| Gene symbol |
Time course plot |
Accession |
Description |
Treatment |
15min |
30min |
1hr |
2hr |
4hr |
8hr |
20hr |
| Adcy1 | 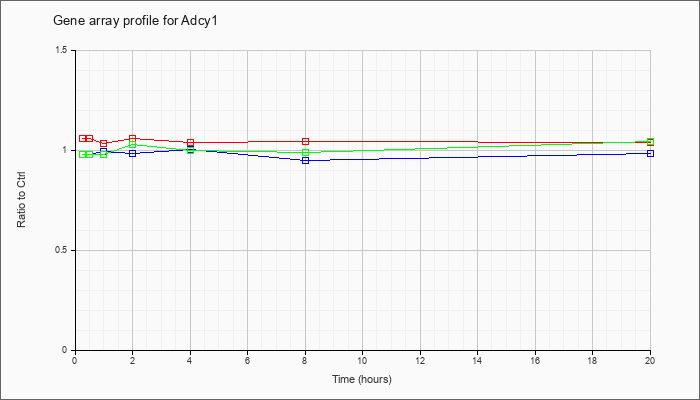 |
BC050125 |
adenylate cyclase 1, mRNA (cDNA clone IMAGE:30015692). [BC050125] |
KLA | 1.05 |
1.03 |
.98 |
1.04 |
1.05 |
1.09 |
1.10 |
| ATP | .90 |
1.02 |
.99 |
.97 |
1.03 |
.95 |
1.01 |
| KLA/ATP | .99 |
1.04 |
.90 |
.98 |
.98 |
1.00 |
1.03 |
|
| Adcy1 |  |
NM_009622 |
adenylate cyclase 1 (Adcy1), mRNA [NM_009622] |
KLA | 1.07 |
1.02 |
1.06 |
1.07 |
1.03 |
1.02 |
1.01 |
| ATP | 1.02 |
1.04 |
1.00 |
.99 |
.99 |
.95 |
.97 |
| KLA/ATP | .97 |
1.05 |
1.02 |
1.05 |
1.01 |
.98 |
1.05 |
|
| Adcy2 | 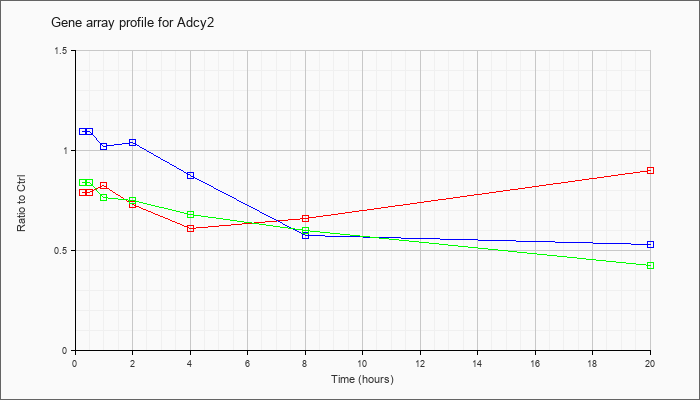 |
NM_153534 |
adenylate cyclase 2 (Adcy2), mRNA [NM_153534] |
KLA | .79 |
.82 |
.83 |
.73 |
.61 |
.66 |
.90 |
| ATP | 1.10 |
1.09 |
1.02 |
1.04 |
.88 |
.58 |
.53 |
| KLA/ATP | .84 |
.87 |
.77 |
.75 |
.68 |
.60 |
.43 |
|
| Adcy3 | 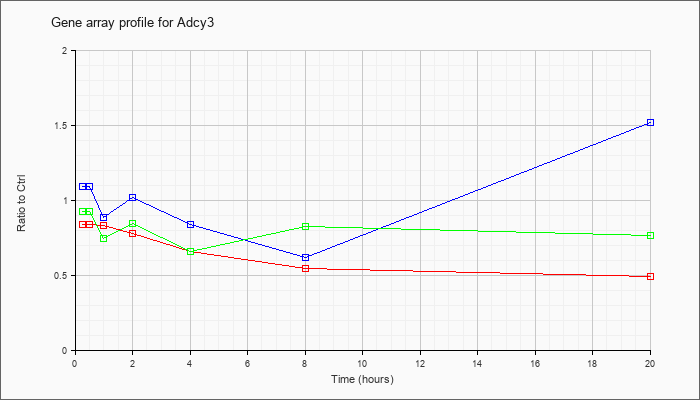 |
NM_138305 |
adenylate cyclase 3 (Adcy3), mRNA [NM_138305] |
KLA | .84 |
.84 |
.83 |
.78 |
.66 |
.55 |
.49 |
| ATP | 1.09 |
1.05 |
.89 |
1.02 |
.84 |
.62 |
1.52 |
| KLA/ATP | .93 |
.87 |
.75 |
.85 |
.66 |
.83 |
.77 |
|
| Adcy4 | 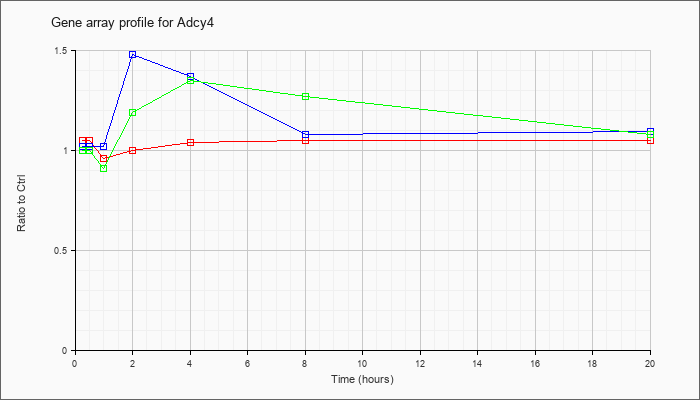 |
NM_080435 |
adenylate cyclase 4 (Adcy4), mRNA [NM_080435] |
KLA | 1.05 |
1.07 |
.96 |
1.00 |
1.04 |
1.05 |
1.05 |
| ATP | 1.02 |
1.05 |
1.02 |
1.48 |
1.37 |
1.08 |
1.09 |
| KLA/ATP | 1.00 |
1.05 |
.91 |
1.19 |
1.35 |
1.27 |
1.08 |
|
| Adcy5 |  |
NM_001012765 |
adenylate cyclase 5 (Adcy5), mRNA [NM_001012765] |
KLA | 1.06 |
1.00 |
.97 |
1.01 |
1.03 |
.95 |
.93 |
| ATP | .95 |
1.08 |
1.08 |
1.07 |
.97 |
1.03 |
.97 |
| KLA/ATP | .97 |
.92 |
1.04 |
.99 |
.91 |
.96 |
1.04 |
|
| Adcy6 | 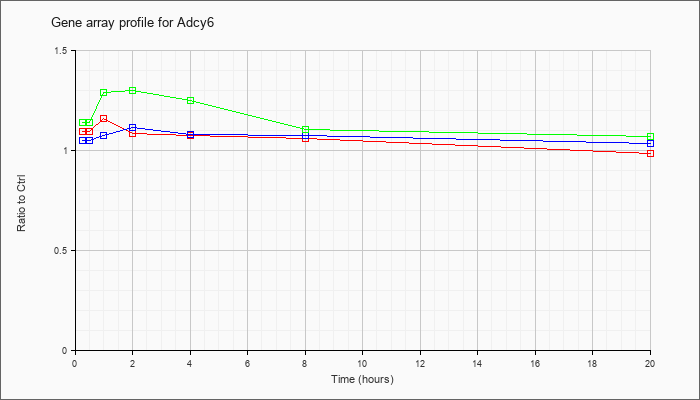 |
NM_007405 |
adenylate cyclase 6 (Adcy6), mRNA [NM_007405] |
KLA | 1.10 |
1.16 |
1.16 |
1.09 |
1.08 |
1.06 |
.99 |
| ATP | 1.05 |
.97 |
1.08 |
1.12 |
1.08 |
1.08 |
1.04 |
| KLA/ATP | 1.14 |
1.20 |
1.29 |
1.30 |
1.25 |
1.11 |
1.07 |
|
| Adcy7 |  |
NM_001037724 |
adenylate cyclase 7 (Adcy7), transcript variant 3, mRNA [NM_001037724] |
KLA | .43 |
.39 |
.35 |
.32 |
.45 |
.72 |
.84 |
| ATP | .90 |
.84 |
.62 |
.41 |
.53 |
.42 |
.94 |
| KLA/ATP | .39 |
.34 |
.23 |
.17 |
.41 |
.34 |
.67 |
|
| Adcy7 |  |
NM_007406 |
adenylate cyclase 7 (Adcy7), transcript variant 1, mRNA [NM_007406] |
KLA | .51 |
.52 |
.48 |
.41 |
.59 |
.81 |
.99 |
| ATP | 1.01 |
1.02 |
.62 |
.52 |
.53 |
.38 |
.98 |
| KLA/ATP | .53 |
.48 |
.30 |
.27 |
.42 |
.36 |
.65 |
|
| Adcy8 | 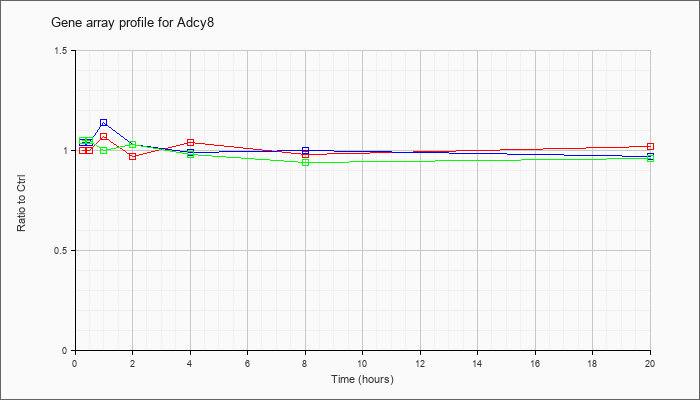 |
NM_009623 |
adenylate cyclase 8 (Adcy8), mRNA [NM_009623] |
KLA | 1.00 |
1.00 |
1.07 |
.97 |
1.04 |
.98 |
1.02 |
| ATP | 1.04 |
1.05 |
1.14 |
1.03 |
.99 |
1.00 |
.97 |
| KLA/ATP | 1.05 |
1.04 |
1.00 |
1.03 |
.98 |
.94 |
.96 |
|
| Adcy9 | 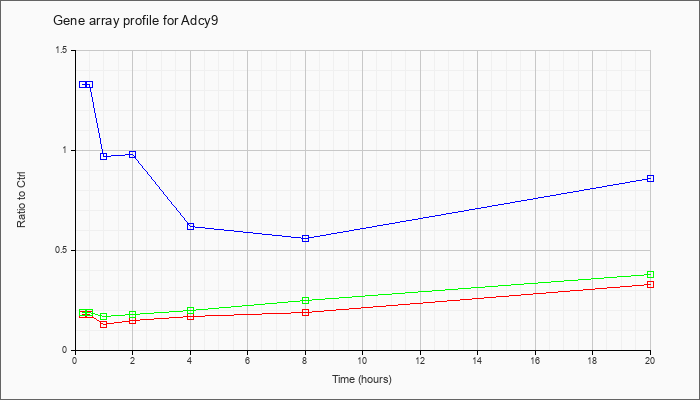 |
NM_009624 |
adenylate cyclase 9 (Adcy9), mRNA [NM_009624] |
KLA | .18 |
.15 |
.13 |
.15 |
.17 |
.19 |
.33 |
| ATP | 1.33 |
1.16 |
.97 |
.98 |
.62 |
.56 |
.86 |
| KLA/ATP | .19 |
.16 |
.17 |
.18 |
.20 |
.25 |
.38 |
|
| Akt1 | 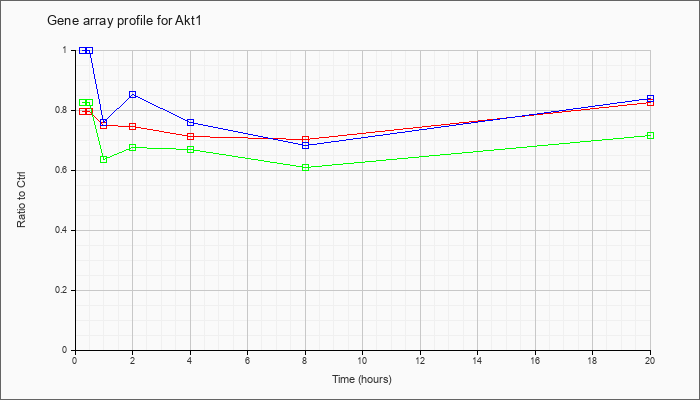 |
NM_009652 |
thymoma viral proto-oncogene 1 (Akt1), mRNA [NM_009652] |
KLA | .80 |
.80 |
.75 |
.75 |
.71 |
.70 |
.82 |
| ATP | 1.00 |
1.02 |
.76 |
.85 |
.76 |
.68 |
.84 |
| KLA/ATP | .82 |
.80 |
.64 |
.68 |
.67 |
.61 |
.72 |
|
| Akt2 | 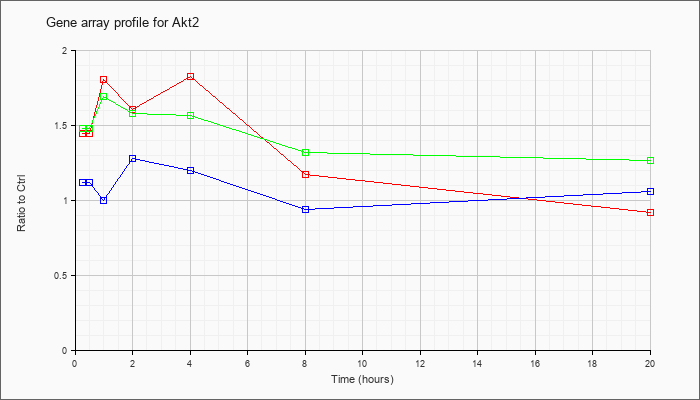 |
NM_007434 |
thymoma viral proto-oncogene 2 (Akt2), transcript variant 2, mRNA [NM_007434] |
KLA | 1.45 |
1.49 |
1.81 |
1.61 |
1.83 |
1.17 |
.92 |
| ATP | 1.12 |
1.12 |
1.00 |
1.28 |
1.20 |
.94 |
1.06 |
| KLA/ATP | 1.48 |
1.70 |
1.69 |
1.58 |
1.57 |
1.32 |
1.27 |
|
| Akt3 | 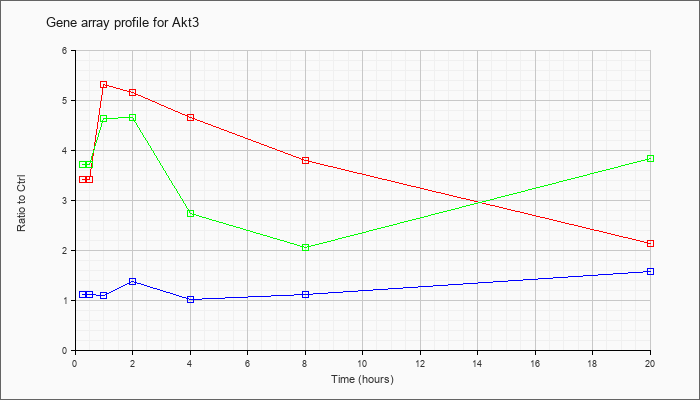 |
NM_011785 |
thymoma viral proto-oncogene 3 (Akt3), mRNA [NM_011785] |
