| Gene symbol |
Time course plot |
Accession |
Description |
Treatment |
15min |
30min |
1hr |
2hr |
4hr |
8hr |
20hr |
| Akt1 | 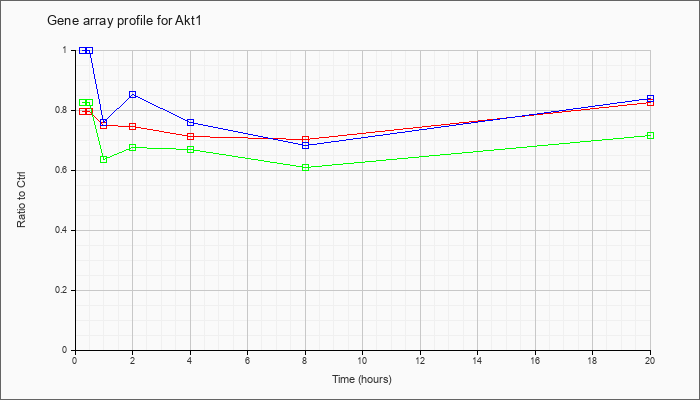 |
NM_009652 |
thymoma viral proto-oncogene 1 (Akt1), mRNA [NM_009652] |
KLA | .80 |
.80 |
.75 |
.75 |
.71 |
.70 |
.82 |
| ATP | 1.00 |
1.02 |
.76 |
.85 |
.76 |
.68 |
.84 |
| KLA/ATP | .82 |
.80 |
.64 |
.68 |
.67 |
.61 |
.72 |
|
| Akt2 | 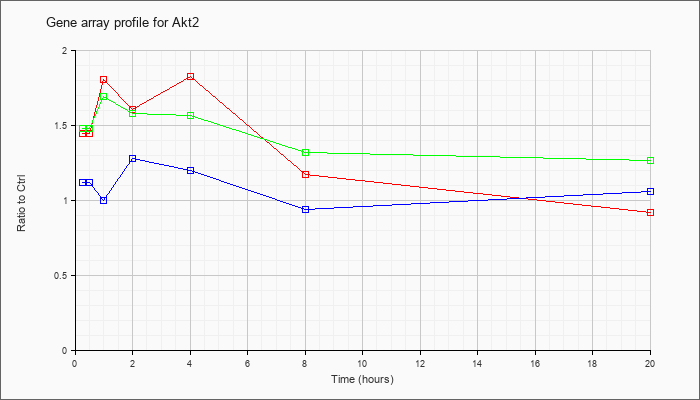 |
NM_007434 |
thymoma viral proto-oncogene 2 (Akt2), transcript variant 2, mRNA [NM_007434] |
KLA | 1.45 |
1.49 |
1.81 |
1.61 |
1.83 |
1.17 |
.92 |
| ATP | 1.12 |
1.12 |
1.00 |
1.28 |
1.20 |
.94 |
1.06 |
| KLA/ATP | 1.48 |
1.70 |
1.69 |
1.58 |
1.57 |
1.32 |
1.27 |
|
| Akt3 | 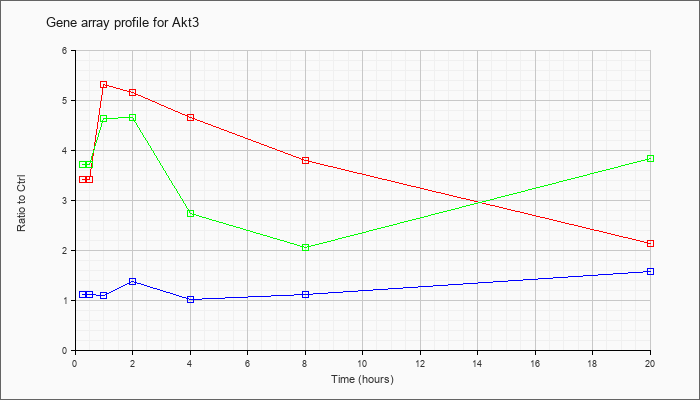 |
NM_011785 |
thymoma viral proto-oncogene 3 (Akt3), mRNA [NM_011785] |
KLA | 3.42 |
3.45 |
5.31 |
5.16 |
4.65 |
3.80 |
2.12 |
| ATP | 1.12 |
1.22 |
1.10 |
1.37 |
1.02 |
1.11 |
1.56 |
| KLA/ATP | 3.72 |
4.17 |
4.62 |
4.64 |
2.74 |
2.05 |
3.82 |
|
| Atf2 | 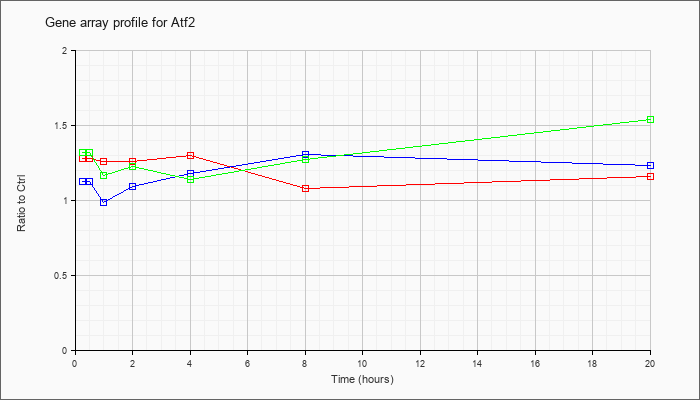 |
AK047405 |
10 days neonate cerebellum cDNA, RIKEN full-length enriched library, clone:B930056P03 product:activating transcription factor 2, full insert sequence. [AK047405] |
KLA | 1.01 |
1.00 |
1.02 |
1.04 |
1.03 |
1.03 |
1.00 |
| ATP | .99 |
1.00 |
.95 |
1.01 |
1.04 |
.98 |
1.03 |
| KLA/ATP | 1.07 |
1.00 |
.98 |
.99 |
1.00 |
1.03 |
.93 |
|
| Atf2 |  |
NM_001025093 |
activating transcription factor 2 (Atf2), transcript variant 1, mRNA [NM_001025093] |
KLA | 1.35 |
1.30 |
1.32 |
1.31 |
1.37 |
1.09 |
1.20 |
| ATP | 1.15 |
1.17 |
.99 |
1.11 |
1.22 |
1.39 |
1.28 |
| KLA/ATP | 1.38 |
1.42 |
1.21 |
1.28 |
1.17 |
1.33 |
1.69 |
|
| Atf4 | 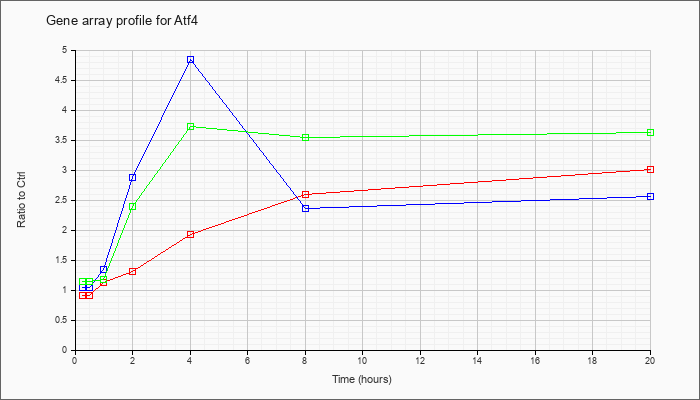 |
NM_009716 |
activating transcription factor 4 (Atf4), mRNA [NM_009716] |
KLA | .91 |
1.06 |
1.13 |
1.31 |
1.92 |
2.59 |
3.01 |
| ATP | 1.05 |
1.14 |
1.34 |
2.88 |
4.84 |
2.36 |
2.56 |
| KLA/ATP | 1.14 |
1.08 |
1.17 |
2.40 |
3.72 |
3.54 |
3.62 |
|
| Bag4 | 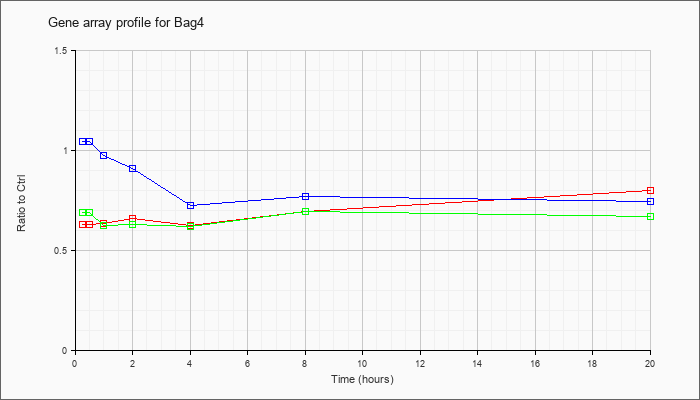 |
NM_026121 |
BCL2-associated athanogene 4 (Bag4), mRNA [NM_026121] |
KLA | .63 |
.63 |
.64 |
.66 |
.63 |
.70 |
.80 |
| ATP | 1.05 |
1.14 |
.98 |
.91 |
.73 |
.77 |
.75 |
| KLA/ATP | .69 |
.70 |
.63 |
.63 |
.62 |
.70 |
.67 |
|
| Bcl3 | 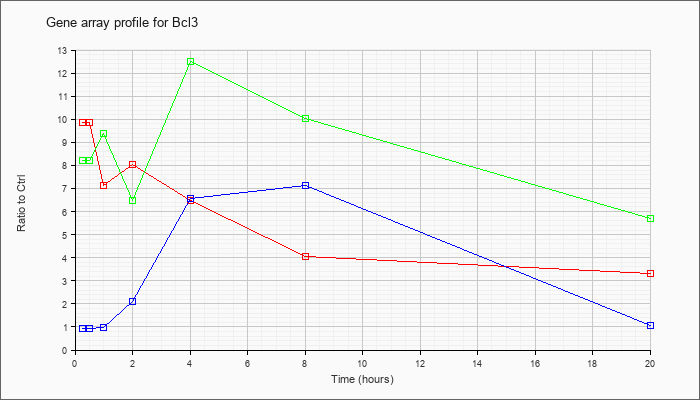 |
NM_033601 |
B-cell leukemia/lymphoma 3 (Bcl3), mRNA [NM_033601] |
KLA | 9.86 |
8.97 |
7.13 |
8.05 |
6.48 |
4.07 |
3.33 |
| ATP | .93 |
.84 |
.99 |
2.12 |
6.57 |
7.15 |
1.07 |
| KLA/ATP | 8.21 |
9.05 |
9.37 |
6.50 |
12.52 |
10.04 |
5.68 |
|
| Birc2 | 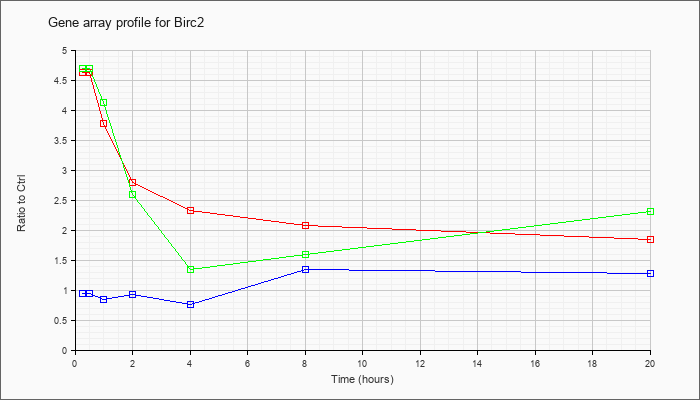 |
NM_007465 |
baculoviral IAP repeat-containing 2 (Birc2), mRNA [NM_007465] |
KLA | 4.63 |
4.59 |
3.77 |
2.79 |
2.32 |
2.07 |
1.84 |
| ATP | .95 |
.88 |
.84 |
.92 |
.76 |
1.34 |
1.28 |
| KLA/ATP | 4.70 |
4.93 |
4.13 |
2.60 |
1.34 |
1.60 |
2.31 |
|
| Birc3 | 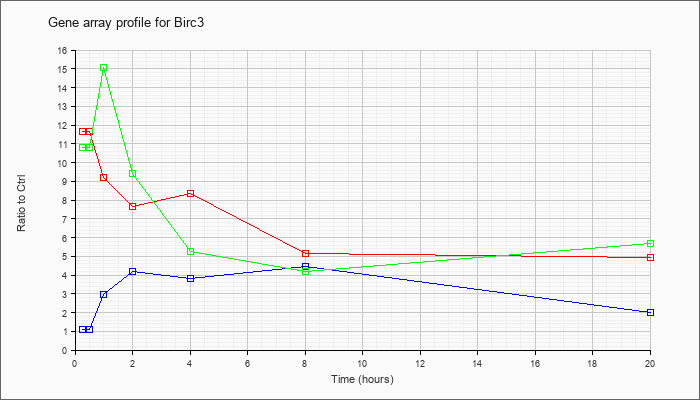 |
BC011338 |
baculoviral IAP repeat-containing 3, mRNA (cDNA clone MGC:18386 IMAGE:3661563), complete cds [BC011338] |
KLA | 5.48 |
5.46 |
4.94 |
4.16 |
4.91 |
3.36 |
3.74 |
| ATP | 1.05 |
1.08 |
4.21 |
4.28 |
2.59 |
2.55 |
1.57 |
| KLA/ATP | 6.78 |
8.91 |
11.01 |
4.87 |
2.28 |
2.12 |
3.66 |
|
| Birc3 |  |
NM_007464 |
baculoviral IAP repeat-containing 3 (Birc3), mRNA [NM_007464] |
KLA | 17.83 |
15.88 |
13.48 |
11.14 |
11.79 |
6.90 |
6.10 |
| ATP | 1.14 |
1.16 |
1.70 |
4.07 |
5.00 |
6.33 |
2.41 |
| KLA/ATP | 14.77 |
16.44 |
19.09 |
13.91 |
8.27 |
6.22 |
7.73 |
|
| Casp3 | 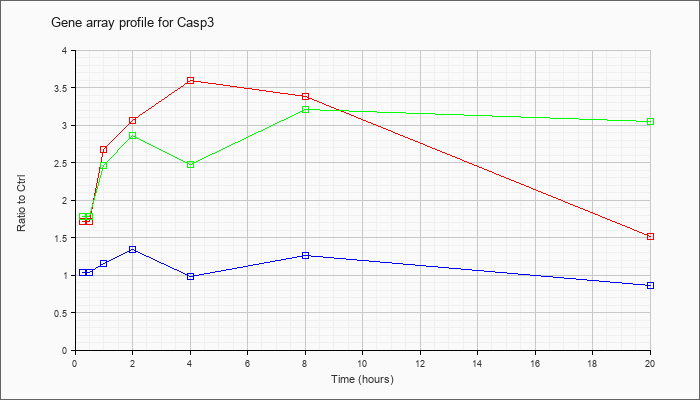 |
NM_009810 |
caspase 3 (Casp3), mRNA [NM_009810] |
KLA | 1.66 |
1.94 |
2.47 |
2.61 |
3.01 |
2.96 |
1.44 |
| ATP | .90 |
.99 |
1.17 |
1.30 |
1.01 |
1.17 |
.86 |
| KLA/ATP | 1.55 |
1.87 |
2.40 |
2.35 |
2.27 |
2.84 |
2.52 |
|
| Casp3 |  |
U49929 |
ICE-like cysteine protease (Lice) mRNA, complete cds. [U49929] |
KLA | 1.71 |
1.87 |
2.69 |
3.11 |
3.64 |
3.42 |
1.52 |
| ATP | 1.04 |
1.07 |
1.16 |
1.34 |
.98 |
1.27 |
.86 |
| KLA/ATP | 1.80 |
1.96 |
2.46 |
2.91 |
2.50 |
3.24 |
3.10 |
|
| Casp7 | 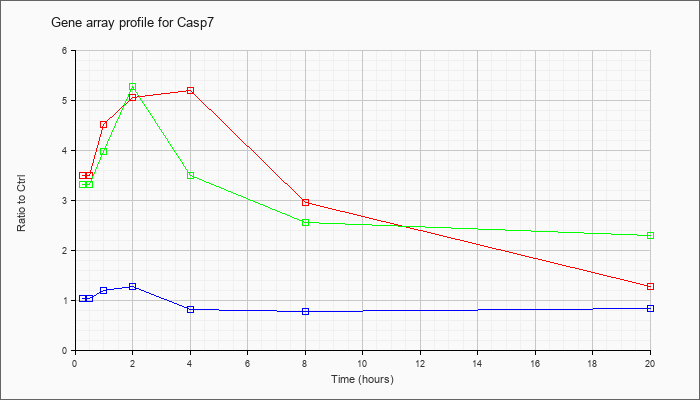 |
NM_007611 |
caspase 7 (Casp7), mRNA [NM_007611] |
KLA | 3.50 |
3.60 |
4.52 |
5.05 |
5.20 |
2.96 |
1.27 |
| ATP | 1.03 |
1.14 |
1.20 |
1.27 |
.82 |
.77 |
.84 |
| KLA/ATP | 3.32 |
3.86 |
3.98 |
5.27 |
3.49 |
2.55 |
2.30 |
|
| Casp8 | 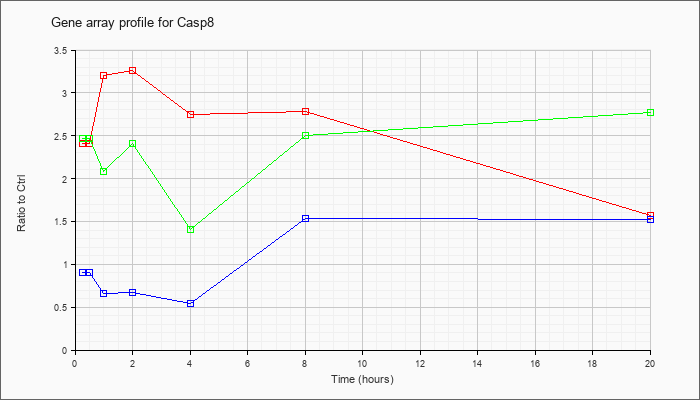 |
NM_009812 |
caspase 8 (Casp8), transcript variant 1, mRNA [NM_009812] |
KLA | 2.41 |
2.53 |
3.20 |
3.26 |
2.75 |
2.78 |
1.57 |
| ATP | .91 |
.96 |
.66 |
.67 |
.54 |
1.53 |
1.52 |
| KLA/ATP | 2.47 |
2.52 |
2.08 |
2.41 |
1.41 |
2.50 |
2.77 |
|
| Ccl12 | 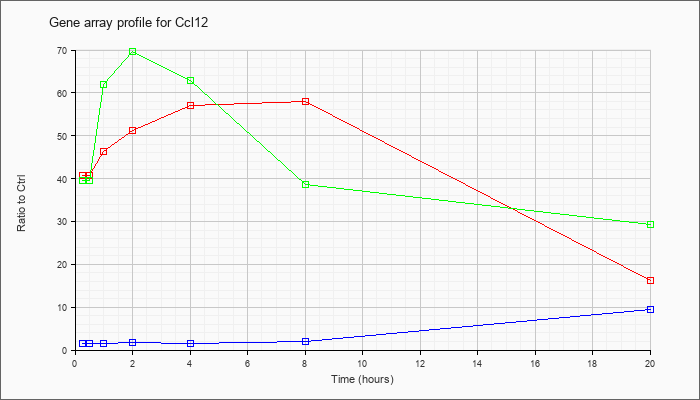 |
NM_011331 |
chemokine (C-C motif) ligand 12 (Ccl12), mRNA [NM_011331] |
KLA | 40.77 |
33.71 |
46.23 |
51.22 |
57.13 |
57.92 |
16.26 |
| ATP | 1.41 |
1.14 |
1.54 |
1.79 |
1.41 |
1.89 |
9.34 |
| KLA/ATP | 39.52 |
43.18 |
61.96 |
69.72 |
62.95 |
38.70 |
29.25 |
|
| Ccl2 | 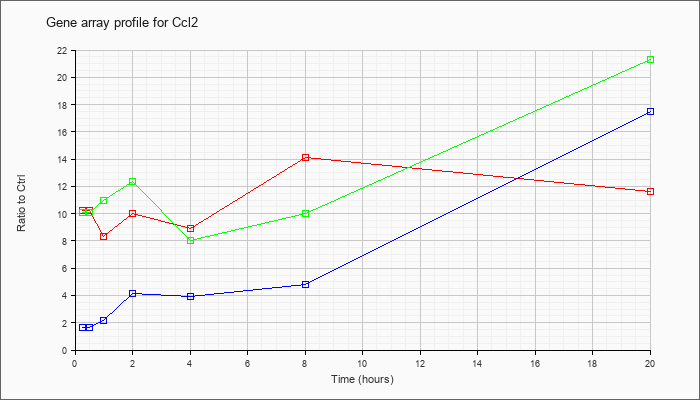 |
NM_011333 |
chemokine (C-C motif) ligand 2 (Ccl2), mRNA [NM_011333] |
KLA | 10.26 |
9.30 |
8.35 |
10.02 |
8.92 |
14.08 |
11.59 |
| ATP | 1.62 |
1.68 |
2.20 |
4.16 |
3.92 |
4.84 |
17.45 |
| KLA/ATP | 10.09 |
10.31 |
10.95 |
12.32 |
8.00 |
10.04 |
21.29 |
|
| Ccl20 | 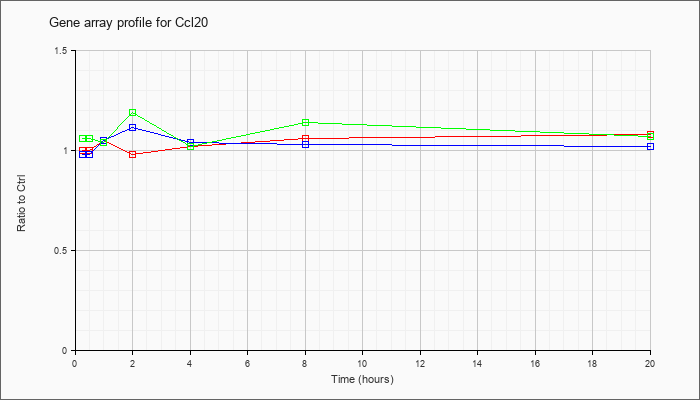 |
NM_016960 |
chemokine (C-C motif) ligand 20 (Ccl20), mRNA [NM_016960] |
KLA | 1.00 |
1.03 |
1.05 |
.98 |
1.02 |
1.06 |
1.08 |
| ATP | .98 |
1.01 |
1.05 |
1.11 |
1.04 |
1.03 |
1.02 |
| KLA/ATP | 1.06 |
.97 |
1.04 |
1.19 |
1.02 |
1.14 |
1.07 |
|
| Ccl5 | 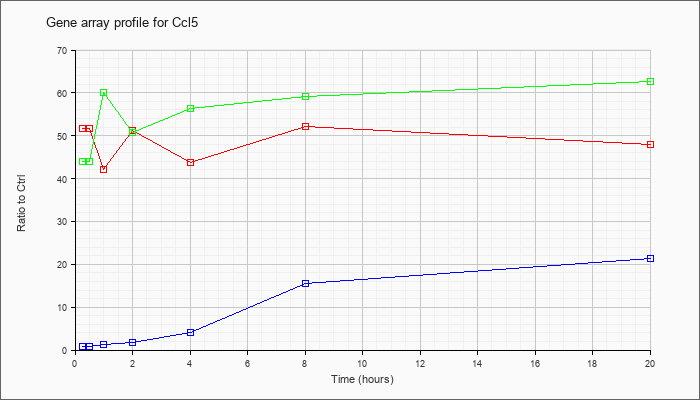 |
M77747 |
rantes mRNA, complete cds. [M77747] |
KLA | 63.18 |
60.41 |
56.25 |
63.63 |
58.00 |
66.51 |
67.20 |
| ATP | .91 |
