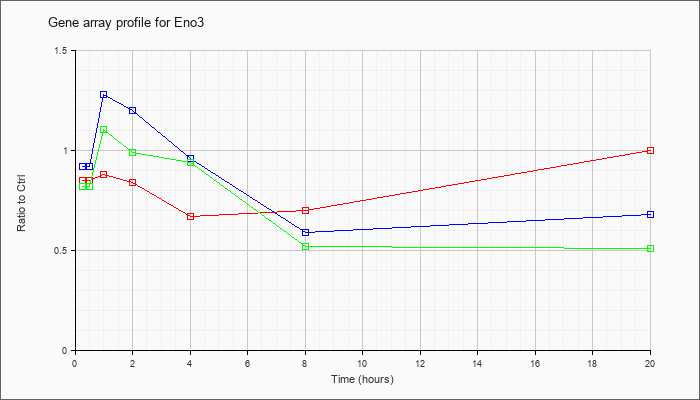 |
NM_007933 |
enolase 3, beta muscle (Eno3), mRNA [NM_007933] |
KLA | .85 |
.86 |
.88 |
.84 |
.67 |
.70 |
1.00 |
| ATP | .92 |
.90 |
1.28 |
1.20 |
.96 |
.59 |
.68 |
| KLA/ATP | .82 |
.90 |
1.10 |
.99 |
.94 |
.52 |
.51 |
|
| Ep300 | 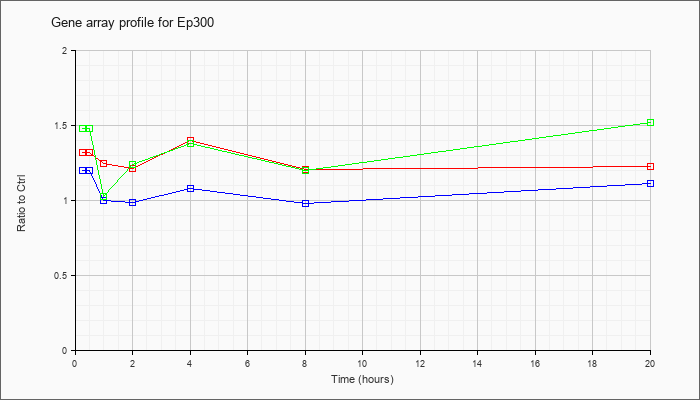 |
AK042627 |
7 days neonate cerebellum cDNA, RIKEN full-length enriched library, clone:A730011L11 product:P300 TRANSCRIPTION COACTIVATOR (FRAGMENT) homolog [Mus sp], full insert sequence. [AK042627] |
KLA | 1.02 |
.95 |
.92 |
1.04 |
1.01 |
1.00 |
1.05 |
| ATP | 1.06 |
.89 |
.88 |
.89 |
1.00 |
.90 |
1.07 |
| KLA/ATP | 1.10 |
1.02 |
.87 |
.91 |
1.04 |
.97 |
.98 |
|
| Ep300 |  |
NM_177821 |
E1A binding protein p300 (Ep300), mRNA [NM_177821] |
KLA | 1.61 |
1.26 |
1.57 |
1.38 |
1.78 |
1.41 |
1.40 |
| ATP | 1.34 |
1.60 |
1.11 |
1.08 |
1.16 |
1.05 |
1.15 |
| KLA/ATP | 1.86 |
1.94 |
1.18 |
1.56 |
1.71 |
1.42 |
2.05 |
|
| Epo | 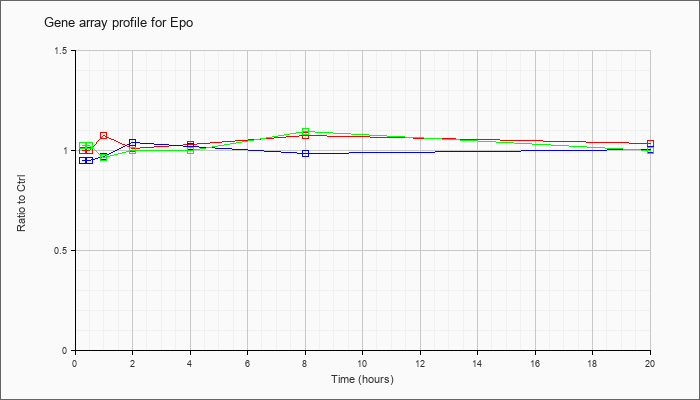 |
NM_007942 |
erythropoietin (Epo), mRNA [NM_007942] |
KLA | 1.00 |
1.05 |
1.08 |
1.01 |
1.03 |
1.08 |
1.04 |
| ATP | .95 |
.97 |
.97 |
1.04 |
1.02 |
.99 |
1.01 |
| KLA/ATP | 1.03 |
1.01 |
.97 |
1.00 |
1.00 |
1.10 |
1.00 |
|
| Erbb2 | 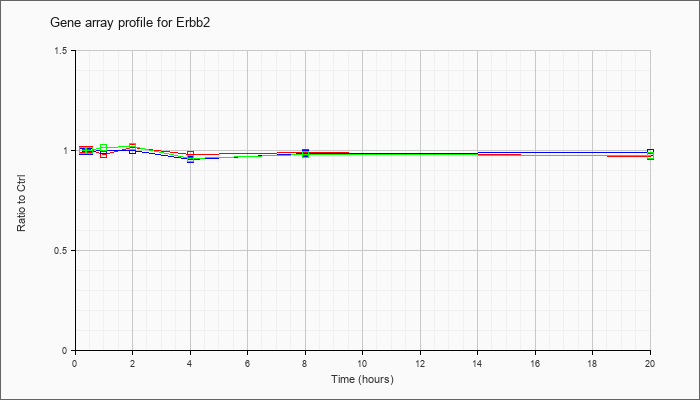 |
NM_001003817 |
v-erb-b2 erythroblastic leukemia viral oncogene homolog 2, neuro/glioblastoma derived oncogene homolog (avian) (Erbb2), mRNA [NM_001003817] |
KLA | 1.01 |
1.00 |
.98 |
1.01 |
.98 |
.99 |
.97 |
| ATP | .99 |
1.00 |
1.00 |
1.00 |
.96 |
.98 |
.99 |
| KLA/ATP | 1.00 |
1.01 |
1.01 |
1.02 |
.96 |
.98 |
.97 |
|
| Flt1 | 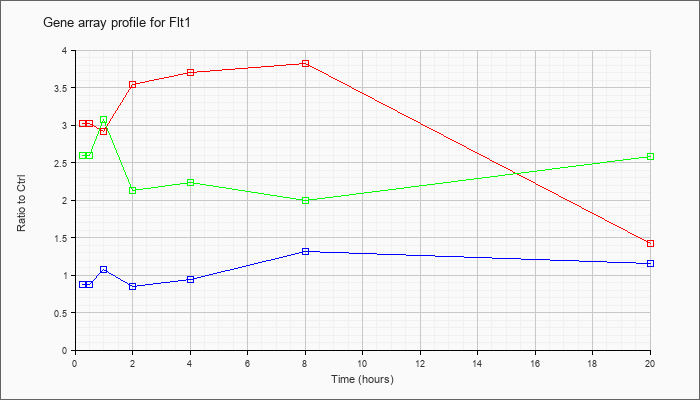 |
NM_010228 |
FMS-like tyrosine kinase 1 (Flt1), mRNA [NM_010228] |
KLA | 3.02 |
2.74 |
2.92 |
3.54 |
3.71 |
3.82 |
1.43 |
| ATP | .88 |
.85 |
1.07 |
.85 |
.94 |
1.31 |
1.16 |
| KLA/ATP | 2.60 |
2.99 |
3.08 |
2.13 |
2.23 |
2.00 |
2.59 |
|
| Frap1 | 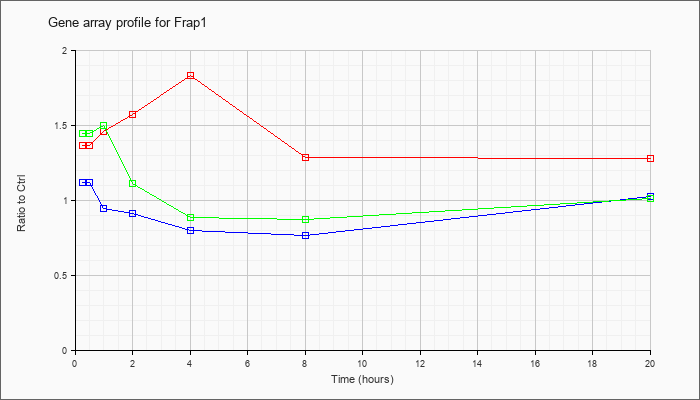 |
NM_020009 |
FK506 binding protein 12-rapamycin associated protein 1 (Frap1), mRNA [NM_020009] |
KLA | 1.37 |
1.32 |
1.46 |
1.57 |
1.83 |
1.29 |
1.28 |
| ATP | 1.12 |
1.07 |
.95 |
.91 |
.80 |
.77 |
1.03 |
| KLA/ATP | 1.45 |
1.51 |
1.50 |
1.11 |
.89 |
.87 |
1.01 |
|
| Gapdh | 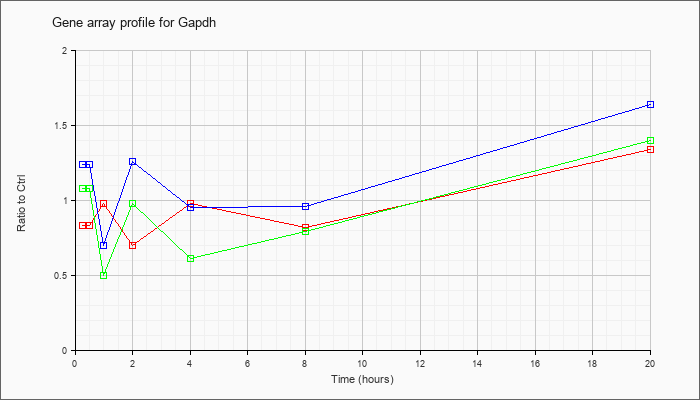 |
NM_008084 |
glyceraldehyde-3-phosphate dehydrogenase (Gapdh), mRNA [NM_008084] |
KLA | .83 |
.87 |
.98 |
.70 |
.98 |
.82 |
1.34 |
| ATP | 1.24 |
1.33 |
.70 |
1.26 |
.95 |
.96 |
1.64 |
| KLA/ATP | 1.08 |
1.18 |
.50 |
.98 |
.61 |
.79 |
1.40 |
|
| Hif1a | 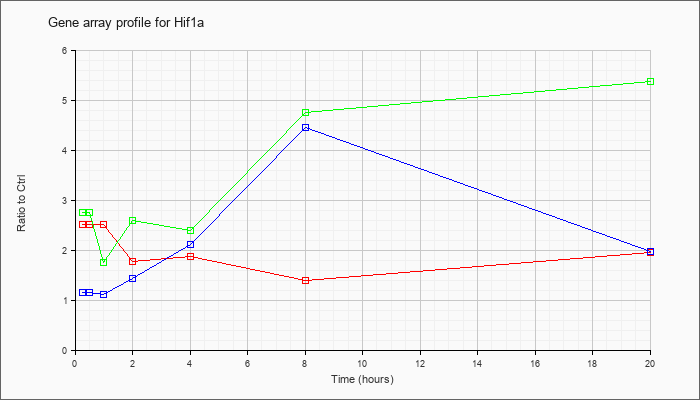 |
NM_010431 |
hypoxia inducible factor 1, alpha subunit (Hif1a), mRNA [NM_010431] |
KLA | 2.52 |
2.50 |
2.52 |
1.78 |
1.87 |
1.39 |
1.96 |
| ATP | 1.16 |
1.37 |
1.11 |
1.44 |
2.11 |
4.46 |
1.97 |
| KLA/ATP | 2.76 |
2.92 |
1.75 |
2.60 |
2.39 |
4.75 |
5.37 |
|
| Hk1 | 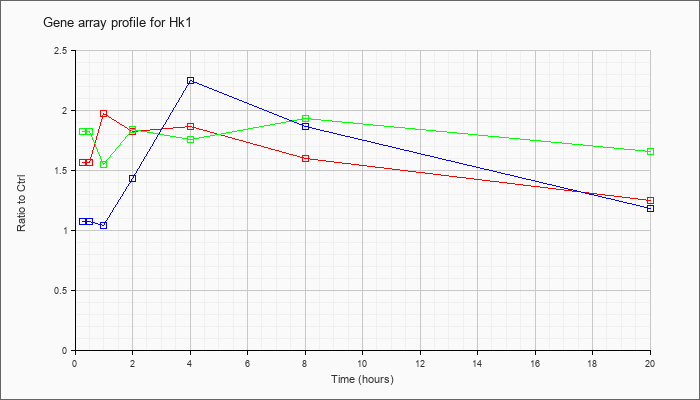 |
J05277 |
gb|Mouse hexokinase mRNA, complete cds [J05277] |
KLA | 1.49 |
1.45 |
1.66 |
1.78 |
1.51 |
1.29 |
.99 |
| ATP | 1.11 |
1.05 |
1.15 |
1.20 |
2.31 |
2.17 |
1.09 |
| KLA/ATP | 1.63 |
1.51 |
1.70 |
1.47 |
1.94 |
1.92 |
1.40 |
|
| Hk1 |  |
NM_010438 |
hexokinase 1 (Hk1), nuclear gene encoding mitochondrial protein, mRNA [NM_010438] |
KLA | 1.58 |
1.75 |
2.05 |
1.84 |
1.96 |
1.67 |
1.31 |
| ATP | 1.06 |
1.25 |
1.01 |
1.49 |
2.23 |
1.79 |
1.20 |
| KLA/ATP | 1.87 |
1.96 |
1.51 |
1.93 |
1.71 |
1.94 |
1.72 |
|
| Hk2 | 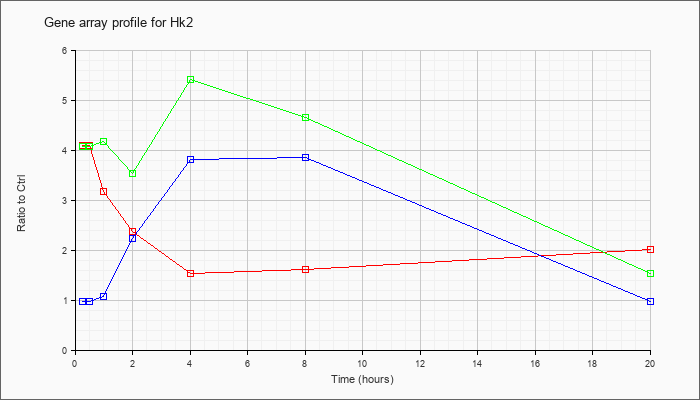 |
NM_013820 |
hexokinase 2 (Hk2), mRNA [NM_013820] |
KLA | 4.09 |
4.09 |
3.17 |
2.37 |
1.53 |
1.62 |
2.02 |
| ATP | .98 |
.92 |
1.08 |
2.24 |
3.82 |
3.86 |
.97 |
| KLA/ATP | 4.07 |
3.94 |
4.17 |
3.53 |
5.42 |
4.65 |
1.54 |
|
| Hk3 | 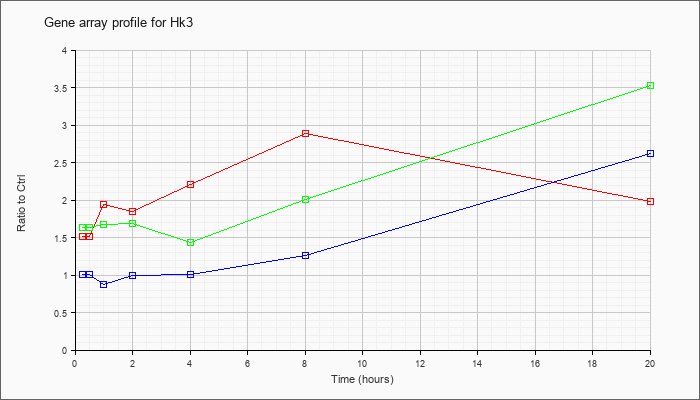 |
NM_001033245 |
hexokinase 3 (Hk3), nuclear gene encoding mitochondrial protein, mRNA [NM_001033245] |
KLA | 1.52 |
1.68 |
1.94 |
1.85 |
2.21 |
2.89 |