KLA | 3.42 |
3.45 |
5.31 |
5.16 |
4.65 |
3.80 |
2.12 |
| ATP | 1.12 |
1.22 |
1.10 |
1.37 |
1.02 |
1.11 |
1.56 |
| KLA/ATP | 3.72 |
4.17 |
4.62 |
4.64 |
2.74 |
2.05 |
3.82 |
|
| Anapc1 | 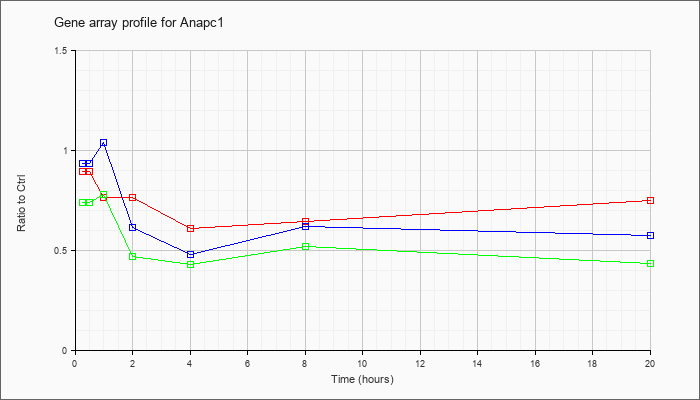 |
AK090134 |
nullipotent stem cell CRL-2070 NE cDNA, RIKEN full-length enriched library, clone:G431003J08 product:meiotic check point regulator, full insert sequence [AK090134] |
KLA | .79 |
.76 |
.72 |
.64 |
.56 |
.54 |
.76 |
| ATP | .99 |
1.04 |
1.11 |
.77 |
.48 |
.31 |
.39 |
| KLA/ATP | .72 |
.75 |
.73 |
.57 |
.42 |
.30 |
.25 |
|
| Anapc1 |  |
NM_008569 |
anaphase promoting complex subunit 1 (Anapc1), mRNA [NM_008569] |
KLA | 1.00 |
.89 |
.81 |
.89 |
.66 |
.75 |
.74 |
| ATP | .88 |
.79 |
.97 |
.46 |
.48 |
.93 |
.76 |
| KLA/ATP | .76 |
.77 |
.83 |
.37 |
.44 |
.74 |
.62 |
|
| Anapc10 | 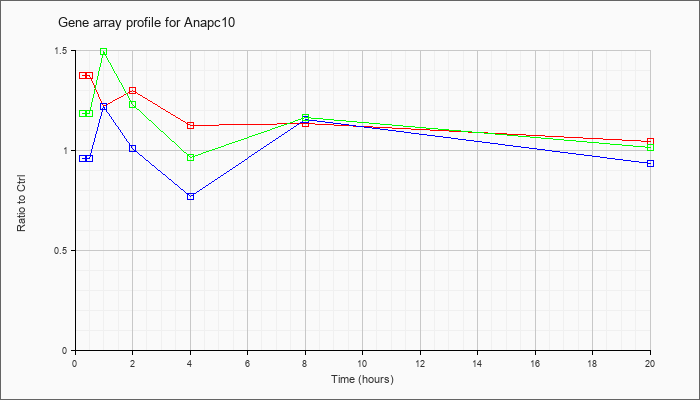 |
NM_026904 |
anaphase promoting complex subunit 10 (Anapc10), mRNA [NM_026904] |
KLA | 1.38 |
1.24 |
1.22 |
1.30 |
1.13 |
1.14 |
1.05 |
| ATP | .96 |
.99 |
1.22 |
1.01 |
.77 |
1.16 |
.94 |
| KLA/ATP | 1.19 |
1.24 |
1.50 |
1.23 |
.97 |
1.17 |
1.02 |
|
| Anapc11 | 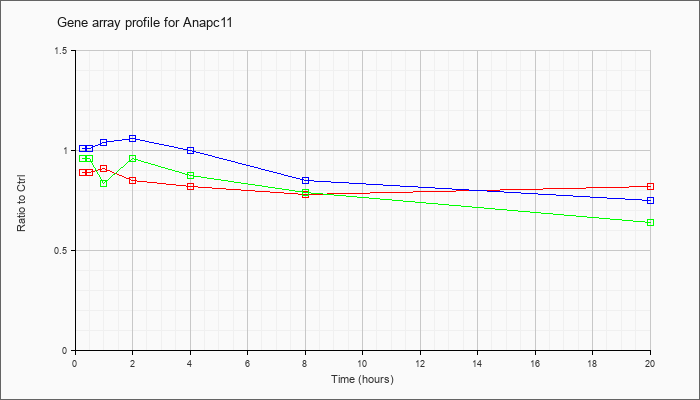 |
NM_001038230 |
anaphase promoting complex subunit 11 homolog (yeast) (Anapc11), transcript variant 1, mRNA [NM_001038230] |
KLA | .89 |
.90 |
.91 |
.85 |
.82 |
.78 |
.82 |
| ATP | 1.01 |
1.06 |
1.04 |
1.06 |
1.00 |
.85 |
.75 |
| KLA/ATP | .96 |
.93 |
.83 |
.96 |
.87 |
.79 |
.64 |
|
| Anapc13 | 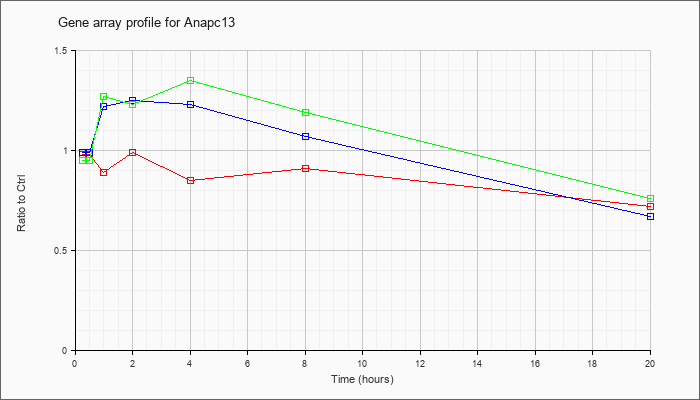 |
NM_181394 |
anaphase promoting complex subunit 13 (Anapc13), mRNA [NM_181394] |
KLA | .98 |
.96 |
.89 |
.99 |
.85 |
.91 |
.72 |
| ATP | .99 |
.95 |
1.22 |
1.25 |
1.23 |
1.07 |
.67 |
| KLA/ATP | .95 |
1.01 |
1.27 |
1.23 |
1.35 |
1.19 |
.76 |
|
| Anapc2 | 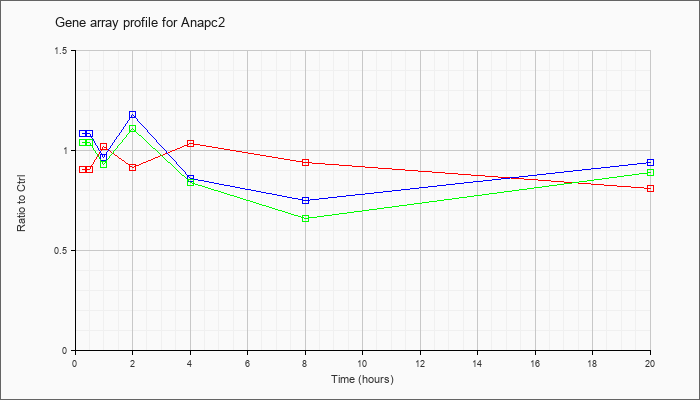 |
NM_175300 |
anaphase promoting complex subunit 2 (Anapc2), mRNA [NM_175300] |
KLA | .90 |
.92 |
1.02 |
.91 |
1.03 |
.94 |
.81 |
| ATP | 1.08 |
1.16 |
.96 |
1.18 |
.86 |
.75 |
.94 |
| KLA/ATP | 1.04 |
1.07 |
.93 |
1.11 |
.84 |
.66 |
.89 |
|
| Anapc4 | 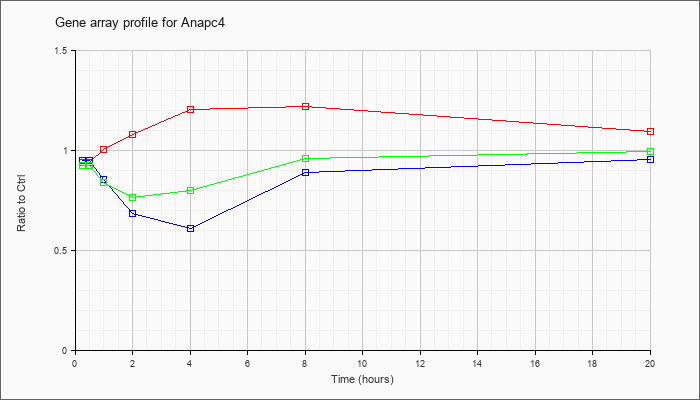 |
NM_024213 |
anaphase promoting complex subunit 4 (Anapc4), mRNA [NM_024213] |
KLA | .94 |
.99 |
1.01 |
1.08 |
1.21 |
1.22 |
1.09 |
| ATP | .95 |
.91 |
.86 |
.69 |
.61 |
.89 |
.96 |
| KLA/ATP | .93 |
.95 |
.84 |
.77 |
.80 |
.96 |
1.00 |
|
| Anapc5 |  |
NM_021505 |
anaphase-promoting complex subunit 5 (Anapc5), transcript variant 1, mRNA [NM_021505] |
KLA | .80 |
.82 |
.83 |
.82 |
.79 |
.84 |
1.10 |
| ATP | .99 |
.99 |
.89 |
.81 |
.79 |
.70 |
.71 |
| KLA/ATP | .82 |
.81 |
.72 |
.62 |
.63 |
.73 |
.68 |
|
| Anapc7 | 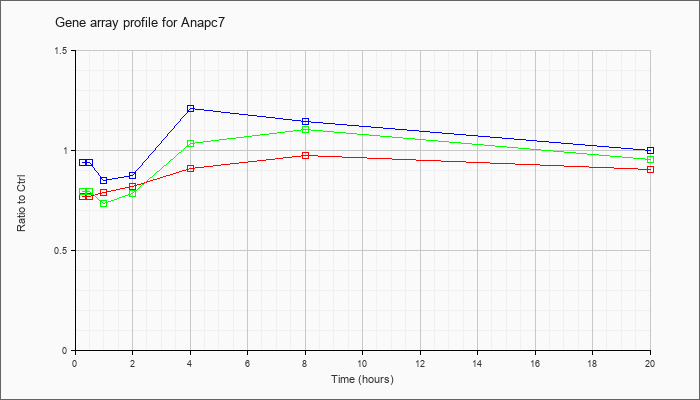 |
AK049968 |
adult male hippocampus cDNA, RIKEN full-length enriched library, clone:C630034H22 product:anaphase-promoting complex subunit 7, full insert sequence. [AK049968] |
KLA | .89 |
.86 |
.88 |
.85 |
.91 |
.95 |
.89 |
| ATP | .92 |
.89 |
.89 |
.86 |
.96 |
.99 |
1.00 |
| KLA/ATP | .89 |
.79 |
.83 |
.85 |
.93 |
.92 |
.87 |
|
| Anapc7 |  |
NM_019805 |
anaphase promoting complex subunit 7 (Anapc7), mRNA [NM_019805] |
KLA | .65 |
.69 |
.70 |
.79 |
.91 |
1.00 |
.92 |
| ATP | .96 |
.99 |
.81 |
.89 |
1.46 |
1.30 |
1.00 |
| KLA/ATP | .70 |
.73 |
.64 |
.72 |
1.14 |
1.28 |
1.04 |
|
| Araf | 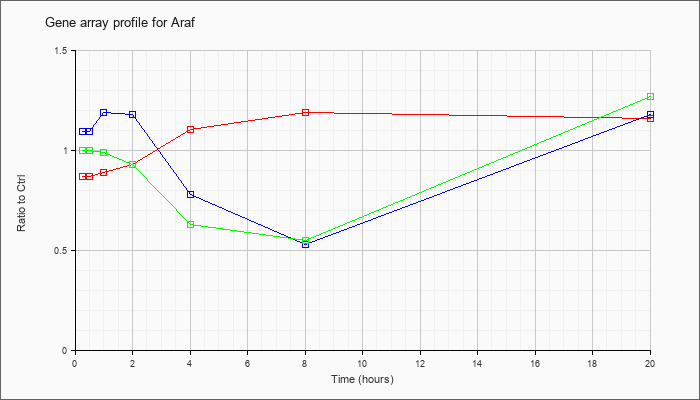 |
NM_009703 |
v-raf murine sarcoma 3611 viral oncogene homolog (Araf), mRNA [NM_009703] |
KLA | .87 |
.80 |
.89 |
.93 |
1.10 |
1.19 |
1.16 |
| ATP | 1.09 |
1.11 |
1.19 |
1.18 |
.78 |
.53 |
1.18 |
| KLA/ATP | 1.00 |
.95 |
.99 |
.93 |
.63 |
.55 |
1.27 |
|
| Braf |  |
NM_139294 |
Braf transforming gene (Braf), mRNA [NM_139294] |
KLA | 1.52 |
1.57 |
1.55 |
1.33 |
1.83 |
1.52 |
1.60 |
| ATP | 1.08 |
1.17 |
1.02 |
1.37 |
.95 |
1.15 |
.95 |
| KLA/ATP | 1.64 |
1.69 |
1.39 |
1.51 |
1.04 |
1.31 |
1.26 |