1.08 |
1.25 |
1.90 |
4.34 |
16.10 |
26.07 |
| KLA/ATP | 55.92 |
63.34 |
72.91 |
65.68 |
70.81 |
71.97 |
81.34 |
|
| Ccl5 |  |
NM_013653 |
chemokine (C-C motif) ligand 5 (Ccl5), mRNA [NM_013653] |
KLA | 40.31 |
35.22 |
27.92 |
38.59 |
29.29 |
37.98 |
28.83 |
| ATP | .86 |
.92 |
1.43 |
1.66 |
3.86 |
15.09 |
16.66 |
| KLA/ATP | 32.14 |
31.72 |
47.19 |
35.89 |
42.00 |
46.15 |
43.89 |
|
| Cebpb | 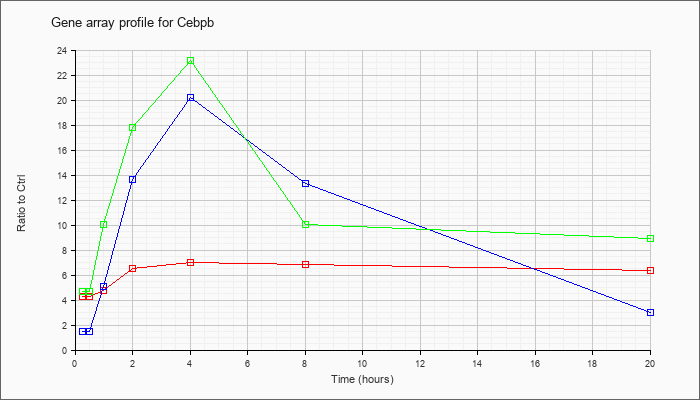 |
NM_009883 |
CCAAT/enhancer binding protein (C/EBP), beta (Cebpb), mRNA [NM_009883] |
KLA | 4.23 |
5.09 |
5.13 |
5.70 |
7.11 |
6.62 |
5.94 |
| ATP | 1.52 |
2.60 |
4.55 |
14.19 |
16.93 |
10.86 |
3.06 |
| KLA/ATP | 4.97 |
7.06 |
8.37 |
18.16 |
20.66 |
8.78 |
8.37 |
|
| Cebpb |  |
X62600 |
gb|M.musculus mRNA for C/EBP beta. [X62600] |
KLA | 4.28 |
4.63 |
4.35 |
7.32 |
6.89 |
7.11 |
6.82 |
| ATP | 1.41 |
2.03 |
5.60 |
13.16 |
23.55 |
15.76 |
3.02 |
| KLA/ATP | 4.35 |
6.27 |
11.70 |
17.46 |
25.69 |
11.34 |
9.42 |
|
| Cflar | 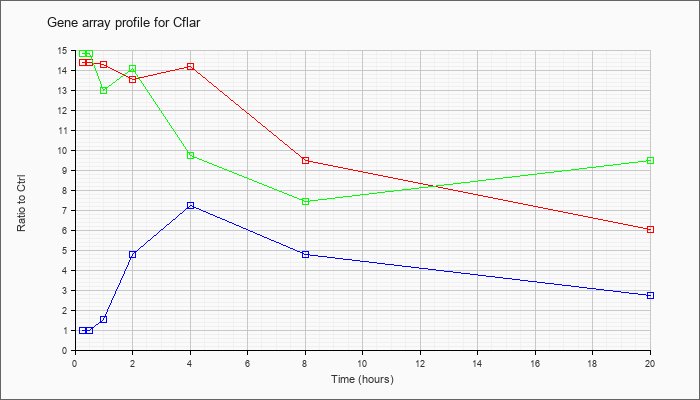 |
NM_009805 |
CASP8 and FADD-like apoptosis regulator (Cflar), transcript variant 2, mRNA [NM_009805] |
KLA | 18.63 |
20.03 |
20.30 |
17.44 |
17.99 |
11.88 |
7.25 |
| ATP | .95 |
1.03 |
1.20 |
2.78 |
4.04 |
3.45 |
3.30 |
| KLA/ATP | 19.31 |
19.09 |
11.65 |
13.57 |
7.62 |
6.20 |
11.21 |
|
| Cflar |  |
NM_207653 |
CASP8 and FADD-like apoptosis regulator (Cflar), transcript variant 1, mRNA [NM_207653] |
KLA | 12.32 |
12.86 |
11.83 |
11.79 |
12.14 |
8.52 |
5.42 |
| ATP | .92 |
.92 |
1.63 |
4.97 |
7.95 |
5.18 |
2.39 |
| KLA/ATP | 12.27 |
13.57 |
13.15 |
12.79 |
10.27 |
7.71 |
8.32 |
|
| Cflar |  |
Y14041 |
mRNA for CASH alpha protein. [Y14041] |
KLA | 14.31 |
13.82 |
13.05 |
13.06 |
14.36 |
9.02 |
5.92 |
| ATP | 1.18 |
1.15 |
1.66 |
6.44 |
8.99 |
5.20 |
2.73 |
| KLA/ATP | 15.53 |
17.07 |
14.00 |
17.10 |
10.76 |
8.02 |
9.97 |
|
| Chuk | 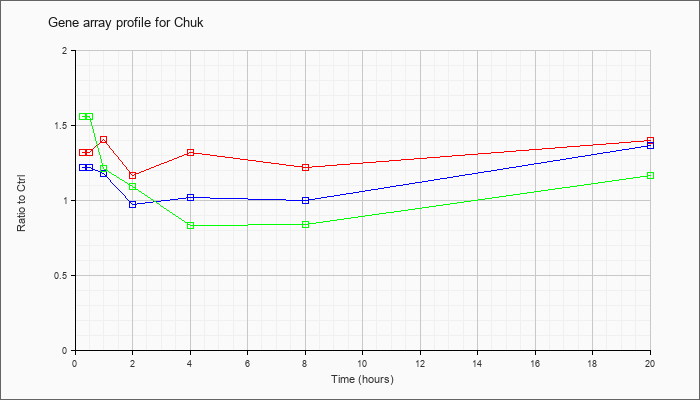 |
NM_007700 |
conserved helix-loop-helix ubiquitous kinase (Chuk), mRNA [NM_007700] |
KLA | 1.32 |
1.42 |
1.41 |
1.17 |
1.32 |
1.22 |
1.40 |
| ATP | 1.22 |
1.30 |
1.18 |
.97 |
1.02 |
1.00 |
1.37 |
| KLA/ATP | 1.56 |
1.61 |
1.21 |
1.09 |
.83 |
.84 |
1.17 |
|
| Creb1 | 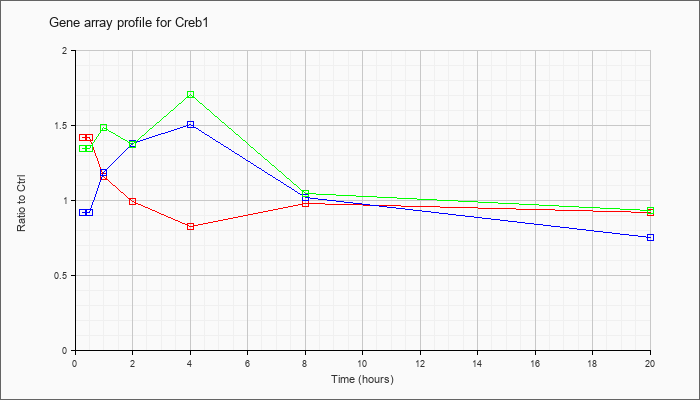 |
NM_009952 |
cAMP responsive element binding protein 1 (Creb1), transcript variant B, mRNA [NM_009952] |
KLA | 1.42 |
1.33 |
1.16 |
.99 |
.83 |
.97 |
.92 |
| ATP | .92 |
.91 |
1.18 |
1.38 |
1.51 |
1.02 |
.75 |
| KLA/ATP | 1.34 |
1.31 |
1.49 |
1.37 |
1.71 |
1.05 |
.93 |
|
| Creb3 | 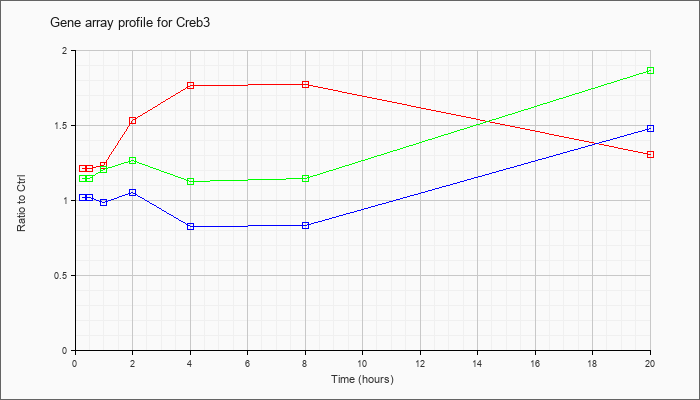 |
NM_013497 |
cAMP responsive element binding protein 3 (Creb3), mRNA [NM_013497] |
KLA | 1.21 |
1.21 |
1.23 |
1.53 |
1.77 |
1.77 |
1.31 |
| ATP | 1.02 |
1.03 |
.99 |
1.05 |
.82 |
.83 |
1.48 |
| KLA/ATP | 1.15 |
1.18 |
1.20 |
1.26 |
1.13 |
1.15 |
1.86 |
|
| Creb3l1 | 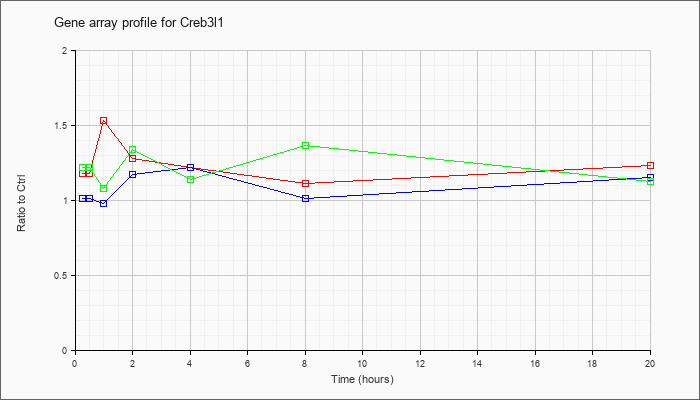 |
NM_011957 |
cAMP responsive element binding protein 3-like 1 (Creb3l1), mRNA [NM_011957] |
KLA | 1.18 |
1.17 |
1.53 |
1.28 |
1.22 |
1.11 |
1.23 |
| ATP | 1.01 |
1.09 |
.98 |
1.17 |
1.22 |
1.01 |
1.15 |
| KLA/ATP | 1.22 |
1.29 |
1.08 |
1.34 |
1.14 |
1.36 |
1.12 |
|
| Creb3l2 | 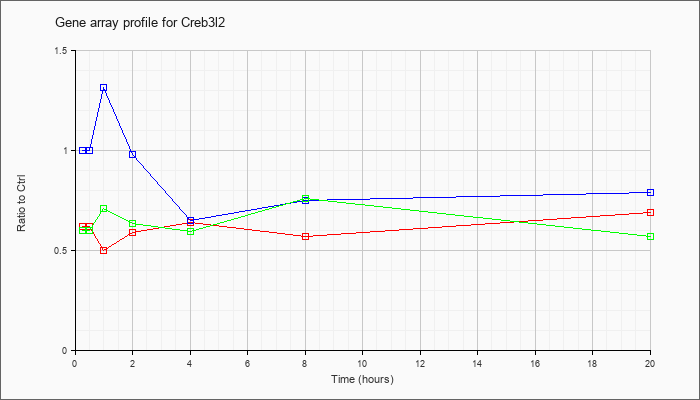 |
AK082964 |
12 days embryo spinal cord cDNA, RIKEN full-length enriched library, clone:C530025K05 product:unclassifiable, full insert sequence [AK082964] |
KLA | .44 |
.38 |
.37 |
.40 |
.45 |
.42 |
.47 |
| ATP | .90 |
.78 |
1.22 |
.79 |
.54 |
.46 |
.44 |
| KLA/ATP | .41 |
.41 |
.49 |
.43 |
.50 |
.51 |
.23 |
|
| Creb3l2 |  |
NM_178661 |
cAMP responsive element binding protein 3-like 2 (Creb3l2), mRNA [NM_178661] |
KLA | .98 |
.87 |
.76 |
.98 |
1.03 |
.87 |
1.12 |
| ATP | 1.20 |
1.18 |
1.50 |
1.35 |
.86 |
1.32 |
1.50 |
| KLA/ATP | .98 |
1.04 |
1.15 |
1.05 |
.79 |
1.27 |
1.25 |
|
| Creb3l3 | 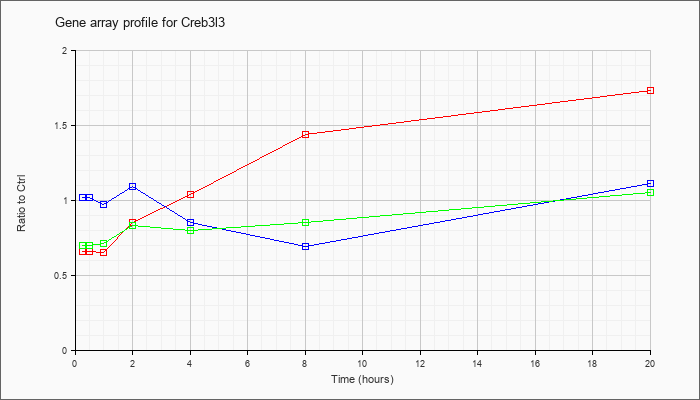 |
NM_145365 |
cAMP responsive element binding protein 3-like 3 (Creb3l3), mRNA [NM_145365] |
KLA | .66 |
.75 |
.65 |
.85 |
1.04 |
1.44 |
1.73 |
| ATP | 1.02 |
.98 |
.97 |
1.09 |
.85 |
.69 |
1.11 |
| KLA/ATP | .70 |
.66 |
.71 |
.83 |
.80 |
.85 |
1.05 |
|
| Creb3l4 | 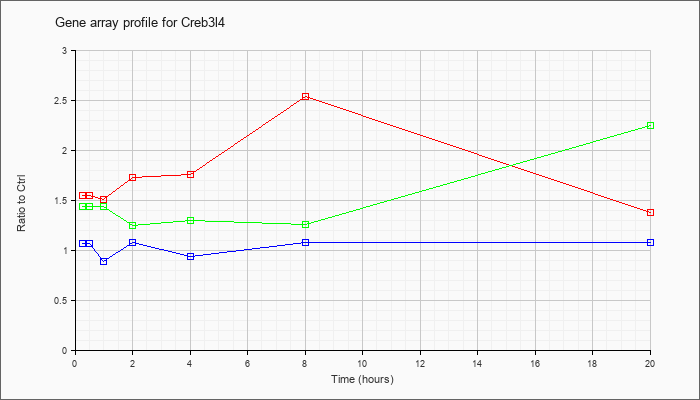 |
NM_030080 |
cAMP responsive element binding protein 3-like 4 (Creb3l4), mRNA [NM_030080] |
KLA | 1.55 |
1.42 |
1.51 |
1.73 |
1.76 |
2.54 |
1.38 |
| ATP | 1.07 |
.81 |
.89 |
1.08 |
.94 |
1.08 |
1.08 |
| KLA/ATP | 1.44 |
1.38 |
1.44 |
1.25 |
1.30 |
1.26 |
2.24 |
|
| Creb5 | 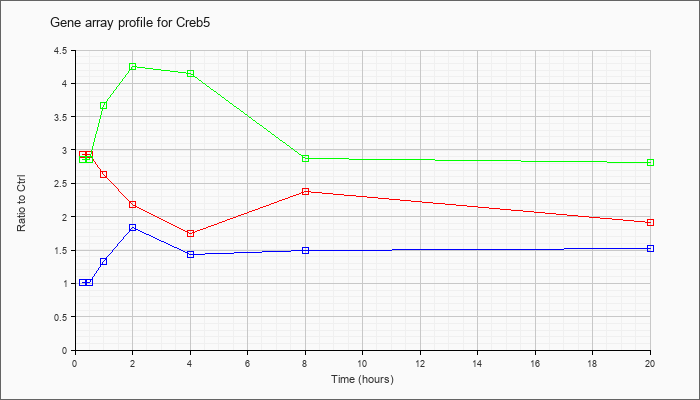 |
AK051306 |
12 days embryo spinal ganglion cDNA, RIKEN full-length enriched library, clone:D130032G03 product:unclassifiable, full insert sequence. [AK051306] |
KLA | 2.18 |
2.13 |
2.42 |
1.97 |
1.57 |
1.94 |
1.98 |
| ATP | 1.11 |
1.09 |
1.47 |
1.74 |
1.36 |
1.44 |
1.62 |
| KLA/ATP | 2.55 |
2.77 |
3.02 |
3.15 |
2.51 |
2.52 |
2.61 |
|
| Creb5 |  |
NM_172728 |
cAMP responsive element binding protein 5 (Creb5), mRNA [NM_172728] |
KLA | 3.70 |
3.60 |
2.84 |
2.39 |
1.92 |
2.81 |
1.84 |
| ATP | .92 |
.89 |
1.19 |
1.93 |
1.51 |
1.55 |
1.44 |
| KLA/ATP | 3.16 |
3.51 |
4.33 |
5.35 |
5.78 |
3.23 |
3.03 |
|
| Crebl1 | 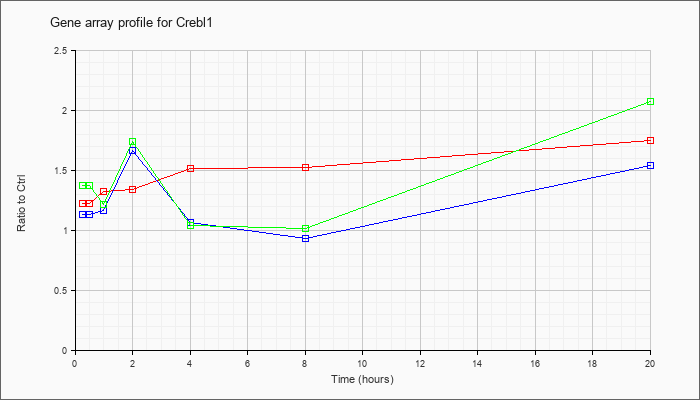 |
NM_017406 |
cAMP responsive element binding protein-like 1 (Crebl1), mRNA [NM_017406] |
KLA | 1.22 |
1.30 |
1.32 |
1.34 |
1.51 |
1.52 |
1.75 |
| ATP | 1.13 |
1.32 |
1.16 |
1.66 |
1.06 |
.93 |
1.54 |
| KLA/ATP | 1.37 |
1.25 |
1.21 |
1.74 |
1.04 |
1.01 |
2.07 |
|
| Csf1 | 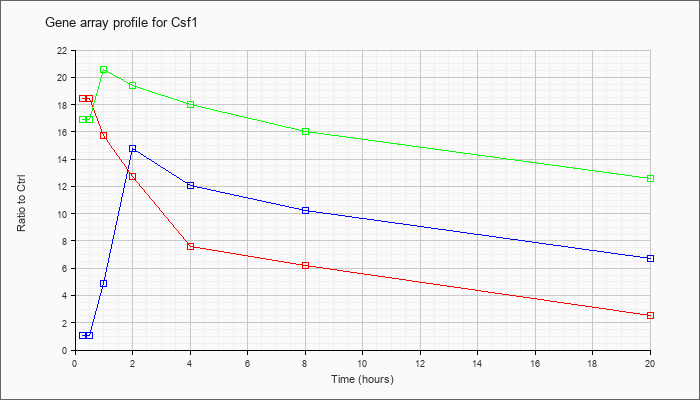 |
NM_007778 |
colony stimulating factor 1 (macrophage) (Csf1), transcript variant 1, mRNA [NM_007778] |
KLA | 18.44 |
16.94 |
15.74 |
12.71 |
7.63 |
6.20 |
2.51 |
| ATP | 1.04 |
1.22 |
4.90 |
14.80 |
12.10 |
10.26 |
6.73 |
| KLA/ATP | 16.93 |
18.17 |
20.56 |
19.41 |
18.03 |
15.99 |
12.59 |
|