1.98 |
| ATP | 1.01 |
1.07 |
.87 |
.99 |
1.01 |
1.26 |
2.62 |
| KLA/ATP | 1.63 |
1.74 |
1.67 |
1.69 |
1.44 |
2.01 |
3.53 |
|
| Hkdc1 | 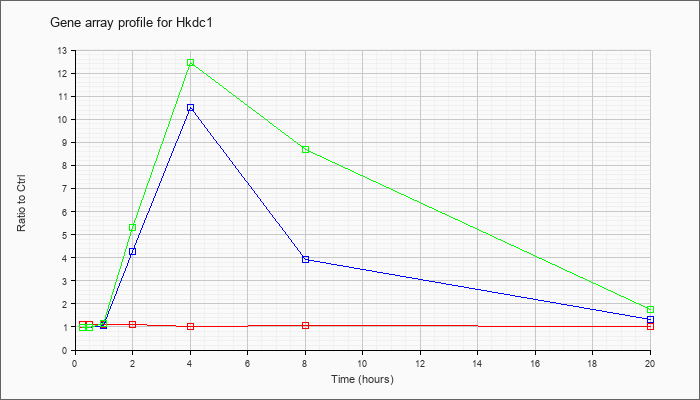 |
NM_145419 |
hexokinase domain containing 1 (Hkdc1), mRNA [NM_145419] |
KLA | 1.10 |
1.08 |
1.12 |
1.12 |
1.02 |
1.07 |
1.04 |
| ATP | .96 |
1.04 |
1.06 |
4.28 |
10.50 |
3.91 |
1.34 |
| KLA/ATP | .98 |
1.04 |
1.14 |
5.33 |
12.44 |
8.68 |
1.77 |
|
| Hmox1 | 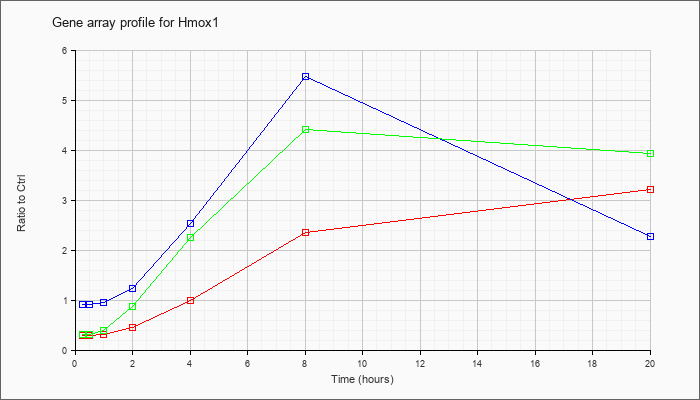 |
NM_010442 |
heme oxygenase (decycling) 1 (Hmox1), mRNA [NM_010442] |
KLA | .30 |
.31 |
.32 |
.45 |
1.00 |
2.36 |
3.22 |
| ATP | .91 |
.98 |
.96 |
1.24 |
2.53 |
5.47 |
2.27 |
| KLA/ATP | .31 |
.30 |
.40 |
.87 |
2.26 |
4.40 |
3.94 |
|
| Ifng | 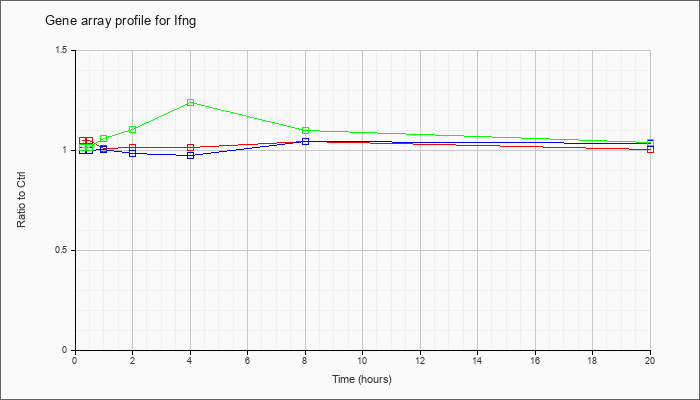 |
NM_008337 |
interferon gamma (Ifng), mRNA [NM_008337] |
KLA | 1.05 |
1.02 |
1.01 |
1.01 |
1.01 |
1.04 |
1.00 |
| ATP | 1.00 |
1.01 |
1.01 |
.98 |
.97 |
1.04 |
1.03 |
| KLA/ATP | 1.01 |
1.01 |
1.06 |
1.10 |
1.24 |
1.10 |
1.04 |
|
| Ifngr1 | 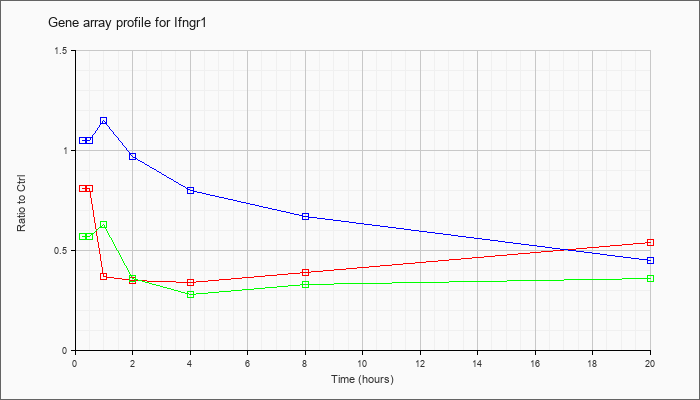 |
NM_010511 |
interferon gamma receptor 1 (Ifngr1), mRNA [NM_010511] |
KLA | .81 |
.60 |
.37 |
.35 |
.34 |
.39 |
.54 |
| ATP | 1.05 |
.96 |
1.15 |
.97 |
.80 |
.67 |
.45 |
| KLA/ATP | .57 |
.57 |
.63 |
.36 |
.28 |
.33 |
.36 |
|
| Ifngr2 | 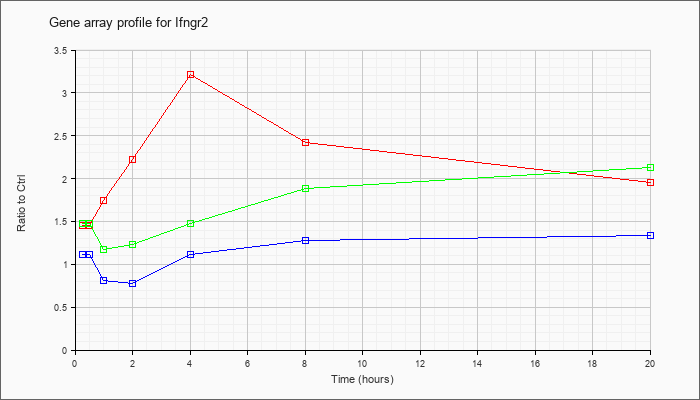 |
NM_008338 |
interferon gamma receptor 2 (Ifngr2), mRNA [NM_008338] |
KLA | 1.45 |
1.49 |
1.74 |
2.22 |
3.22 |
2.42 |
1.96 |
| ATP | 1.11 |
1.03 |
.81 |
.78 |
1.12 |
1.28 |
1.34 |
| KLA/ATP | 1.48 |
1.41 |
1.17 |
1.23 |
1.48 |
1.89 |
2.13 |
|
| Igf1 | 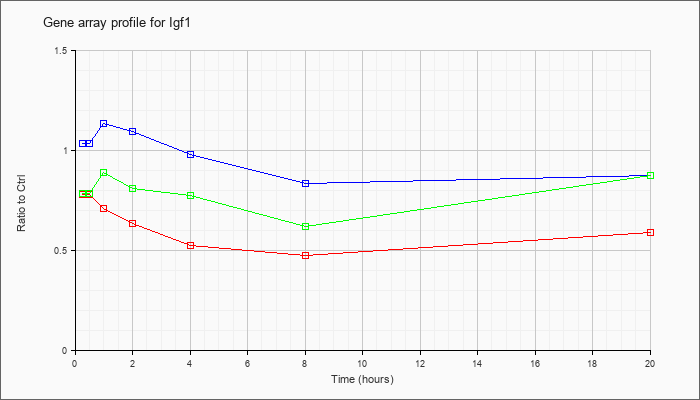 |
AK050118 |
adult male liver tumor cDNA, RIKEN full-length enriched library, clone:C730016P09 product:unclassifiable, full insert sequence. [AK050118] |
KLA | .13 |
.14 |
.15 |
.16 |
.22 |
.34 |
.48 |
| ATP | 1.78 |
2.05 |
1.37 |
.72 |
.34 |
.41 |
.58 |
| KLA/ATP | .24 |
.45 |
.50 |
.24 |
.20 |
.26 |
.68 |
|
| Igf1 |  |
NM_010512 |
insulin-like growth factor 1 (Igf1), transcript variant 1, mRNA [NM_010512] |
KLA | .84 |
.78 |
.76 |
.67 |
.54 |
.48 |
.59 |
| ATP | .97 |
.98 |
1.13 |
1.13 |
1.03 |
.88 |
.89 |
| KLA/ATP | .82 |
.81 |
.93 |
.86 |
.83 |
.65 |
.89 |
|
| Igf1 |  |
NM_184052 |
insulin-like growth factor 1 (Igf1), transcript variant 2, mRNA [NM_184052] |
KLA | .77 |
.82 |
.76 |
.65 |
.60 |
.52 |
.69 |
| ATP | 1.01 |
1.07 |
.98 |
1.06 |
1.03 |
.78 |
.94 |
| KLA/ATP | .89 |
.86 |
.77 |
.80 |
.75 |
.65 |
.89 |
|
| Igf1r | 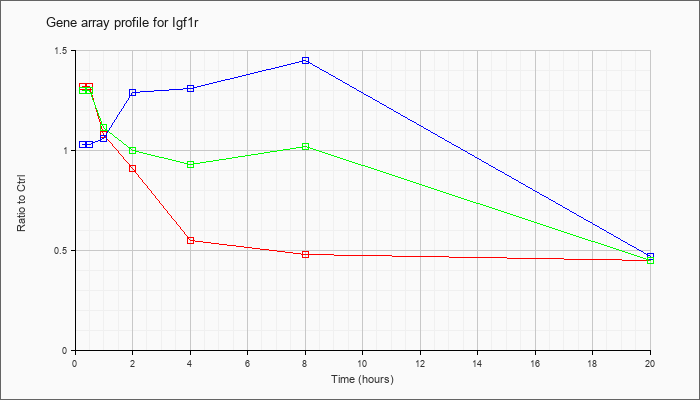 |
NM_010513 |
insulin-like growth factor I receptor (Igf1r), mRNA [NM_010513] |
KLA | 1.32 |
1.32 |
1.08 |
.91 |
.55 |
.48 |
.45 |
| ATP | 1.03 |
1.06 |
1.06 |
1.29 |
1.31 |
1.45 |
.47 |
| KLA/ATP | 1.30 |
1.37 |
1.11 |
1.00 |
.93 |
1.02 |
.45 |
|
| Il6 | 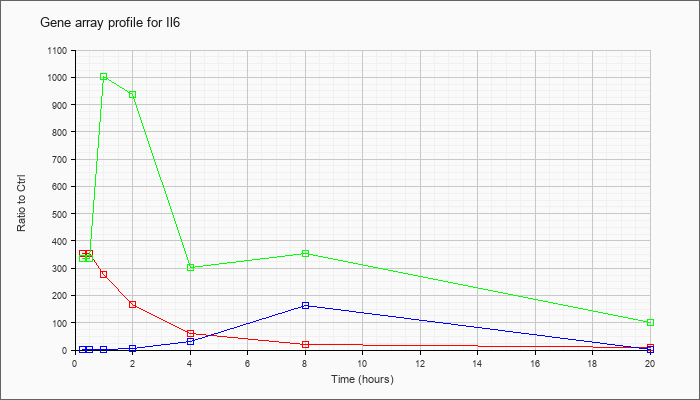 |
NM_031168 |
interleukin 6 (Il6), mRNA [NM_031168] |
KLA | 352.99 |
378.70 |
276.81 |
168.47 |
59.99 |
18.66 |
8.27 |
| ATP | 1.04 |
1.08 |
2.28 |
3.69 |
30.40 |
161.93 |
2.11 |
| KLA/ATP | 334.28 |
528.66 |
###.## |
935.86 |
302.45 |
355.36 |
99.65 |
|
| Il6ra | 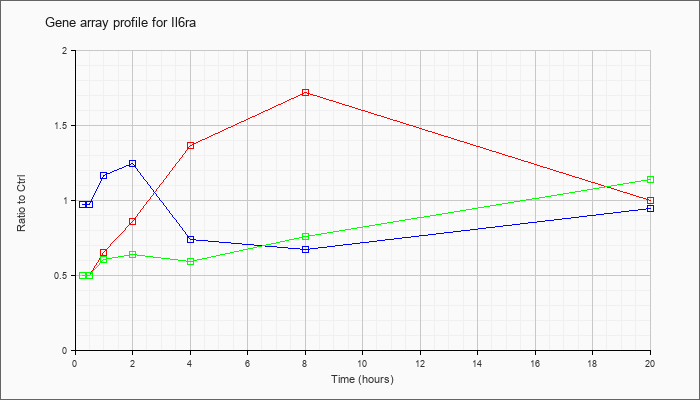 |
NM_010559 |
interleukin 6 receptor, alpha (Il6ra), mRNA [NM_010559] |
KLA | .50 |
.53 |
.65 |
.86 |
1.36 |
1.72 |
1.00 |
| ATP | .97 |
1.20 |
1.17 |
1.25 |
.74 |
.67 |
.95 |
| KLA/ATP | .50 |
.60 |
.61 |
.64 |
.59 |
.76 |
1.14 |
|
| Ins1 | 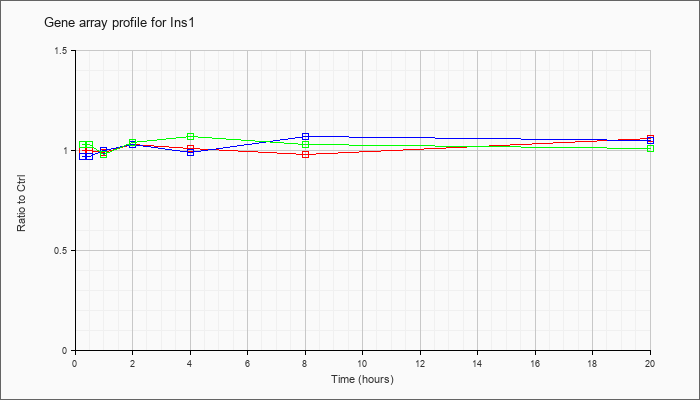 |
NM_008386 |
insulin I (Ins1), mRNA [NM_008386] |
KLA | 1.00 |
.99 |
.99 |
1.03 |
1.01 |
.98 |
1.06 |
| ATP | .97 |
1.03 |
1.00 |
1.03 |
.99 |
1.07 |
1.05 |
| KLA/ATP | 1.03 |
.93 |
.98 |
1.04 |
1.07 |
1.03 |
1.01 |
|
| Ins2 | 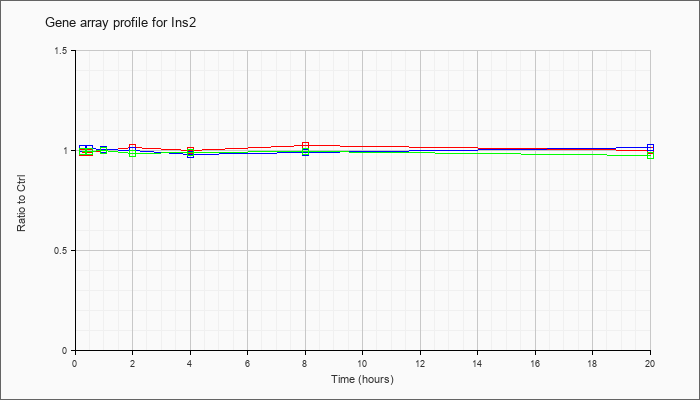 |
NM_008387 |
insulin II (Ins2), mRNA [NM_008387] |
KLA | .99 |
1.00 |
1.00 |
1.01 |
1.00 |
1.02 |
1.00 |
| ATP | 1.01 |
.99 |
1.00 |
1.00 |
.98 |
.99 |
1.01 |
| KLA/ATP | .99 |