|
| Bub1 | 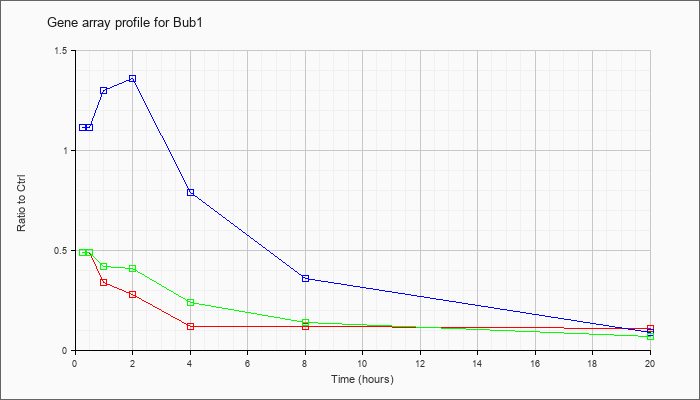 |
NM_009772 |
budding uninhibited by benzimidazoles 1 homolog (S. cerevisiae) (Bub1), transcript variant 2, mRNA [NM_009772] |
KLA | .49 |
.43 |
.34 |
.28 |
.12 |
.12 |
.11 |
| ATP | 1.11 |
1.16 |
1.30 |
1.36 |
.79 |
.36 |
.09 |
| KLA/ATP | .49 |
.40 |
.42 |
.41 |
.24 |
.14 |
.07 |
|
| Ccna1 | 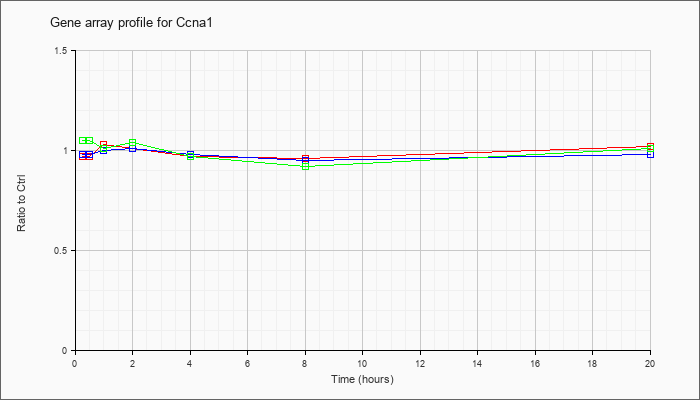 |
NM_007628 |
cyclin A1 (Ccna1), mRNA [NM_007628] |
KLA | .97 |
.96 |
1.03 |
1.01 |
.97 |
.96 |
1.02 |
| ATP | .98 |
.99 |
1.00 |
1.01 |
.98 |
.95 |
.98 |
| KLA/ATP | 1.05 |
.96 |
1.01 |
1.04 |
.97 |
.92 |
1.01 |
|
| Ccna2 | 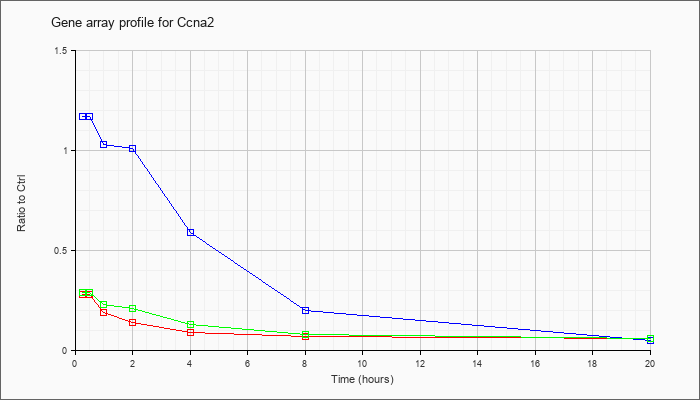 |
NM_009828 |
cyclin A2 (Ccna2), mRNA [NM_009828] |
KLA | .28 |
.21 |
.19 |
.14 |
.09 |
.07 |
.06 |
| ATP | 1.17 |
1.11 |
1.03 |
1.01 |
.59 |
.20 |
.05 |
| KLA/ATP | .29 |
.22 |
.23 |
.21 |
.13 |
.08 |
.06 |
|
| Ccnb1 | 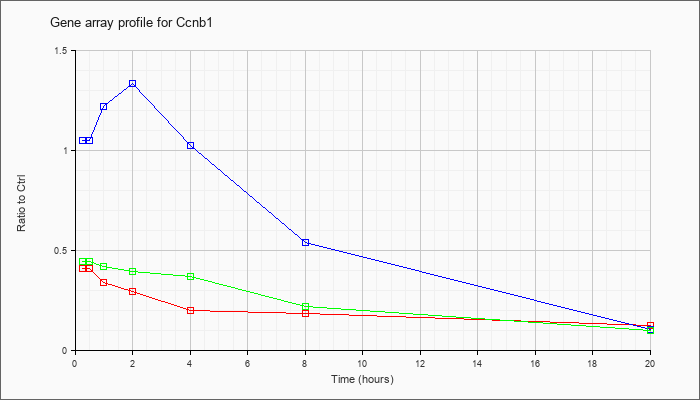 |
NM_172301 |
cyclin B1 (Ccnb1), mRNA [NM_172301] |
KLA | .41 |
.37 |
.34 |
.29 |
.20 |
.18 |
.12 |
| ATP | 1.05 |
1.04 |
1.22 |
1.33 |
1.02 |
.54 |
.10 |
| KLA/ATP | .44 |
.37 |
.42 |
.39 |
.37 |
.22 |
.10 |
|
| Ccnb2 | 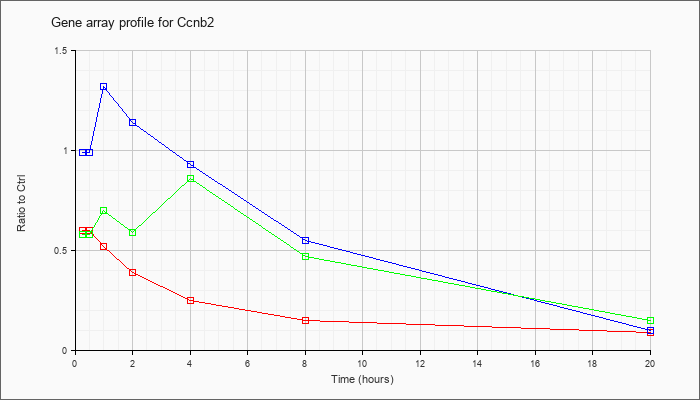 |
NM_007630 |
cyclin B2 (Ccnb2), mRNA [NM_007630] |
KLA | .60 |
.51 |
.52 |
.39 |
.25 |
.15 |
.09 |
| ATP | .99 |
.89 |
1.32 |
1.14 |
.93 |
.55 |
.10 |
| KLA/ATP | .58 |
.52 |
.70 |
.59 |
.86 |
.47 |
.15 |
|
| Ccnb3 | 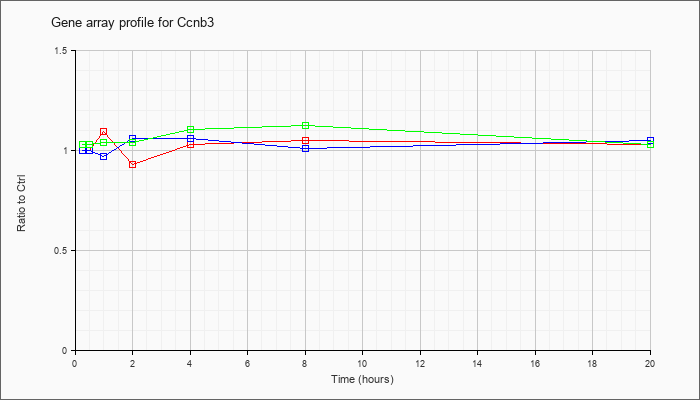 |
NM_183015 |
cyclin B3 (Ccnb3), mRNA [NM_183015] |
KLA | 1.00 |
1.04 |
1.09 |
.93 |
1.03 |
1.05 |
1.03 |
| ATP | 1.00 |
1.02 |
.97 |
1.06 |
1.06 |
1.01 |
1.05 |
| KLA/ATP | 1.03 |
.99 |
1.04 |
1.04 |
1.10 |
1.12 |
1.03 |
|
| Cdc16 | 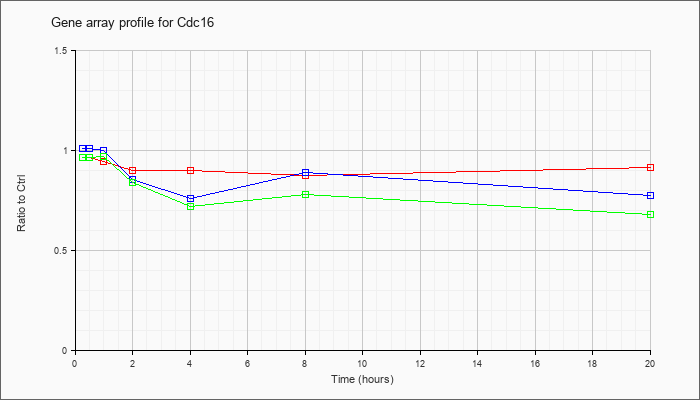 |
AK012511 |
11 days embryo whole body cDNA, RIKEN full-length enriched library, clone:2700071J12 product:unclassifiable, full insert sequence. [AK012511] |
KLA | .97 |
.92 |
.89 |
.90 |
.93 |
.92 |
.99 |
| ATP | .97 |
.94 |
.89 |
.92 |
.89 |
.94 |
.91 |
| KLA/ATP | .98 |
.88 |
.99 |
.93 |
.80 |
.89 |
.82 |
|
| Cdc16 |  |
NM_027276 |
CDC16 cell division cycle 16 homolog (S. cerevisiae) (Cdc16), mRNA [NM_027276] |
KLA | .96 |
.98 |
.97 |
.90 |
.89 |
.85 |
.88 |
| ATP | 1.03 |
1.02 |
1.06 |
.82 |
.70 |
.87 |
.71 |
| KLA/ATP | .96 |
.93 |
.96 |
.79 |
.68 |
.73 |
.61 |
|
| Cdc23 |  |
NM_178347 |
CDC23 (cell division cycle 23, yeast, homolog) (Cdc23), mRNA [NM_178347] |
KLA | .74 |
.76 |
.75 |
.74 |
.72 |
.79 |
.79 |
| ATP | 1.06 |
1.04 |
.91 |
.71 |
.48 |
.60 |
.71 |
| KLA/ATP | .76 |
.77 |
.73 |
.66 |
.46 |
.51 |
.59 |
|
| Cdc25a | 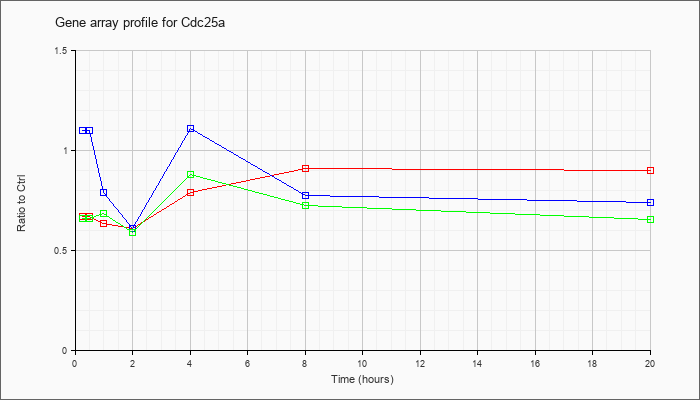 |
NM_007658 |
cell division cycle 25 homolog A (S. pombe) (Cdc25a), mRNA [NM_007658] |
KLA | .76 |
.77 |
.75 |
.68 |
.83 |
1.00 |
.95 |
| ATP | .97 |
1.03 |
.83 |
.69 |
1.01 |
.84 |
.79 |
| KLA/ATP | .69 |
.71 |
.80 |
.70 |
.91 |
.81 |
.81 |
|
| Cdc25a |  |
U27323 |
Cdc25a (cdc25a) mRNA, complete cds. [U27323] |
KLA | .62 |
.60 |
.58 |
.58 |
.77 |
.87 |
.87 |
| ATP | 1.16 |
1.04 |
.77 |
.57 |
1.16 |
.74 |
.72 |
| KLA/ATP | .65 |
.64 |
.63 |
.54 |
.86 |
.68 |
.58 |
|
| Cdc25b | 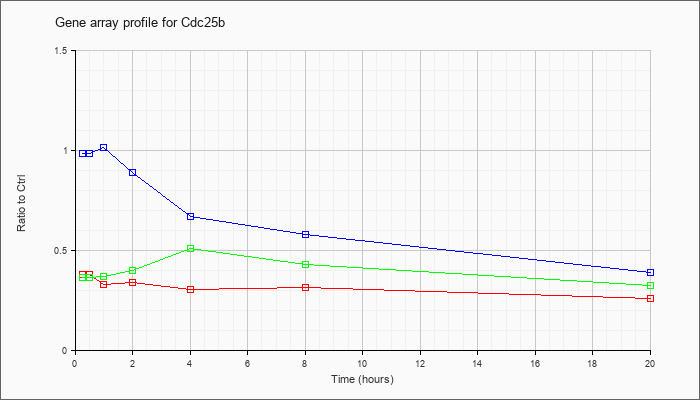 |
NM_023117 |
cell division cycle 25 homolog B (S. pombe) (Cdc25b), transcript variant 1, mRNA [NM_023117] |
KLA | .38 |
.34 |
.33 |
.34 |
.30 |
.31 |
.26 |
| ATP | .98 |
1.04 |
1.01 |
.89 |
.67 |
.58 |
.39 |
| KLA/ATP | .36 |
.33 |
.37 |
.40 |
.51 |
.43 |
.32 |
|
| Cdc25c | 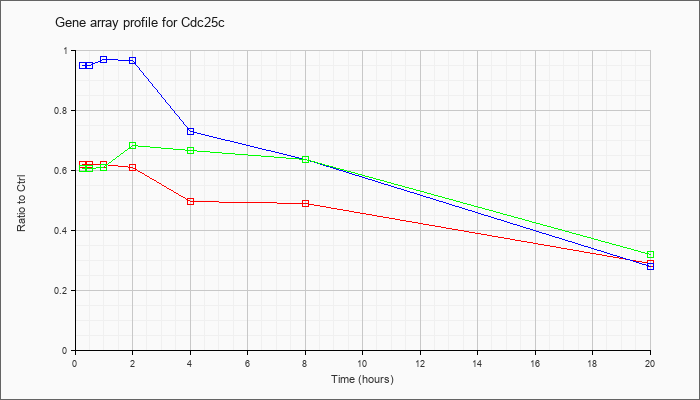 |
NM_009860 |
cell division cycle 25 homolog C (S. pombe) (Cdc25c), mRNA [NM_009860] |
KLA | .62 |
.63 |
.62 |
.61 |
.50 |
.49 |
.29 |
| ATP | .95 |
1.00 |
.97 |
.97 |
.73 |
.64 |
.28 |
| KLA/ATP | .61 |
.62 |
.61 |
.68 |
.67 |
.64 |
.32 |
|
| Cdc26 | 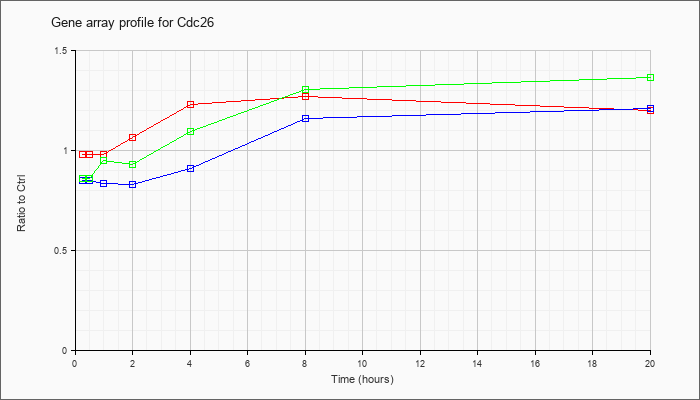 |
AK041840 |