| Csf2 | 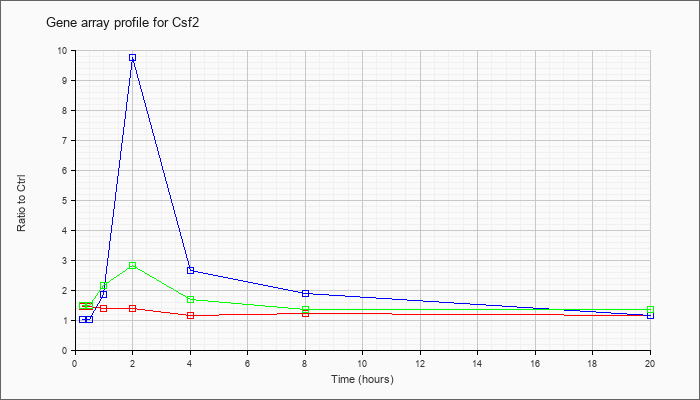 |
NM_009969 |
colony stimulating factor 2 (granulocyte-macrophage) (Csf2), mRNA [NM_009969] |
KLA | 1.44 |
1.53 |
1.38 |
1.37 |
1.15 |
1.21 |
1.14 |
| ATP | 1.01 |
1.02 |
1.85 |
9.73 |
2.64 |
1.90 |
1.14 |
| KLA/ATP | 1.49 |
1.84 |
2.14 |
2.80 |
1.67 |
1.36 |
1.35 |
|
| Cx3cl1 | 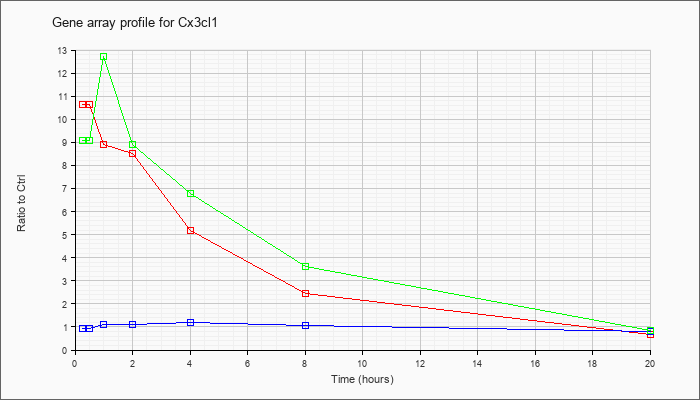 |
AB030188 |
mRNA, complete cds, clone:1-53. [AB030188] |
KLA | 1.04 |
.94 |
1.03 |
1.04 |
.92 |
1.05 |
1.00 |
| ATP | .94 |
.96 |
.98 |
.96 |
1.02 |
.91 |
1.04 |
| KLA/ATP | 1.03 |
.99 |
.96 |
.98 |
.89 |
1.03 |
.99 |
|
| Cx3cl1 |  |
NM_009142 |
chemokine (C-X3-C motif) ligand 1 (Cx3cl1), mRNA [NM_009142] |
KLA | 20.22 |
19.73 |
16.74 |
16.02 |
9.46 |
3.82 |
.33 |
| ATP | .91 |
.89 |
1.19 |
1.28 |
1.33 |
1.21 |
.55 |
| KLA/ATP | 17.11 |
18.99 |
24.51 |
16.85 |
12.70 |
6.17 |
.69 |
|
| Cxcl1 | 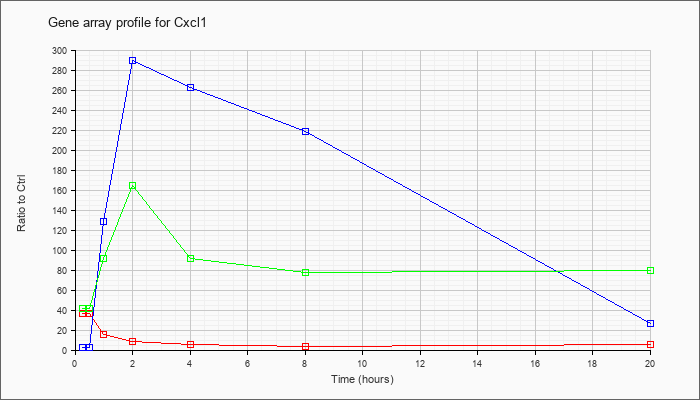 |
NM_008176 |
chemokine (C-X-C motif) ligand 1 (Cxcl1), mRNA [NM_008176] |
KLA | 36.87 |
30.25 |
15.74 |
8.84 |
5.41 |
3.01 |
5.84 |
| ATP | 2.77 |
8.26 |
128.38 |
289.44 |
262.38 |
218.83 |
26.91 |
| KLA/ATP | 41.06 |
56.76 |
91.82 |
164.83 |
91.44 |
77.89 |
79.05 |
|
| Cxcl10 | 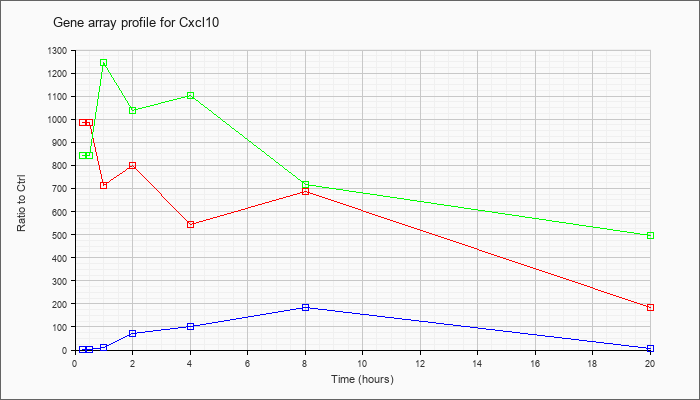 |
NM_021274 |
chemokine (C-X-C motif) ligand 10 (Cxcl10), mRNA [NM_021274] |
KLA | 986.69 |
854.98 |
711.49 |
799.39 |
542.62 |
686.82 |
183.27 |
| ATP | .98 |
1.71 |
9.85 |
73.04 |
100.42 |
184.63 |
8.33 |
| KLA/ATP | 842.13 |
793.12 |
###.## |
###.## |
###.## |
715.22 |
496.10 |
|
| Cxcl2 | 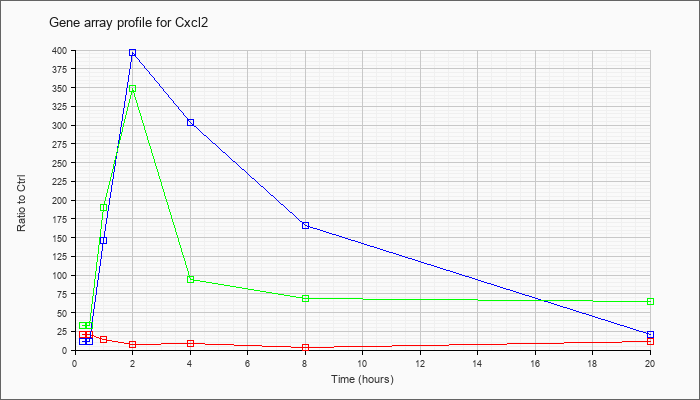 |
NM_009140 |
chemokine (C-X-C motif) ligand 2 (Cxcl2), mRNA [NM_009140] |
KLA | 20.11 |
17.63 |
13.50 |
7.78 |
9.14 |
3.98 |
10.85 |
| ATP | 10.96 |
27.84 |
145.37 |
396.18 |
303.88 |
165.35 |
21.04 |
| KLA/ATP | 33.28 |
74.00 |
189.99 |
348.56 |
93.84 |
68.75 |
64.87 |
|
| Cxcl3 | 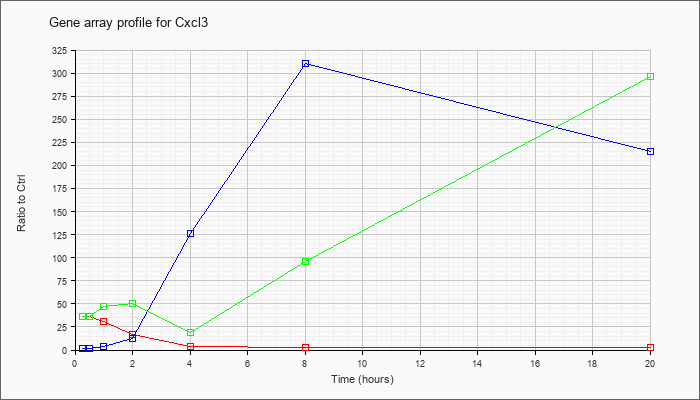 |
NM_203320 |
chemokine (C-X-C motif) ligand 3 (Cxcl3), mRNA [NM_203320] |
KLA | 35.82 |
43.91 |
31.09 |
16.74 |
3.98 |
2.27 |
2.93 |
| ATP | 1.13 |
1.23 |
3.43 |
12.43 |
126.40 |
310.60 |
215.03 |
| KLA/ATP | 36.32 |
44.70 |
46.93 |
50.61 |
18.91 |
95.89 |
296.64 |
|
| Dnm1l | 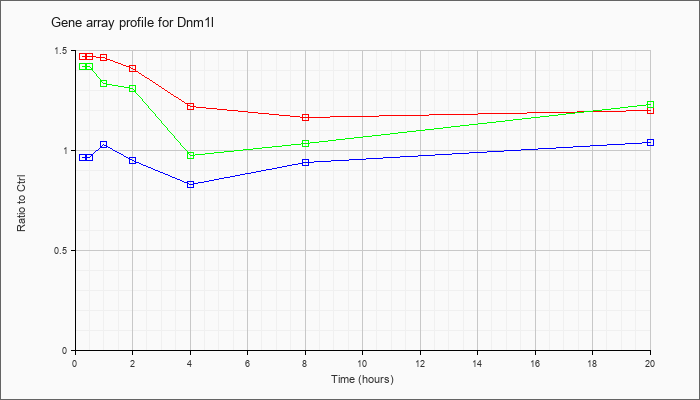 |
BC027538 |
dynamin 1-like, mRNA (cDNA clone IMAGE:1395338), complete cds. [BC027538] |
KLA | 1.08 |
1.11 |
1.09 |
1.05 |
1.00 |
1.00 |
1.09 |
| ATP | .92 |
.87 |
.83 |
.91 |
.94 |
.87 |
1.02 |
| KLA/ATP | 1.08 |
.96 |
.89 |
.92 |
.87 |
.80 |
1.02 |
|
| Dnm1l |  |
NM_152816 |
dynamin 1-like (Dnm1l), transcript variant 1, mRNA [NM_152816] |
KLA | 1.66 |
1.72 |
1.65 |
1.59 |
1.33 |
1.25 |
1.25 |
| ATP | .99 |
1.04 |
1.13 |
.97 |
.77 |
.97 |
1.05 |
| KLA/ATP | 1.59 |
1.65 |
1.56 |
1.50 |
1.03 |
1.15 |
1.33 |
|
| Edn1 | 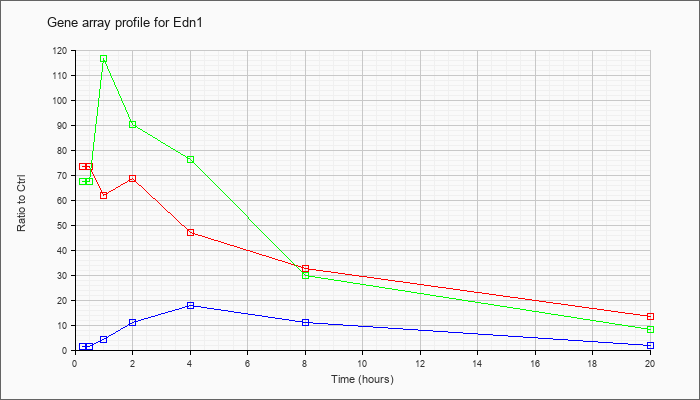 |
NM_010104 |
endothelin 1 (Edn1), mRNA [NM_010104] |
KLA | 73.36 |
71.33 |
61.92 |
68.69 |
46.94 |
32.60 |
13.54 |
| ATP | 1.23 |
1.89 |
4.18 |
10.86 |
17.85 |
11.03 |
1.91 |
| KLA/ATP | 67.27 |
76.78 |
116.70 |
90.29 |
76.38 |
29.71 |
8.04 |
|
EG432555
| 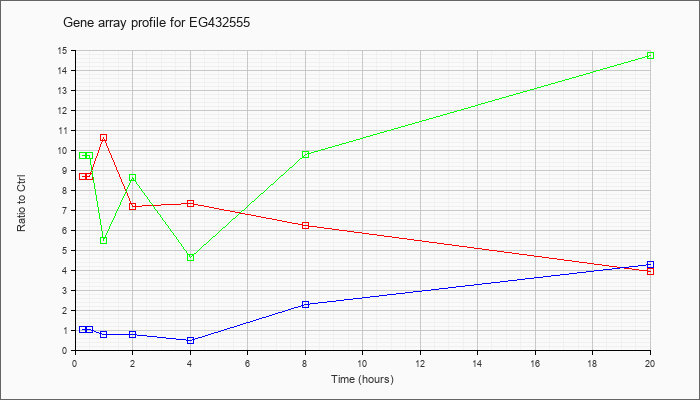 |
NM_001024230 |
predicted gene, EG432555 (EG432555), mRNA [NM_001024230] |
KLA | 8.68 |
9.07 |
10.61 |
7.17 |
7.31 |
6.24 |
3.94 |
| ATP | 1.03 |
1.09 |
.78 |
.77 |
.48 |
2.27 |
4.30 |
| KLA/ATP | 9.73 |
9.16 |
5.47 |
8.65 |
4.63 |
9.80 |
14.74 |
|
| Fadd | 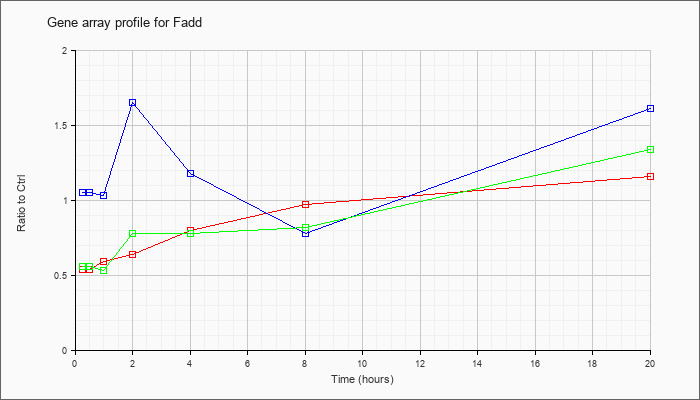 |
NM_010175 |
Fas (TNFRSF6)-associated via death domain (Fadd), mRNA [NM_010175] |
KLA | .54 |
.54 |
.59 |
.64 |
.80 |
.97 |
1.16 |
| ATP | 1.05 |
1.09 |
1.03 |
1.65 |
1.18 |
.78 |
1.61 |
| KLA/ATP | .56 |
.56 |
.53 |
.78 |
.78 |
.82 |
1.34 |
|
| Fas | 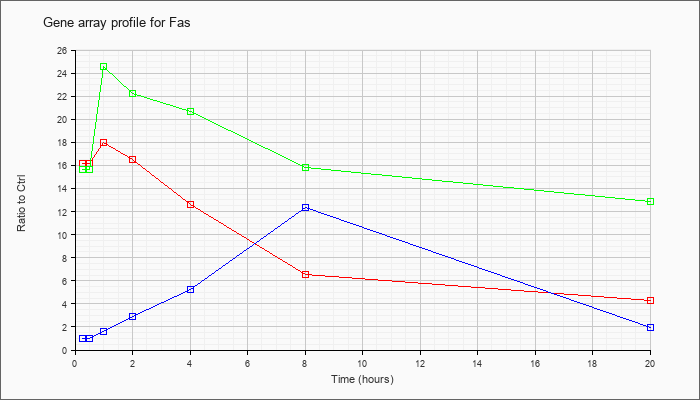 |
AK086933 |
0 day neonate lung cDNA, RIKEN full-length enriched library, clone:E030012O08 product:tumor necrosis factor receptor superfamily, member 6, full insert sequence. [AK086933] |
KLA | 6.31 |
6.56 |
6.33 |
6.70 |
4.93 |
3.74 |
2.48 |
| ATP | .95 |
1.04 |
1.33 |
1.55 |
1.85 |
6.00 |
2.77 |
| KLA/ATP | 5.67 |
7.16 |
7.71 |
7.73 |
6.02 |
5.79 |
7.85 |
|
| Fas |  |
NM_007987 |
Fas (TNF receptor superfamily member 6) (Fas), mRNA [NM_007987] |
KLA | 17.08 |
17.45 |
19.08 |
17.38 |
13.33 |
6.78 |
4.46 |
| ATP | .98 |
.96 |
1.64 |
3.05 |
5.58 |
12.91 |
1.89 |
| KLA/ATP | 16.57 |
17.25 |
26.06 |
23.57 |
22.03 |
16.74 |
13.29 |
|
| Fos | 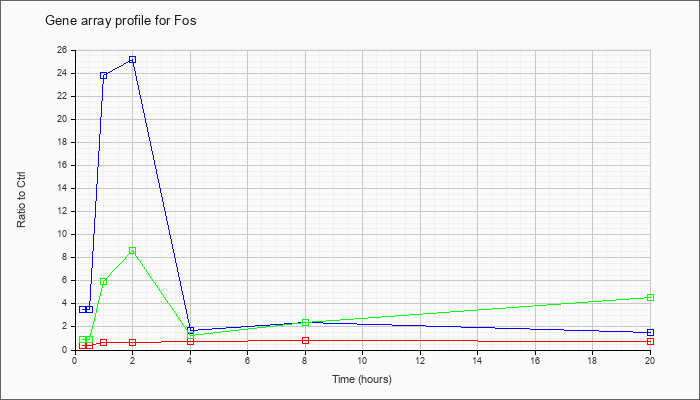 |
NM_010234 |
FBJ osteosarcoma oncogene (Fos), mRNA [NM_010234] |
KLA | .42 |
.33 |
.64 |
.61 |
.71 |
.85 |
.71 |
| ATP | 3.48 |
5.46 |
23.78 |
25.22 |
1.71 |
2.36 |
1.50 |
| KLA/ATP | .90 |
1.74 |
5.97 |
8.62 |
1.28 |
2.42 |
4.52 |
|
| Icam1 | 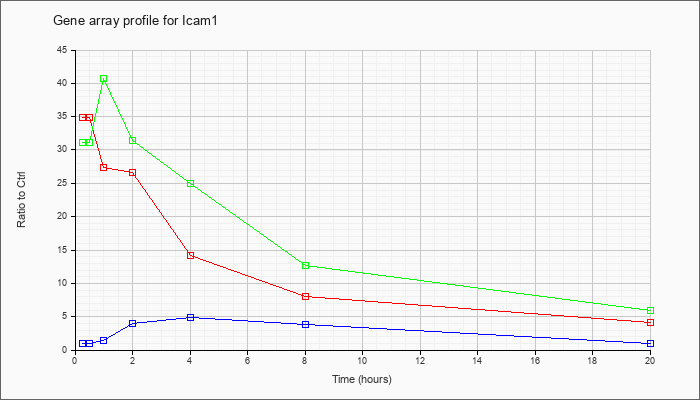 |
BC008626 |
intercellular adhesion molecule, mRNA (cDNA clone MGC:6195 IMAGE:3588949), complete cds [BC008626] |
KLA | 36.67 |
32.93 |
28.63 |
27.91 |
14.65 |
8.16 |
4.15 |
| ATP | .96 |
.88 |
1.50 |
3.88 |
4.98 |
3.90 |
1.01 |
| KLA/ATP | 32.58 |
31.60 |
42.74 |
32.60 |
26.00 |
13.02 |
6.01 |
|
| Icam1 |  |
NM_010493 |
intercellular adhesion molecule 1 (Icam1), mRNA [NM_010493] |
KLA | 15.66 |
15.20 |
12.96 |
12.31 |
9.39 |
6.17 |
3.81 |
| ATP | .96 |
1.06 |
1.21 |
4.19 |
4.33 |
3.16 |
1.06 |
| KLA/ATP | 14.31 |
15.46 |
17.84 |
17.60 |
13.77 |
8.36 |
5.42 |
|
| Ifi47 | 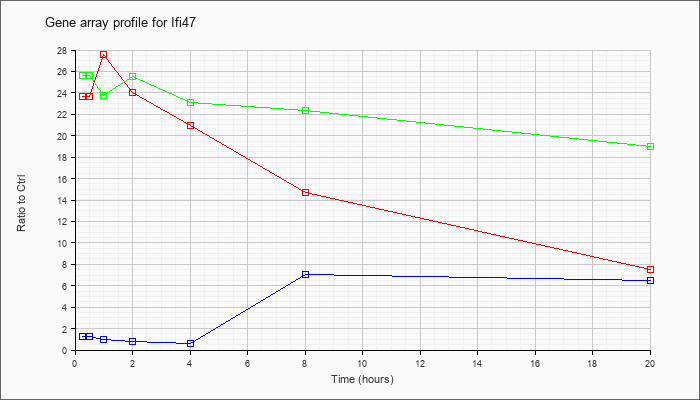 |
NM_008330 |
interferon gamma inducible protein 47 (Ifi47), mRNA [NM_008330] |
KLA | 23.67 |
20.20 |
27.55 |
23.99 |
20.93 |
14.71 |
7.52 |
| ATP | 1.31 |
1.13 |
.96 |
.75 |
.63 |
7.06 |
6.50 |
| KLA/ATP | 25.59 |
27.19 |
23.80 |
25.56 |
23.08 |
22.33 |
19.02 |
|
| Ikbkb | 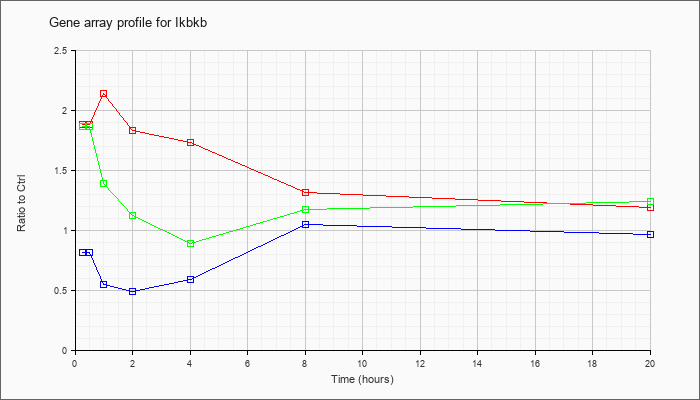 |
NM_010546 |
inhibitor of kappaB kinase beta (Ikbkb), mRNA [NM_010546] |
KLA | 1.88 |
2.10 |
2.14 |
1.83 |