1.01 |
1.00 |
.98 |
.99 |
.99 |
.98 |
|
| Insr | 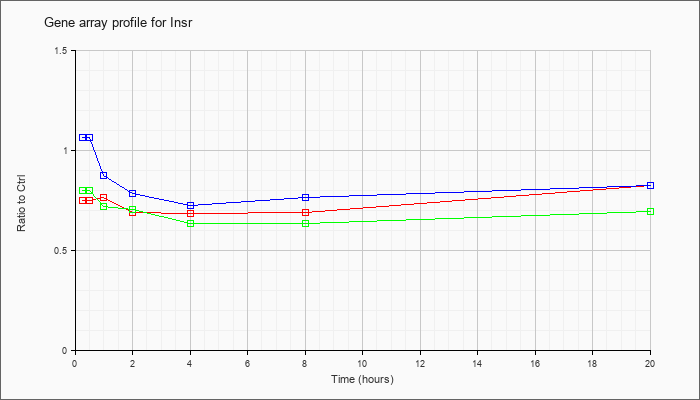 |
NM_010568 |
insulin receptor (Insr), mRNA [NM_010568] |
KLA | .75 |
.81 |
.77 |
.69 |
.69 |
.69 |
.83 |
| ATP | 1.07 |
1.06 |
.88 |
.79 |
.73 |
.77 |
.83 |
| KLA/ATP | .80 |
.80 |
.72 |
.71 |
.64 |
.64 |
.70 |
|
| Ldha | 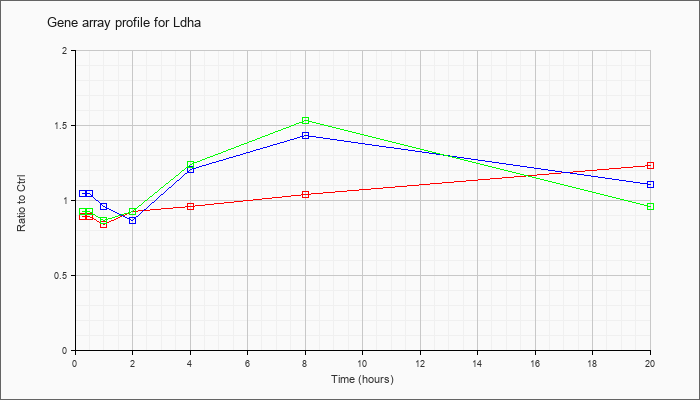 |
NM_010699 |
lactate dehydrogenase A (Ldha), mRNA [NM_010699] |
KLA | .89 |
.80 |
.84 |
.93 |
.96 |
1.04 |
1.23 |
| ATP | 1.05 |
1.04 |
.96 |
.87 |
1.21 |
1.43 |
1.11 |
| KLA/ATP | .93 |
.87 |
.87 |
.93 |
1.24 |
1.53 |
.96 |
|
| Ltbr | 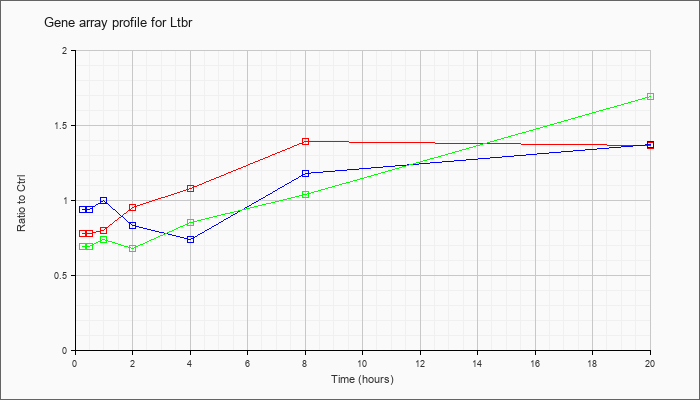 |
NM_010736 |
lymphotoxin B receptor (Ltbr), mRNA [NM_010736] |
KLA | .78 |
.77 |
.80 |
.95 |
1.08 |
1.39 |
1.36 |
| ATP | .94 |
.86 |
1.00 |
.83 |
.74 |
1.18 |
1.37 |
| KLA/ATP | .69 |
.71 |
.74 |
.68 |
.85 |
1.04 |
1.69 |
|
| Map2k1 | 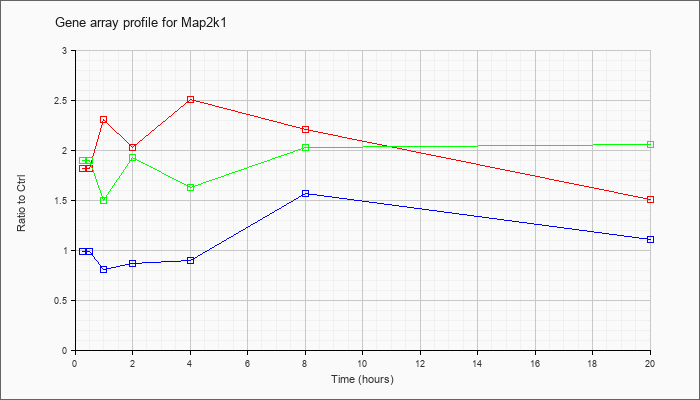 |
NM_008927 |
mitogen-activated protein kinase kinase 1 (Map2k1), mRNA [NM_008927] |
KLA | 1.82 |
2.00 |
2.31 |
2.03 |
2.51 |
2.20 |
1.51 |
| ATP | .99 |
1.14 |
.81 |
.87 |
.90 |
1.57 |
1.11 |
| KLA/ATP | 1.90 |
2.03 |
1.50 |
1.93 |
1.63 |
2.03 |
2.06 |
|
| Map2k2 | 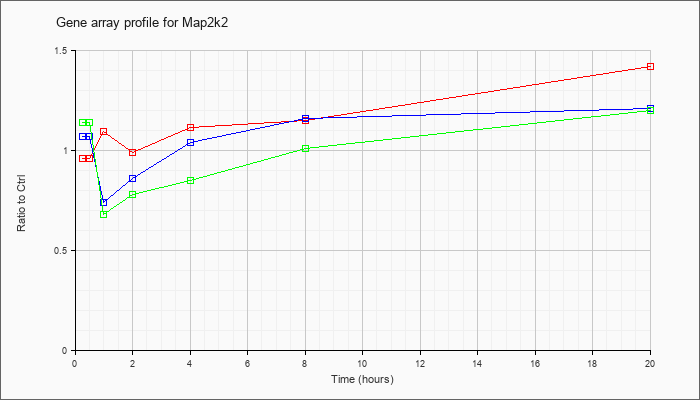 |
NM_023138 |
mitogen-activated protein kinase kinase 2 (Map2k2), mRNA [NM_023138] |
KLA | .96 |
1.01 |
1.09 |
.99 |
1.11 |
1.15 |
1.42 |
| ATP | 1.07 |
1.05 |
.74 |
.86 |
1.04 |
1.16 |
1.21 |
| KLA/ATP | 1.14 |
1.03 |
.68 |
.78 |
.85 |
1.01 |
1.20 |
|
| Mapk1 | 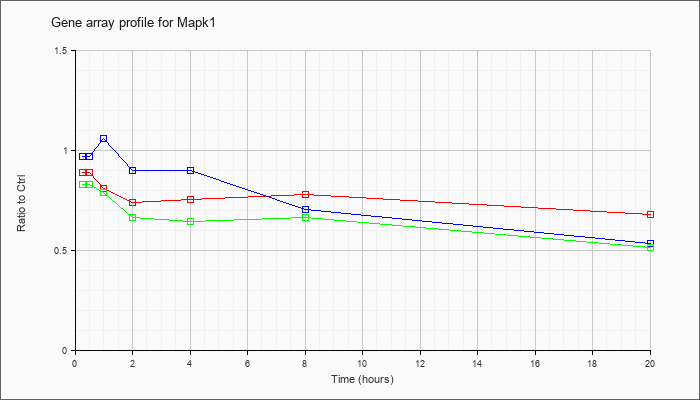 |
NM_001038663 |
mitogen-activated protein kinase 1 (Mapk1), transcript variant 2, mRNA [NM_001038663] |
KLA | .97 |
1.03 |
.93 |
.86 |
.79 |
1.01 |
1.04 |
| ATP | .96 |
1.18 |
1.01 |
1.13 |
.84 |
1.04 |
1.16 |
| KLA/ATP | .99 |
.99 |
.84 |
.87 |
.63 |
.96 |
1.26 |
|
| Mapk1 |  |
NM_011949 |
mitogen-activated protein kinase 1 (Mapk1), transcript variant 1, mRNA [NM_011949] |
KLA | .88 |
.80 |
.80 |
.73 |
.75 |
.76 |
.65 |
| ATP | .97 |
.96 |
1.06 |
.88 |
.90 |
.68 |
.48 |
| KLA/ATP | .82 |
.81 |
.78 |
.65 |
.64 |
.64 |
.45 |
|
| Mapk3 | 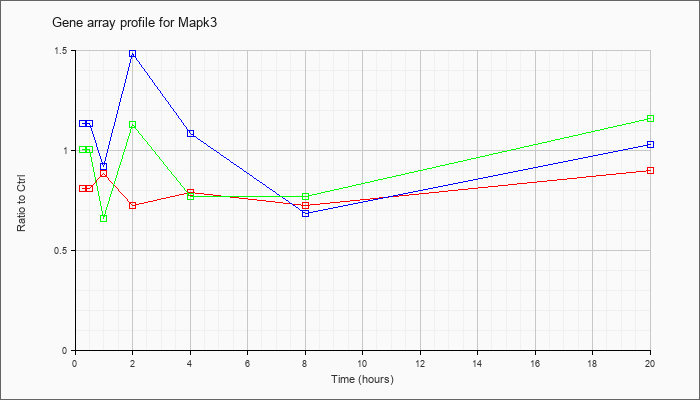 |
NM_011952 |
mitogen-activated protein kinase 3 (Mapk3), mRNA [NM_011952] |
KLA | .81 |
.85 |
.88 |
.72 |
.79 |
.73 |
.90 |
| ATP | 1.13 |
1.30 |
.92 |
1.48 |
1.08 |
.68 |
1.03 |
| KLA/ATP | 1.01 |
1.01 |
.66 |
1.13 |
.77 |
.77 |
1.16 |
|
| Mknk1 | 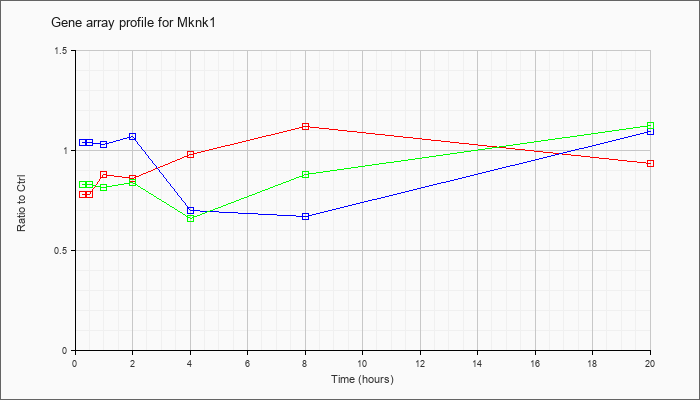 |
NM_021461 |
MAP kinase-interacting serine/threonine kinase 1 (Mknk1), mRNA [NM_021461] |
KLA | .78 |
.81 |
.88 |
.86 |
.98 |
1.12 |
.93 |
| ATP | 1.04 |
1.03 |
1.03 |
1.07 |
.70 |
.67 |
1.09 |
| KLA/ATP | .83 |
.83 |
.81 |
.84 |
.66 |
.88 |
1.12 |
|
| Mknk2 | 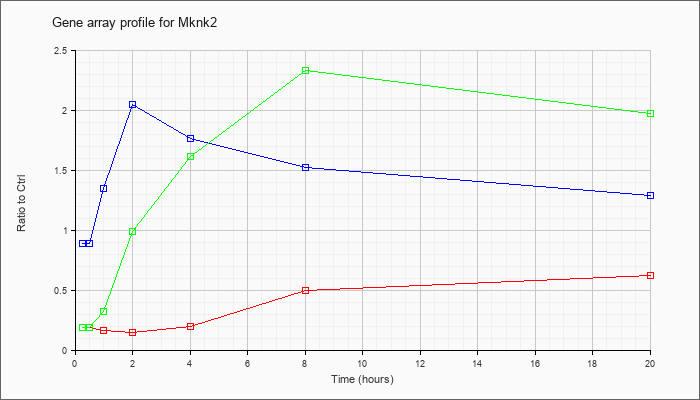 |
NM_021462 |
MAP kinase-interacting serine/threonine kinase 2 (Mknk2), mRNA [NM_021462] |
KLA | .19 |
.18 |
.16 |
.15 |
.20 |
.50 |
.62 |
| ATP | .89 |
.86 |
1.35 |
2.05 |
1.76 |
1.52 |
1.29 |
| KLA/ATP | .19 |
.17 |
.32 |
.99 |
1.61 |
2.33 |
1.97 |
|
Mm.13153
0 | 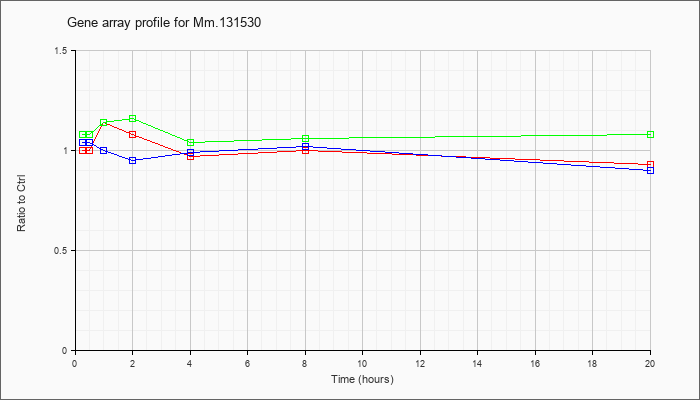 |
161086915 |
Unknown |
KLA | 1.00 |
1.07 |
1.14 |
1.08 |
.97 |
1.00 |
.93 |
| ATP | 1.04 |
.99 |
1.00 |
.95 |
.99 |
1.02 |
.90 |
| KLA/ATP | 1.08 |
1.18 |
1.14 |
1.16 |
1.04 |
1.06 |
1.08 |
|
Mm.19961
| 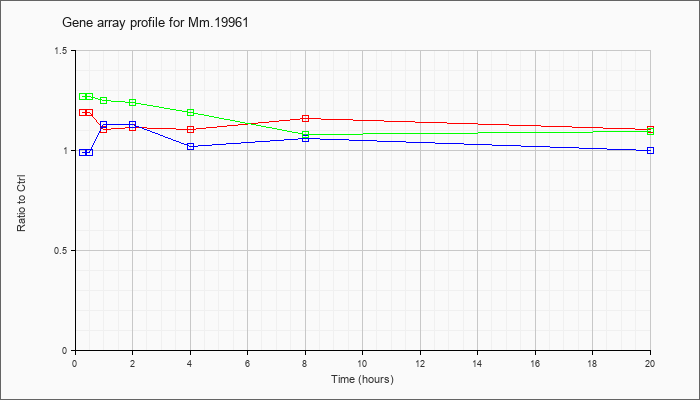 |
142378163 |
Unknown |
KLA | 1.19 |
1.17 |
1.10 |
1.11 |
1.10 |
1.16 |
1.10 |
| ATP | .99 |
1.04 |
1.13 |
1.13 |
1.02 |
1.06 |
1.00 |
| KLA/ATP | 1.27 |
1.24 |
1.25 |
1.24 |
1.19 |
1.08 |
1.09 |
|
Mm.21312
8 | 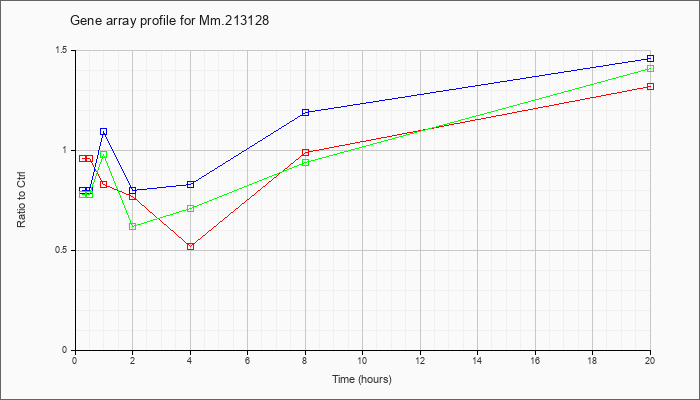 |
141801344 |
Unknown |
KLA | .96 |
.95 |
.83 |
.77 |
.52 |
.99 |
1.32 |
| ATP | .80 |
.82 |
1.09 |
.80 |
.83 |
1.19 |
1.46 |
| KLA/ATP | .78 |
.92 |
.98 |
.62 |
.71 |
.94 |
1.41 |
|
Mm.24936
4 | 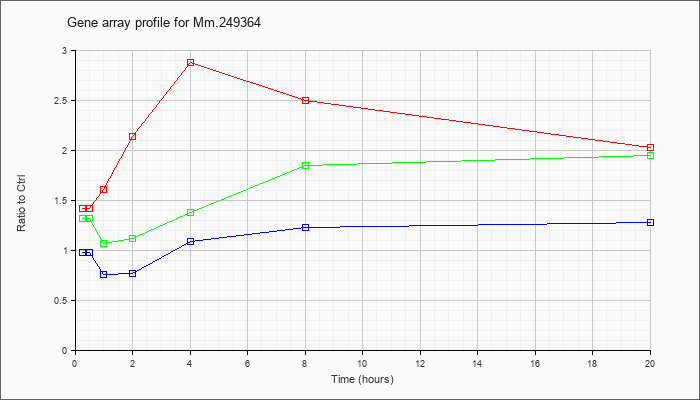 |
133892283 |
Unknown |
KLA | 1.42 |
1.41 |
1.61 |
2.14 |
2.88 |
2.49 |
2.03 |
| ATP | .98 |