3 days neonate thymus cDNA, RIKEN full-length enriched library, clone:A630041F01 product:unclassifiable, full insert sequence. [AK041840] |
KLA | .92 |
.88 |
.95 |
.88 |
1.08 |
.99 |
1.15 |
| ATP | .72 |
.64 |
.61 |
.75 |
.91 |
.85 |
1.04 |
| KLA/ATP | .84 |
.75 |
.66 |
.87 |
.94 |
.95 |
.99 |
|
| Cdc26 |  |
NM_139291 |
cell division cycle 26 (Cdc26), mRNA [NM_139291] |
KLA | 1.04 |
1.07 |
1.01 |
1.25 |
1.38 |
1.55 |
1.25 |
| ATP | .98 |
.97 |
1.06 |
.91 |
.91 |
1.47 |
1.38 |
| KLA/ATP | .88 |
1.02 |
1.24 |
.99 |
1.25 |
1.66 |
1.74 |
|
| Cdc27 | 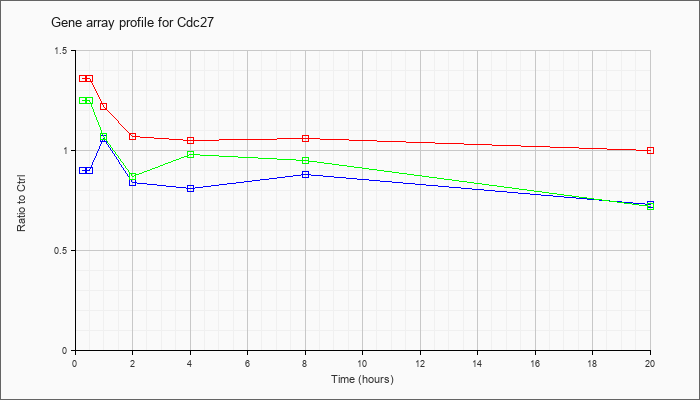 |
NM_145436 |
cell division cycle 27 homolog (S. cerevisiae) (Cdc27), mRNA [NM_145436] |
KLA | 1.36 |
1.25 |
1.22 |
1.07 |
1.05 |
1.06 |
1.00 |
| ATP | .90 |
.97 |
1.06 |
.84 |
.81 |
.88 |
.73 |
| KLA/ATP | 1.25 |
1.22 |
1.07 |
.87 |
.98 |
.95 |
.72 |
|
| Cdc2a | 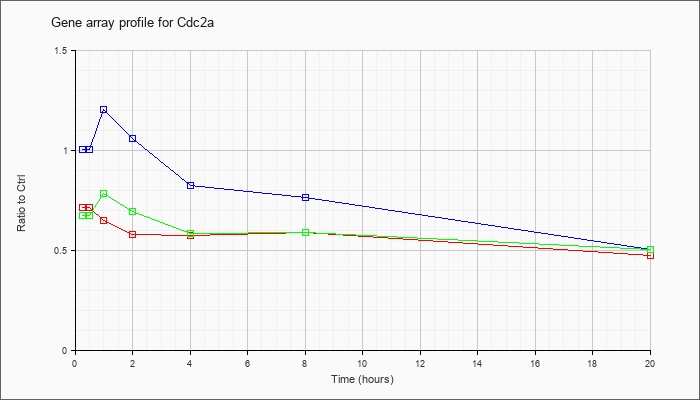 |
NM_007659 |
cell division cycle 2 homolog A (S. pombe) (Cdc2a), mRNA [NM_007659] |
KLA | .72 |
.68 |
.65 |
.58 |
.58 |
.59 |
.48 |
| ATP | 1.01 |
1.05 |
1.21 |
1.06 |
.83 |
.77 |
.51 |
| KLA/ATP | .68 |
.63 |
.79 |
.70 |
.59 |
.59 |
.51 |
|
| Cdk2 | 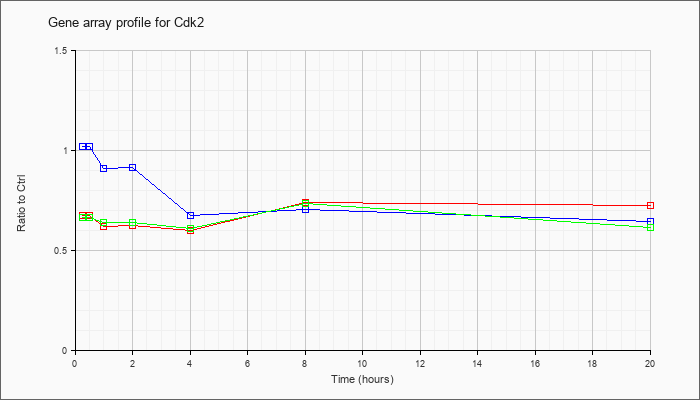 |
NM_016756 |
cyclin-dependent kinase 2 (Cdk2), transcript variant 2, mRNA [NM_016756] |
KLA | .68 |
.67 |
.62 |
.63 |
.60 |
.74 |
.73 |
| ATP | 1.02 |
1.14 |
.91 |
.92 |
.68 |
.71 |
.65 |
| KLA/ATP | .67 |
.64 |
.64 |
.64 |
.61 |
.74 |
.62 |
|
| Cpeb1 | 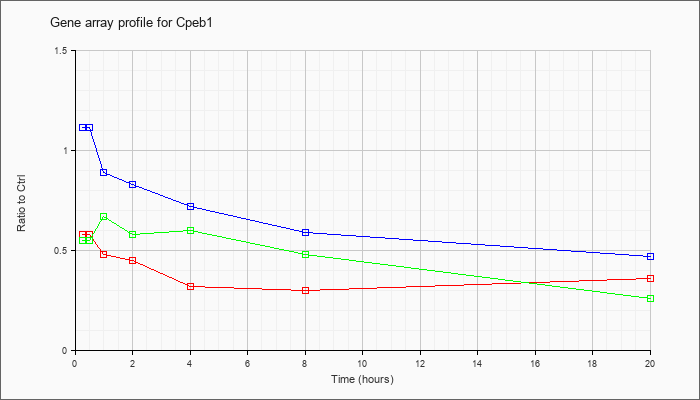 |
NM_007755 |
cytoplasmic polyadenylation element binding protein 1 (Cpeb1), mRNA [NM_007755] |
KLA | .58 |
.50 |
.48 |
.45 |
.32 |
.30 |
.36 |
| ATP | 1.11 |
1.07 |
.89 |
.83 |
.72 |
.59 |
.47 |
| KLA/ATP | .55 |
.56 |
.67 |
.58 |
.60 |
.48 |
.26 |
|
| Fzr1 | 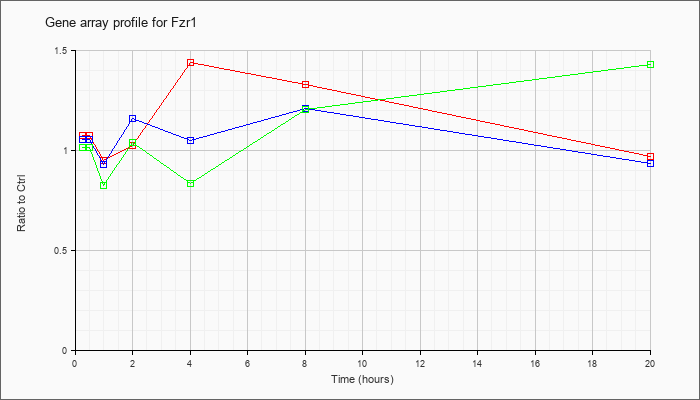 |
NM_019757 |
fizzy/cell division cycle 20 related 1 (Drosophila) (Fzr1), mRNA [NM_019757] |
KLA | 1.08 |
.98 |
.95 |
1.03 |
1.44 |
1.33 |
.97 |
| ATP | 1.06 |
1.19 |
.93 |
1.16 |
1.05 |
1.21 |
.94 |
| KLA/ATP | 1.02 |
1.05 |
.83 |
1.04 |
.84 |
1.21 |
1.43 |
|
| Gnai1 | 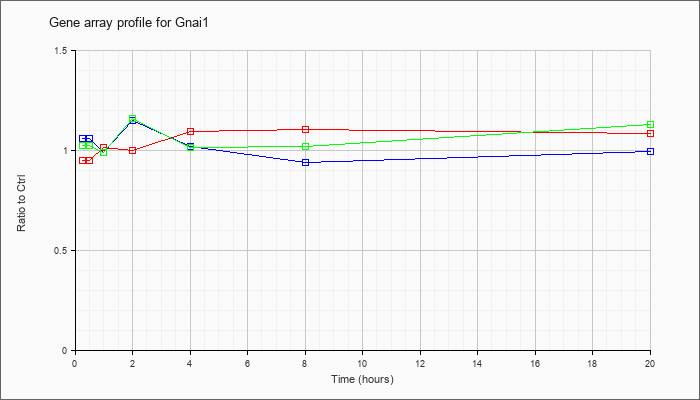 |
NM_010305 |
guanine nucleotide binding protein (G protein), alpha inhibiting 1 (Gnai1), mRNA [NM_010305] |
KLA | .95 |
1.01 |
1.01 |
1.00 |
1.09 |
1.10 |
1.08 |
| ATP | 1.06 |
1.06 |
.99 |
1.15 |
1.02 |
.94 |
.99 |
| KLA/ATP | 1.02 |
1.04 |
.99 |
1.16 |
1.01 |
1.02 |
1.13 |
|
| Gnai2 | 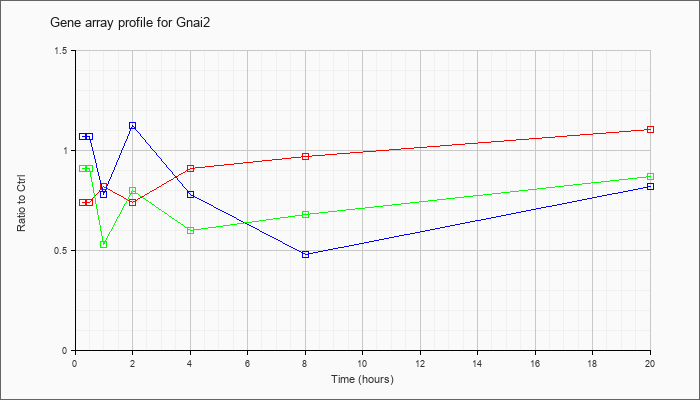 |
NM_008138 |
guanine nucleotide binding protein (G protein), alpha inhibiting 2 (Gnai2), mRNA [NM_008138] |
KLA | .74 |
.79 |
.82 |
.74 |
.91 |
.97 |
1.10 |
| ATP | 1.07 |
1.16 |
.78 |
1.12 |
.78 |
.48 |
.82 |
| KLA/ATP | .91 |
.81 |
.53 |
.80 |
.60 |
.68 |
.87 |
|
| Gnai3 | 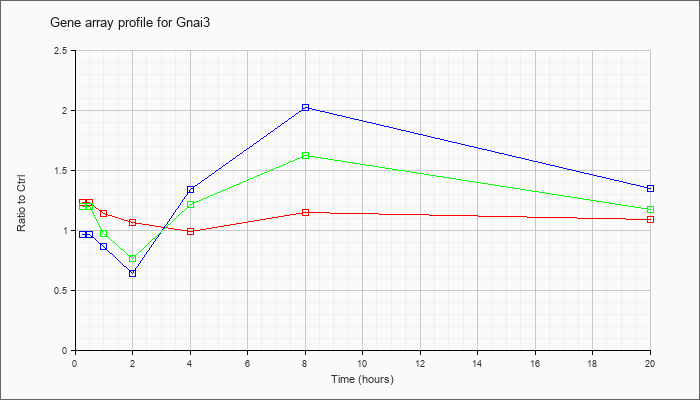 |
NM_010306 |
guanine nucleotide binding protein (G protein), alpha inhibiting 3 (Gnai3), mRNA [NM_010306] |
KLA | 1.23 |
1.24 |
1.14 |
1.06 |
.99 |
1.15 |
1.09 |
| ATP | .96 |
.93 |
.86 |
.64 |
1.34 |
2.02 |
1.35 |
| KLA/ATP | 1.20 |
1.24 |
.97 |
.76 |
1.21 |
1.62 |
1.17 |
|
Hsp90aa1
| 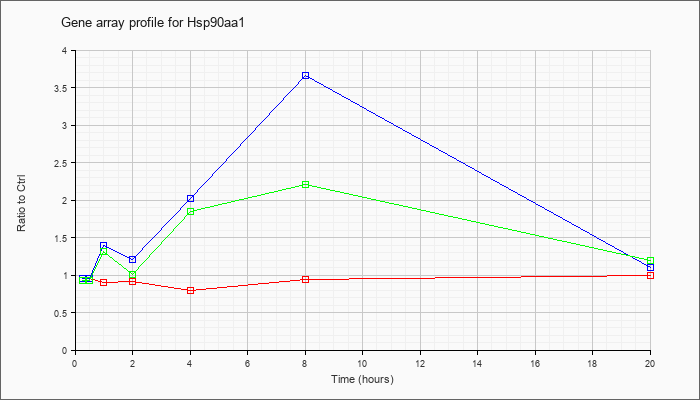 |
NM_010480 |
heat shock protein 90, alpha (cytosolic), class A member 1 (Hsp90aa1), mRNA [NM_010480] |
KLA | .95 |
1.03 |
.90 |
.92 |
.80 |
.94 |
.99 |
| ATP | .96 |
.92 |
1.39 |
1.21 |
2.02 |
3.66 |
1.10 |
| KLA/ATP | .93 |
.92 |
1.32 |
1.01 |
1.85 |
2.21 |
1.20 |
|
Hsp90ab1
| 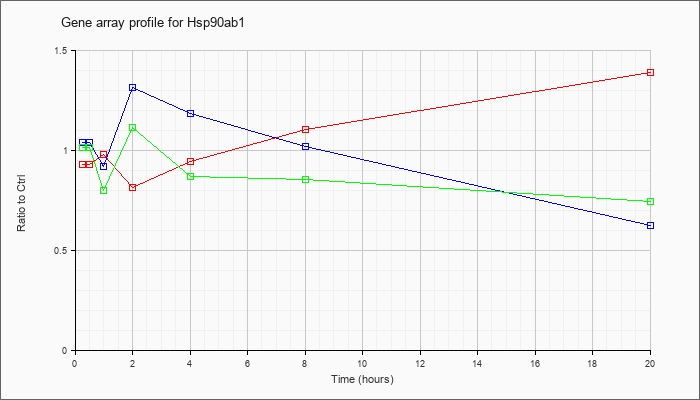 |
NM_008302 |
heat shock protein 90kDa alpha (cytosolic), class B member 1 (Hsp90ab1), mRNA [NM_008302] |
KLA | .93 |
1.02 |
.98 |
.82 |
.95 |
1.10 |
1.39 |
| ATP | 1.04 |
1.14 |
.92 |
1.32 |
1.19 |
1.02 |
.63 |
| KLA/ATP | 1.02 |
1.04 |
.80 |
1.12 |
.87 |
.86 |
.75 |
|
| Igf1 | 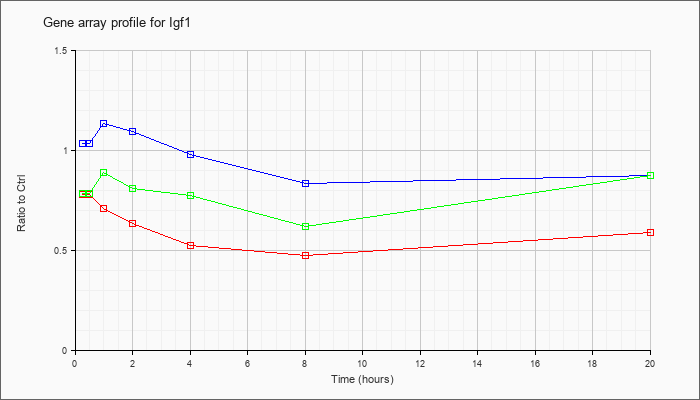 |
AK050118 |
adult male liver tumor cDNA, RIKEN full-length enriched library, clone:C730016P09 product:unclassifiable, full insert sequence. [AK050118] |