1.73 |
1.31 |
1.19 |
| ATP | .81 |
.74 |
.55 |
.49 |
.59 |
1.05 |
.96 |
| KLA/ATP | 1.86 |
1.73 |
1.39 |
1.12 |
.89 |
1.17 |
1.24 |
|
| Ikbkg | 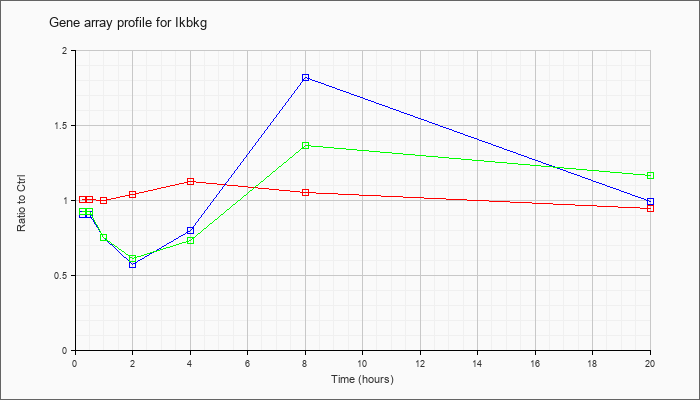 |
AK042138 |
3 days neonate thymus cDNA, RIKEN full-length enriched library, clone:A630062P04 product:unclassifiable, full insert sequence. [AK042138] |
KLA | .92 |
.87 |
.78 |
.78 |
.87 |
.88 |
.62 |
| ATP | .84 |
.71 |
.80 |
.50 |
1.15 |
2.33 |
.61 |
| KLA/ATP | .72 |
.69 |
.74 |
.48 |
1.06 |
1.97 |
.68 |
|
| Ikbkg |  |
NM_010547 |
inhibitor of kappaB kinase gamma (Ikbkg), transcript variant 1, mRNA [NM_010547] |
KLA | 1.51 |
1.63 |
1.74 |
1.83 |
1.83 |
1.73 |
1.72 |
| ATP | .80 |
.79 |
.82 |
.75 |
.66 |
2.58 |
2.37 |
| KLA/ATP | 1.43 |
1.42 |
1.17 |
1.04 |
.71 |
1.54 |
3.17 |
|
| Ikbkg |  |
NM_178590 |
inhibitor of kappaB kinase gamma (Ikbkg), transcript variant 2, mRNA [NM_178590] |
KLA | .84 |
.93 |
.86 |
.91 |
1.03 |
.89 |
.88 |
| ATP | 1.03 |
.98 |
.68 |
.56 |
.51 |
.93 |
.68 |
| KLA/ATP | .89 |
.81 |
.55 |
.52 |
.41 |
.67 |
.65 |
|
| Il15 | 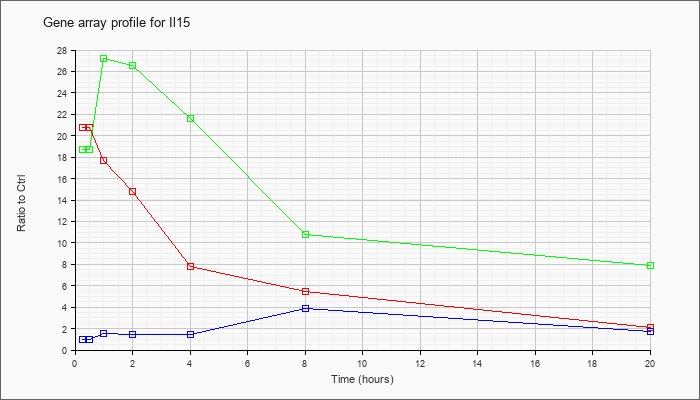 |
AB022307 |
mRNA for shorter isoform of interleukin 15, partial cds. [AB022307] |
KLA | 13.11 |
12.14 |
12.05 |
8.72 |
4.50 |
3.62 |
1.70 |
| ATP | 1.04 |
1.05 |
1.94 |
1.72 |
1.74 |
3.48 |
1.69 |
| KLA/ATP | 13.23 |
13.71 |
21.01 |
21.43 |
21.51 |
10.85 |
4.55 |
|
| Il15 |  |
NM_008357 |
interleukin 15 (Il15), mRNA [NM_008357] |
KLA | 28.44 |
24.75 |
23.34 |
20.83 |
11.02 |
7.27 |
2.51 |
| ATP | 1.00 |
.96 |
1.22 |
1.21 |
1.24 |
4.27 |
1.85 |
| KLA/ATP | 24.24 |
23.93 |
33.47 |
31.76 |
21.73 |
10.79 |
11.23 |
|
| Il18r1 | 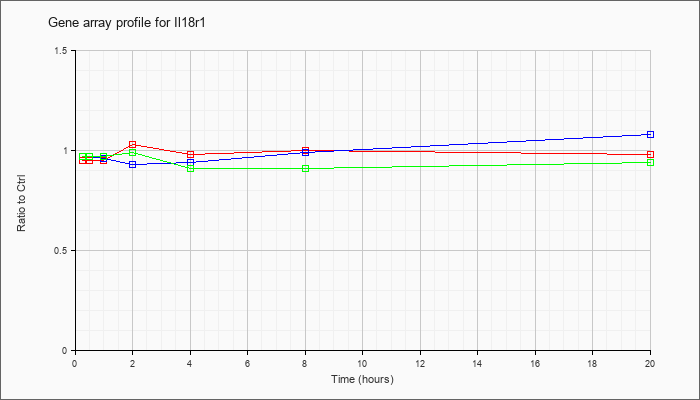 |
NM_008365 |
interleukin 18 receptor 1 (Il18r1), mRNA [NM_008365] |
KLA | .95 |
1.02 |
.95 |
1.03 |
.98 |
1.00 |
.98 |
| ATP | .97 |
1.02 |
.96 |
.93 |
.94 |
.99 |
1.08 |
| KLA/ATP | .97 |
.97 |
.97 |
.99 |
.91 |
.91 |
.94 |
|
| Il1b | 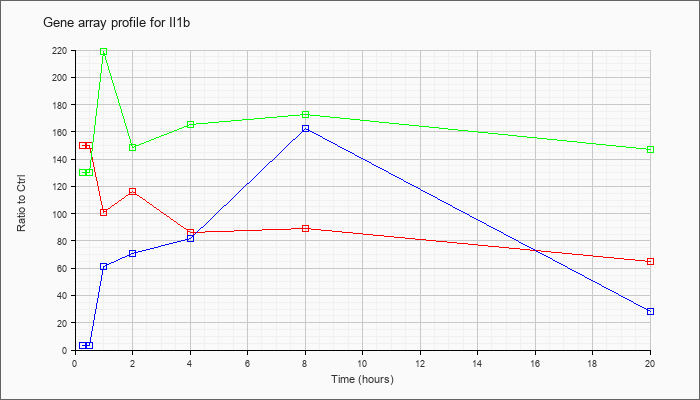 |
NM_008361 |
interleukin 1 beta (Il1b), mRNA [NM_008361] |
KLA | 149.78 |
133.59 |
100.91 |
116.58 |
86.48 |
89.37 |
64.72 |
| ATP | 3.49 |
13.96 |
61.44 |
70.91 |
82.12 |
162.79 |
28.18 |
| KLA/ATP | 130.32 |
139.00 |
219.12 |
148.17 |
165.44 |
172.57 |
146.98 |
|
| Il6 | 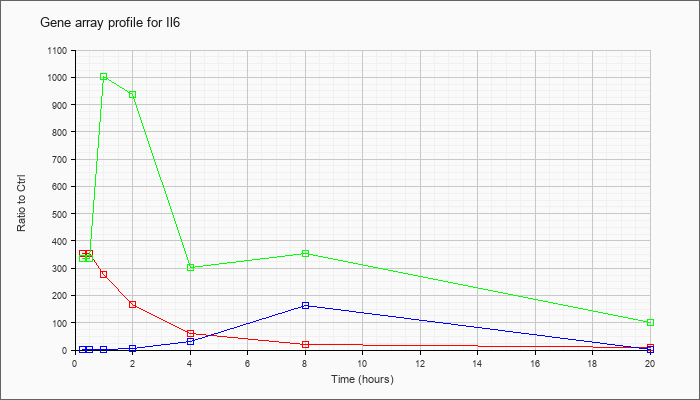 |
NM_031168 |
interleukin 6 (Il6), mRNA [NM_031168] |
KLA | 352.99 |
378.70 |
276.81 |
168.47 |
59.99 |
18.66 |
8.27 |
| ATP | 1.04 |
1.08 |
2.28 |
3.69 |
30.40 |
161.93 |
2.11 |
| KLA/ATP | 334.28 |
528.66 |
###.## |
935.86 |
302.45 |
355.36 |
99.65 |
|
| Itch | 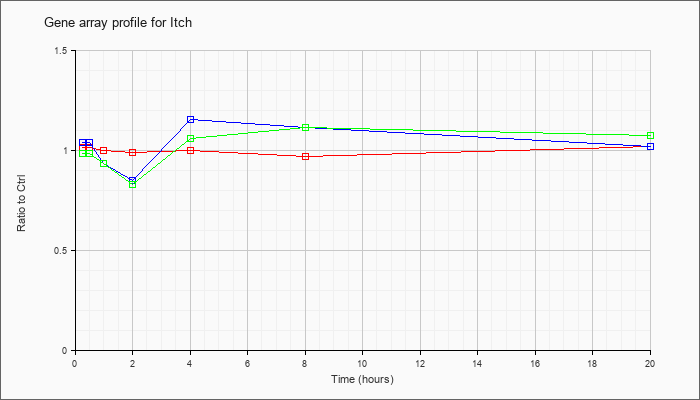 |
AK033318 |
15 days embryo male testis cDNA, RIKEN full-length enriched library, clone:8030492O04 product:unclassifiable, full insert sequence. [AK033318] |
KLA | .94 |
.82 |
.94 |
.88 |
1.04 |
.93 |
.95 |
| ATP | 1.15 |
1.14 |
.93 |
.75 |
1.42 |
1.04 |
.99 |
| KLA/ATP | .98 |
.95 |
.86 |
.75 |
1.25 |
1.01 |
.96 |
|
| Itch |  |
AK038333 |
16 days neonate thymus cDNA, RIKEN full-length enriched library, clone:A130096D14 product:similar to TYROSINE-PROTEIN KINASE JAK3 (EC 2.7.1.112) (JANUS KINASE 3) (JAK-3) [Mus musculus], full insert sequence. [AK038333] |
KLA | .97 |
.89 |
.88 |
.89 |
.97 |
.87 |
.94 |
| ATP | 1.03 |
.87 |
.72 |
.90 |
1.16 |
1.01 |
.93 |
| KLA/ATP | .94 |
.89 |
.83 |
.80 |
.94 |
.93 |
.93 |
|
| Itch |  |
AK220164 |
mRNA for mKIAA4011 protein [AK220164] |
KLA | 1.08 |
.99 |
1.04 |
1.13 |
.99 |
.99 |
1.06 |
| ATP | .95 |
.92 |
1.13 |
.73 |
.96 |
1.33 |
.93 |
| KLA/ATP | .94 |
.97 |
1.12 |
.75 |
1.10 |
1.40 |
1.11 |
|
| Itch |  |
NM_008395 |
itchy, E3 ubiquitin protein ligase (Itch), mRNA [NM_008395] |
KLA | 1.06 |
1.01 |
1.13 |
1.05 |
1.00 |
1.08 |
1.13 |
| ATP | 1.02 |
1.10 |
.96 |
1.02 |
1.07 |
1.06 |
1.22 |
| KLA/ATP | 1.07 |
1.00 |
.93 |
1.01 |
.95 |
1.10 |
1.29 |
|
| Jun | 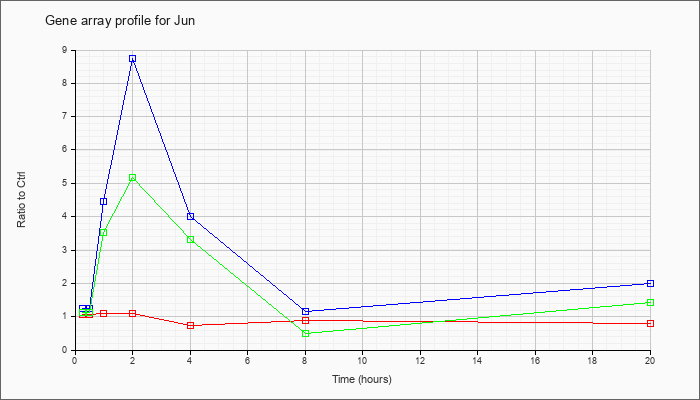 |
NM_010591 |
Jun oncogene (Jun), mRNA [NM_010591] |
KLA | 1.08 |
.96 |
1.09 |
1.09 |
.75 |
.88 |
.81 |
| ATP | 1.26 |
1.48 |
4.45 |
8.73 |
4.02 |
1.16 |
1.99 |
| KLA/ATP | 1.12 |
1.38 |
3.52 |
5.17 |
3.32 |
.49 |
1.42 |
|
| Junb | 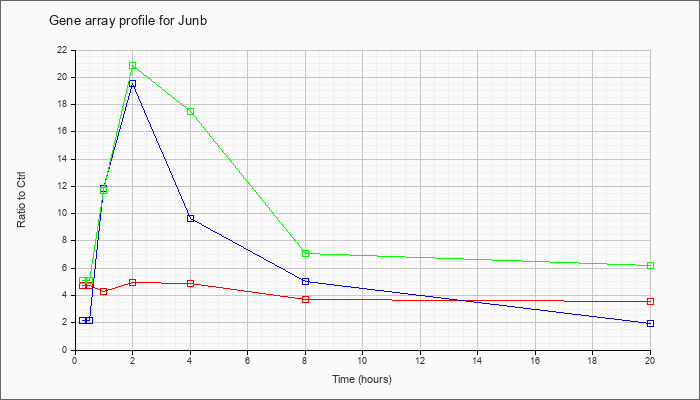 |
NM_008416 |
Jun-B oncogene (Junb), mRNA [NM_008416] |
KLA | 4.71 |
4.37 |
4.30 |
4.98 |
4.85 |
3.70 |
3.53 |
| ATP | 2.20 |
3.40 |
11.86 |
19.56 |
9.63 |
5.04 |
1.94 |
| KLA/ATP | 5.09 |
6.66 |
11.69 |
20.90 |
17.53 |
7.06 |
6.21 |
|
| Lta | 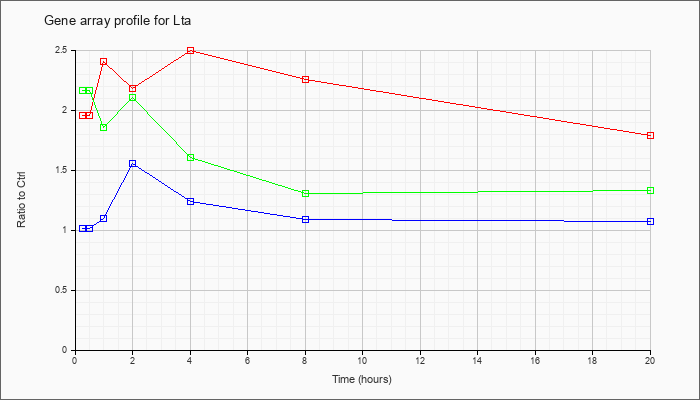 |
NM_010735 |
lymphotoxin A (Lta), mRNA [NM_010735] |
KLA | 1.95 |
2.18 |
2.41 |
2.18 |
2.50 |
2.26 |
1.79 |
| ATP | 1.01 |
1.00 |
1.10 |
1.55 |
1.24 |
1.09 |
1.07 |
| KLA/ATP | 2.16 |
2.23 |
1.85 |
2.11 |
1.60 |
1.31 |
1.33 |
|
| Magi2 | 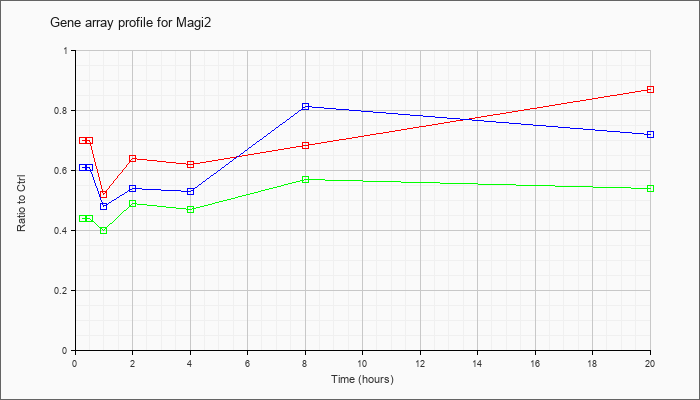 |
NM_015823 |
membrane associated guanylate kinase, WW and PDZ domain containing 2 (Magi2), mRNA [NM_015823] |
KLA | .70 |
.66 |
.52 |
.64 |
.62 |
.68 |
.87 |
| ATP | .61 |
.56 |
.48 |
.54 |
.53 |
.81 |
.72 |
| KLA/ATP | .44 |
.46 |
.40 |
.49 |
.47 |
.57 |
.54 |
|
| Map2k1 | 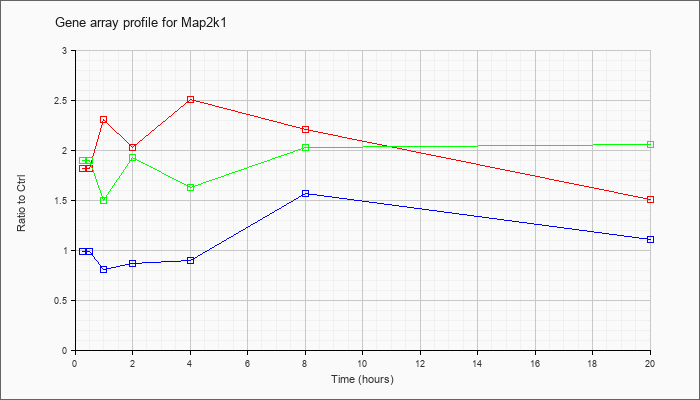 |
NM_008927 |
mitogen-activated protein kinase kinase 1 (Map2k1), mRNA [NM_008927] |
KLA | 1.82 |
2.00 |
2.31 |
2.03 |
2.51 |
2.20 |
1.51 |
| ATP | .99 |
1.14 |
.81 |
.87 |
.90 |
1.57 |
1.11 |
| KLA/ATP | 1.90 |
2.03 |
1.50 |
1.93 |
1.63 |
2.03 |
2.06 |
|
| Map2k3 | 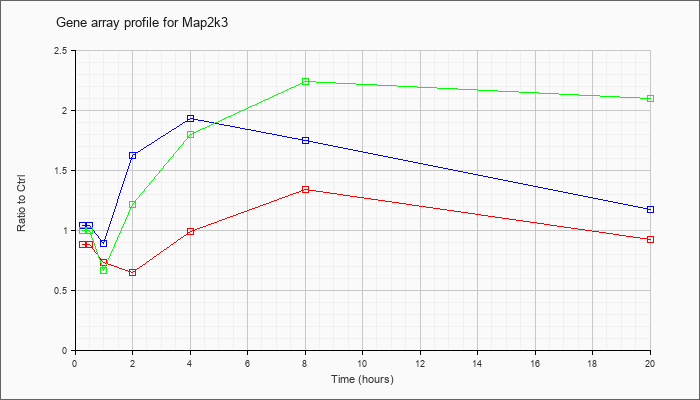 |
NM_008928 |
mitogen-activated protein kinase kinase 3 (Map2k3), mRNA [NM_008928] |
KLA | .88 |
.88 |
.73 |
.65 |
.99 |
1.34 |
.92 |
| ATP | 1.04 |
1.09 |
.89 |
1.62 |
1.93 |
1.75 |
1.17 |
| KLA/ATP | 1.00 |
.92 |
.66 |
1.21 |
1.80 |
2.24 |
2.10 |
|
| Map2k4 | 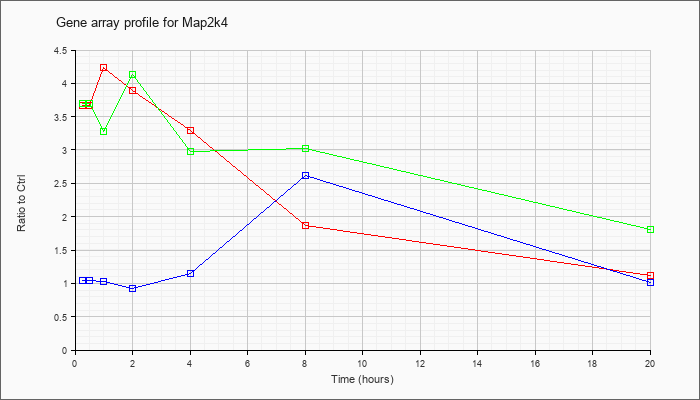 |
NM_009157 |
mitogen-activated protein kinase kinase 4 (Map2k4), mRNA [NM_009157] |
KLA | 3.67 |
3.75 |