.98 |
.76 |
.77 |
1.09 |
1.23 |
1.28 |
| KLA/ATP | 1.32 |
1.32 |
1.07 |
1.12 |
1.38 |
1.85 |
1.95 |
|
Mm.25381
9 | 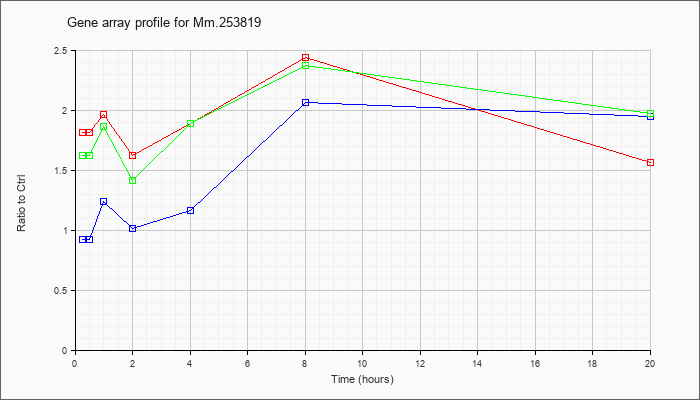 |
118130676 |
Unknown |
KLA | 1.81 |
1.88 |
1.96 |
1.62 |
1.89 |
2.44 |
1.56 |
| ATP | .92 |
.91 |
1.24 |
1.01 |
1.16 |
2.06 |
1.95 |
| KLA/ATP | 1.62 |
1.86 |
1.86 |
1.41 |
1.89 |
2.37 |
1.97 |
|
Mm.27574
2 | 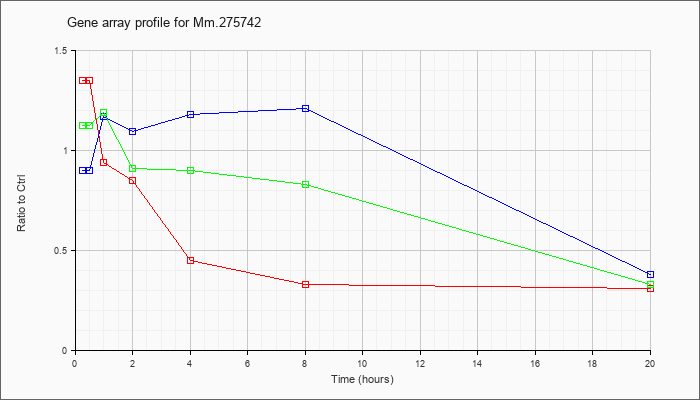 |
112983655 |
Unknown |
KLA | 1.35 |
1.16 |
.94 |
.85 |
.45 |
.33 |
.31 |
| ATP | .90 |
.86 |
1.17 |
1.09 |
1.18 |
1.21 |
.38 |
| KLA/ATP | 1.12 |
1.22 |
1.19 |
.91 |
.90 |
.83 |
.33 |
|
Mm.28924
8 | 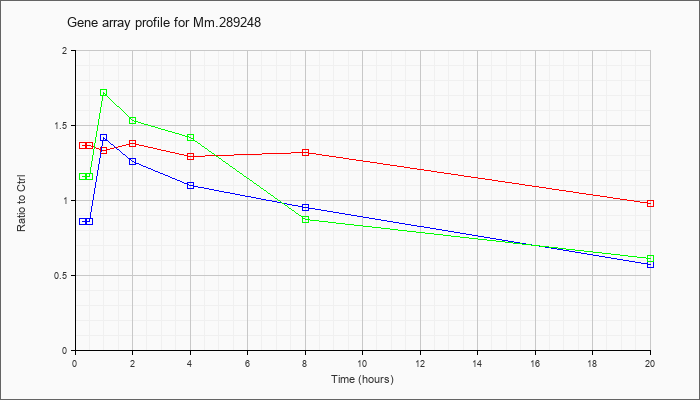 |
148747277 |
Unknown |
KLA | 1.36 |
1.32 |
1.33 |
1.38 |
1.29 |
1.32 |
.98 |
| ATP | .86 |
.92 |
1.42 |
1.26 |
1.10 |
.95 |
.57 |
| KLA/ATP | 1.16 |
1.34 |
1.72 |
1.53 |
1.42 |
.87 |
.61 |
|
Mm.38049
| 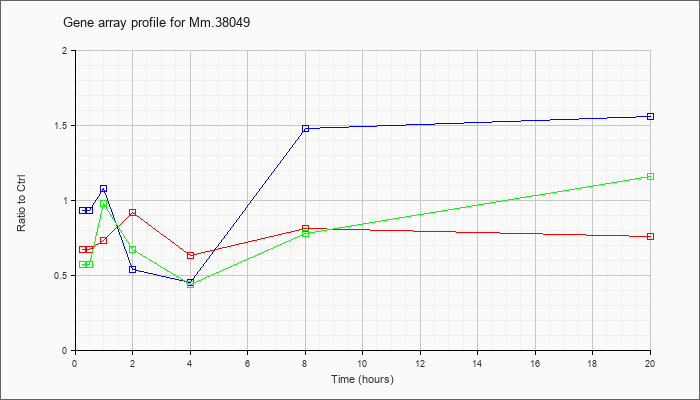 |
118130391 |
Unknown |
KLA | .67 |
.65 |
.73 |
.92 |
.63 |
.81 |
.76 |
| ATP | .93 |
.92 |
1.08 |
.54 |
.45 |
1.48 |
1.56 |
| KLA/ATP | .57 |
.65 |
.98 |
.67 |
.44 |
.78 |
1.16 |
|
Mm.44463
|  |
118129851 |
Unknown |
KLA | 1.01 |
.98 |
.97 |
1.02 |
.93 |
1.06 |
1.05 |
| ATP | .97 |
1.01 |
1.04 |
1.06 |
1.02 |
1.08 |
.97 |
| KLA/ATP | 1.02 |
1.06 |
.95 |
1.04 |
1.09 |
1.14 |
1.03 |
|
Mm.47351
0 | 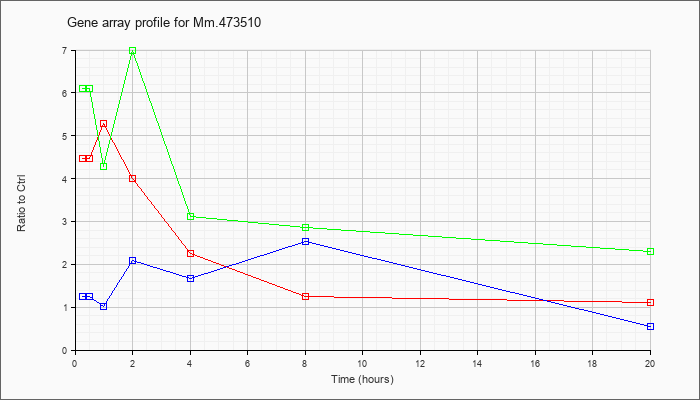 |
162287332 |
Unknown |
KLA | 4.46 |
4.82 |
5.29 |
4.00 |
2.26 |
1.25 |
1.10 |
| ATP | 1.24 |
1.72 |
1.02 |
2.08 |
1.66 |
2.53 |
.55 |
| KLA/ATP | 6.11 |
7.67 |
4.29 |
6.99 |
3.11 |
2.86 |
2.30 |
|
| Nfkb1 | 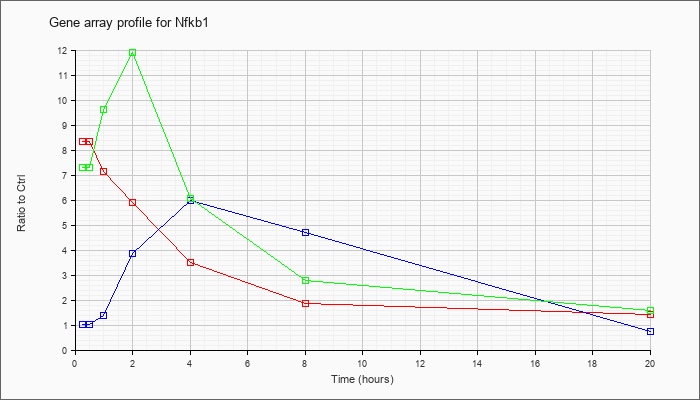 |
BC050841 |
nuclear factor of kappa light chain gene enhancer in B-cells 1, p105, mRNA (cDNA clone IMAGE:6392793), complete cds. [BC050841] |
KLA | 5.81 |
4.96 |
4.77 |
3.21 |
3.37 |
2.00 |
1.71 |
| ATP | 1.39 |
1.52 |
3.25 |
12.63 |
13.64 |
4.80 |
1.08 |
| KLA/ATP | 7.21 |
8.69 |
19.53 |
23.93 |
11.70 |
2.85 |
2.01 |
|
| Nfkb1 |  |
M57999 |
gb|Mouse transcription factor NF-kappa-B DNA binding subunit mRNA, complete cds. [M57999] |
KLA | 8.55 |
7.87 |
7.35 |
6.13 |
3.51 |
1.85 |
1.38 |
| ATP | .98 |
.96 |
1.21 |
3.06 |
5.29 |
4.67 |
.69 |
| KLA/ATP | 7.32 |
7.20 |
8.71 |
10.82 |
5.55 |
2.77 |
1.54 |
|
| Nos2 | 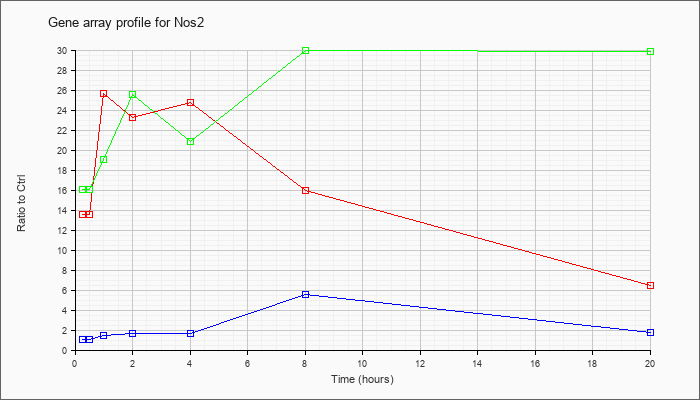 |
NM_010927 |
nitric oxide synthase 2, inducible, macrophage (Nos2), mRNA [NM_010927] |
KLA | 13.50 |
16.36 |
25.65 |
23.25 |
24.78 |
15.99 |
6.48 |
| ATP | 1.03 |
1.03 |
1.48 |
1.66 |
1.65 |
5.55 |
1.70 |
| KLA/ATP | 16.01 |
17.41 |
19.06 |
25.50 |
20.84 |
29.92 |
29.80 |
|
| Nos3 | 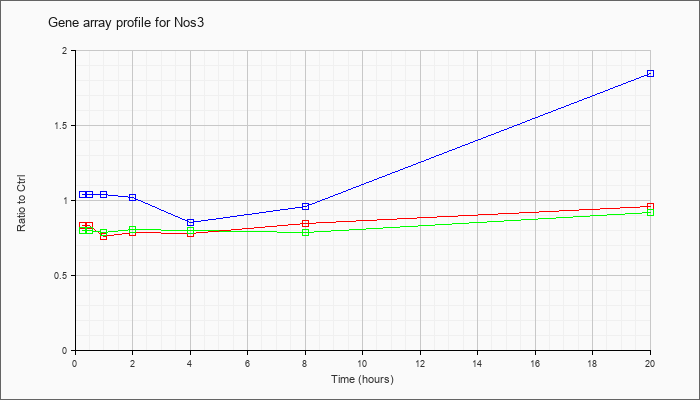 |
NM_008713 |
nitric oxide synthase 3, endothelial cell (Nos3), mRNA [NM_008713] |
KLA | .83 |
.79 |
.76 |
.78 |
.77 |
.85 |
.96 |
| ATP | 1.03 |
1.09 |
1.03 |
1.02 |
.85 |
.96 |
1.84 |
| KLA/ATP | .80 |
.78 |
.79 |
.80 |
.79 |
.78 |
.91 |
|
| Nox1 | 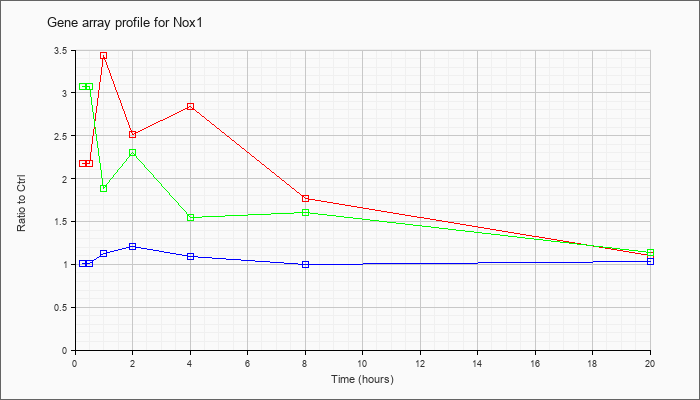 |
NM_172203 |
NADPH oxidase 1 (Nox1), mRNA [NM_172203] |
KLA | 2.18 |
2.93 |
3.44 |
2.51 |
2.84 |
1.77 |
1.10 |
| ATP | 1.01 |
.99 |
1.13 |
1.21 |
1.09 |
1.00 |
1.03 |
| KLA/ATP | 3.07 |
3.07 |
1.88 |
2.30 |
1.55 |
1.60 |
1.14 |
|
| Nox3 | 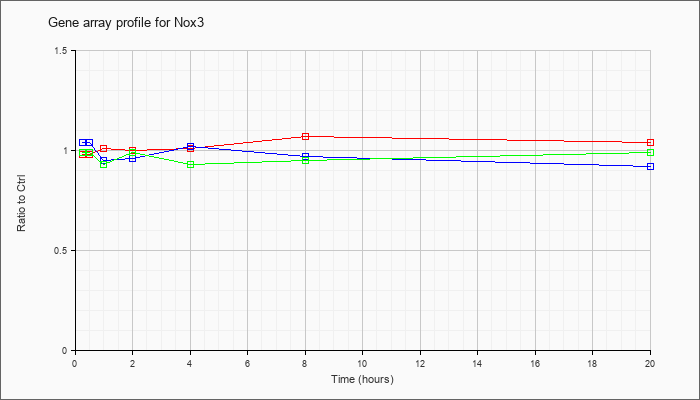 |
NM_198958 |
NADPH oxidase 3 (Nox3), mRNA [NM_198958] |
KLA | .98 |
.99 |
1.01 |
1.00 |
1.01 |
1.07 |
1.04 |
| ATP | 1.04 |
.95 |
.95 |
.96 |
1.02 |
.97 |
.92 |
| KLA/ATP | .99 |
1.03 |
.93 |
.99 |
.93 |
.95 |
.99 |
|
| Nppa | 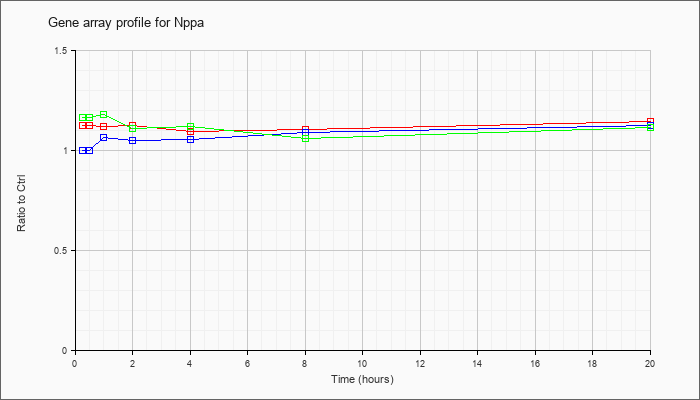 |
NM_008725 |
natriuretic peptide precursor type A (Nppa), mRNA [NM_008725] |
KLA | 1.12 |
1.09 |
1.12 |
1.12 |
1.09 |
1.10 |
1.14 |
| ATP | 1.00 |
1.01 |
1.06 |
1.05 |
1.05 |
1.09 |
1.12 |
| KLA/ATP | 1.16 |
1.17 |
1.18 |
1.11 |
1.12 |
1.06 |
1.11 |
|
| Pdha1 | 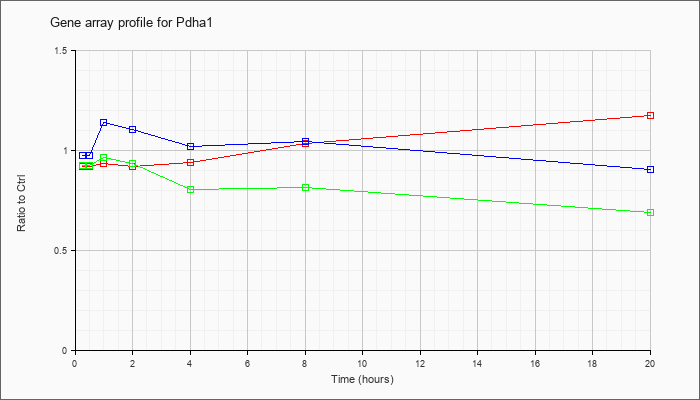 |
NM_008810 |
pyruvate dehydrogenase E1 alpha 1 (Pdha1), nuclear gene encoding mitochondrial protein, mRNA [NM_008810] |