KLA | .13 |
.14 |
.15 |
.16 |
.22 |
.34 |
.48 |
| ATP | 1.78 |
2.05 |
1.37 |
.72 |
.34 |
.41 |
.58 |
| KLA/ATP | .24 |
.45 |
.50 |
.24 |
.20 |
.26 |
.68 |
|
| Igf1 |  |
NM_010512 |
insulin-like growth factor 1 (Igf1), transcript variant 1, mRNA [NM_010512] |
KLA | .84 |
.78 |
.76 |
.67 |
.54 |
.48 |
.59 |
| ATP | .97 |
.98 |
1.13 |
1.13 |
1.03 |
.88 |
.89 |
| KLA/ATP | .82 |
.81 |
.93 |
.86 |
.83 |
.65 |
.89 |
|
| Igf1 |  |
NM_184052 |
insulin-like growth factor 1 (Igf1), transcript variant 2, mRNA [NM_184052] |
KLA | .77 |
.82 |
.76 |
.65 |
.60 |
.52 |
.69 |
| ATP | 1.01 |
1.07 |
.98 |
1.06 |
1.03 |
.78 |
.94 |
| KLA/ATP | .89 |
.86 |
.77 |
.80 |
.75 |
.65 |
.89 |
|
| Igf1r | 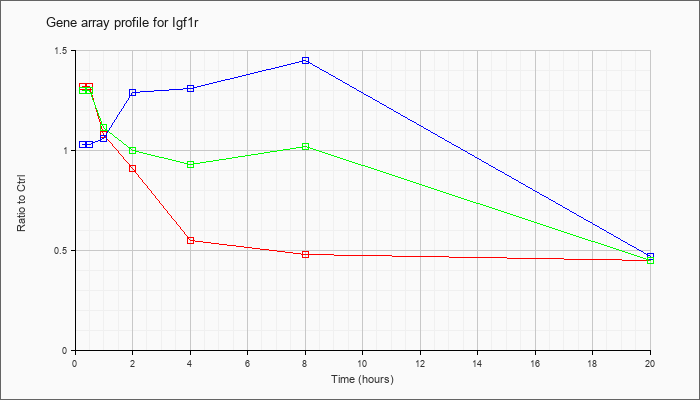 |
NM_010513 |
insulin-like growth factor I receptor (Igf1r), mRNA [NM_010513] |
KLA | 1.32 |
1.32 |
1.08 |
.91 |
.55 |
.48 |
.45 |
| ATP | 1.03 |
1.06 |
1.06 |
1.29 |
1.31 |
1.45 |
.47 |
| KLA/ATP | 1.30 |
1.37 |
1.11 |
1.00 |
.93 |
1.02 |
.45 |
|
| Ins1 | 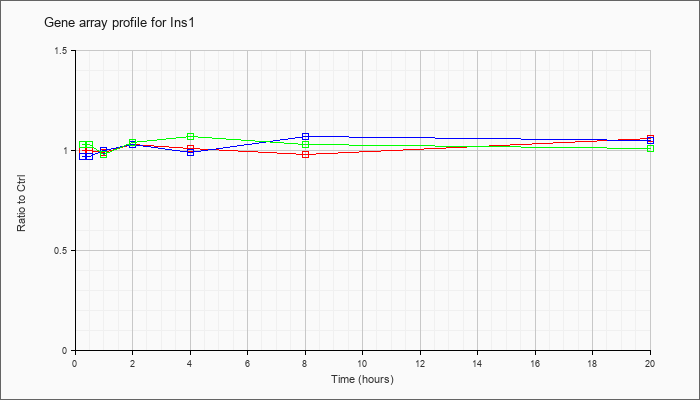 |
NM_008386 |
insulin I (Ins1), mRNA [NM_008386] |
KLA | 1.00 |
.99 |
.99 |
1.03 |
1.01 |
.98 |
1.06 |
| ATP | .97 |
1.03 |
1.00 |
1.03 |
.99 |
1.07 |
1.05 |
| KLA/ATP | 1.03 |
.93 |
.98 |
1.04 |
1.07 |
1.03 |
1.01 |
|
| Ins2 | 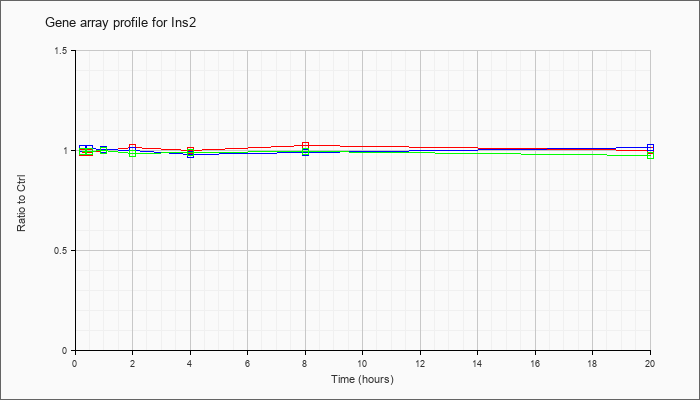 |
NM_008387 |
insulin II (Ins2), mRNA [NM_008387] |
KLA | .99 |
1.00 |
1.00 |
1.01 |
1.00 |
1.02 |
1.00 |
| ATP | 1.01 |
.99 |
1.00 |
1.00 |
.98 |
.99 |
1.01 |
| KLA/ATP | .99 |
1.01 |
1.00 |
.98 |
.99 |
.99 |
.98 |
|
| Kras | 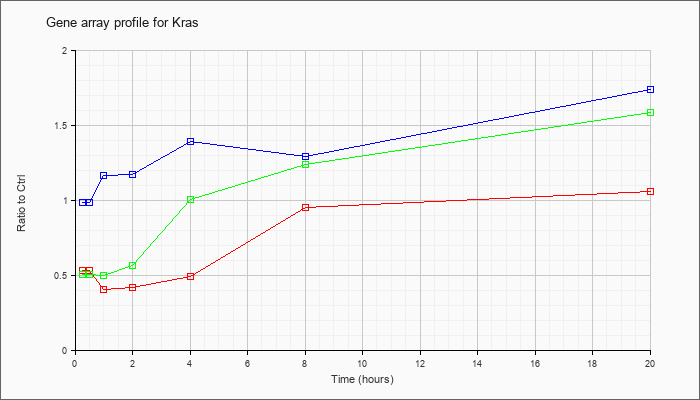 |
NM_021284 |
v-Ki-ras2 Kirsten rat sarcoma viral oncogene homolog (Kras), mRNA [NM_021284] |
KLA | .53 |
.49 |
.40 |
.42 |
.49 |
.95 |
1.05 |
| ATP | .98 |
1.01 |
1.16 |
1.17 |
1.39 |
1.29 |
1.74 |
| KLA/ATP | .50 |
.47 |
.50 |
.56 |
1.00 |
1.24 |
1.58 |
|
| Mad1l1 | 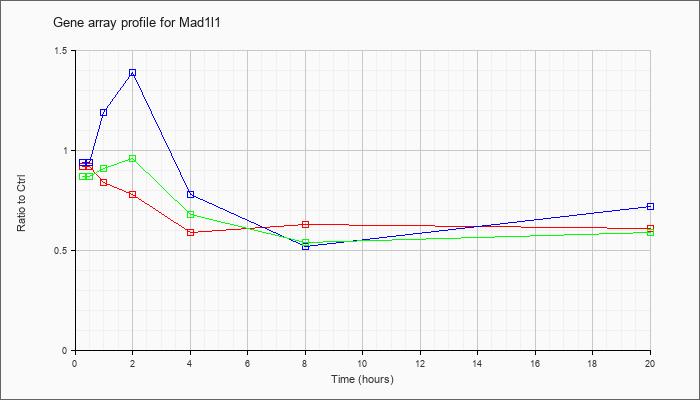 |
NM_010752 |
mitotic arrest deficient 1-like 1 (Mad1l1), mRNA [NM_010752] |
KLA | .92 |
.84 |
.84 |
.78 |
.59 |
.63 |
.61 |
| ATP | .94 |
.99 |
1.19 |
1.39 |
.78 |
.52 |
.72 |
| KLA/ATP | .87 |
.92 |
.91 |
.96 |
.68 |
.54 |
.59 |
|
| Mad2l1 | 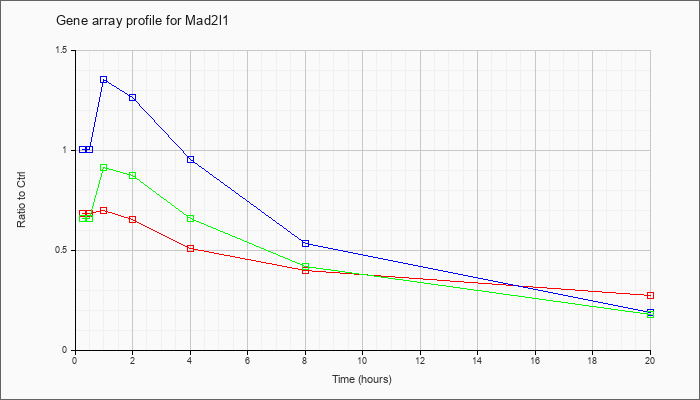 |
NM_019499 |
MAD2 (mitotic arrest deficient, homolog)-like 1 (yeast) (Mad2l1), mRNA [NM_019499] |
KLA | .70 |
.68 |
.73 |
.71 |
.52 |
.41 |
.28 |
| ATP | 1.06 |
1.02 |
1.43 |
1.25 |
1.03 |
.53 |
.19 |
| KLA/ATP | .70 |
.69 |
.99 |
.84 |
.70 |
.43 |
.18 |
|
| Mad2l1 |  |
U83902 |
mitotic checkpoint component Mad2 mRNA, complete cds. [U83902] |
KLA | .67 |
.68 |
.67 |
.60 |
.50 |
.39 |
.27 |
| ATP | .95 |
.97 |
1.28 |
1.28 |
.88 |
.54 |
.19 |
| KLA/ATP | .62 |
.66 |
.84 |
.91 |
.62 |
.41 |
.18 |
|
| Mad2l2 | 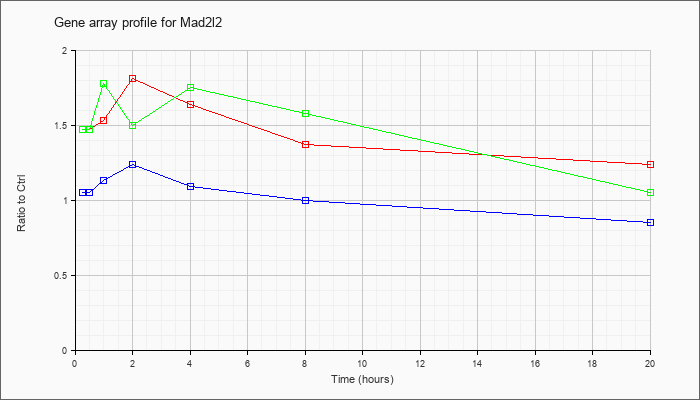 |
NM_027985 |
MAD2 mitotic arrest deficient-like 2 (yeast) (Mad2l2), mRNA [NM_027985] |
KLA | 1.47 |
1.43 |
1.53 |
1.81 |
1.64 |
1.37 |
1.24 |
| ATP | 1.05 |
.92 |
1.13 |
1.24 |
1.09 |
1.00 |
.85 |
| KLA/ATP | 1.47 |
1.50 |
1.78 |
1.50 |
1.75 |
1.58 |
1.05 |
|
| Map2k1 | 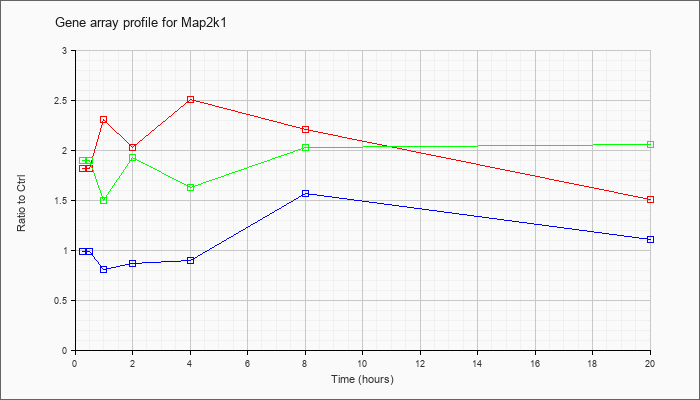 |
NM_008927 |
mitogen-activated protein kinase kinase 1 (Map2k1), mRNA [NM_008927] |
KLA | 1.82 |
2.00 |
2.31 |
2.03 |
2.51 |
2.20 |
1.51 |
| ATP | .99 |
1.14 |
.81 |
.87 |
.90 |
1.57 |
1.11 |
| KLA/ATP | 1.90 |
2.03 |
1.50 |
1.93 |
1.63 |
2.03 |
2.06 |
|
| Mapk1 | 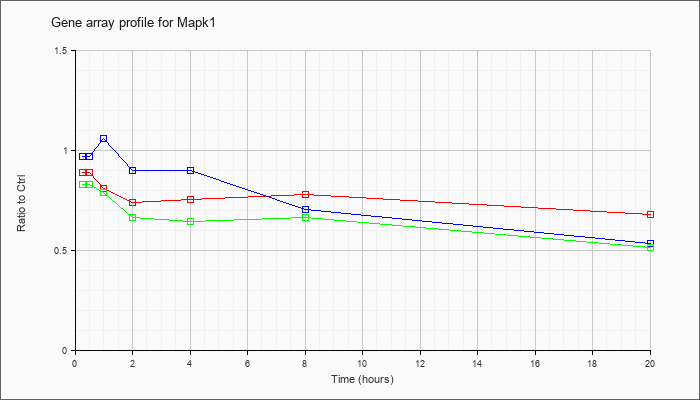 |
NM_001038663 |
mitogen-activated protein kinase 1 (Mapk1), transcript variant 2, mRNA [NM_001038663] |
KLA | .97 |
1.03 |
.93 |
.86 |
.79 |
1.01 |
1.04 |
| ATP | .96 |
1.18 |
1.01 |
1.13 |
.84 |
1.04 |
1.16 |
| KLA/ATP | .99 |
.99 |
.84 |
.87 |
.63 |
.96 |
1.26 |
|
| Mapk1 |  |
NM_011949 |
mitogen-activated protein kinase 1 (Mapk1), transcript variant 1, mRNA [NM_011949] |
KLA | .88 |
.80 |
.80 |
.73 |
.75 |
.76 |