4.23 |
3.90 |
3.30 |
1.87 |
1.12 |
| ATP | 1.05 |
1.11 |
1.04 |
.93 |
1.16 |
2.62 |
1.02 |
| KLA/ATP | 3.70 |
4.13 |
3.27 |
4.13 |
2.98 |
3.03 |
1.82 |
|
| Map2k6 | 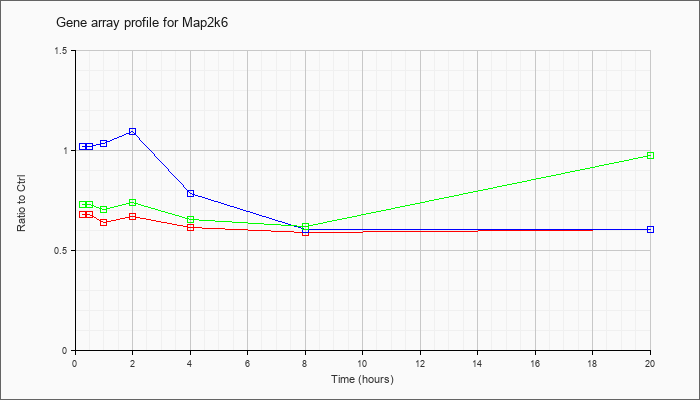 |
NM_011943 |
mitogen-activated protein kinase kinase 6 (Map2k6), mRNA [NM_011943] |
KLA | .68 |
.67 |
.64 |
.67 |
.62 |
.59 |
.61 |
| ATP | 1.02 |
1.12 |
1.04 |
1.10 |
.79 |
.61 |
.61 |
| KLA/ATP | .73 |
.69 |
.71 |
.74 |
.66 |
.62 |
.98 |
|
| Map2k7 | 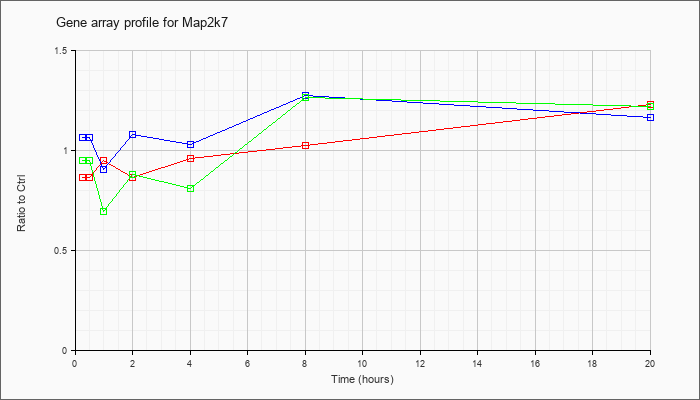 |
BC070467 |
mitogen activated protein kinase kinase 7, mRNA (cDNA clone MGC:96654 IMAGE:30615302), complete cds [BC070467] |
KLA | .77 |
.79 |
.88 |
.77 |
.97 |
.96 |
1.27 |
| ATP | 1.11 |
1.26 |
.93 |
1.19 |
1.13 |
1.45 |
1.16 |
| KLA/ATP | .89 |
.88 |
.59 |
.89 |
.86 |
1.32 |
1.28 |
|
| Map2k7 |  |
NM_001042557 |
mitogen-activated protein kinase kinase 7 (Map2k7), transcript variant 1, mRNA [NM_001042557] |
KLA | .96 |
.95 |
1.02 |
.96 |
.95 |
1.09 |
1.19 |
| ATP | 1.02 |
1.07 |
.88 |
.97 |
.93 |
1.10 |
1.17 |
| KLA/ATP | 1.01 |
.99 |
.80 |
.87 |
.76 |
1.21 |
1.16 |
|
| Map3k14 | 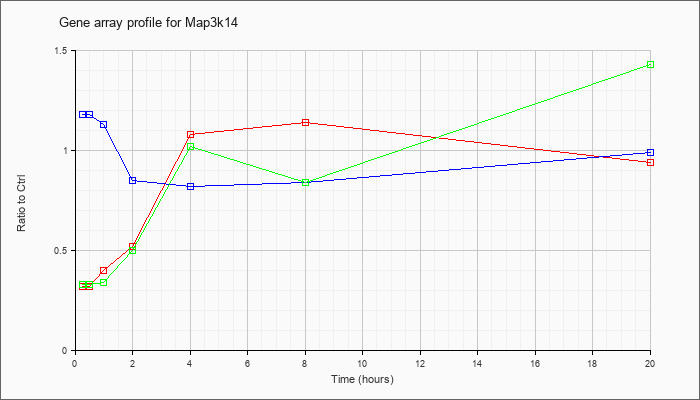 |
NM_016896 |
mitogen-activated protein kinase kinase kinase 14 (Map3k14), mRNA [NM_016896] |
KLA | .32 |
.30 |
.40 |
.52 |
1.08 |
1.14 |
.94 |
| ATP | 1.18 |
1.24 |
1.13 |
.85 |
.82 |
.84 |
.99 |
| KLA/ATP | .33 |
.30 |
.34 |
.50 |
1.02 |
.84 |
1.43 |
|
| Map3k5 | 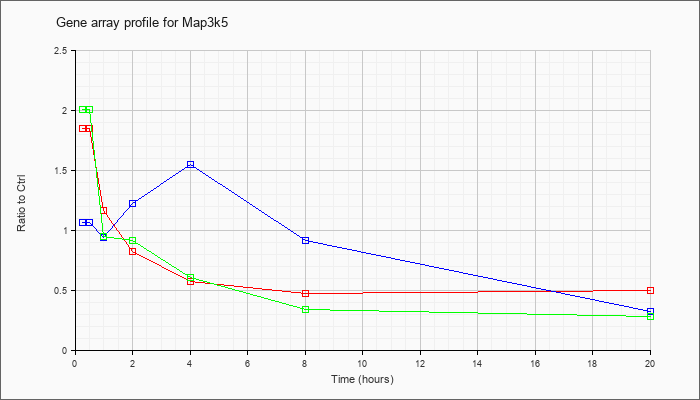 |
NM_008580 |
mitogen-activated protein kinase kinase kinase 5 (Map3k5), mRNA [NM_008580] |
KLA | 1.85 |
1.97 |
1.17 |
.82 |
.57 |
.47 |
.49 |
| ATP | 1.06 |
1.05 |
.94 |
1.22 |
1.55 |
.91 |
.32 |
| KLA/ATP | 2.00 |
1.82 |
.95 |
.91 |
.60 |
.34 |
.28 |
|
| Map3k7 | 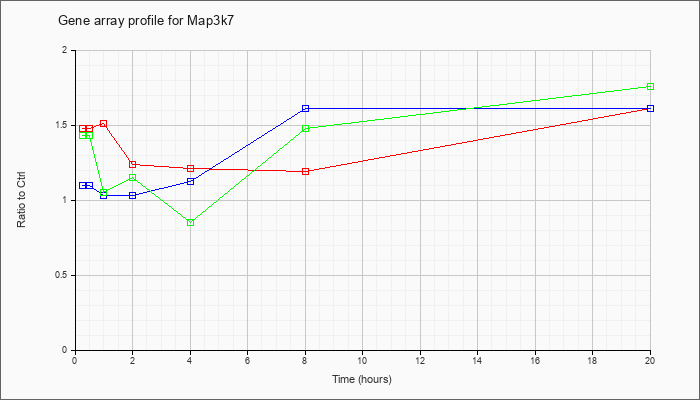 |
NM_172688 |
mitogen-activated protein kinase kinase kinase 7 (Map3k7), mRNA [NM_172688] |
KLA | 1.48 |
1.49 |
1.51 |
1.24 |
1.21 |
1.19 |
1.61 |
| ATP | 1.10 |
1.16 |
1.03 |
1.03 |
1.12 |
1.61 |
1.61 |
| KLA/ATP | 1.43 |
1.56 |
1.05 |
1.15 |
.85 |
1.48 |
1.76 |
|
Map3k7ip
1 | 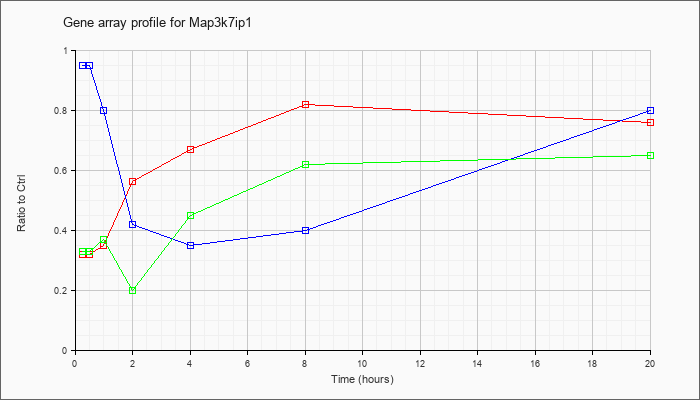 |
NM_025609 |
mitogen-activated protein kinase kinase kinase 7 interacting protein 1 (Map3k7ip1), mRNA [NM_025609] |
KLA | .32 |
.30 |
.35 |
.56 |
.67 |
.82 |
.76 |
| ATP | .95 |
.81 |
.80 |
.42 |
.35 |
.40 |
.80 |
| KLA/ATP | .33 |
.30 |
.37 |
.20 |
.45 |
.62 |
.65 |
|
Map3k7ip
2 | 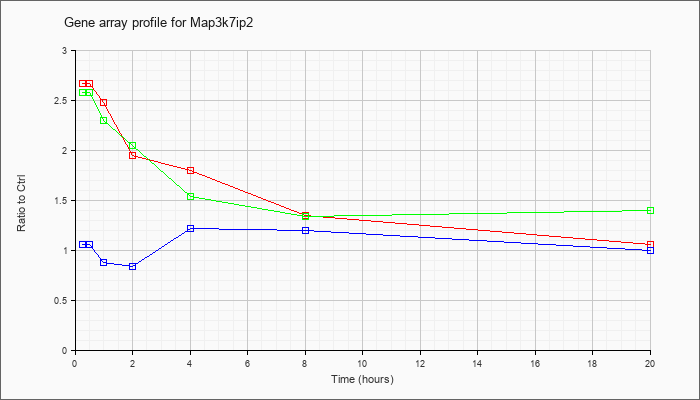 |
AK086230 |
15 days embryo head cDNA, RIKEN full-length enriched library, clone:D930015C19 product:inferred: Mus musculus, Similar to TAK1-binding protein 2; KIAA0733 protein, clone MGC:8010 IMAGE:3586132, mRNA, complete cds, full insert sequence. [A |
KLA | 1.14 |
1.02 |
1.07 |
1.03 |
1.13 |
1.02 |
.96 |
| ATP | 1.00 |
.88 |
.92 |
.90 |
1.06 |
.96 |
.94 |
| KLA/ATP | 1.05 |
1.00 |
.90 |
.98 |
1.02 |
.97 |
.97 |
|
Map3k7ip
2 |  |
NM_138667 |
mitogen-activated protein kinase kinase kinase 7 interacting protein 2 (Map3k7ip2), mRNA [NM_138667] |
KLA | 3.43 |
3.22 |
3.19 |
2.41 |
2.13 |
1.51 |
1.10 |
| ATP | 1.08 |
1.08 |
.86 |
.81 |
1.30 |
1.32 |
1.02 |
| KLA/ATP | 3.34 |
3.66 |
3.00 |
2.58 |
1.80 |
1.52 |
1.61 |
|
Map3k7ip
3 | 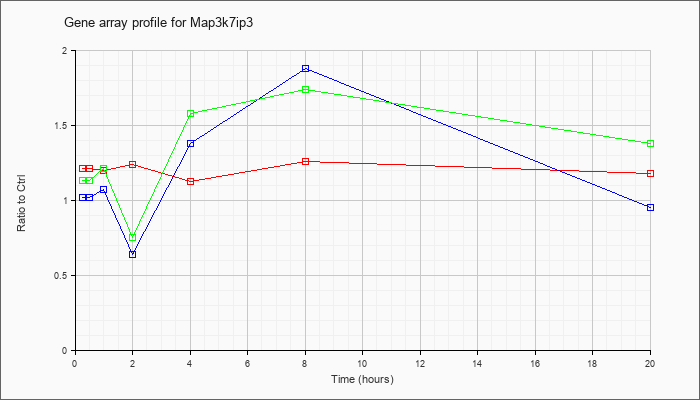 |
NM_025729 |
mitogen-activated protein kinase kinase kinase 7 interacting protein 3 (Map3k7ip3), mRNA [NM_025729] |
KLA | 1.21 |
1.05 |
1.20 |
1.24 |
1.12 |
1.26 |
1.18 |
| ATP | 1.02 |
.88 |
1.07 |
.64 |
1.38 |
1.88 |
.95 |
| KLA/ATP | 1.13 |
1.11 |
1.21 |
.75 |
1.58 |
1.74 |
1.38 |
|
| Map3k8 | 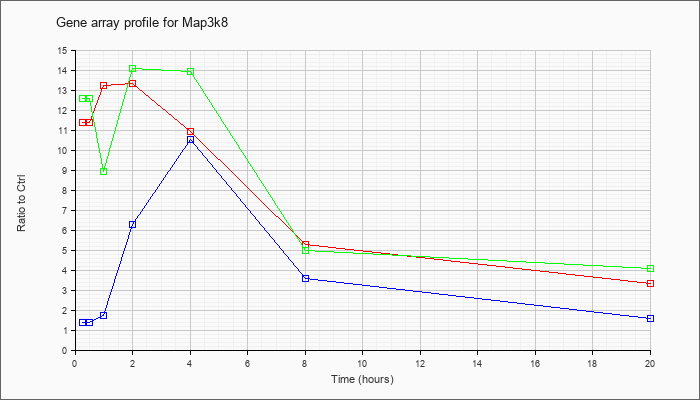 |
D13759 |
cot mRNA for proto-oncogene protein, complete cds. [D13759] |
KLA | 11.38 |
11.64 |
13.24 |
13.31 |
10.92 |
5.26 |
3.31 |
| ATP | 1.36 |
1.09 |
1.75 |
6.27 |
10.51 |
3.58 |
1.60 |
| KLA/ATP | 12.58 |
11.57 |
8.93 |
14.10 |
13.92 |
4.98 |
4.09 |
|
| Mapk1 | 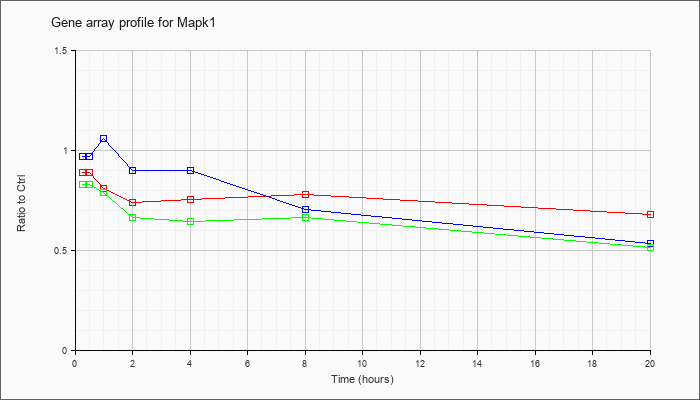 |
NM_001038663 |
mitogen-activated protein kinase 1 (Mapk1), transcript variant 2, mRNA [NM_001038663] |
KLA | .97 |
1.03 |
.93 |
.86 |
.79 |
1.01 |
1.04 |
| ATP | .96 |
1.18 |
1.01 |
1.13 |
.84 |
1.04 |
1.16 |
| KLA/ATP | .99 |
.99 |
.84 |
.87 |
.63 |
.96 |
1.26 |
|
| Mapk1 |  |
NM_011949 |
mitogen-activated protein kinase 1 (Mapk1), transcript variant 1, mRNA [NM_011949] |
KLA | .88 |
.80 |
.80 |
.73 |
.75 |
.76 |
.65 |
| ATP | .97 |
.96 |
1.06 |
.88 |
.90 |
.68 |
.48 |
| KLA/ATP | .82 |
.81 |
.78 |
.65 |
.64 |
.64 |
.45 |
|
| Mapk10 | 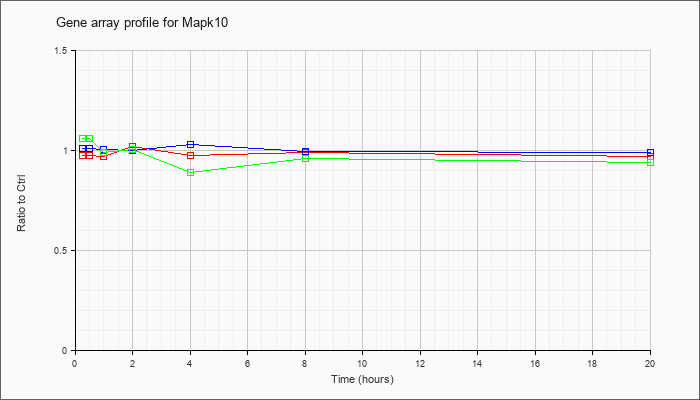 |
AK076990 |
adult male testis cDNA, RIKEN full-length enriched library, clone:4930597C01 product:mitogen activated protein kinase 10, full insert sequence. [AK076990] |
KLA | 1.03 |
1.05 |
.98 |
1.04 |
.94 |
.95 |
1.02 |
| ATP | 1.00 |
.99 |
1.06 |
.97 |
1.01 |
.96 |
.99 |
| KLA/ATP | 1.07 |
1.01 |
.95 |
1.05 |
.82 |
.95 |
.87 |
|
| Mapk10 |  |
NM_009158 |
mitogen-activated protein kinase 10 (Mapk10), transcript variant 1, mRNA [NM_009158] |
KLA | .95 |
1.00 |
.97 |
1.01 |
.99 |
1.01 |
.95 |
| ATP | 1.01 |
1.02 |
.98 |
1.01 |
1.04 |
1.01 |
.99 |
| KLA/ATP | 1.05 |
.99 |
1.01 |
.98 |
.92 |
.97 |
.98 |
|
| Mapk11 | 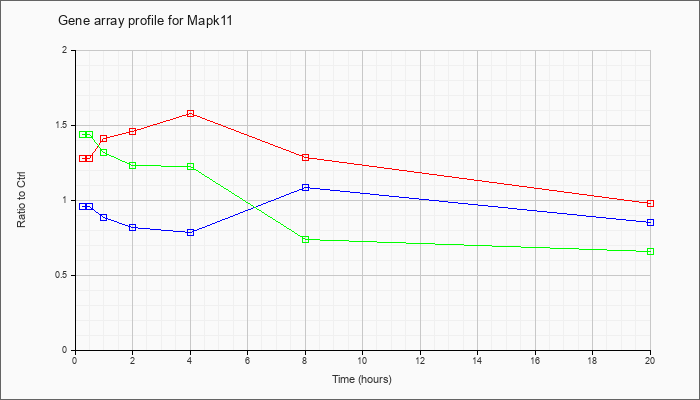 |
NM_011161 |
mitogen-activated protein kinase 11 (Mapk11), mRNA [NM_011161] |
KLA | 1.28 |
1.30 |
1.41 |
1.46 |
1.58 |
1.29 |
.98 |
| ATP | .96 |
.95 |
.89 |
.82 |
.79 |
1.09 |
.85 |
| KLA/ATP | 1.44 |
1.42 |
1.32 |
1.23 |
1.23 |
.74 |
.66 |
|
| Mapk12 | 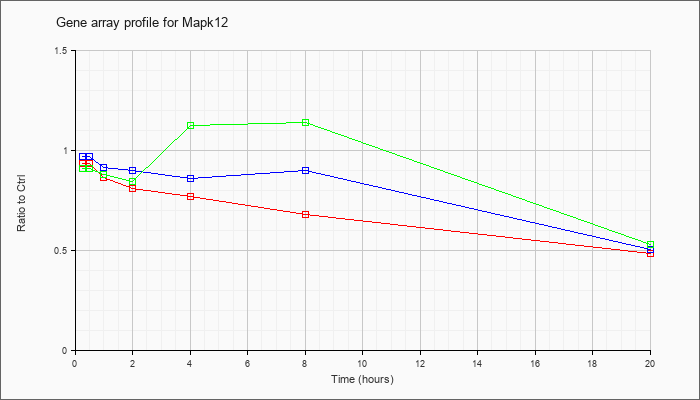 |
NM_013871 |
mitogen-activated protein kinase 12 (Mapk12), mRNA [NM_013871] |
KLA | .94 |
.86 |
.87 |
.81 |
.77 |
.68 |
.49 |
| ATP | .97 |
.83 |
.92 |
.90 |
.86 |
.90 |
.51 |
| KLA/ATP | .91 |
.89 |
.88 |
.85 |
1.12 |
1.14 |
.53 |
|
| Mapk13 | 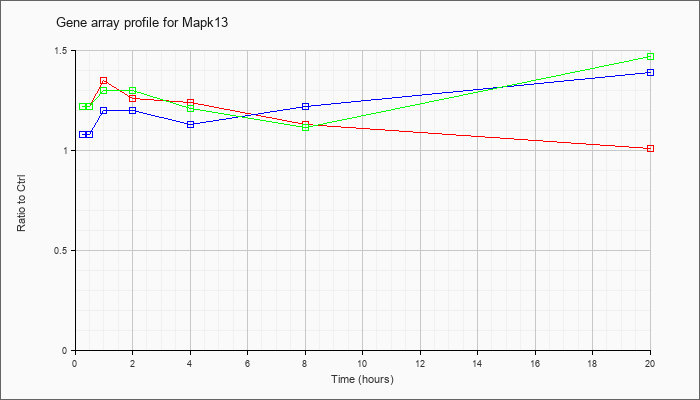 |
NM_011950 |