KLA | .92 |
.96 |
.94 |
.92 |
.94 |
1.04 |
1.18 |
| ATP | .98 |
1.03 |
1.14 |
1.10 |
1.02 |
1.05 |
.91 |
| KLA/ATP | .93 |
.93 |
.97 |
.94 |
.81 |
.82 |
.69 |
|
| Pdha2 | 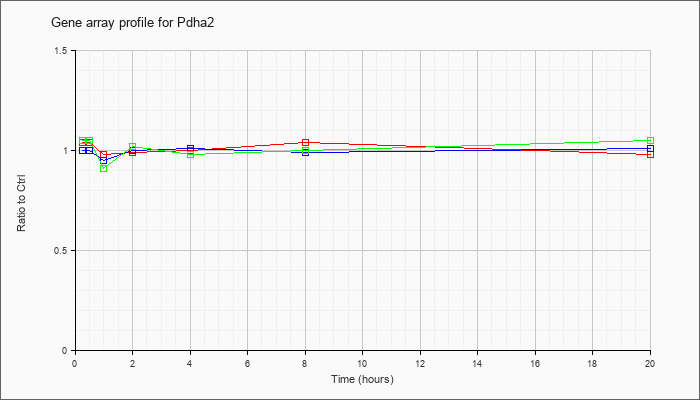 |
NM_008811 |
pyruvate dehydrogenase E1 alpha 2 (Pdha2), mRNA [NM_008811] |
KLA | 1.04 |
.95 |
.98 |
.99 |
1.00 |
1.04 |
.98 |
| ATP | 1.00 |
.97 |
.95 |
1.00 |
1.01 |
.99 |
1.01 |
| KLA/ATP | 1.05 |
1.07 |
.91 |
1.02 |
.98 |
1.00 |
1.05 |
|
| Pdhb | 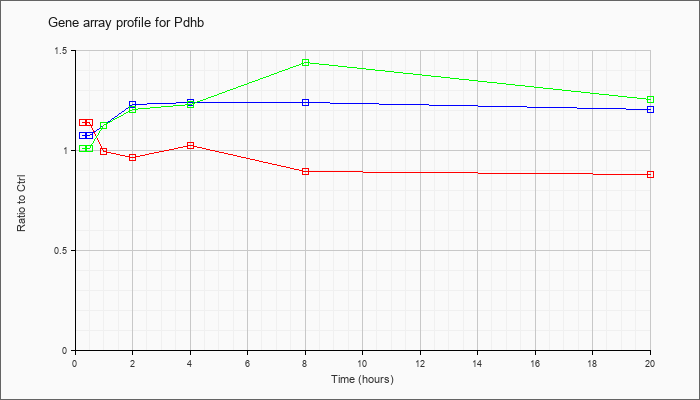 |
NM_024221 |
pyruvate dehydrogenase (lipoamide) beta (Pdhb), mRNA [NM_024221] |
KLA | 1.14 |
1.05 |
1.00 |
.97 |
1.03 |
.90 |
.88 |
| ATP | 1.08 |
1.10 |
1.12 |
1.23 |
1.24 |
1.24 |
1.21 |
| KLA/ATP | 1.01 |
1.16 |
1.13 |
1.21 |
1.23 |
1.44 |
1.26 |
|
| Pdk1 | 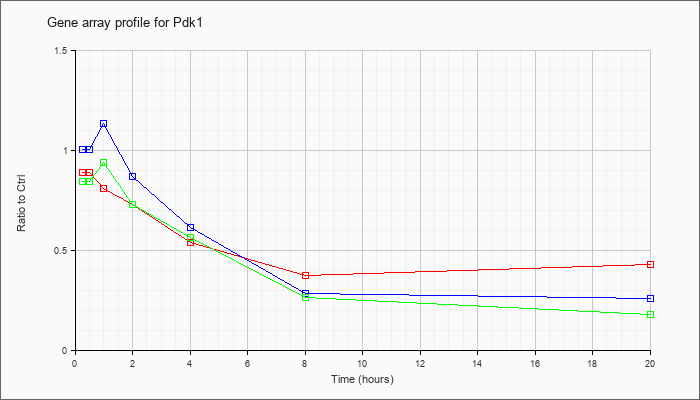 |
AK165090 |
2 days pregnant adult female ovary cDNA, RIKEN full-length enriched library, clone:E330005M21 product:hypothetical protein, full insert sequence. [AK165090] |
KLA | .86 |
.77 |
.71 |
.67 |
.45 |
.29 |
.35 |
| ATP | 1.06 |
.88 |
1.21 |
.72 |
.60 |
.26 |
.15 |
| KLA/ATP | .81 |
.74 |
.97 |
.57 |
.51 |
.22 |
.09 |
|
| Pdk1 |  |
NM_172665 |
pyruvate dehydrogenase kinase, isoenzyme 1 (Pdk1), nuclear gene encoding mitochondrial protein, mRNA [NM_172665] |
KLA | .92 |
.94 |
.91 |
.79 |
.63 |
.46 |
.51 |
| ATP | .95 |
1.03 |
1.06 |
1.02 |
.63 |
.31 |
.37 |
| KLA/ATP | .88 |
.91 |
.91 |
.89 |
.62 |
.31 |
.27 |
|
| Pfkfb1 | 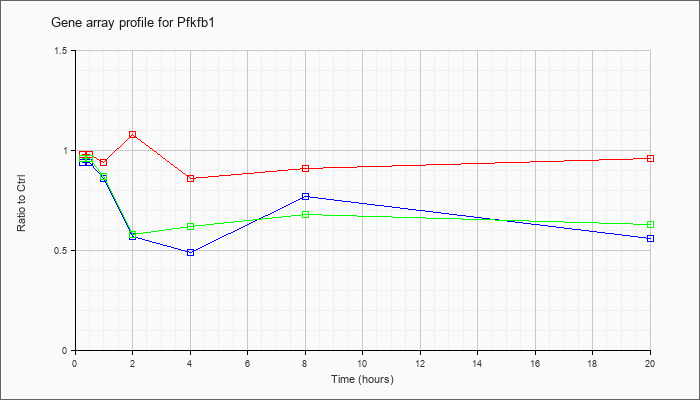 |
NM_008824 |
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 (Pfkfb1), mRNA [NM_008824] |
KLA | .98 |
.94 |
.94 |
1.08 |
.86 |
.91 |
.96 |
| ATP | .94 |
.86 |
.86 |
.57 |
.49 |
.77 |
.56 |
| KLA/ATP | .96 |
.90 |
.87 |
.58 |
.62 |
.68 |
.63 |
|
| Pfkfb2 | 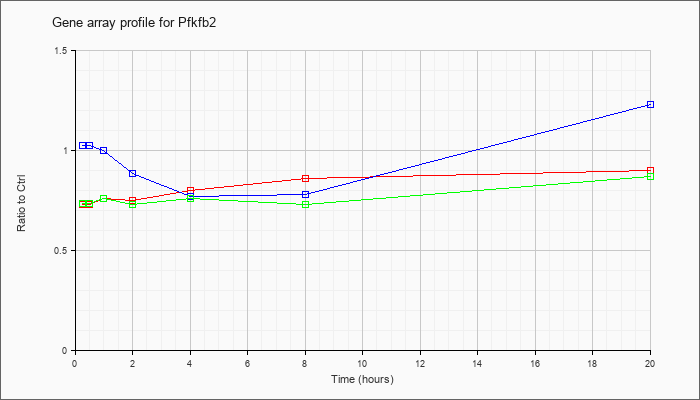 |
BC018418 |
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 2, mRNA (cDNA clone MGC:25723 IMAGE:3979400), complete cds. [BC018418] |
KLA | .88 |
1.00 |
.92 |
.91 |
1.00 |
.99 |
1.01 |
| ATP | 1.13 |
1.04 |
.99 |
.92 |
.94 |
.95 |
1.37 |
| KLA/ATP | .86 |
.87 |
.88 |
.84 |
.90 |
.89 |
1.05 |
|
| Pfkfb2 |  |
NM_008825 |
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 2 (Pfkfb2), mRNA [NM_008825] |
KLA | .65 |
.68 |
.68 |
.67 |
.70 |
.79 |
.84 |
| ATP | .97 |
.99 |
1.01 |
.87 |
.68 |
.70 |
1.16 |
| KLA/ATP | .67 |
.69 |
.70 |
.67 |
.69 |
.65 |
.78 |
|
| Pfkfb3 | 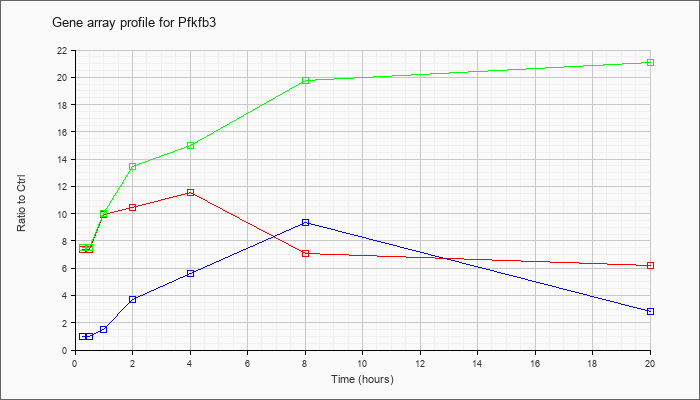 |
NM_133232 |
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 3 (Pfkfb3), mRNA [NM_133232] |
KLA | 7.38 |
8.57 |
9.97 |
10.42 |
11.55 |
7.07 |
6.18 |
| ATP | .99 |
1.07 |
1.47 |
3.74 |
5.65 |
9.34 |
2.84 |
| KLA/ATP | 7.53 |
8.77 |
9.99 |
13.46 |
15.00 |
19.74 |
21.11 |
|
| Pfkfb4 | 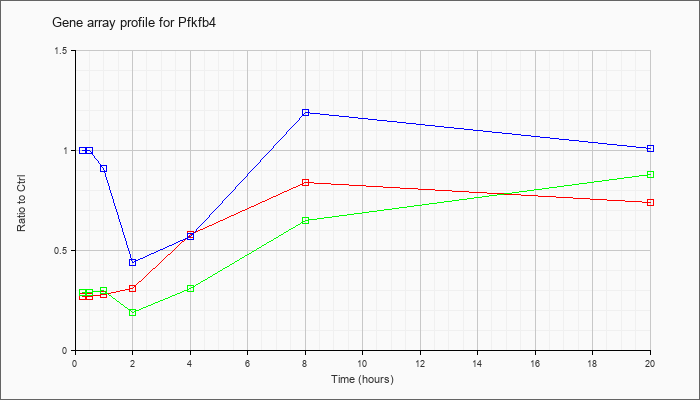 |
NM_173019 |
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 4 (Pfkfb4), mRNA [NM_173019] |
KLA | .27 |
.28 |
.28 |
.31 |
.58 |
.84 |
.74 |
| ATP | 1.00 |
.79 |
.91 |
.44 |
.57 |
1.19 |
1.01 |
| KLA/ATP | .29 |
.25 |
.30 |
.19 |
.31 |
.65 |
.88 |
|
| Pfkl | 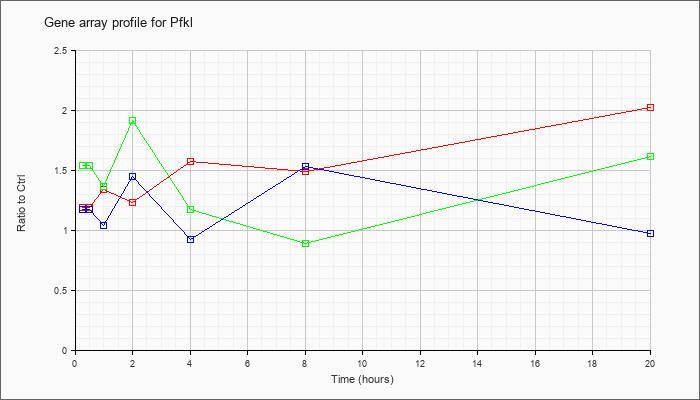 |
NM_008826 |
phosphofructokinase, liver, B-type (Pfkl), mRNA [NM_008826] |
KLA | 1.19 |
1.33 |
1.34 |
1.23 |
1.57 |
1.49 |
2.02 |
| ATP | 1.17 |
1.34 |
1.04 |
1.45 |
.92 |
1.53 |
.97 |
| KLA/ATP | 1.54 |
1.64 |
1.36 |
1.91 |
1.17 |
.89 |
1.61 |
|
| Pgk1 | 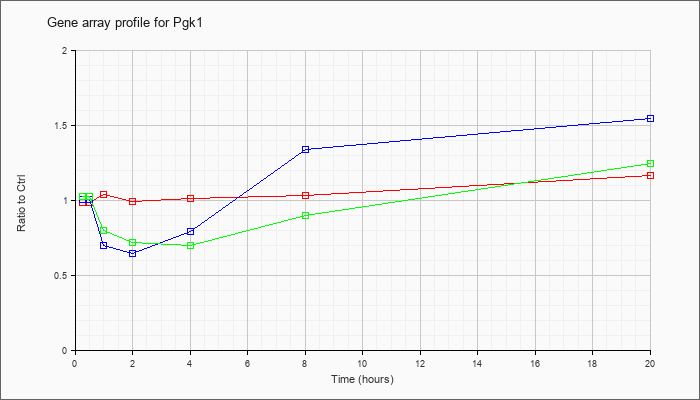 |
NM_008828 |
phosphoglycerate kinase 1 (Pgk1), mRNA [NM_008828] |
KLA | .98 |
1.04 |
1.04 |
.99 |
1.01 |
1.03 |
1.16 |
| ATP | 1.00 |
.94 |
.70 |
.64 |
.79 |
1.34 |
1.54 |
| KLA/ATP | 1.02 |
.96 |
.80 |
.72 |
.70 |
.90 |
1.24 |
|
| Pik3ca | 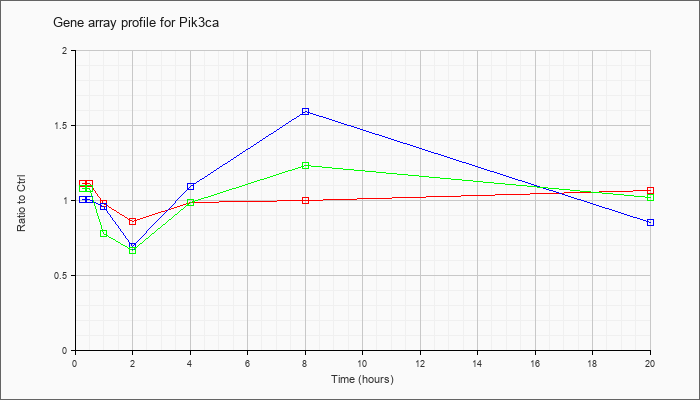 |
ENSMUST00000108243 |
ens|Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha isoform (EC 2.7.1.153) (PI3-kinase p110 subunit alpha) (PtdIns-3- kinase p110) (PI3K). [Source:Uniprot/SWISSPROT;Acc:P42337] [ENSMUST00000108243] |