.65 |
| ATP | .97 |
.96 |
1.06 |
.88 |
.90 |
.68 |
.48 |
| KLA/ATP | .82 |
.81 |
.78 |
.65 |
.64 |
.64 |
.45 |
|
| Mapk10 | 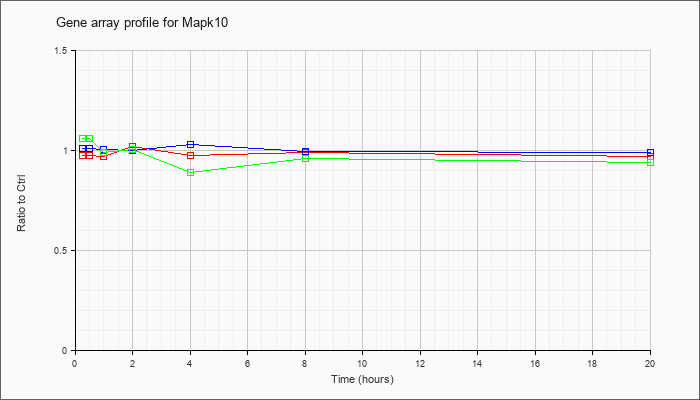 |
AK076990 |
adult male testis cDNA, RIKEN full-length enriched library, clone:4930597C01 product:mitogen activated protein kinase 10, full insert sequence. [AK076990] |
KLA | 1.03 |
1.05 |
.98 |
1.04 |
.94 |
.95 |
1.02 |
| ATP | 1.00 |
.99 |
1.06 |
.97 |
1.01 |
.96 |
.99 |
| KLA/ATP | 1.07 |
1.01 |
.95 |
1.05 |
.82 |
.95 |
.87 |
|
| Mapk10 |  |
NM_009158 |
mitogen-activated protein kinase 10 (Mapk10), transcript variant 1, mRNA [NM_009158] |
KLA | .95 |
1.00 |
.97 |
1.01 |
.99 |
1.01 |
.95 |
| ATP | 1.01 |
1.02 |
.98 |
1.01 |
1.04 |
1.01 |
.99 |
| KLA/ATP | 1.05 |
.99 |
1.01 |
.98 |
.92 |
.97 |
.98 |
|
| Mapk11 | 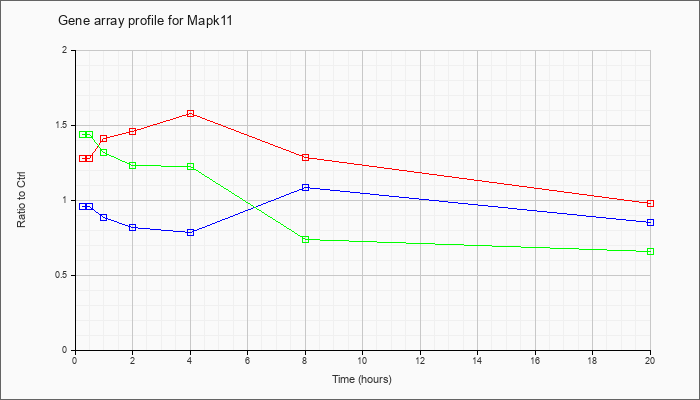 |
NM_011161 |
mitogen-activated protein kinase 11 (Mapk11), mRNA [NM_011161] |
KLA | 1.28 |
1.30 |
1.41 |
1.46 |
1.58 |
1.29 |
.98 |
| ATP | .96 |
.95 |
.89 |
.82 |
.79 |
1.09 |
.85 |
| KLA/ATP | 1.44 |
1.42 |
1.32 |
1.23 |
1.23 |
.74 |
.66 |
|
| Mapk12 | 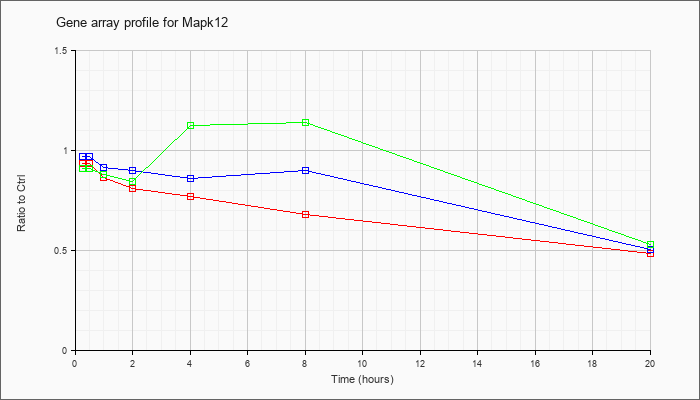 |
NM_013871 |
mitogen-activated protein kinase 12 (Mapk12), mRNA [NM_013871] |
KLA | .94 |
.86 |
.87 |
.81 |
.77 |
.68 |
.49 |
| ATP | .97 |
.83 |
.92 |
.90 |
.86 |
.90 |
.51 |
| KLA/ATP | .91 |
.89 |
.88 |
.85 |
1.12 |
1.14 |
.53 |
|
| Mapk13 | 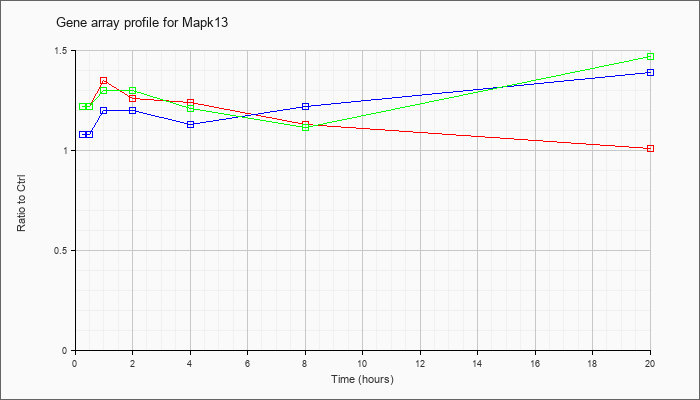 |
NM_011950 |
mitogen-activated protein kinase 13 (Mapk13), mRNA [NM_011950] |
KLA | 1.22 |
1.21 |
1.35 |
1.26 |
1.24 |
1.13 |
1.01 |
| ATP | 1.08 |
1.00 |
1.20 |
1.20 |
1.13 |
1.22 |
1.39 |
| KLA/ATP | 1.22 |
1.27 |
1.30 |
1.30 |
1.21 |
1.11 |
1.47 |
|
| Mapk14 | 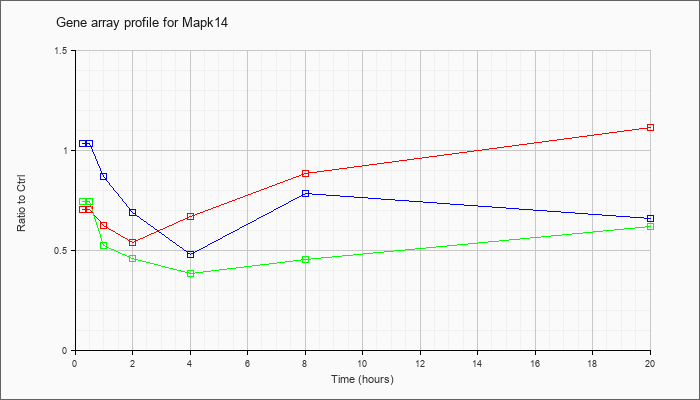 |
NM_011951 |
mitogen-activated protein kinase 14 (Mapk14), mRNA [NM_011951] |
KLA | .70 |
.70 |
.62 |
.54 |
.67 |
.88 |
1.11 |
| ATP | 1.03 |
.98 |
.87 |
.69 |
.48 |
.78 |
.66 |
| KLA/ATP | .74 |
.68 |
.52 |
.46 |
.38 |
.45 |
.62 |
|
| Mapk3 | 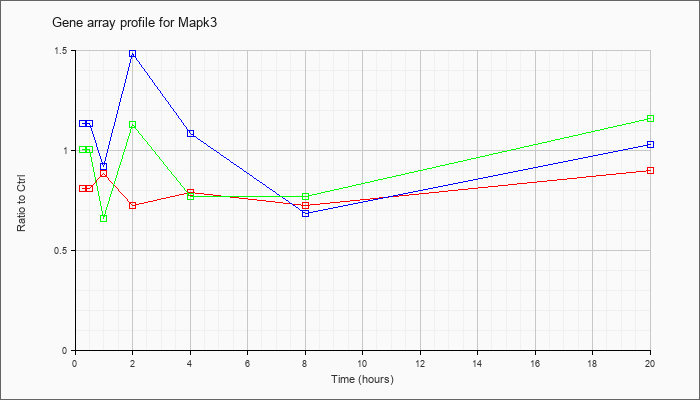 |
NM_011952 |
mitogen-activated protein kinase 3 (Mapk3), mRNA [NM_011952] |
KLA | .81 |
.85 |
.88 |
.72 |
.79 |
.73 |
.90 |
| ATP | 1.13 |
1.30 |
.92 |
1.48 |
1.08 |
.68 |
1.03 |
| KLA/ATP | 1.01 |
1.01 |
.66 |
1.13 |
.77 |
.77 |
1.16 |
|
| Mapk8 |  |
AK162915 |
adult male spinal cord cDNA, RIKEN full-length enriched library, clone:A330031O19 product:mitogen activated protein kinase 8, full insert sequence [AK162915] |
KLA | .94 |
.86 |
.80 |
.83 |
.77 |
.85 |
.83 |
| ATP | 1.08 |
1.12 |
.94 |
.80 |
.83 |
.71 |
.75 |
| KLA/ATP | .83 |
.81 |
.80 |
.74 |
.74 |
.75 |
.67 |
|
| Mapk8 |  |
AK163829 |
16 days embryo head cDNA, RIKEN full-length enriched library, clone:C130070A06 product:mitogen activated protein kinase 8, full insert sequence [AK163829] |
KLA | .65 |
.61 |
.66 |
.70 |
.90 |
1.02 |
1.03 |
| ATP | .99 |
.94 |
1.68 |
1.44 |
1.75 |
1.84 |
1.10 |
| KLA/ATP | .63 |
.67 |
1.11 |
1.00 |
2.01 |
1.68 |
1.15 |
|
| Mapk8 |  |
NM_016700 |
mitogen-activated protein kinase 8 (Mapk8), mRNA [NM_016700] |
KLA | .96 |
1.00 |
.94 |
.96 |
.94 |
.99 |
1.04 |
| ATP | 1.01 |
1.05 |
1.01 |
.99 |
1.03 |
.98 |
1.02 |
| KLA/ATP | .96 |
.97 |
.96 |
.97 |
.93 |
.94 |
.96 |
|
| Mapk9 | 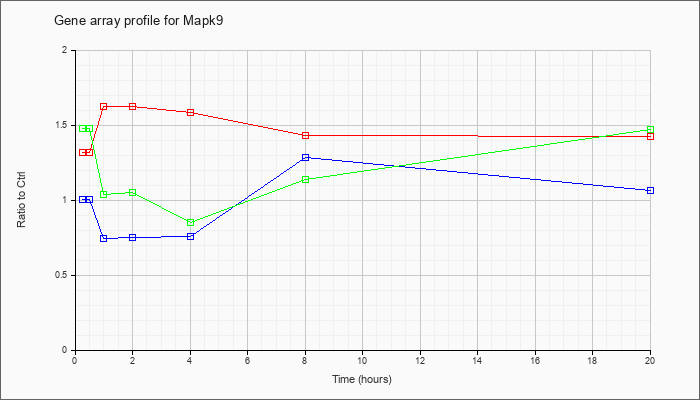 |
NM_016961 |
mitogen-activated protein kinase 9 (Mapk9), transcript variant 2, mRNA [NM_016961] |
KLA | 1.28 |
1.37 |
1.59 |
1.60 |
1.61 |
1.43 |
1.38 |
| ATP | 1.03 |
.99 |
.74 |
.71 |
.79 |
1.20 |
1.01 |
| KLA/ATP | 1.53 |
1.30 |
1.07 |
.99 |
.83 |
1.10 |
1.40 |
|
| Mapk9 |  |
NM_207692 |
mitogen-activated protein kinase 9 (Mapk9), transcript variant 1, mRNA [NM_207692] |
KLA | 1.39 |
1.53 |
1.69 |
1.69 |
1.55 |
1.44 |
1.51 |
| ATP | .95 |
.92 |
.75 |
.85 |
.70 |
1.46 |
1.17 |
| KLA/ATP | 1.37 |
1.35 |
.99 |
1.18 |
.91 |
1.22 |
1.62 |
|
Mm.18219