mitogen-activated protein kinase 13 (Mapk13), mRNA [NM_011950] |
KLA | 1.22 |
1.21 |
1.35 |
1.26 |
1.24 |
1.13 |
1.01 |
| ATP | 1.08 |
1.00 |
1.20 |
1.20 |
1.13 |
1.22 |
1.39 |
| KLA/ATP | 1.22 |
1.27 |
1.30 |
1.30 |
1.21 |
1.11 |
1.47 |
|
| Mapk14 | 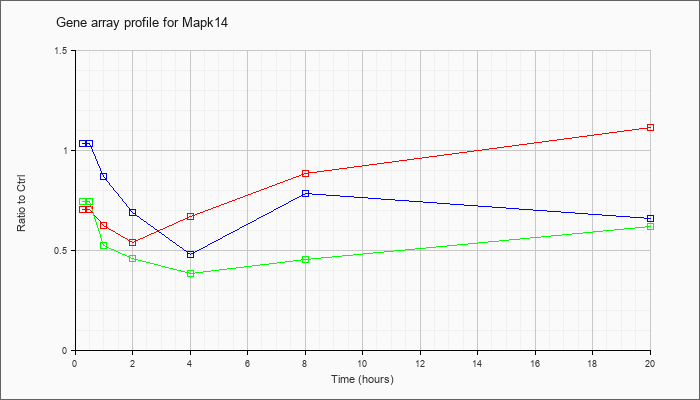 |
NM_011951 |
mitogen-activated protein kinase 14 (Mapk14), mRNA [NM_011951] |
KLA | .70 |
.70 |
.62 |
.54 |
.67 |
.88 |
1.11 |
| ATP | 1.03 |
.98 |
.87 |
.69 |
.48 |
.78 |
.66 |
| KLA/ATP | .74 |
.68 |
.52 |
.46 |
.38 |
.45 |
.62 |
|
| Mapk3 | 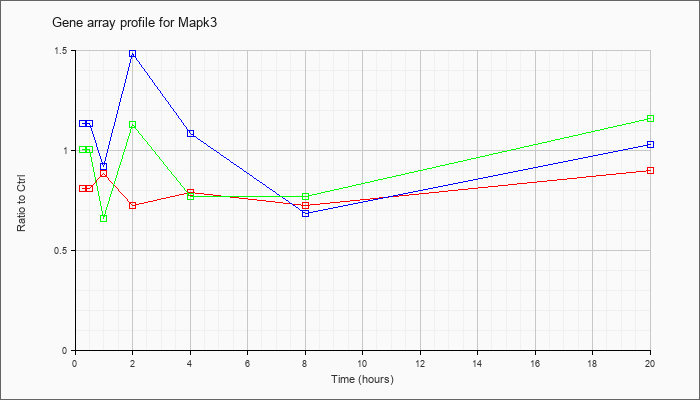 |
NM_011952 |
mitogen-activated protein kinase 3 (Mapk3), mRNA [NM_011952] |
KLA | .81 |
.85 |
.88 |
.72 |
.79 |
.73 |
.90 |
| ATP | 1.13 |
1.30 |
.92 |
1.48 |
1.08 |
.68 |
1.03 |
| KLA/ATP | 1.01 |
1.01 |
.66 |
1.13 |
.77 |
.77 |
1.16 |
|
| Mapk8 |  |
AK162915 |
adult male spinal cord cDNA, RIKEN full-length enriched library, clone:A330031O19 product:mitogen activated protein kinase 8, full insert sequence [AK162915] |
KLA | .94 |
.86 |
.80 |
.83 |
.77 |
.85 |
.83 |
| ATP | 1.08 |
1.12 |
.94 |
.80 |
.83 |
.71 |
.75 |
| KLA/ATP | .83 |
.81 |
.80 |
.74 |
.74 |
.75 |
.67 |
|
| Mapk8 |  |
AK163829 |
16 days embryo head cDNA, RIKEN full-length enriched library, clone:C130070A06 product:mitogen activated protein kinase 8, full insert sequence [AK163829] |
KLA | .65 |
.61 |
.66 |
.70 |
.90 |
1.02 |
1.03 |
| ATP | .99 |
.94 |
1.68 |
1.44 |
1.75 |
1.84 |
1.10 |
| KLA/ATP | .63 |
.67 |
1.11 |
1.00 |
2.01 |
1.68 |
1.15 |
|
| Mapk8 |  |
NM_016700 |
mitogen-activated protein kinase 8 (Mapk8), mRNA [NM_016700] |
KLA | .96 |
1.00 |
.94 |
.96 |
.94 |
.99 |
1.04 |
| ATP | 1.01 |
1.05 |
1.01 |
.99 |
1.03 |
.98 |
1.02 |
| KLA/ATP | .96 |
.97 |
.96 |
.97 |
.93 |
.94 |
.96 |
|
| Mapk9 | 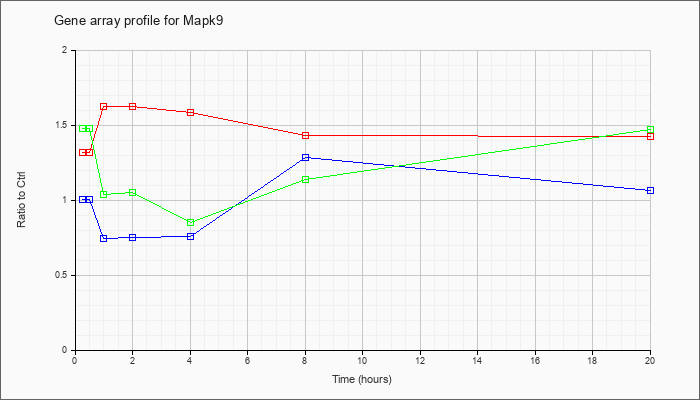 |
NM_016961 |
mitogen-activated protein kinase 9 (Mapk9), transcript variant 2, mRNA [NM_016961] |
KLA | 1.28 |
1.37 |
1.59 |
1.60 |
1.61 |
1.43 |
1.38 |
| ATP | 1.03 |
.99 |
.74 |
.71 |
.79 |
1.20 |
1.01 |
| KLA/ATP | 1.53 |
1.30 |
1.07 |
.99 |
.83 |
1.10 |
1.40 |
|
| Mapk9 |  |
NM_207692 |
mitogen-activated protein kinase 9 (Mapk9), transcript variant 1, mRNA [NM_207692] |
KLA | 1.39 |
1.53 |
1.69 |
1.69 |
1.55 |
1.44 |
1.51 |
| ATP | .95 |
.92 |
.75 |
.85 |
.70 |
1.46 |
1.17 |
| KLA/ATP | 1.37 |
1.35 |
.99 |
1.18 |
.91 |
1.22 |
1.62 |
|
| Mlkl | 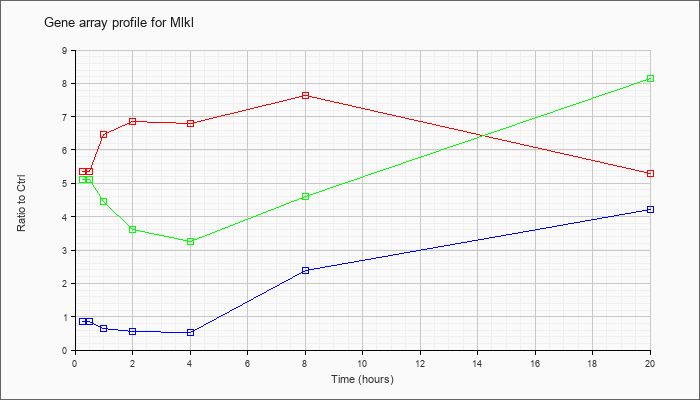 |
NM_029005 |
mixed lineage kinase domain-like (Mlkl), mRNA [NM_029005] |
KLA | 5.36 |
5.21 |
6.46 |
6.85 |
6.78 |
7.62 |
5.29 |
| ATP | .86 |
.75 |
.65 |
.56 |
.53 |
2.40 |
4.22 |
| KLA/ATP | 5.12 |
4.58 |
4.45 |
3.63 |
3.26 |
4.60 |
8.15 |
|
Mm.15898
1 | 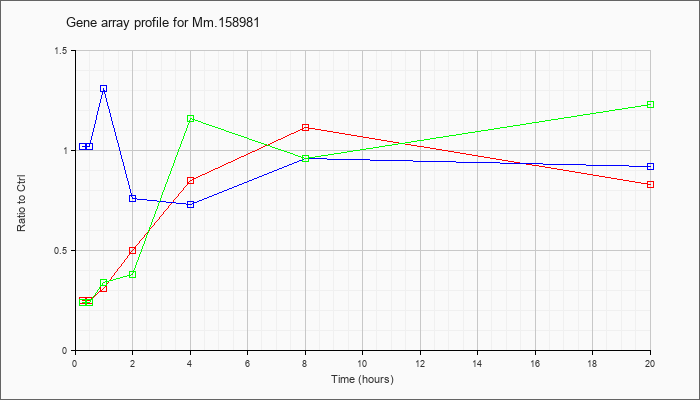 |
142388182 |
Unknown |
KLA | .25 |
.22 |
.31 |
.50 |
.85 |
1.11 |
.83 |
| ATP | 1.02 |
.94 |
1.31 |
.76 |
.73 |
.96 |
.92 |
| KLA/ATP | .24 |
.22 |
.34 |
.38 |
1.16 |
.96 |
1.23 |
|
Mm.21312
8 | 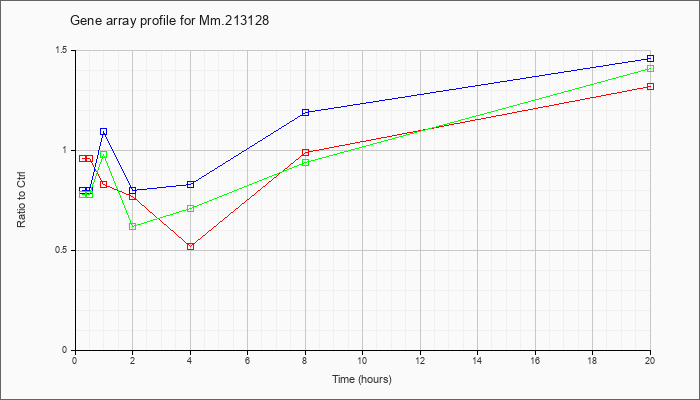 |
141801344 |
Unknown |
KLA | .96 |
.95 |
.83 |
.77 |
.52 |
.99 |
1.32 |
| ATP | .80 |
.82 |
1.09 |
.80 |
.83 |
1.19 |
1.46 |
| KLA/ATP | .78 |
.92 |
.98 |
.62 |
.71 |
.94 |
1.41 |
|
Mm.22263
3 | 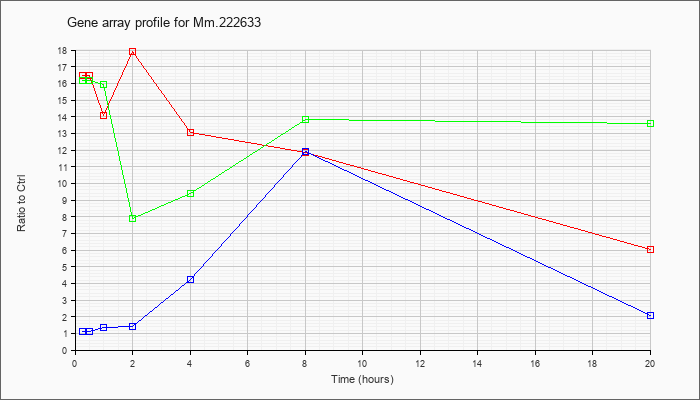 |
83977453 |
Unknown |
KLA | 16.47 |
14.58 |
14.05 |
17.89 |
13.04 |
11.85 |
6.01 |
| ATP | 1.14 |
.97 |
1.34 |
1.39 |
4.26 |
11.88 |
2.10 |
| KLA/ATP | 16.16 |
16.61 |
15.95 |
7.90 |
9.42 |
13.83 |
13.56 |
|
Mm.25366
4 | 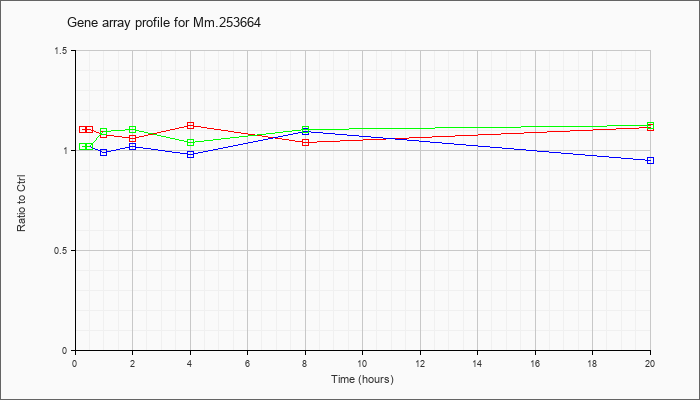 |
6680422 |
Unknown |
KLA | 1.10 |
1.06 |
1.08 |
1.06 |
1.12 |
1.04 |
1.11 |
| ATP | 1.02 |
.98 |
.99 |
1.02 |
.98 |
1.09 |
.95 |
| KLA/ATP | 1.02 |
1.09 |
1.09 |
1.10 |
1.04 |
1.10 |
1.12 |
|
Mm.25381
9 | 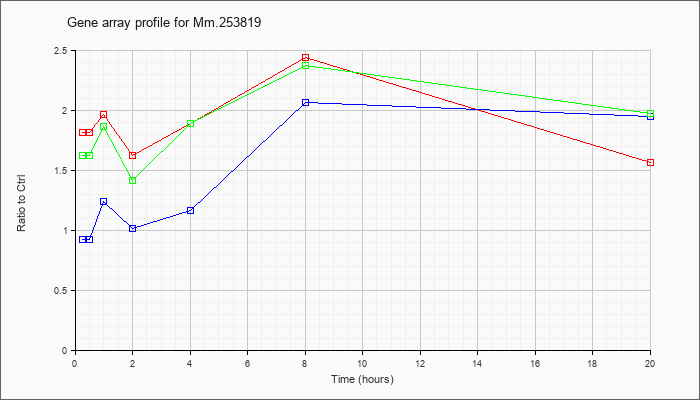 |
118130676 |
Unknown |
KLA | 1.81 |
1.88 |
1.96 |
1.62 |
1.89 |
2.44 |
1.56 |
| ATP | .92 |
.91 |
1.24 |
1.01 |
1.16 |
2.06 |
1.95 |
| KLA/ATP | 1.62 |
1.86 |
1.86 |
1.41 |
1.89 |
2.37 |
1.97 |
|
Mm.27970
| 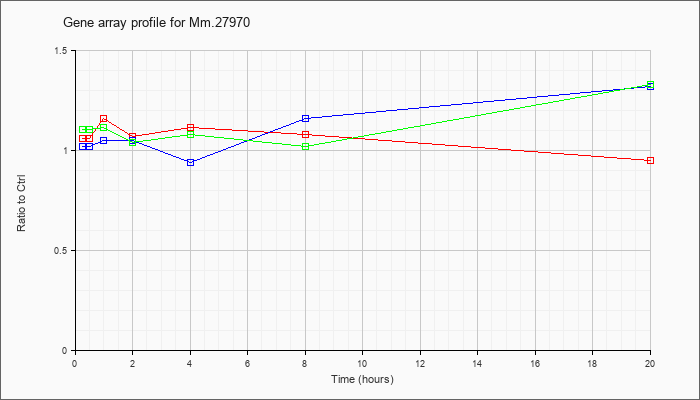 |
13994135 |
Unknown |
KLA | 1.06 |
1.11 |
1.16 |
1.07 |
1.11 |
1.08 |
.95 |
| ATP | 1.02 |
1.09 |
1.05 |
1.05 |
.94 |
1.16 |
1.32 |
| KLA/ATP | 1.10 |
1.19 |
1.11 |
1.04 |
1.08 |
1.02 |
1.33 |
|
Mm.33565
9 | 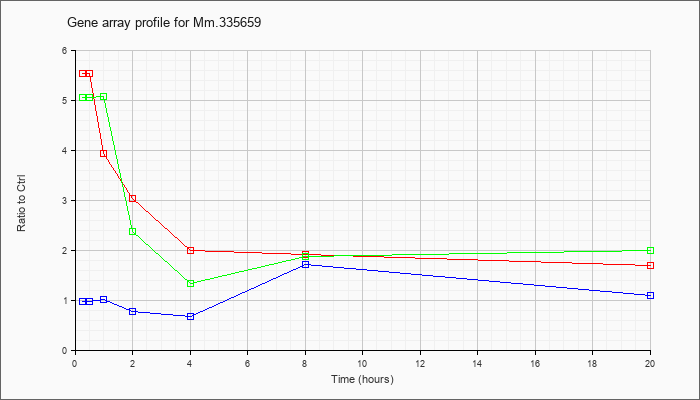 |
160333365 |
Unknown |
KLA | 5.54 |
4.69 |
3.93 |
3.03 |
2.00 |
1.91 |
1.69 |
| ATP | .98 |
.81 |
1.01 |
.78 |
.68 |
1.71 |
1.10 |
| KLA/ATP | 5.05 |
4.44 |
5.07 |
2.38 |
1.34 |
1.88 |
2.00 |
|
Mm.41292
2 | 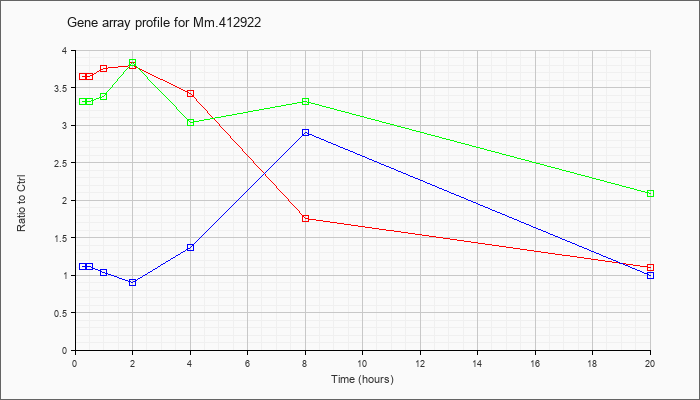 |
145966829 |
Unknown |
KLA | 3.65 |
3.76 |
3.75 |
3.79 |
3.42 |
1.75 |
1.10 |
| ATP | 1.11 |
1.17 |
1.04 |
.90 |
1.37 |
2.90 |
1.00 |
| KLA/ATP | 3.31 |
3.97 |
3.38 |
3.84 |
3.04 |
3.31 |
2.09 |
|
Mm.44128
3 | 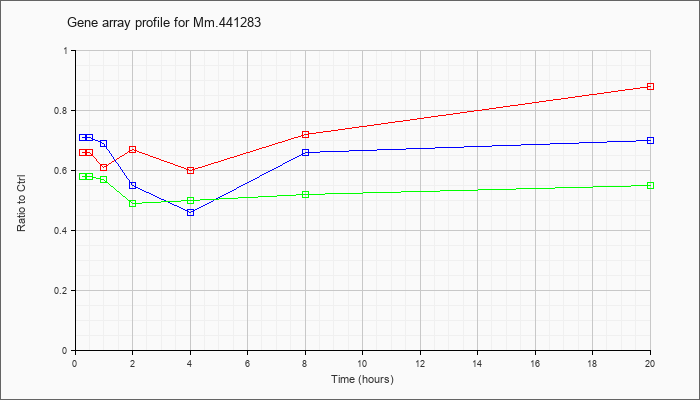 |
149253604 |
Unknown |
KLA | .66 |
.64 |
.61 |
.67 |
.60 |
.72 |
.88 |
| ATP | .71 |
.62 |
.69 |
.55 |
.46 |
.66 |
.70 |
| KLA/ATP | .58 |
.48 |
.57 |
.49 |
.50 |
.52 |
.55 |
|
Mm.45038
3 | 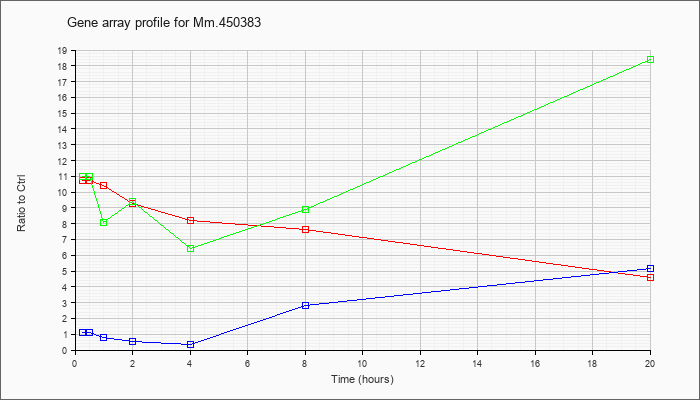 |
184186089 |
Unknown |
KLA | 10.75 |
9.72 |
10.45 |
9.31 |
8.19 |
7.64 |
4.58 |
| ATP | 1.13 |
1.10 |
.81 |
.53 |
.34 |
2.79 |
5.19 |
| KLA/ATP | 10.98 |
9.61 |
8.08 |
9.42 |
6.42 |
8.91 |
18.41 |
|
| Mm.5126 | 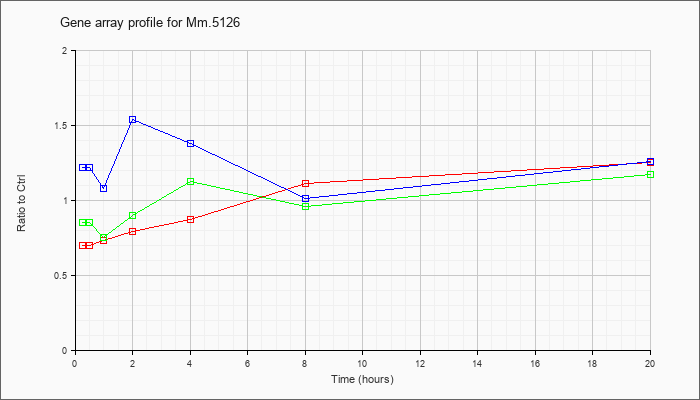 |
158508678 |
Unknown |
KLA | .70 |
.71 |
.73 |
.79 |
.87 |
1.11 |
1.25 |
| ATP | 1.22 |
1.37 |
1.08 |
1.54 |
1.38 |
1.01 |
1.26 |
| KLA/ATP | .85 |
.83 |
.75 |
.90 |
1.12 |
.96 |
1.17 |
|
| Mm.5245 | 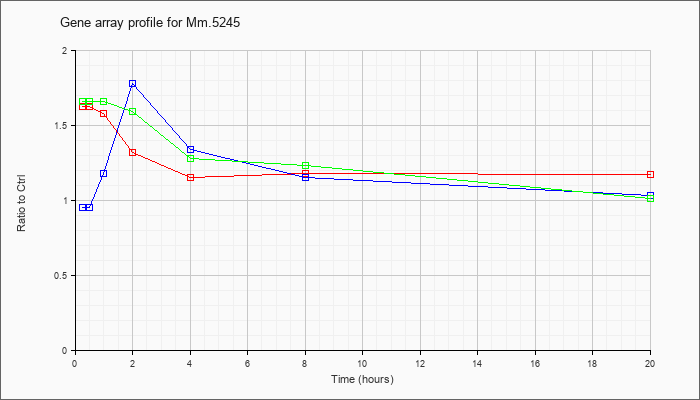 |
118130193 |
Unknown |
KLA | 1.62 |
1.58 |
1.58 |
1.32 |
1.15 |
1.18 |
1.17 |
| ATP | .95 |
.99 |
1.18 |
1.78 |
1.34 |
1.15 |
1.03 |
| KLA/ATP | 1.66 |
1.70 |
1.66 |
1.59 |
1.28 |
1.23 |
1.01 |
|
| Mmp14 | 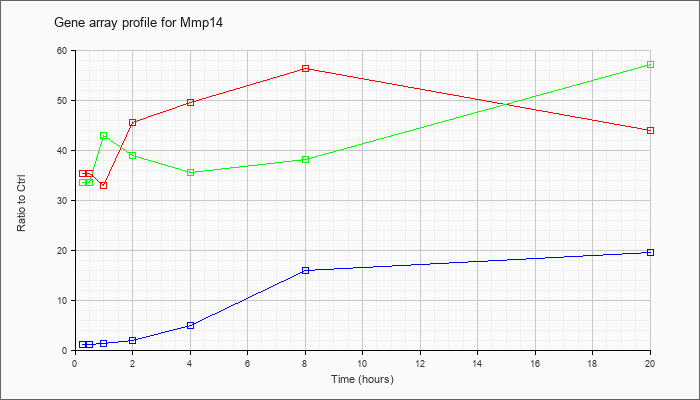 |
NM_008608 |