KLA | 1.07 |
1.01 |
.91 |
.90 |
.98 |
.96 |
.86 |
| ATP | 1.03 |
.87 |
1.12 |
.63 |
1.22 |
1.86 |
.60 |
| KLA/ATP | 1.05 |
.82 |
.91 |
.61 |
1.17 |
1.26 |
.63 |
|
| Pik3ca |  |
NM_008839 |
phosphatidylinositol 3-kinase, catalytic, alpha polypeptide (Pik3ca), mRNA [NM_008839] |
KLA | 1.15 |
1.11 |
1.05 |
.82 |
.99 |
1.03 |
1.27 |
| ATP | .98 |
1.00 |
.79 |
.75 |
.96 |
1.32 |
1.10 |
| KLA/ATP | 1.10 |
.98 |
.64 |
.72 |
.80 |
1.20 |
1.41 |
|
| Pik3cb | 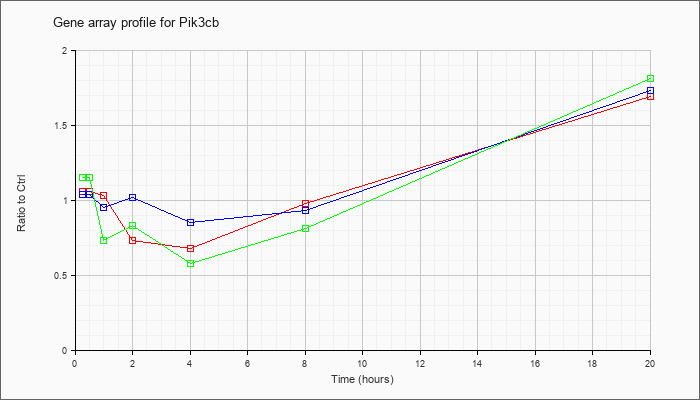 |
NM_029094 |
phosphatidylinositol 3-kinase, catalytic, beta polypeptide (Pik3cb), mRNA [NM_029094] |
KLA | 1.06 |
1.06 |
1.03 |
.73 |
.68 |
.98 |
1.69 |
| ATP | 1.04 |
1.21 |
.95 |
1.02 |
.85 |
.93 |
1.73 |
| KLA/ATP | 1.15 |
1.27 |
.73 |
.83 |
.58 |
.81 |
1.81 |
|
| Pik3cd | 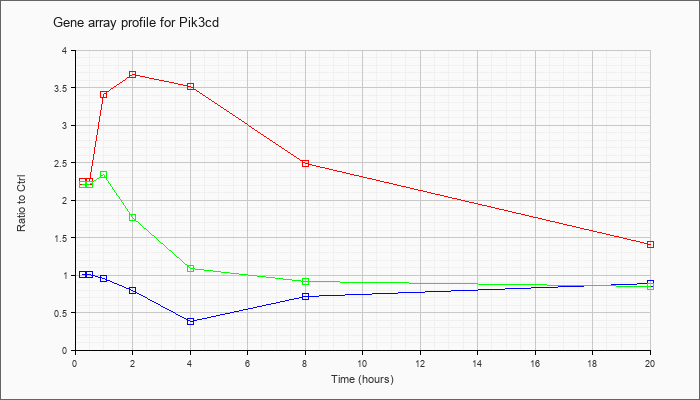 |
NM_008840 |
phosphatidylinositol 3-kinase catalytic delta polypeptide (Pik3cd), transcript variant 1, mRNA [NM_008840] |
KLA | 2.28 |
2.42 |
3.10 |
4.17 |
3.35 |
2.31 |
1.34 |
| ATP | 1.02 |
.91 |
.99 |
.63 |
.30 |
.67 |
.80 |
| KLA/ATP | 2.14 |
2.23 |
2.70 |
1.51 |
1.17 |
.85 |
.72 |
|
| Pik3cd |  |
U86587 |
phosphatidylinositol 3-kinase catalytic subunit p110 delta mRNA, complete cds. [U86587] |
KLA | 2.21 |
2.24 |
3.71 |
3.17 |
3.67 |
2.66 |
1.48 |
| ATP | 1.00 |
1.03 |
.93 |
.96 |
.46 |
.75 |
.97 |
| KLA/ATP | 2.27 |
2.32 |
1.97 |
2.02 |
1.01 |
.99 |
.98 |
|
| Pik3cg | 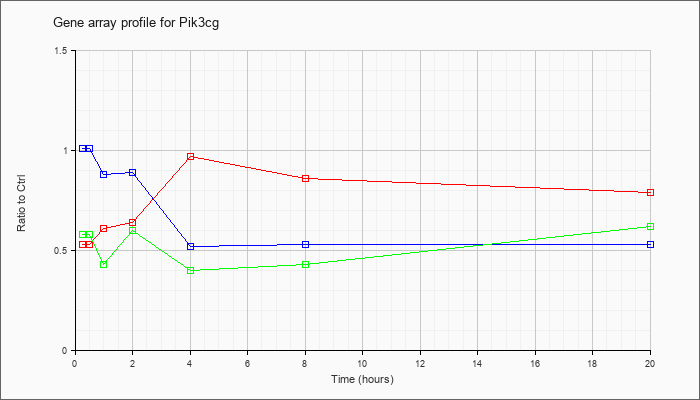 |
NM_020272 |
phosphoinositide-3-kinase, catalytic, gamma polypeptide (Pik3cg), mRNA [NM_020272] |
KLA | .53 |
.53 |
.61 |
.64 |
.97 |
.86 |
.79 |
| ATP | 1.01 |
1.13 |
.88 |
.89 |
.52 |
.53 |
.53 |
| KLA/ATP | .58 |
.53 |
.43 |
.60 |
.40 |
.43 |
.62 |
|
| Pik3r1 | 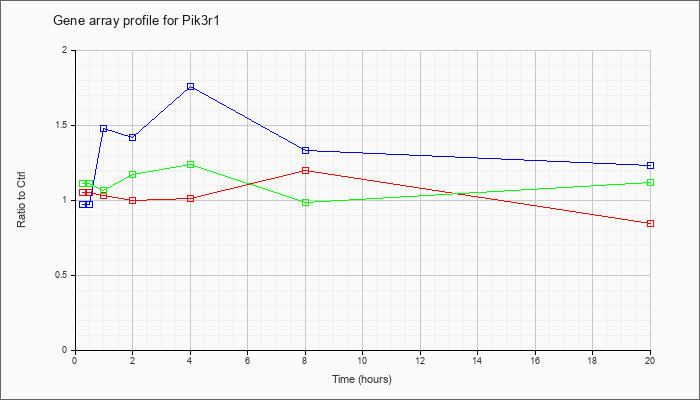 |
NM_001077495 |
phosphatidylinositol 3-kinase, regulatory subunit, polypeptide 1 (p85 alpha) (Pik3r1), transcript variant 2, mRNA [NM_001077495] |
KLA | 1.05 |
1.07 |
1.03 |
1.00 |
1.01 |
1.20 |
.85 |
| ATP | .97 |
1.02 |
1.48 |
1.42 |
1.76 |
1.33 |
1.23 |
| KLA/ATP | 1.11 |
1.12 |
1.06 |
1.17 |
1.24 |
.99 |
1.12 |
|
| Pik3r2 | 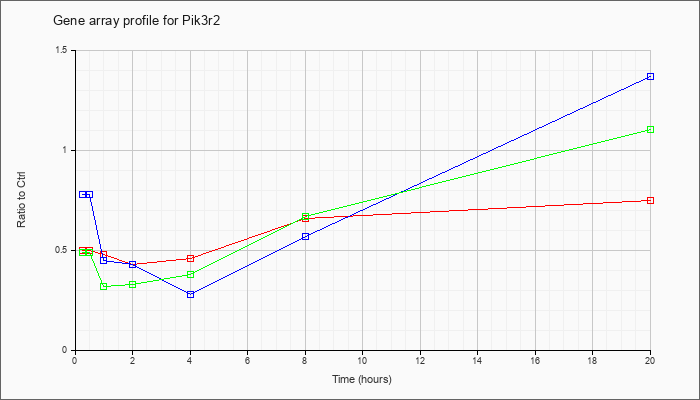 |
NM_008841 |
phosphatidylinositol 3-kinase, regulatory subunit, polypeptide 2 (p85 beta) (Pik3r2), mRNA [NM_008841] |
KLA | .50 |
.50 |
.48 |
.43 |
.46 |
.66 |
.75 |
| ATP | .78 |
.68 |
.45 |
.43 |
.28 |
.57 |
1.37 |
| KLA/ATP | .49 |
.38 |
.32 |
.33 |
.38 |
.67 |
1.10 |
|
| Pik3r3 | 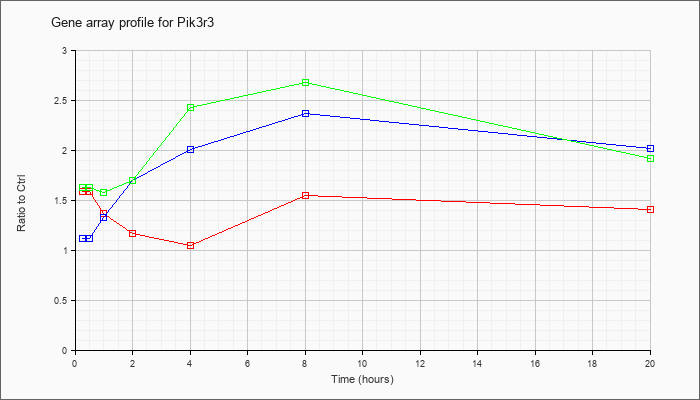 |
NM_181585 |
phosphatidylinositol 3 kinase, regulatory subunit, polypeptide 3 (p55) (Pik3r3), mRNA [NM_181585] |
KLA | 1.59 |
1.49 |
1.37 |
1.17 |
1.05 |
1.55 |
1.41 |
| ATP | 1.12 |
1.00 |
1.33 |
1.70 |
2.01 |
2.37 |
2.02 |
| KLA/ATP | 1.63 |
1.54 |
1.58 |
1.70 |
2.43 |
2.68 |
1.92 |
|
| Pik3r5 | 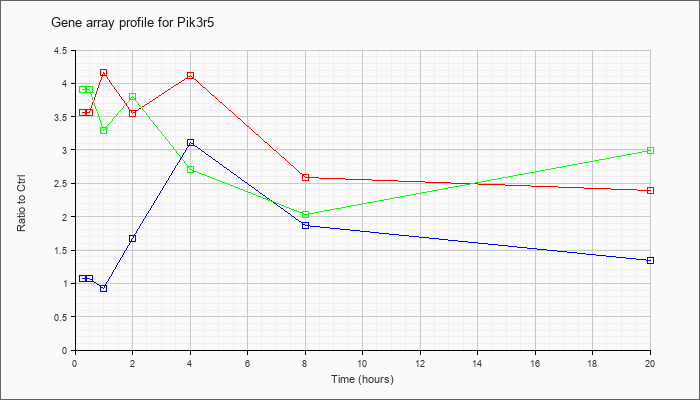 |
NM_177320 |
phosphoinositide-3-kinase, regulatory subunit 5, p101 (Pik3r5), mRNA [NM_177320] |
KLA | 3.57 |
3.58 |
4.16 |
3.54 |
4.11 |
2.59 |
2.39 |
| ATP | 1.08 |
1.08 |
.93 |
1.67 |
3.11 |
1.88 |
1.34 |
| KLA/ATP | 3.91 |
3.70 |
3.30 |
3.80 |
2.70 |
2.04 |
3.00 |
|
| Plcg2 | 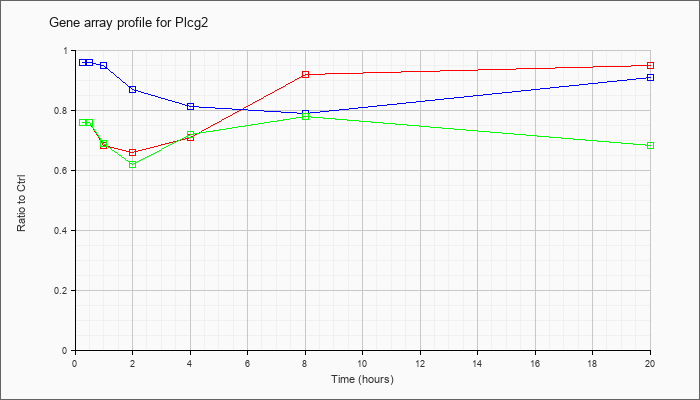 |
NM_172285 |
phospholipase C, gamma 2 (Plcg2), mRNA [NM_172285] |
KLA | .76 |
.74 |
.68 |
.66 |
.71 |
.92 |
.95 |
| ATP | .96 |
.90 |
.95 |
.87 |
.81 |
.79 |
.91 |
| KLA/ATP | .76 |
.71 |
.69 |
.62 |
.72 |
.78 |
.68 |
|
| Prkca | 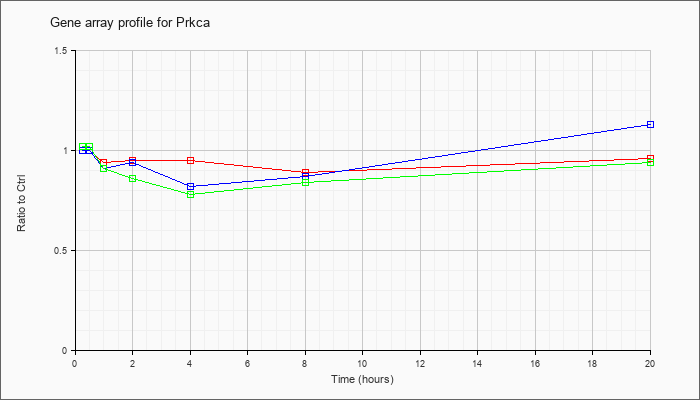 |
NM_011101 |
protein kinase C, alpha (Prkca), mRNA [NM_011101] |
KLA | 1.00 |
.96 |
.94 |
.95 |
.95 |
.89 |
.96 |
| ATP | 1.00 |
1.03 |
.91 |
.94 |
.82 |
.87 |
1.13 |
| KLA/ATP | 1.02 |
.95 |
.91 |
.86 |
.78 |
.84 |
.94 |
|
| Prkcb1 | 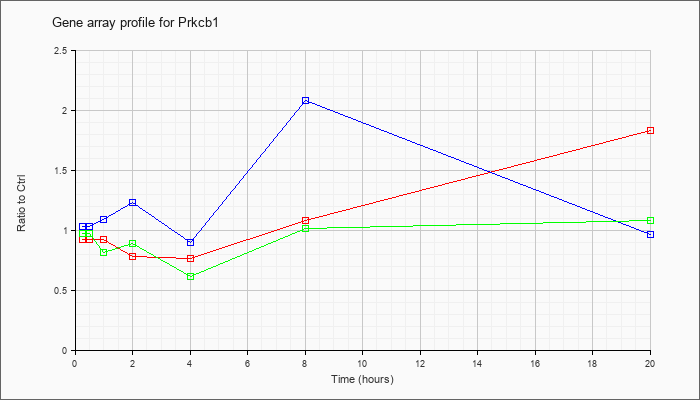 |
NM_008855 |
protein kinase C, beta 1 (Prkcb1), mRNA [NM_008855] |
KLA | .92 |
.95 |
.93 |
.78 |
.77 |
1.08 |
1.83 |
| ATP | 1.03 |
1.12 |
1.09 |
1.23 |