3 | 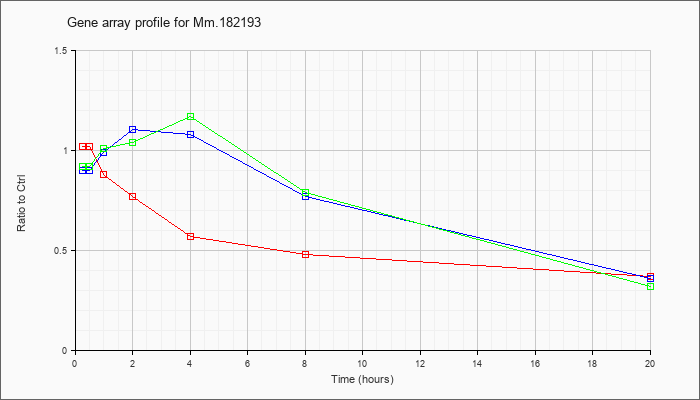 |
142389041 |
Unknown |
KLA | 1.02 |
.95 |
.88 |
.77 |
.57 |
.48 |
.37 |
| ATP | .90 |
.88 |
.99 |
1.10 |
1.08 |
.77 |
.36 |
| KLA/ATP | .92 |
.88 |
1.01 |
1.04 |
1.17 |
.79 |
.32 |
|
Mm.21312
8 | 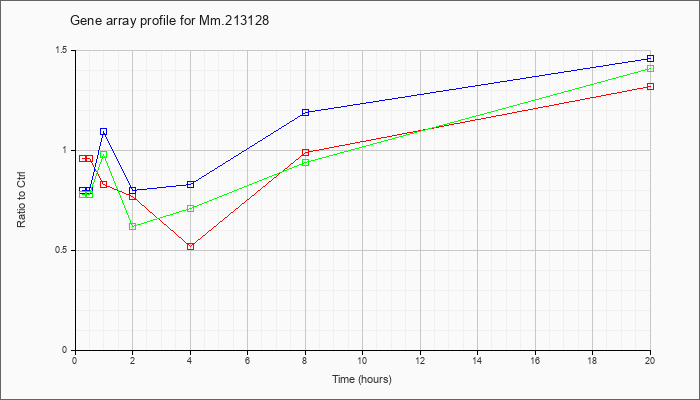 |
141801344 |
Unknown |
KLA | .96 |
.95 |
.83 |
.77 |
.52 |
.99 |
1.32 |
| ATP | .80 |
.82 |
1.09 |
.80 |
.83 |
1.19 |
1.46 |
| KLA/ATP | .78 |
.92 |
.98 |
.62 |
.71 |
.94 |
1.41 |
|
Mm.22094
6 | 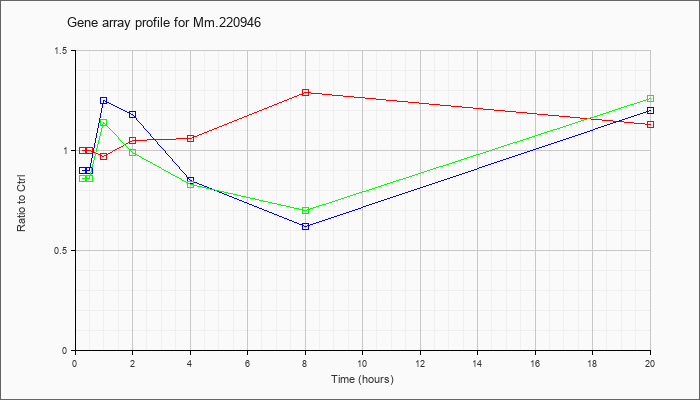 |
27545180 |
Unknown |
KLA | 1.00 |
.93 |
.97 |
1.05 |
1.06 |
1.29 |
1.13 |
| ATP | .90 |
.94 |
1.25 |
1.18 |
.85 |
.62 |
1.20 |
| KLA/ATP | .86 |
.99 |
1.14 |
.99 |
.83 |
.70 |
1.26 |
|
Mm.25381
9 | 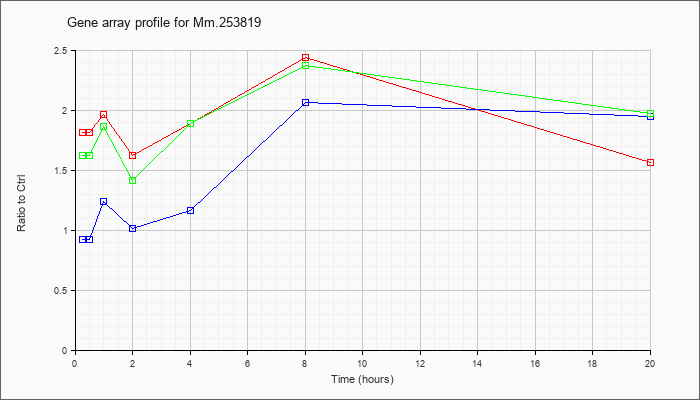 |
118130676 |
Unknown |
KLA | 1.81 |
1.88 |
1.96 |
1.62 |
1.89 |
2.44 |
1.56 |
| ATP | .92 |
.91 |
1.24 |
1.01 |
1.16 |
2.06 |
1.95 |
| KLA/ATP | 1.62 |
1.86 |
1.86 |
1.41 |
1.89 |
2.37 |
1.97 |
|
Mm.27574
2 | 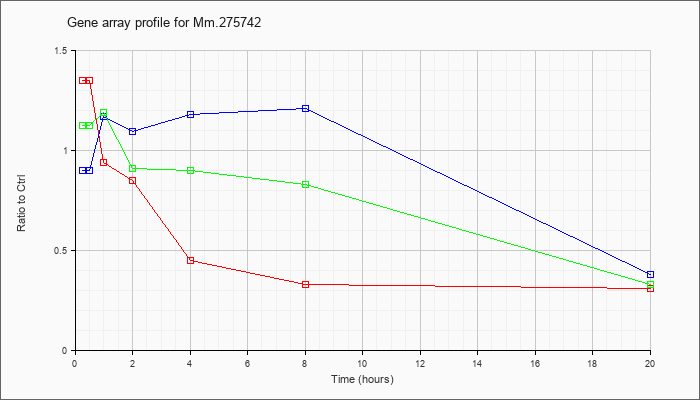 |
112983655 |
Unknown |
KLA | 1.35 |
1.16 |
.94 |
.85 |
.45 |
.33 |
.31 |
| ATP | .90 |
.86 |
1.17 |
1.09 |
1.18 |
1.21 |
.38 |
| KLA/ATP | 1.12 |
1.22 |
1.19 |
.91 |
.90 |
.83 |
.33 |
|
Mm.27970
| 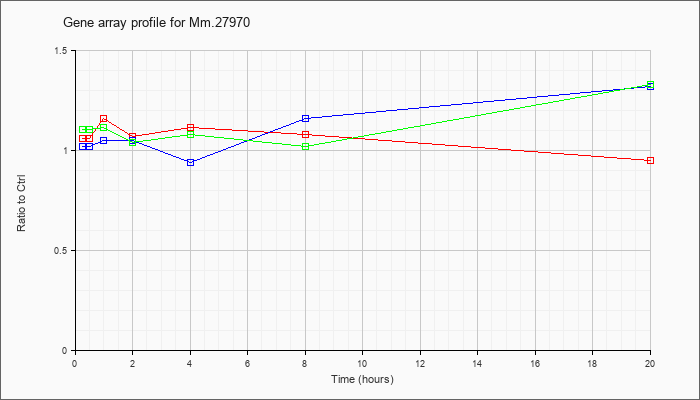 |
13994135 |
Unknown |
KLA | 1.06 |
1.11 |
1.16 |
1.07 |
1.11 |
1.08 |
.95 |
| ATP | 1.02 |
1.09 |
1.05 |
1.05 |
.94 |
1.16 |
1.32 |
| KLA/ATP | 1.10 |
1.19 |
1.11 |
1.04 |
1.08 |
1.02 |
1.33 |
|
Mm.41137
| 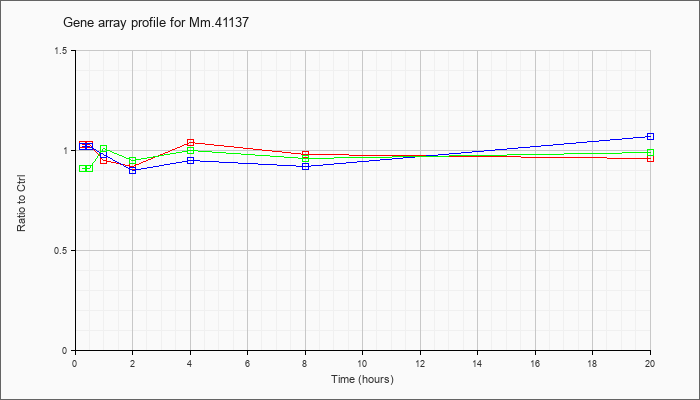 |
148747308 |
Unknown |
KLA | 1.03 |
1.00 |
.95 |
.92 |
1.04 |
.98 |
.96 |
| ATP | 1.02 |
.98 |
.98 |
.90 |
.95 |
.92 |
1.07 |
| KLA/ATP | .91 |
.89 |
1.01 |
.95 |
1.00 |
.96 |
.99 |
|
| Mos | 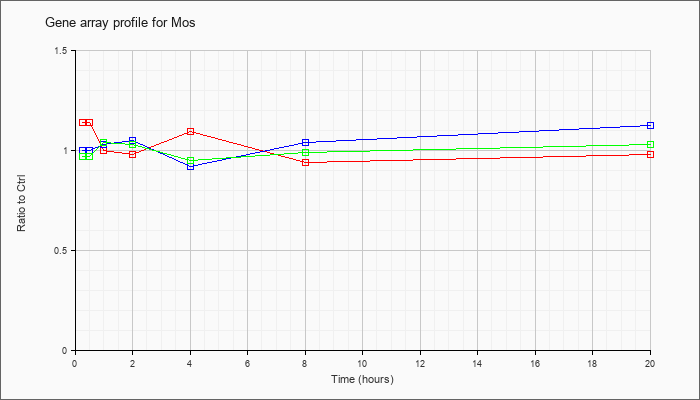 |
NM_020021 |
Moloney sarcoma oncogene (Mos), mRNA [NM_020021] |
KLA | 1.14 |
1.04 |
1.00 |
.98 |
1.09 |
.94 |
.98 |
| ATP | 1.00 |
1.01 |
1.03 |
1.05 |
.92 |
1.04 |
1.12 |
| KLA/ATP | .97 |
.96 |
1.04 |
1.03 |
.95 |
.99 |
1.03 |
|
| Pde3a | 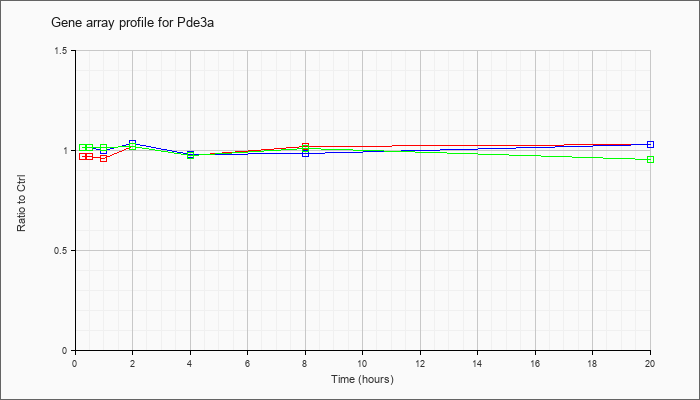 |
AK004638 |
adult male lung cDNA, RIKEN full-length enriched library, clone:1200007J10 product:phosphodiesterase 3A, cGMP inhibited, full insert sequence. [AK004638] |
KLA | .96 |
.94 |
.95 |
1.07 |
1.01 |
1.00 |
1.03 |
| ATP | 1.02 |
.95 |
.96 |
1.07 |
.97 |
.96 |
1.01 |
| KLA/ATP | 1.00 |
1.00 |
1.02 |
.94 |
.98 |
.99 |
1.01 |
|
| Pde3a |  |
AK052718 |
0 day neonate kidney cDNA, RIKEN full-length enriched library, clone:D630035A07 product:inferred: phosphodiesterase 3A, cGMP inhibited, full insert sequence. [AK052718] |
KLA | .93 |
1.02 |
.94 |
1.00 |
.98 |
1.03 |
.96 |
| ATP | 1.03 |
.96 |
1.00 |
.98 |
1.00 |
1.03 |
1.01 |
| KLA/ATP | .98 |
1.06 |
1.08 |
1.03 |
.94 |
.98 |
.89 |
|
| Pde3a |  |
NM_018779 |
phosphodiesterase 3A, cGMP inhibited (Pde3a), mRNA [NM_018779] |
KLA | 1.01 |
1.09 |
.98 |
.98 |
.93 |
1.03 |
1.09 |
| ATP | .99 |
.97 |
1.04 |
1.05 |
.96 |
.96 |
1.06 |
| KLA/ATP | 1.06 |
.97 |
.94 |
1.09 |
1.00 |
1.06 |
.96 |
|
| Pde3b | 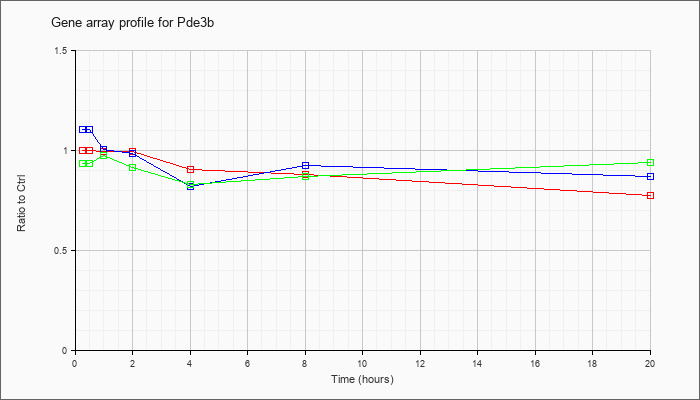 |
AK044183 |
10 days neonate cortex cDNA, RIKEN full-length enriched library, clone:A830097M14 product:phosphodiesterase 3B, cGMP-inhibited, full insert sequence. [AK044183] |
KLA | .98 |
.99 |
1.04 |
.99 |
.93 |
1.01 |
.96 |
| ATP | 1.08 |
.96 |
.92 |
.93 |
.93 |
.90 |
.93 |
| KLA/ATP | .97 |
.94 |
.95 |
.89 |
.92 |
.89 |
.95 |
|
| Pde3b |  |
NM_011055 |
phosphodiesterase 3B, cGMP-inhibited (Pde3b), mRNA [NM_011055] |
KLA | 1.02 |
1.01 |
.95 |
1.00 |
.88 |
.75 |
.59 |
| ATP | 1.12 |
1.20 |
1.09 |
1.04 |
.71 |
.95 |
.81 |
| KLA/ATP | .90 |
.92 |
1.00 |
.94 |
.74 |
.85 |
.93 |
|
| Pgr | 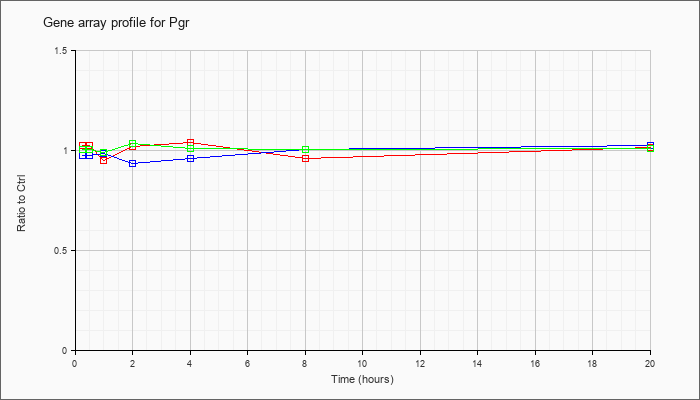 |
AK036862 |
adult female vagina cDNA, RIKEN full-length enriched library, clone:9930019P03 product:PROGESTERONE RECEPTOR (PR) homolog [Mus musculus], full insert sequence. [AK036862] |