matrix metallopeptidase 14 (membrane-inserted) (Mmp14), mRNA [NM_008608] |
KLA | 35.40 |
35.40 |
32.92 |
45.56 |
49.47 |
56.29 |
43.99 |
| ATP | 1.01 |
1.01 |
1.36 |
1.97 |
4.97 |
15.97 |
19.45 |
| KLA/ATP | 33.50 |
33.70 |
42.96 |
38.89 |
35.59 |
38.08 |
57.15 |
|
| Mmp3 | 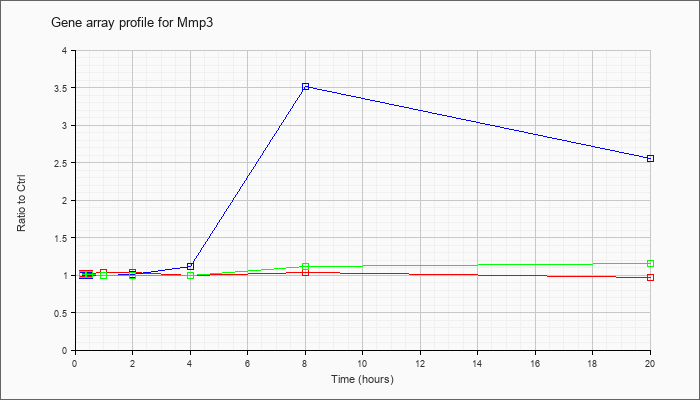 |
NM_010809 |
matrix metallopeptidase 3 (Mmp3), mRNA [NM_010809] |
KLA | 1.02 |
1.04 |
1.04 |
1.04 |
.99 |
1.04 |
.97 |
| ATP | 1.00 |
1.01 |
.99 |
1.01 |
1.12 |
3.52 |
2.55 |
| KLA/ATP | 1.01 |
1.07 |
.99 |
.99 |
.99 |
1.11 |
1.16 |
|
| Mmp9 | 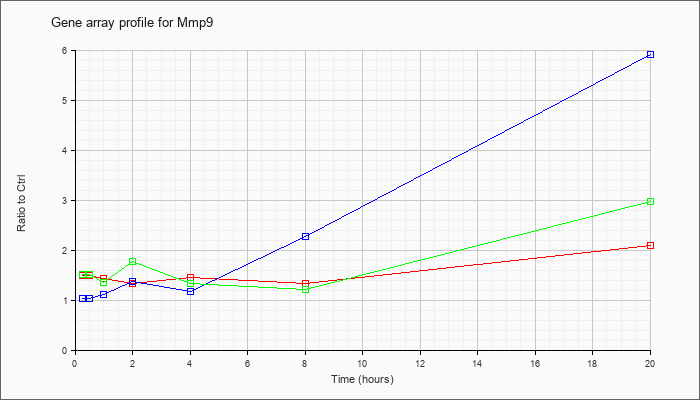 |
NM_013599 |
matrix metallopeptidase 9 (Mmp9), mRNA [NM_013599] |
KLA | 1.49 |
1.65 |
1.43 |
1.32 |
1.46 |
1.33 |
2.10 |
| ATP | 1.03 |
1.05 |
1.12 |
1.36 |
1.17 |
2.27 |
5.92 |
| KLA/ATP | 1.51 |
1.64 |
1.35 |
1.78 |
1.34 |
1.20 |
2.96 |
|
| Nfkb1 | 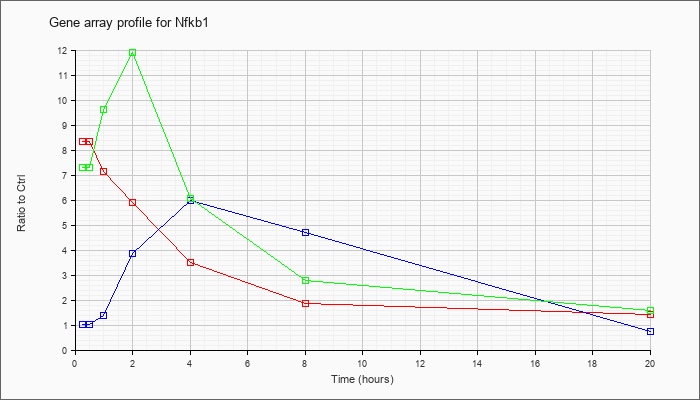 |
BC050841 |
nuclear factor of kappa light chain gene enhancer in B-cells 1, p105, mRNA (cDNA clone IMAGE:6392793), complete cds. [BC050841] |
KLA | 5.81 |
4.96 |
4.77 |
3.21 |
3.37 |
2.00 |
1.71 |
| ATP | 1.39 |
1.52 |
3.25 |
12.63 |
13.64 |
4.80 |
1.08 |
| KLA/ATP | 7.21 |
8.69 |
19.53 |
23.93 |
11.70 |
2.85 |
2.01 |
|
| Nfkb1 |  |
M57999 |
gb|Mouse transcription factor NF-kappa-B DNA binding subunit mRNA, complete cds. [M57999] |
KLA | 8.55 |
7.87 |
7.35 |
6.13 |
3.51 |
1.85 |
1.38 |
| ATP | .98 |
.96 |
1.21 |
3.06 |
5.29 |
4.67 |
.69 |
| KLA/ATP | 7.32 |
7.20 |
8.71 |
10.82 |
5.55 |
2.77 |
1.54 |
|
| Nfkbia | 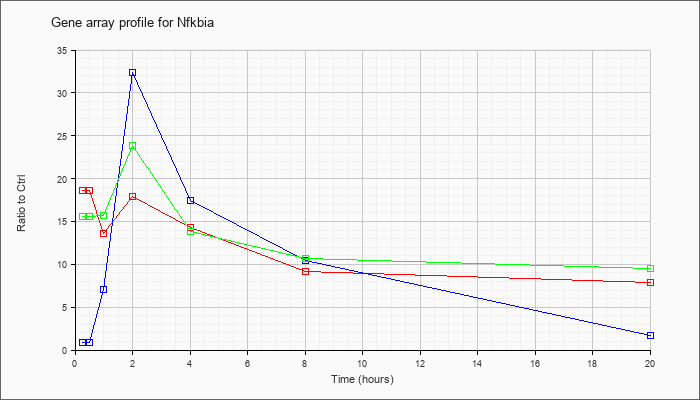 |
NM_010907 |
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha (Nfkbia), mRNA [NM_010907] |
KLA | 18.65 |
18.46 |
13.63 |
17.88 |
14.25 |
9.17 |
7.92 |
| ATP | .91 |
1.09 |
7.11 |
32.39 |
17.44 |
10.49 |
1.72 |
| KLA/ATP | 15.57 |
17.66 |
15.73 |
23.91 |
13.83 |
10.73 |
9.48 |
|
| Nod2 | 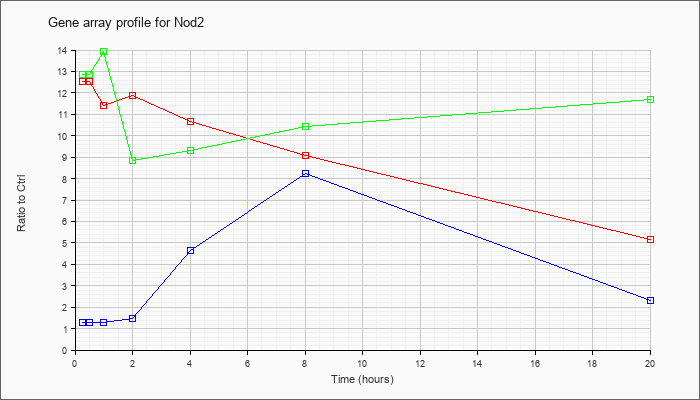 |
NM_145857 |
nucleotide-binding oligomerization domain containing 2 (Nod2), mRNA [NM_145857] |
KLA | 12.51 |
10.83 |
11.39 |
11.89 |
10.66 |
9.06 |
5.17 |
| ATP | 1.27 |
1.10 |
1.30 |
1.48 |
4.63 |
8.23 |
2.33 |
| KLA/ATP | 12.86 |
12.85 |
13.93 |
8.85 |
9.30 |
10.43 |
11.67 |
|
| Pgam5 | 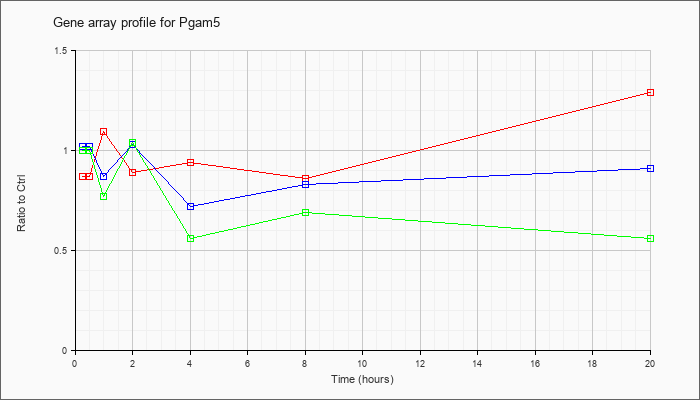 |
NM_028273 |
phosphoglycerate mutase family member 5 (Pgam5), mRNA [NM_028273] |
KLA | .87 |
1.04 |
1.09 |
.89 |
.94 |
.86 |
1.29 |
| ATP | 1.02 |
1.09 |
.87 |
1.03 |
.72 |
.83 |
.91 |
| KLA/ATP | 1.00 |
.97 |
.77 |
1.04 |
.56 |
.69 |
.56 |
|
| Pik3ca | 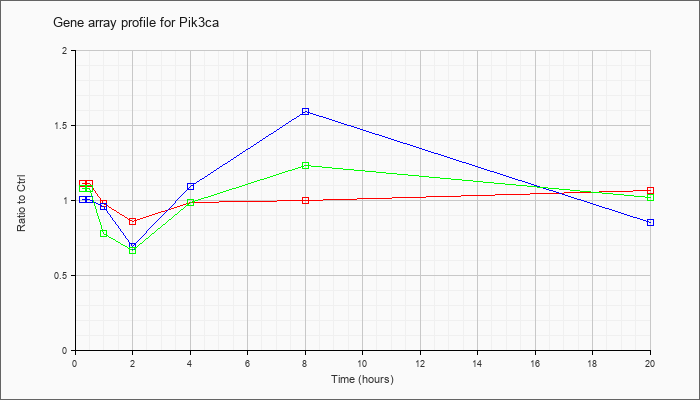 |
ENSMUST00000108243 |
ens|Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha isoform (EC 2.7.1.153) (PI3-kinase p110 subunit alpha) (PtdIns-3- kinase p110) (PI3K). [Source:Uniprot/SWISSPROT;Acc:P42337] [ENSMUST00000108243] |
KLA | 1.07 |
1.01 |
.91 |
.90 |
.98 |
.96 |
.86 |
| ATP | 1.03 |
.87 |
1.12 |
.63 |
1.22 |
1.86 |
.60 |
| KLA/ATP | 1.05 |
.82 |
.91 |
.61 |
1.17 |
1.26 |
.63 |
|
| Pik3ca |  |
NM_008839 |
phosphatidylinositol 3-kinase, catalytic, alpha polypeptide (Pik3ca), mRNA [NM_008839] |
KLA | 1.15 |
1.11 |
1.05 |
.82 |
.99 |
1.03 |
1.27 |
| ATP | .98 |
1.00 |
.79 |
.75 |
.96 |
1.32 |
1.10 |
| KLA/ATP | 1.10 |
.98 |
.64 |
.72 |
.80 |
1.20 |
1.41 |
|
| Pik3cb | 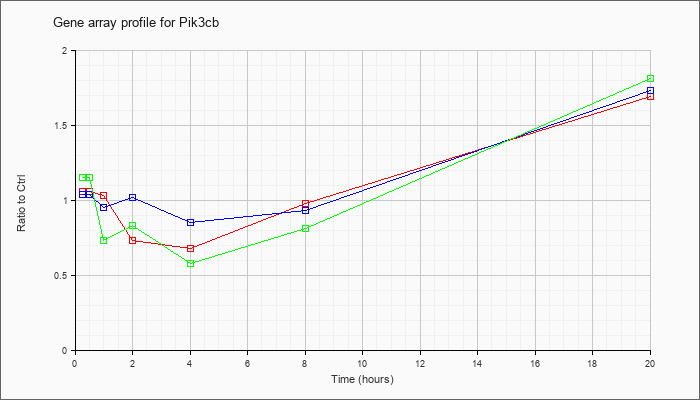 |
NM_029094 |
phosphatidylinositol 3-kinase, catalytic, beta polypeptide (Pik3cb), mRNA [NM_029094] |
KLA | 1.06 |
1.06 |
1.03 |
.73 |
.68 |
.98 |
1.69 |
| ATP | 1.04 |
1.21 |
.95 |
1.02 |
.85 |
.93 |
1.73 |
| KLA/ATP | 1.15 |
1.27 |
.73 |
.83 |
.58 |
.81 |
1.81 |
|
| Pik3cd | 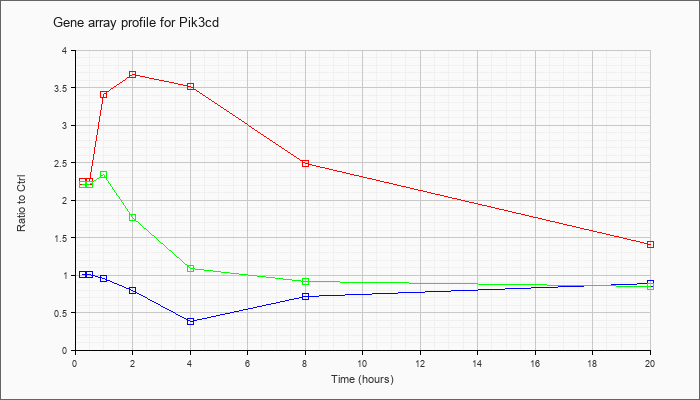 |
NM_008840 |
phosphatidylinositol 3-kinase catalytic delta polypeptide (Pik3cd), transcript variant 1, mRNA [NM_008840] |
KLA | 2.28 |
2.42 |
3.10 |
4.17 |
3.35 |
2.31 |
1.34 |
| ATP | 1.02 |
.91 |
.99 |
.63 |
.30 |
.67 |
.80 |
| KLA/ATP | 2.14 |
2.23 |
2.70 |
1.51 |
1.17 |
.85 |
.72 |
|
| Pik3cd |  |
U86587 |
phosphatidylinositol 3-kinase catalytic subunit p110 delta mRNA, complete cds. [U86587] |
KLA | 2.21 |
2.24 |
3.71 |
3.17 |
3.67 |
2.66 |
1.48 |
| ATP | 1.00 |
1.03 |
.93 |
.96 |
.46 |
.75 |
.97 |
| KLA/ATP | 2.27 |
2.32 |
1.97 |
2.02 |
1.01 |
.99 |
.98 |
|
| Pik3cg | 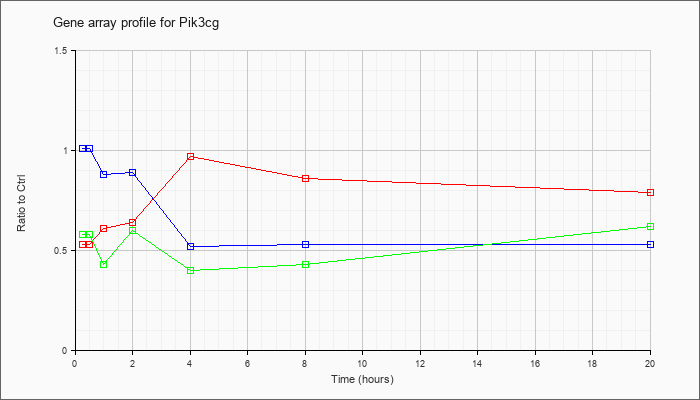 |
NM_020272 |
phosphoinositide-3-kinase, catalytic, gamma polypeptide (Pik3cg), mRNA [NM_020272] |
KLA | .53 |
.53 |
.61 |
.64 |
.97 |
.86 |
.79 |
| ATP | 1.01 |
1.13 |
.88 |
.89 |
.52 |
.53 |
.53 |
| KLA/ATP | .58 |
.53 |
.43 |
.60 |
.40 |
.43 |
.62 |
|
| Pik3r1 | 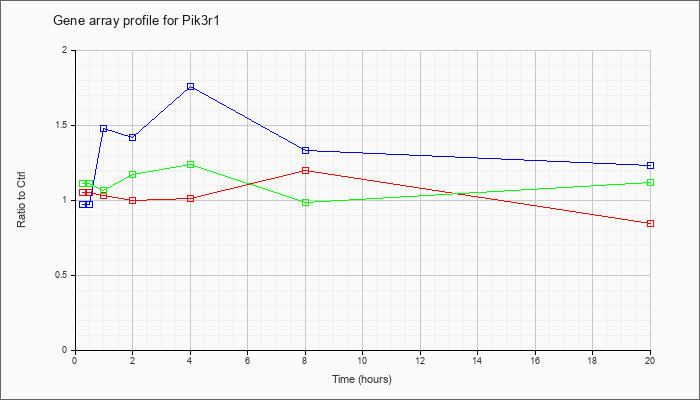 |
NM_001077495 |
phosphatidylinositol 3-kinase, regulatory subunit, polypeptide 1 (p85 alpha) (Pik3r1), transcript variant 2, mRNA [NM_001077495] |
KLA | 1.05 |
1.07 |
1.03 |
1.00 |
1.01 |
1.20 |
.85 |
| ATP | .97 |
1.02 |
1.48 |
1.42 |
1.76 |
1.33 |
1.23 |
| KLA/ATP | 1.11 |
1.12 |
1.06 |
1.17 |
1.24 |
.99 |
1.12 |
|
| Pik3r2 | 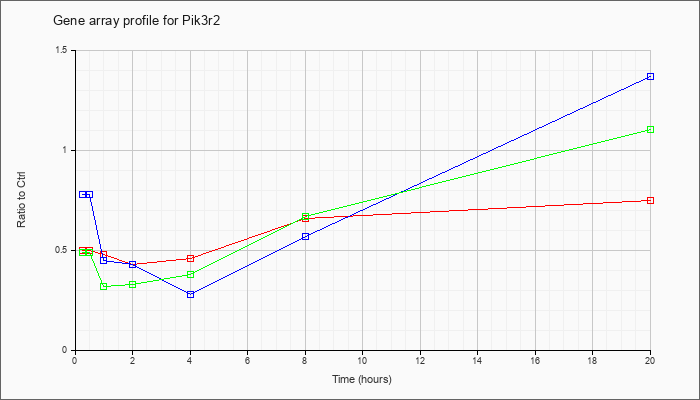 |
NM_008841 |
phosphatidylinositol 3-kinase, regulatory subunit, polypeptide 2 (p85 beta) (Pik3r2), mRNA [NM_008841] |
KLA | .50 |
.50 |
.48 |
.43 |
.46 |
.66 |
.75 |
| ATP | .78 |
.68 |
.45 |
.43 |
.28 |
.57 |
1.37 |
| KLA/ATP | .49 |
.38 |
.32 |
.33 |
.38 |
.67 |
1.10 |
|
| Pik3r3 | 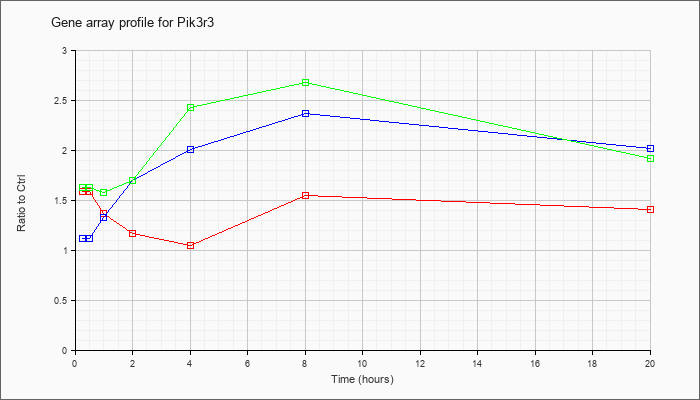 |
NM_181585 |
phosphatidylinositol 3 kinase, regulatory subunit, polypeptide 3 (p55) (Pik3r3), mRNA [NM_181585] |
KLA | 1.59 |
1.49 |
1.37 |
1.17 |
1.05 |
1.55 |
1.41 |
| ATP | 1.12 |
1.00 |
1.33 |
1.70 |
2.01 |
2.37 |
2.02 |
| KLA/ATP | 1.63 |
1.54 |
1.58 |
1.70 |
2.43 |
2.68 |
1.92 |
|
| Pik3r5 | 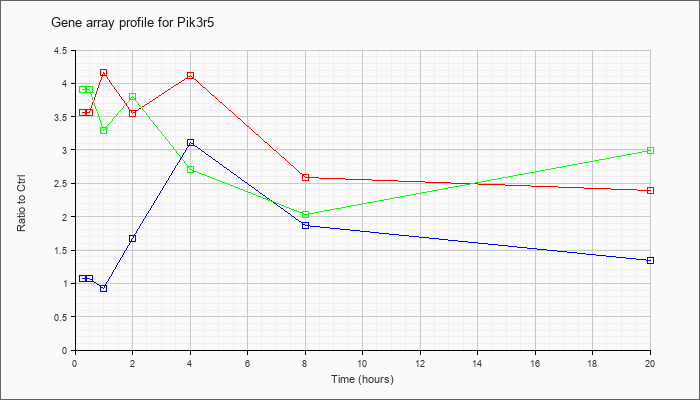 |
NM_177320 |
phosphoinositide-3-kinase, regulatory subunit 5, p101 (Pik3r5), mRNA [NM_177320] |
KLA | 3.57 |
3.58 |
4.16 |
3.54 |
4.11 |
2.59 |
2.39 |
| ATP | 1.08 |