.90 |
2.08 |
.96 |
| KLA/ATP | .98 |
.99 |
.82 |
.89 |
.62 |
1.01 |
1.08 |
|
| Prkcc | 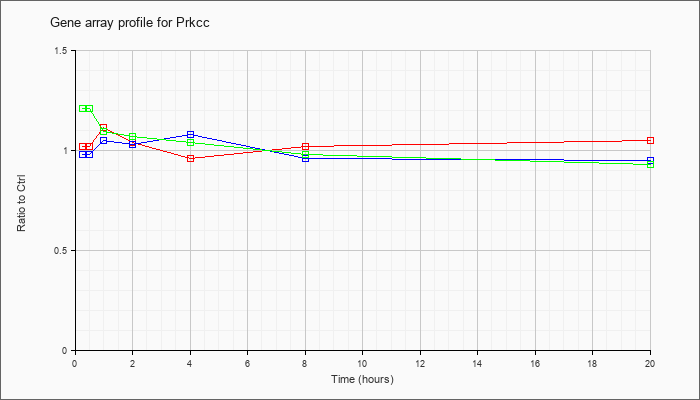 |
NM_011102 |
protein kinase C, gamma (Prkcc), mRNA [NM_011102] |
KLA | 1.02 |
1.04 |
1.11 |
1.04 |
.96 |
1.02 |
1.05 |
| ATP | .98 |
.98 |
1.05 |
1.03 |
1.08 |
.96 |
.95 |
| KLA/ATP | 1.21 |
1.16 |
1.09 |
1.07 |
1.04 |
.98 |
.93 |
|
| Rbx1 | 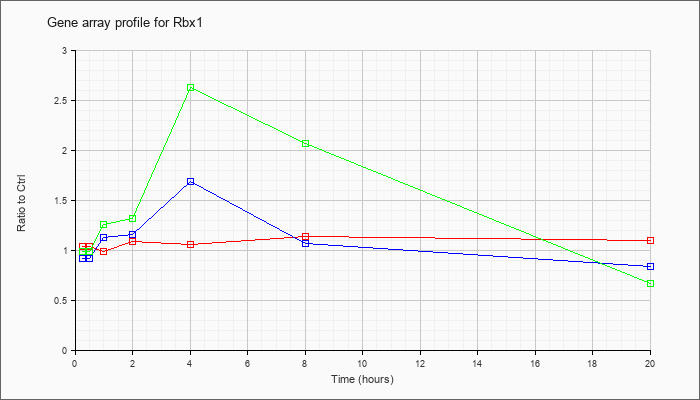 |
NM_019712 |
ring-box 1 (Rbx1), mRNA [NM_019712] |
KLA | 1.04 |
1.01 |
.98 |
1.09 |
1.06 |
1.14 |
1.09 |
| ATP | .92 |
.89 |
1.13 |
1.16 |
1.68 |
1.07 |
.83 |
| KLA/ATP | .99 |
.99 |
1.25 |
1.31 |
2.63 |
2.06 |
.67 |
|
| Rela | 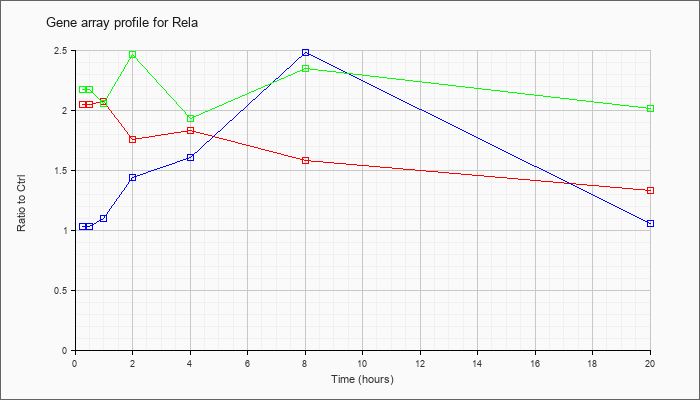 |
NM_009045 |
v-rel reticuloendotheliosis viral oncogene homolog A (avian) (Rela), mRNA [NM_009045] |
KLA | 2.05 |
2.05 |
2.07 |
1.75 |
1.83 |
1.58 |
1.33 |
| ATP | 1.03 |
1.08 |
1.10 |
1.44 |
1.61 |
2.48 |
1.05 |
| KLA/ATP | 2.17 |
2.27 |
2.05 |
2.47 |
1.93 |
2.35 |
2.02 |
|
| Rps6 | 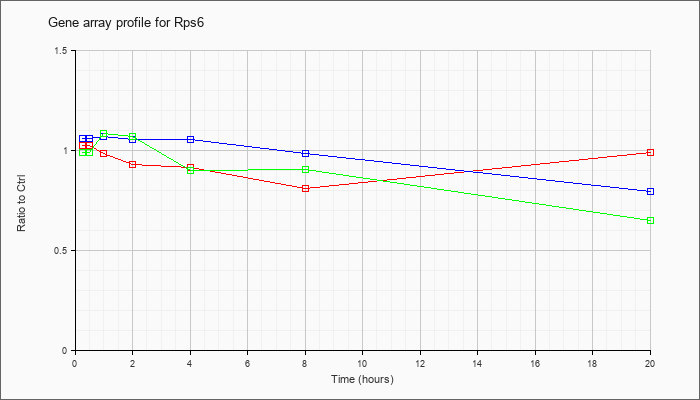 |
AK013825 |
adult male hippocampus cDNA, RIKEN full-length enriched library, clone:2900086I01 product:ribosomal protein S6, full insert sequence. [AK013825] |
KLA | 1.04 |
1.06 |
1.03 |
.99 |
.94 |
.96 |
1.26 |
| ATP | 1.01 |
1.00 |
1.04 |
.99 |
.99 |
.94 |
1.12 |
| KLA/ATP | 1.02 |
1.08 |
.95 |
1.04 |
.94 |
.97 |
.90 |
|
| Rps6 |  |
NM_009096 |
ribosomal protein S6 (Rps6), mRNA [NM_009096] |
KLA | 1.01 |
1.01 |
.94 |
.87 |
.89 |
.66 |
.72 |
| ATP | 1.11 |
1.07 |
1.10 |
1.12 |
1.12 |
1.03 |
.47 |
| KLA/ATP | .96 |
1.04 |
1.22 |
1.10 |
.86 |
.84 |
.40 |
|
| Rps6kb1 | 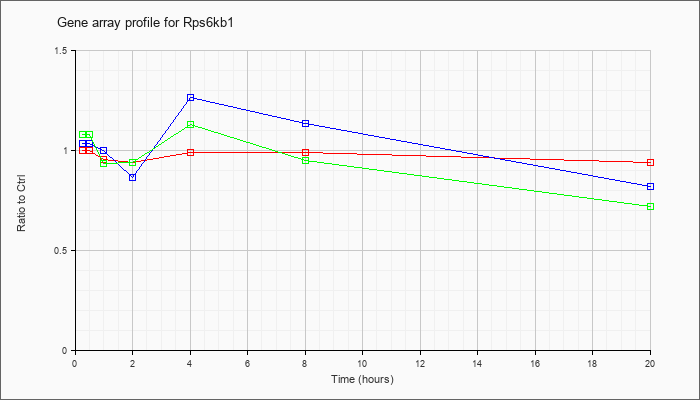 |
NM_001114334 |
ribosomal protein S6 kinase, polypeptide 1 (Rps6kb1), transcript variant 1, mRNA [NM_001114334] |
KLA | .99 |
1.00 |
.90 |
.89 |
.99 |
.99 |
.92 |
| ATP | 1.01 |
.96 |
1.00 |
.79 |
1.36 |
1.19 |
.72 |
| KLA/ATP | 1.08 |
.99 |
.93 |
.92 |
1.18 |
.90 |
.56 |
|
| Rps6kb1 |  |
NM_028259 |
ribosomal protein S6 kinase, polypeptide 1 (Rps6kb1), transcript variant 2, mRNA [NM_028259] |
KLA | 1.01 |
.93 |
1.07 |
1.04 |
.99 |
.99 |
.98 |
| ATP | 1.09 |
1.05 |
1.00 |
1.01 |
1.08 |
1.03 |
1.03 |
| KLA/ATP | 1.07 |
1.04 |
.94 |
.98 |
1.03 |
1.05 |
1.05 |
|
| Rps6kb2 | 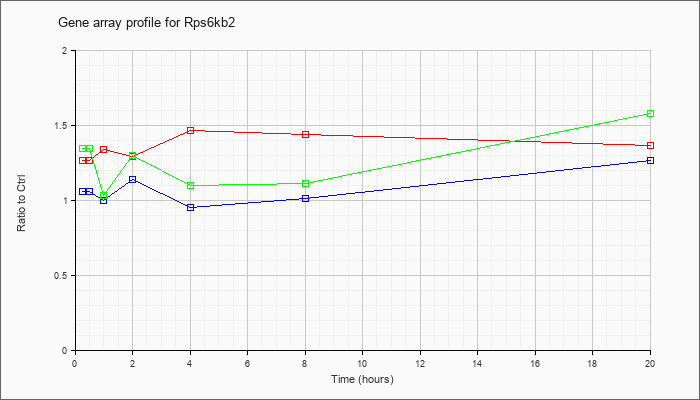 |
AK090093 |
kidney CCL-142 RAG cDNA, RIKEN full-length enriched library, clone:G430107K15 product:ribosomal protein S6 kinase, 70kD, polypeptide 2, full insert sequence [AK090093] |
KLA | .97 |
1.01 |
.98 |
.97 |
.98 |
1.04 |
1.07 |
| ATP | 1.00 |
1.08 |
1.06 |
1.09 |
1.11 |
1.06 |
1.03 |
| KLA/ATP | 1.05 |
.96 |
.86 |
1.03 |
1.40 |
1.07 |
.99 |
|
| Rps6kb2 |  |
NM_021485 |
ribosomal protein S6 kinase, polypeptide 2 (Rps6kb2), mRNA [NM_021485] |
KLA | 1.36 |
1.40 |
1.45 |
1.40 |
1.63 |
1.57 |
1.46 |
| ATP | 1.08 |
1.19 |
.97 |
1.15 |
.90 |
.99 |
1.34 |
| KLA/ATP | 1.44 |
1.55 |
1.09 |
1.39 |
1.00 |
1.12 |
1.77 |
|
Serpine1
| 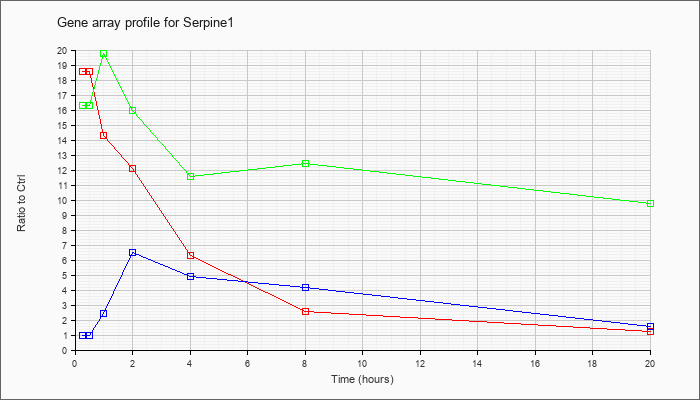 |
AK040662 |
adult male aorta and vein cDNA, RIKEN full-length enriched library, clone:A530009B11 product:serine (or cysteine) proteinase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 1, full insert sequence. [AK040662] |
KLA | 1.12 |
1.08 |
1.10 |
1.03 |
1.00 |
1.01 |
1.05 |
| ATP | 1.00 |
1.09 |
1.39 |
1.44 |
1.05 |
.95 |
.99 |
| KLA/ATP | 1.09 |
1.10 |
1.02 |
1.03 |
1.08 |
1.19 |
1.02 |
|
Serpine1
|  |
NM_008871 |
serine (or cysteine) peptidase inhibitor, clade E, member 1 (Serpine1), mRNA [NM_008871] |
KLA | 36.00 |
31.05 |
27.47 |
23.20 |
11.59 |
4.14 |
1.47 |
| ATP | .97 |
1.15 |
3.43 |
11.58 |
8.77 |
7.42 |
2.16 |
| KLA/ATP | 31.47 |
33.70 |
38.54 |
30.90 |
22.06 |
23.68 |
18.57 |
|
| Slc2a1 | 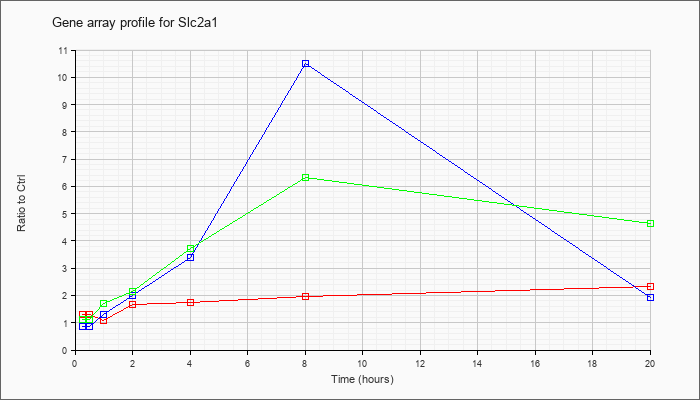 |
NM_011400 |
solute carrier family 2 (facilitated glucose transporter), member 1 (Slc2a1), mRNA [NM_011400] |
KLA | 1.31 |
1.46 |
1.10 |
1.68 |
1.75 |