1.08 |
.93 |
1.67 |
3.11 |
1.88 |
1.34 |
| KLA/ATP | 3.91 |
3.70 |
3.30 |
3.80 |
2.70 |
2.04 |
3.00 |
|
| Ptgs2 | 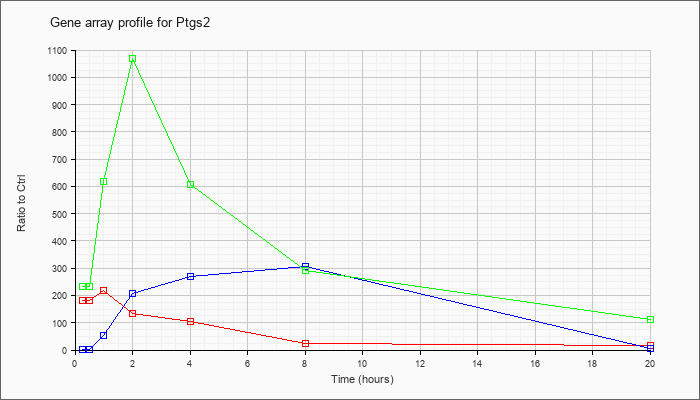 |
NM_011198 |
prostaglandin-endoperoxide synthase 2 (Ptgs2), mRNA [NM_011198] |
KLA | 181.04 |
220.88 |
218.58 |
132.79 |
103.05 |
22.69 |
18.02 |
| ATP | 2.47 |
9.45 |
52.13 |
208.62 |
271.22 |
306.11 |
6.86 |
| KLA/ATP | 233.39 |
387.29 |
617.86 |
###.## |
608.44 |
290.80 |
112.22 |
|
| Rela | 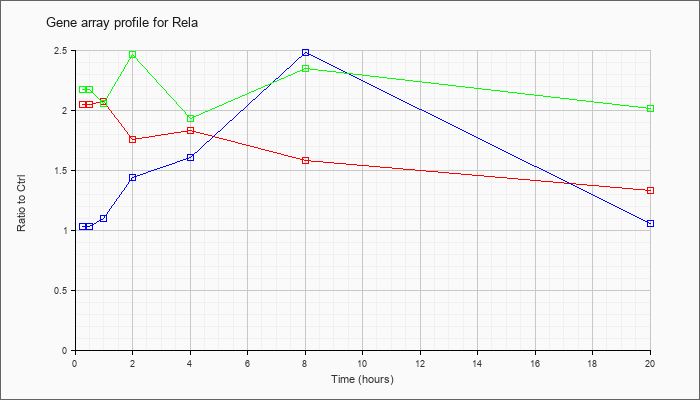 |
NM_009045 |
v-rel reticuloendotheliosis viral oncogene homolog A (avian) (Rela), mRNA [NM_009045] |
KLA | 2.05 |
2.05 |
2.07 |
1.75 |
1.83 |
1.58 |
1.33 |
| ATP | 1.03 |
1.08 |
1.10 |
1.44 |
1.61 |
2.48 |
1.05 |
| KLA/ATP | 2.17 |
2.27 |
2.05 |
2.47 |
1.93 |
2.35 |
2.02 |
|
| Ripk1 | 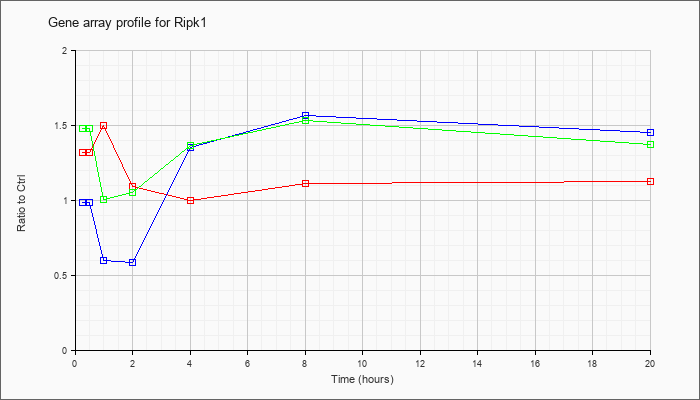 |
NM_009068 |
receptor (TNFRSF)-interacting serine-threonine kinase 1 (Ripk1), mRNA [NM_009068] |
KLA | 1.32 |
1.33 |
1.50 |
1.09 |
1.00 |
1.11 |
1.13 |
| ATP | .98 |
.94 |
.60 |
.59 |
1.35 |
1.56 |
1.45 |
| KLA/ATP | 1.48 |
1.30 |
1.01 |
1.05 |
1.36 |
1.53 |
1.37 |
|
| Ripk3 | 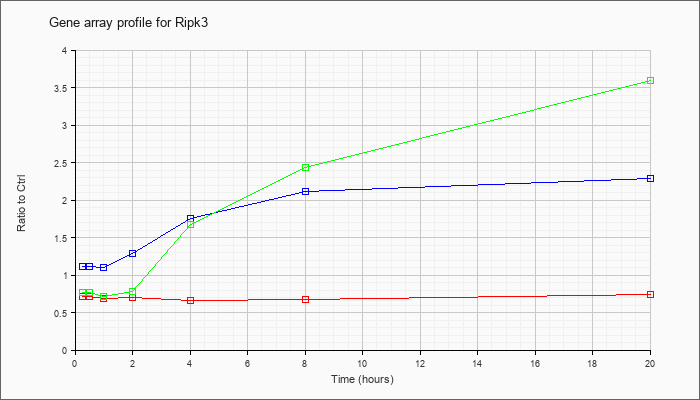 |
NM_019955 |
receptor-interacting serine-threonine kinase 3 (Ripk3), mRNA [NM_019955] |
KLA | .71 |
.68 |
.69 |
.70 |
.66 |
.67 |
.74 |
| ATP | 1.11 |
1.03 |
1.10 |
1.29 |
1.75 |
2.12 |
2.29 |
| KLA/ATP | .77 |
.73 |
.72 |
.78 |
1.67 |
2.43 |
3.60 |
|
| Rps6ka4 | 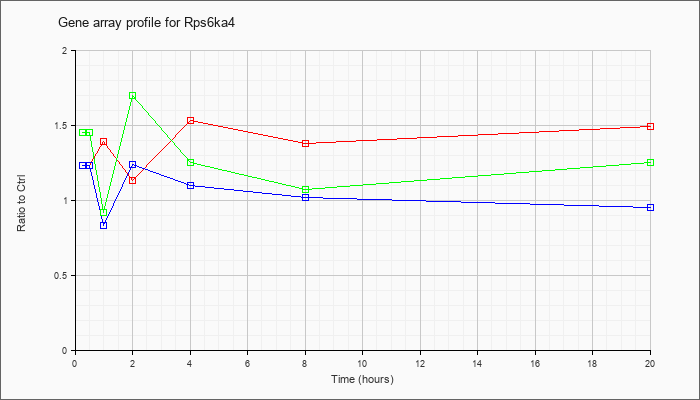 |
NM_019924 |
ribosomal protein S6 kinase, polypeptide 4 (Rps6ka4), mRNA [NM_019924] |
KLA | 1.23 |
1.21 |
1.39 |
1.13 |
1.53 |
1.38 |
1.49 |
| ATP | 1.23 |
1.29 |
.83 |
1.24 |
1.10 |
1.02 |
.95 |
| KLA/ATP | 1.45 |
1.61 |
.92 |
1.70 |
1.25 |
1.07 |
1.25 |
|
| Rps6ka5 | 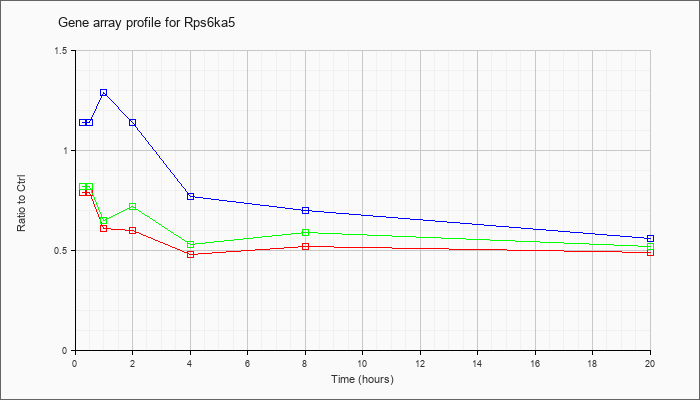 |
NM_153587 |
ribosomal protein S6 kinase, polypeptide 5 (Rps6ka5), mRNA [NM_153587] |
KLA | .79 |
.69 |
.61 |
.60 |
.48 |
.52 |
.49 |
| ATP | 1.14 |
1.25 |
1.29 |
1.14 |
.77 |
.70 |
.56 |
| KLA/ATP | .82 |
.73 |
.65 |
.72 |
.53 |
.59 |
.52 |
|
| Sele | 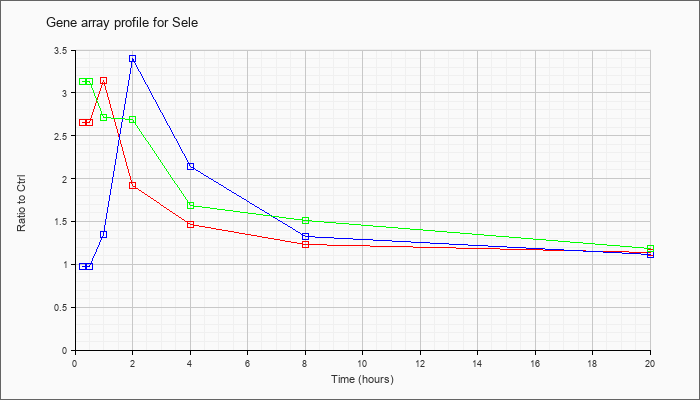 |
NM_011345 |
selectin, endothelial cell (Sele), mRNA [NM_011345] |
KLA | 2.65 |
3.18 |
3.14 |
1.92 |
1.46 |
1.23 |
1.14 |
| ATP | .97 |
1.01 |
1.35 |
3.40 |
2.14 |
1.33 |
1.11 |
| KLA/ATP | 3.13 |
3.40 |
2.71 |
2.69 |
1.69 |
1.51 |
1.18 |
|
| Socs3 | 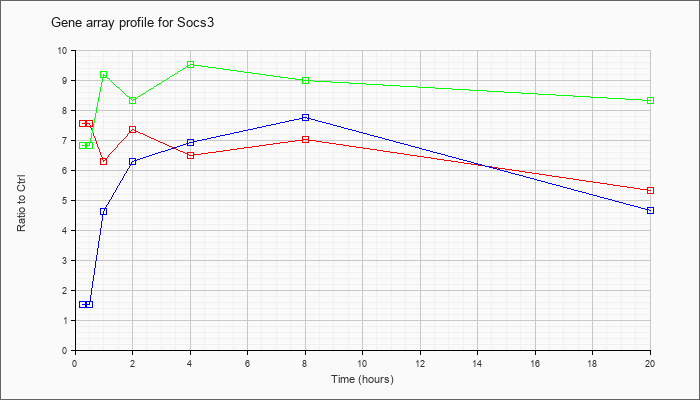 |
NM_007707 |
suppressor of cytokine signaling 3 (Socs3), mRNA [NM_007707] |
KLA | 7.56 |
6.68 |
6.29 |
7.36 |
6.47 |
7.03 |
5.33 |
| ATP | 1.51 |
1.75 |
4.63 |
6.27 |
6.92 |
7.74 |
4.64 |
| KLA/ATP | 6.82 |
6.77 |
9.19 |
8.32 |
9.51 |
8.98 |
8.33 |
|
| Tnf | 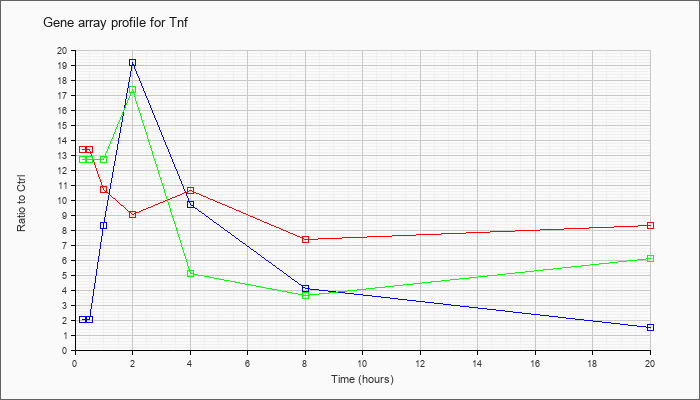 |
NM_013693 |
tumor necrosis factor (Tnf), mRNA [NM_013693] |
KLA | 13.35 |
12.76 |
10.71 |
9.03 |
10.64 |
7.39 |
8.32 |
| ATP | 2.01 |
2.48 |
8.27 |
19.20 |
9.72 |
4.09 |
1.49 |
| KLA/ATP | 12.72 |
15.76 |
12.73 |
17.34 |
5.10 |
3.61 |
6.07 |
|
| Tnfaip3 | 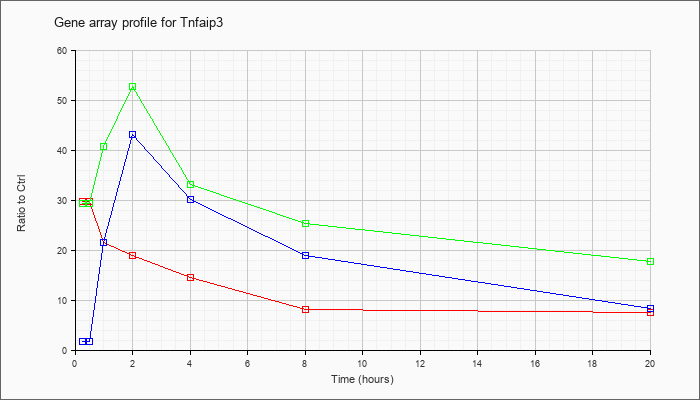 |
NM_009397 |
tumor necrosis factor, alpha-induced protein 3 (Tnfaip3), mRNA [NM_009397] |
KLA | 29.69 |
27.73 |
21.42 |
18.86 |
14.42 |
8.13 |
7.48 |
| ATP | 1.73 |
4.86 |
21.57 |
43.06 |
30.03 |
18.86 |
8.31 |
| KLA/ATP | 29.39 |
35.88 |
40.64 |
52.71 |
33.19 |
25.38 |
17.75 |
|
Tnfrsf1a
| 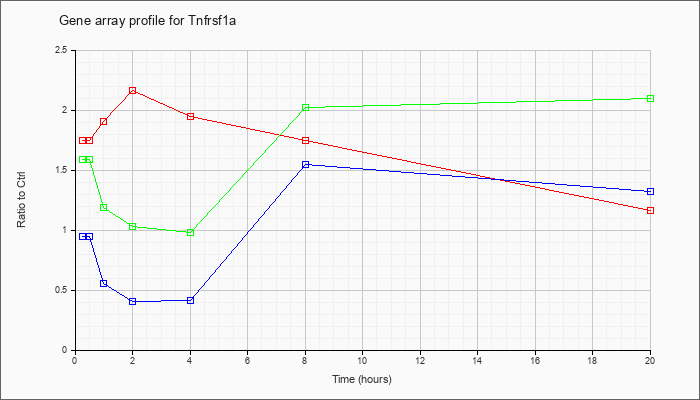 |
NM_011609 |
tumor necrosis factor receptor superfamily, member 1a (Tnfrsf1a), mRNA [NM_011609] |
KLA | 1.74 |
1.71 |
1.90 |
2.16 |
1.95 |
1.74 |
1.16 |
| ATP | .95 |
.83 |
.55 |
.41 |
.41 |
1.55 |
1.32 |
| KLA/ATP | 1.58 |
1.49 |
1.19 |
1.03 |
.98 |
2.02 |
2.10 |
|
Tnfrsf1b
| 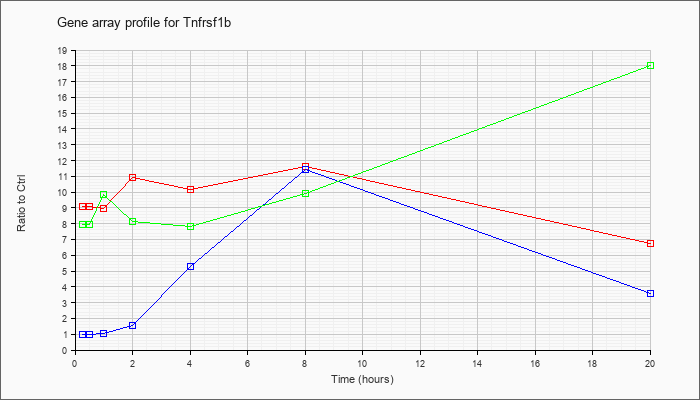 |
NM_011610 |
tumor necrosis factor receptor superfamily, member 1b (Tnfrsf1b), mRNA [NM_011610] |
KLA | 9.07 |
9.05 |
8.93 |
10.89 |
10.14 |
11.61 |
6.74 |
| ATP | .98 |
.95 |
1.03 |
1.55 |
5.28 |
11.41 |
3.60 |
| KLA/ATP | 7.95 |
8.55 |
9.86 |
8.13 |
7.80 |
9.94 |
18.02 |
|
| Tradd | 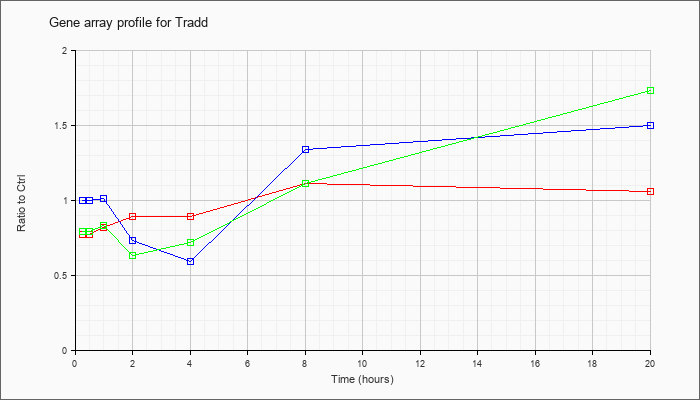 |
NM_001033161 |
TNFRSF1A-associated via death domain (Tradd), mRNA [NM_001033161] |
KLA | .77 |
.71 |
.82 |
.89 |
.89 |
1.11 |
1.06 |
| ATP | 1.00 |
.90 |
1.01 |
.73 |
.59 |
1.34 |
1.50 |
| KLA/ATP | .79 |
.73 |
.83 |
.63 |
.72 |
1.11 |
1.73 |
|
| Traf1 | 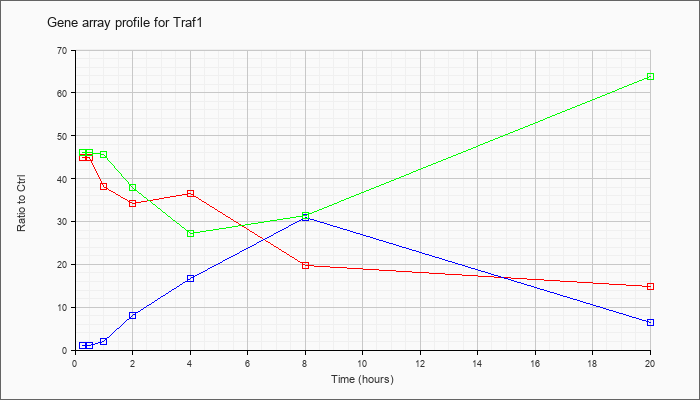 |
NM_009421 |
Tnf receptor-associated factor 1 (Traf1), mRNA [NM_009421] |
KLA | 44.86 |
39.03 |
38.11 |
34.21 |
36.49 |
19.69 |
14.75 |
| ATP | 1.14 |
1.04 |
1.98 |
8.16 |
16.79 |
30.81 |
6.36 |
| KLA/ATP | 46.20 |
45.67 |
45.63 |
38.00 |
27.26 |
31.47 |
63.83 |
|
| Traf2 |  |
NM_009422 |
Tnf receptor-associated factor 2 (Traf2), mRNA [NM_009422] |
KLA | 3.38 |
3.62 |
3.49 |
3.36 |
3.49 |
3.39 |
2.68 |
| ATP | .97 |
1.04 |
1.00 |
1.10 |
1.54 |
2.40 |
1.55 |
| KLA/ATP | 3.38 |
3.16 |
2.93 |
2.74 |
2.67 |
3.11 |
4.70 |
|
| Traf3 | 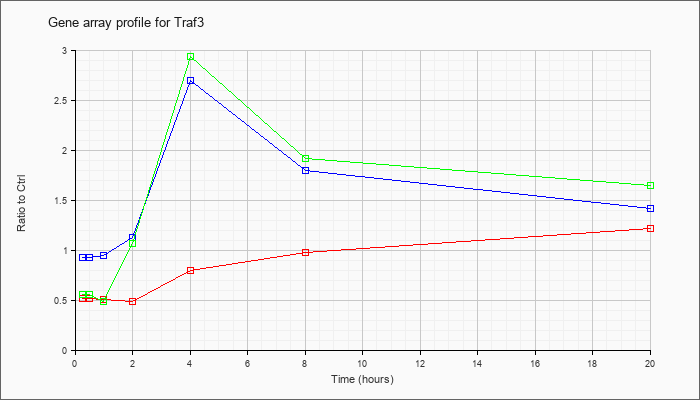 |
NM_011632 |
Tnf receptor-associated factor 3 (Traf3), transcript variant 1, mRNA [NM_011632] |
KLA | .52 |
.49 |
.51 |
.49 |
.80 |
.98 |
1.22 |
| ATP | .93 |
.92 |
.95 |
1.13 |
2.70 |
1.80 |
1.42 |
| KLA/ATP | .56 |
.50 |
.49 |
1.07 |
2.94 |
1.92 |
1.65 |
|
| Traf5 | 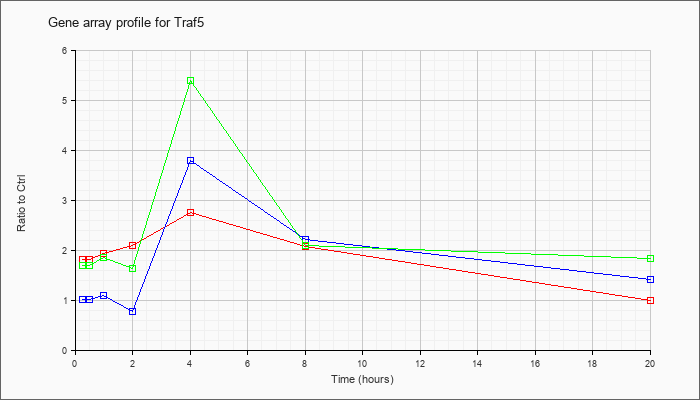 |
NM_011633 |
Tnf receptor-associated factor 5 (Traf5), mRNA [NM_011633] |
KLA | 1.82 |
1.92 |
1.93 |
2.10 |
2.75 |
2.08 |
1.00 |
| ATP | 1.02 |
1.04 |
1.10 |
.78 |
3.79 |
2.22 |
1.42 |
| KLA/ATP | 1.70 |
1.89 |
1.85 |
1.64 |
5.40 |
2.10 |
1.83 |
|
| Vcam1 | 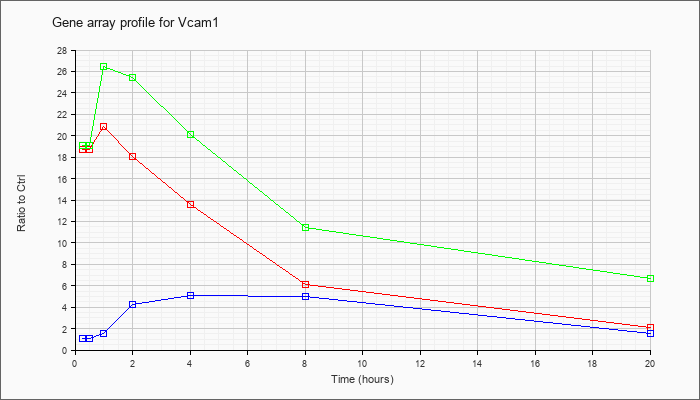 |
NM_011693 |
vascular cell adhesion molecule 1 (Vcam1), mRNA [NM_011693] |
KLA | 18.71 |
22.30 |
20.87 |
18.09 |
13.61 |
6.09 |
2.09 |
| ATP | 1.08 |
1.00 |
1.56 |
4.23 |
5.09 |
5.04 |
1.50 |
| KLA/ATP | 19.09 |
25.02 |
26.43 |
25.42 |
20.08 |
11.45 |
6.67 |
|