1.95 |
2.34 |
| ATP | .88 |
.90 |
1.32 |
1.99 |
3.40 |
10.49 |
1.93 |
| KLA/ATP | 1.13 |
1.25 |
1.71 |
2.14 |
3.71 |
6.31 |
4.63 |
|
| Stat3 | 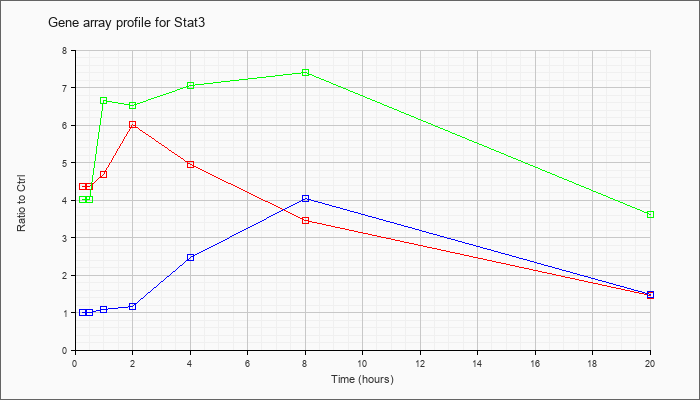 |
NM_213659 |
signal transducer and activator of transcription 3 (Stat3), transcript variant 1, mRNA [NM_213659] |
KLA | 4.36 |
4.42 |
4.67 |
6.02 |
4.95 |
3.45 |
1.46 |
| ATP | 1.01 |
.97 |
1.07 |
1.17 |
2.47 |
4.04 |
1.47 |
| KLA/ATP | 4.02 |
4.57 |
6.65 |
6.51 |
7.06 |
7.39 |
3.61 |
|
| Tceb1 | 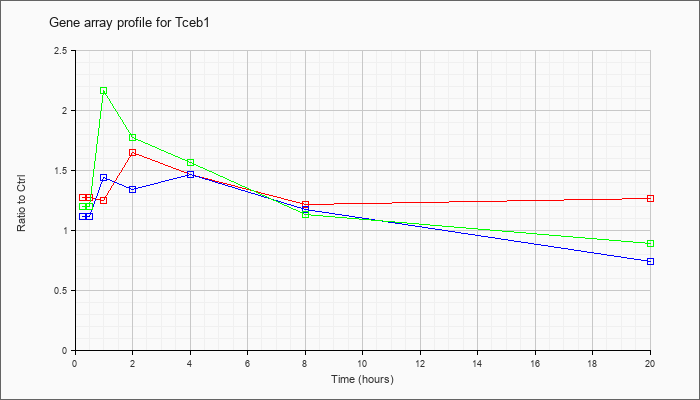 |
NM_026456 |
transcription elongation factor B (SIII), polypeptide 1 (Tceb1), mRNA [NM_026456] |
KLA | 1.27 |
1.21 |
1.25 |
1.65 |
1.46 |
1.21 |
1.26 |
| ATP | 1.11 |
1.01 |
1.44 |
1.34 |
1.46 |
1.17 |
.74 |
| KLA/ATP | 1.20 |
1.48 |
2.16 |
1.77 |
1.56 |
1.13 |
.89 |
|
| Tceb2 | 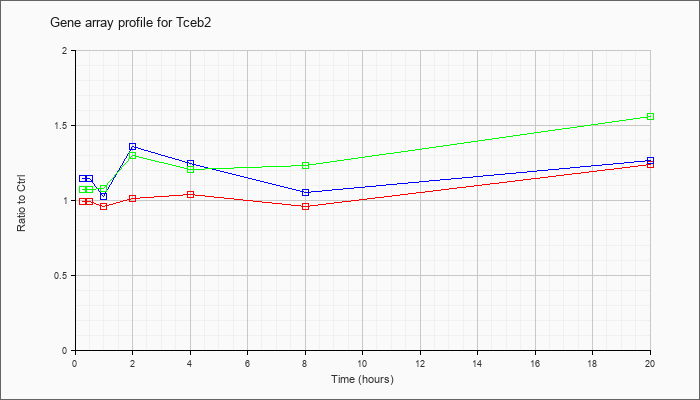 |
NM_026305 |
transcription elongation factor B (SIII), polypeptide 2 (Tceb2), mRNA [NM_026305] |
KLA | .99 |
1.00 |
.96 |
1.01 |
1.04 |
.96 |
1.24 |
| ATP | 1.15 |
1.17 |
1.03 |
1.36 |
1.25 |
1.05 |
1.27 |
| KLA/ATP | 1.07 |
1.13 |
1.08 |
1.30 |
1.21 |
1.23 |
1.56 |
|
| Tek | 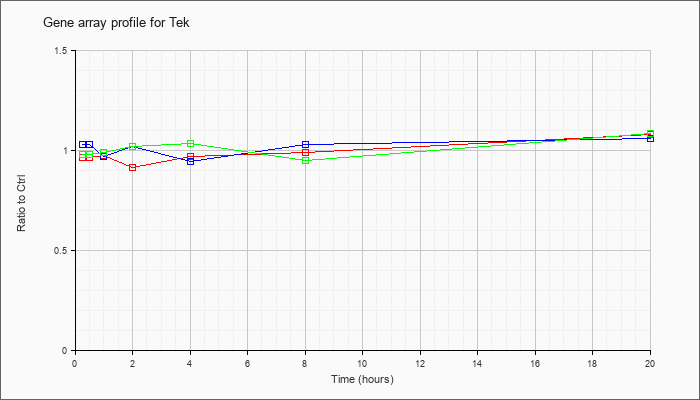 |
AK053336 |
0 day neonate eyeball cDNA, RIKEN full-length enriched library, clone:E130010K24 product:endothelial-specific receptor tyrosine kinase, full insert sequence. [AK053336] |
KLA | .93 |
.98 |
.93 |
.90 |
.96 |
1.01 |
1.10 |
| ATP | 1.01 |
.97 |
.93 |
.99 |
.91 |
.98 |
.99 |
| KLA/ATP | 1.02 |
1.06 |
.95 |
.93 |
.92 |
.91 |
.92 |
|
| Tek |  |
NM_013690 |
endothelial-specific receptor tyrosine kinase (Tek), mRNA [NM_013690] |
KLA | 1.00 |
.95 |
1.02 |
.93 |
.98 |
.97 |
1.06 |
| ATP | 1.05 |
1.07 |
1.01 |
1.05 |
.98 |
1.08 |
1.13 |
| KLA/ATP | .94 |
.96 |
1.03 |
1.11 |
1.15 |
.99 |
1.25 |
|
| Tfrc | 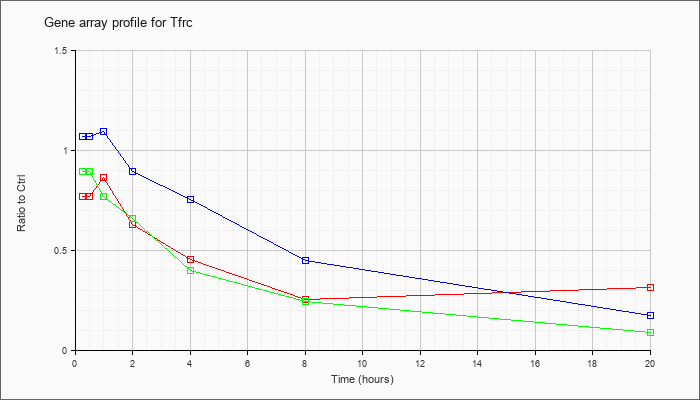 |
NM_011638 |
transferrin receptor (Tfrc), mRNA [NM_011638] |
KLA | .77 |
.79 |
.86 |
.63 |
.45 |
.25 |
.31 |
| ATP | 1.07 |
1.11 |
1.09 |
.89 |
.75 |
.45 |
.17 |
| KLA/ATP | .89 |
.85 |
.77 |
.66 |
.40 |
.24 |
.09 |
|
| Timp1 | 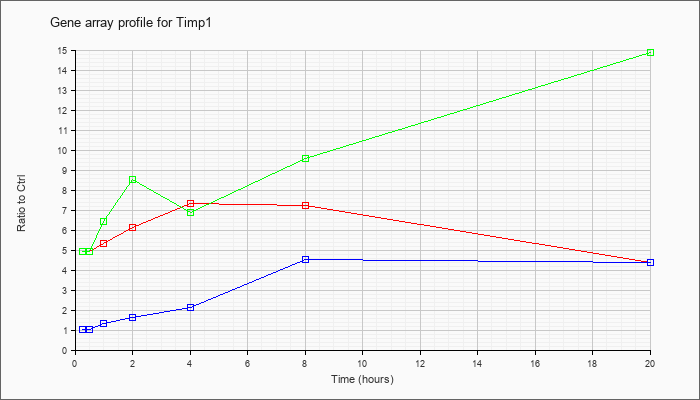 |
NM_001044384 |
tissue inhibitor of metalloproteinase 1 (Timp1), transcript variant 1, mRNA [NM_001044384] |
KLA | 4.91 |
5.11 |
5.35 |
6.12 |
7.31 |
7.24 |
4.37 |
| ATP | 1.02 |
1.10 |
1.31 |
1.62 |
2.13 |
4.52 |
4.40 |
| KLA/ATP | 4.93 |
5.94 |
6.45 |
8.54 |
6.89 |
9.58 |
14.87 |
|
| Tlr4 | 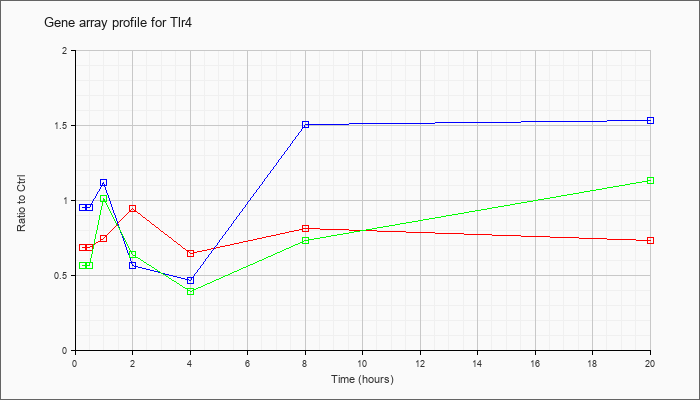 |
NM_021297 |
toll-like receptor 4 (Tlr4), mRNA [NM_021297] |
KLA | .69 |
.61 |
.75 |
.94 |
.64 |
.81 |
.73 |
| ATP | .95 |
.90 |
1.12 |
.56 |
.47 |
1.51 |
1.53 |
| KLA/ATP | .57 |
.62 |
1.01 |
.64 |
.39 |
.73 |
1.13 |
|
| Trf | 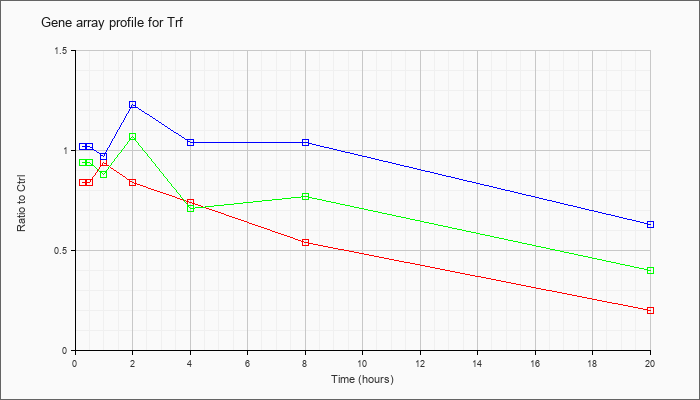 |
NM_133977 |
transferrin (Trf), mRNA [NM_133977] |
KLA | .84 |
.95 |
.94 |
.84 |
.74 |
.54 |
.20 |
| ATP | 1.02 |
1.05 |
.97 |
1.23 |
1.04 |
1.04 |
.63 |
| KLA/ATP | .94 |
.93 |
.88 |
1.07 |
.71 |
.77 |
.40 |
|
| Vegfa | 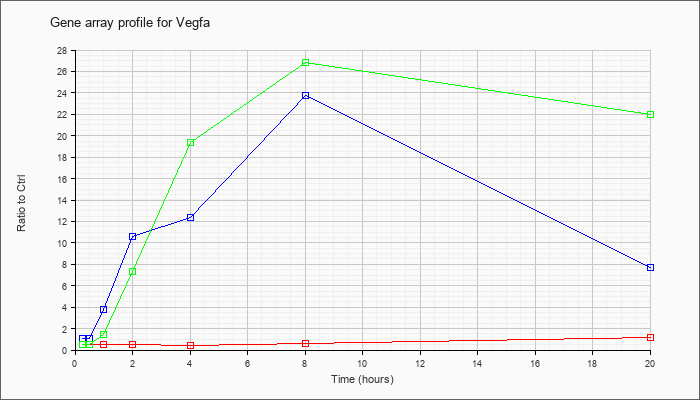 |
NM_001025250 |
vascular endothelial growth factor A (Vegfa), transcript variant 1, mRNA [NM_001025250] |
KLA | .49 |
.48 |
.46 |
.46 |
.39 |
.64 |
1.18 |
| ATP | 1.05 |
1.37 |
3.82 |
10.55 |
12.51 |
24.20 |
7.85 |
| KLA/ATP | .48 |
.59 |
1.45 |
7.42 |
19.75 |
27.42 |
22.35 |
|
| Vegfa |  |
NM_001025257 |
vascular endothelial growth factor A (Vegfa), transcript variant 3, mRNA [NM_001025257] |
KLA | .72 |
.71 |
.66 |
.70 |
.65 |
.80 |
1.22 |
| ATP | 1.01 |
1.21 |
3.24 |
11.12 |
11.09 |
18.59 |
5.43 |
| KLA/ATP | .68 |
.71 |
1.47 |
5.94 |
15.06 |
20.32 |
17.31 |
|
| Vhlh | 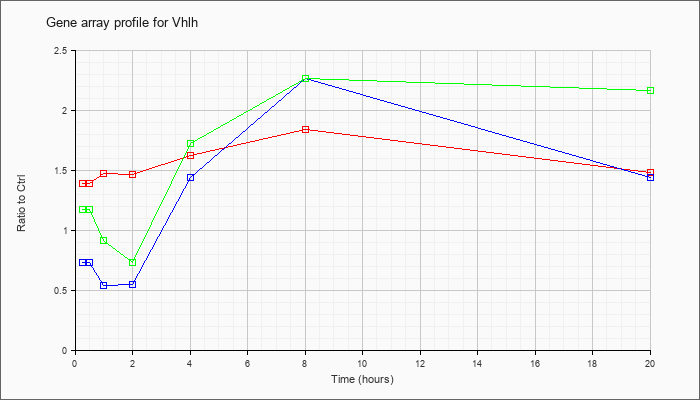 |
S76748 |
gb|mVHLh1=von Hippel-Lindau disease gene [mice, mRNA, 2757 nt]. [S76748] |
KLA | 1.39 |
1.39 |
1.47 |
1.46 |
1.62 |
1.84 |
1.48 |
| ATP | .73 |
.64 |
.54 |
.55 |
1.44 |
2.26 |
1.44 |
| KLA/ATP | 1.17 |
1.13 |
.91 |
.73 |
1.72 |
2.26 |
2.16 |
|