KLA | 1.05 |
1.06 |
.93 |
.99 |
1.10 |
.96 |
.98 |
| ATP | .96 |
1.04 |
.97 |
.94 |
.97 |
1.04 |
1.05 |
| KLA/ATP | .97 |
.96 |
1.00 |
1.06 |
1.01 |
.99 |
1.00 |
|
| Pgr |  |
NM_008829 |
progesterone receptor (Pgr), mRNA [NM_008829] |
KLA | 1.00 |
.95 |
.97 |
1.05 |
.98 |
.96 |
1.05 |
| ATP | .99 |
1.16 |
1.00 |
.93 |
.95 |
.97 |
1.00 |
| KLA/ATP | 1.04 |
1.04 |
.98 |
1.01 |
1.01 |
1.02 |
1.02 |
|
| Pik3ca | 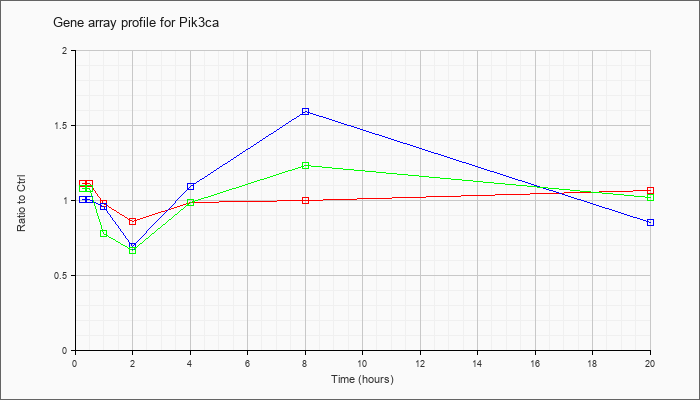 |
ENSMUST00000108243 |
ens|Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha isoform (EC 2.7.1.153) (PI3-kinase p110 subunit alpha) (PtdIns-3- kinase p110) (PI3K). [Source:Uniprot/SWISSPROT;Acc:P42337] [ENSMUST00000108243] |
KLA | 1.07 |
1.01 |
.91 |
.90 |
.98 |
.96 |
.86 |
| ATP | 1.03 |
.87 |
1.12 |
.63 |
1.22 |
1.86 |
.60 |
| KLA/ATP | 1.05 |
.82 |
.91 |
.61 |
1.17 |
1.26 |
.63 |
|
| Pik3ca |  |
NM_008839 |
phosphatidylinositol 3-kinase, catalytic, alpha polypeptide (Pik3ca), mRNA [NM_008839] |
KLA | 1.15 |
1.11 |
1.05 |
.82 |
.99 |
1.03 |
1.27 |
| ATP | .98 |
1.00 |
.79 |
.75 |
.96 |
1.32 |
1.10 |
| KLA/ATP | 1.10 |
.98 |
.64 |
.72 |
.80 |
1.20 |
1.41 |
|
| Pik3cb | 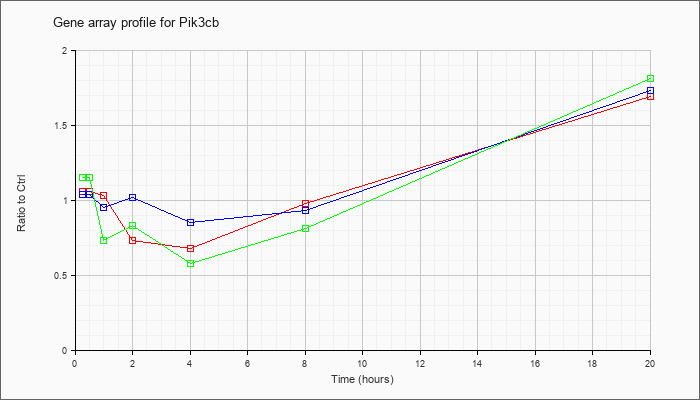 |
NM_029094 |
phosphatidylinositol 3-kinase, catalytic, beta polypeptide (Pik3cb), mRNA [NM_029094] |
KLA | 1.06 |
1.06 |
1.03 |
.73 |
.68 |
.98 |
1.69 |
| ATP | 1.04 |
1.21 |
.95 |
1.02 |
.85 |
.93 |
1.73 |
| KLA/ATP | 1.15 |
1.27 |
.73 |
.83 |
.58 |
.81 |
1.81 |
|
| Pik3cd | 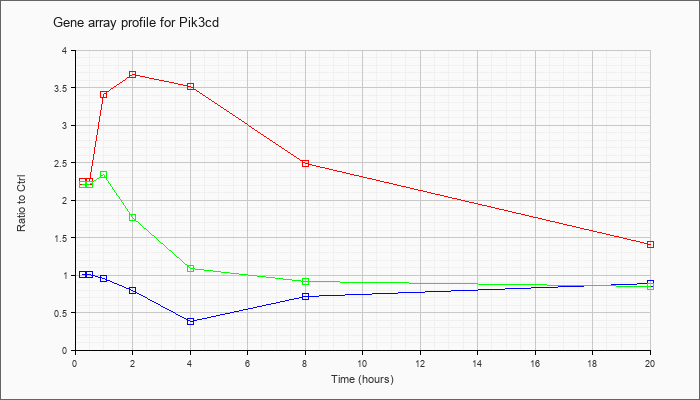 |
NM_008840 |
phosphatidylinositol 3-kinase catalytic delta polypeptide (Pik3cd), transcript variant 1, mRNA [NM_008840] |
KLA | 2.28 |
2.42 |
3.10 |
4.17 |
3.35 |
2.31 |
1.34 |
| ATP | 1.02 |
.91 |
.99 |
.63 |
.30 |
.67 |
.80 |
| KLA/ATP | 2.14 |
2.23 |
2.70 |
1.51 |
1.17 |
.85 |
.72 |
|
| Pik3cd |  |
U86587 |
phosphatidylinositol 3-kinase catalytic subunit p110 delta mRNA, complete cds. [U86587] |
KLA | 2.21 |
2.24 |
3.71 |
3.17 |
3.67 |
2.66 |
1.48 |
| ATP | 1.00 |
1.03 |
.93 |
.96 |
.46 |
.75 |
.97 |
| KLA/ATP | 2.27 |
2.32 |
1.97 |
2.02 |
1.01 |
.99 |
.98 |
|
| Pik3cg | 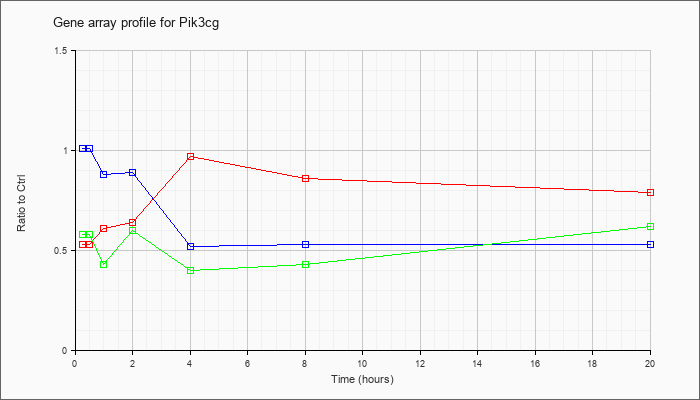 |
NM_020272 |
phosphoinositide-3-kinase, catalytic, gamma polypeptide (Pik3cg), mRNA [NM_020272] |
KLA | .53 |
.53 |
.61 |
.64 |
.97 |
.86 |
.79 |
| ATP | 1.01 |
1.13 |
.88 |
.89 |
.52 |
.53 |
.53 |
| KLA/ATP | .58 |
.53 |
.43 |
.60 |
.40 |
.43 |
.62 |
|
| Pik3r1 | 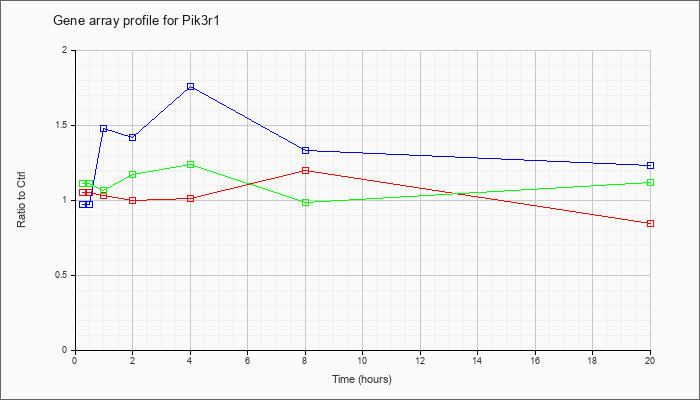 |
NM_001077495 |
phosphatidylinositol 3-kinase, regulatory subunit, polypeptide 1 (p85 alpha) (Pik3r1), transcript variant 2, mRNA [NM_001077495] |
KLA | 1.05 |
1.07 |
1.03 |
1.00 |
1.01 |
1.20 |
.85 |
| ATP | .97 |
1.02 |
1.48 |
1.42 |
1.76 |
1.33 |
1.23 |
| KLA/ATP | 1.11 |
1.12 |
1.06 |
1.17 |
1.24 |
.99 |
1.12 |
|
| Pik3r2 | 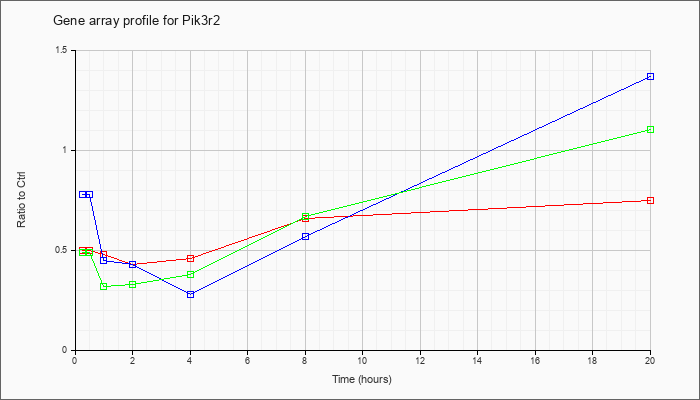 |
NM_008841 |
phosphatidylinositol 3-kinase, regulatory subunit, polypeptide 2 (p85 beta) (Pik3r2), mRNA [NM_008841] |
KLA | .50 |
.50 |
.48 |
.43 |
.46 |
.66 |
.75 |
| ATP | .78 |
.68 |
.45 |
.43 |
.28 |
.57 |
1.37 |
| KLA/ATP | .49 |
.38 |
.32 |
.33 |
.38 |
.67 |
1.10 |
|
| Pik3r3 | 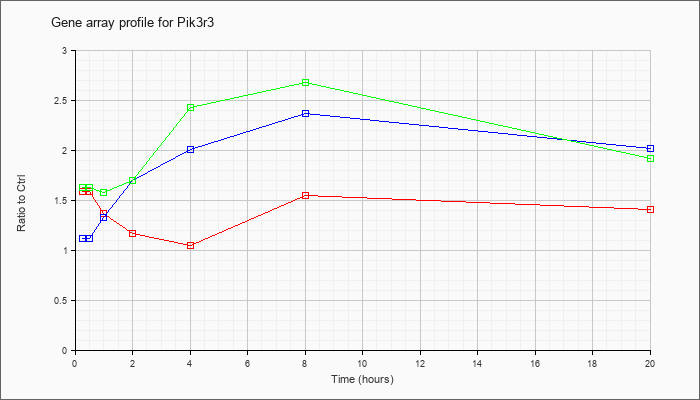 |
NM_181585 |
phosphatidylinositol 3 kinase, regulatory subunit, polypeptide 3 (p55) (Pik3r3), mRNA [NM_181585] |
KLA | 1.59 |
1.49 |
1.37 |
1.17 |
1.05 |
1.55 |
1.41 |
| ATP | 1.12 |
1.00 |
1.33 |
1.70 |
2.01 |
2.37 |
2.02 |
| KLA/ATP | 1.63 |
1.54 |
1.58 |
1.70 |
2.43 |
2.68 |
1.92 |
|
| Pik3r5 | 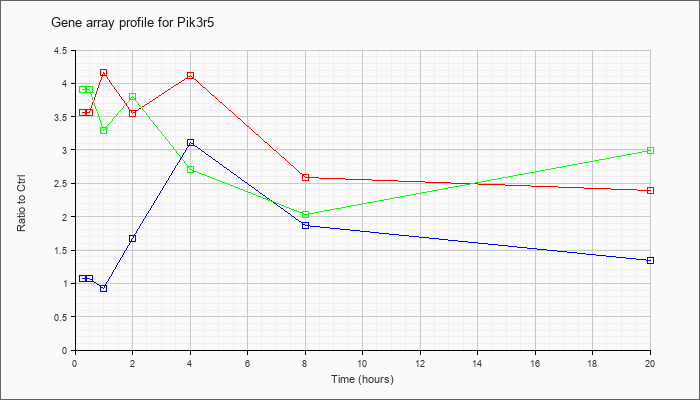 |
NM_177320 |
phosphoinositide-3-kinase, regulatory subunit 5, p101 (Pik3r5), mRNA [NM_177320] |
KLA | 3.57 |
3.58 |
4.16 |
3.54 |
4.11 |
2.59 |
2.39 |
| ATP | 1.08 |
1.08 |
.93 |
1.67 |
3.11 |
1.88 |
1.34 |
| KLA/ATP | 3.91 |
3.70 |
3.30 |
3.80 |
2.70 |
2.04 |
3.00 |
|
| Plk1 | 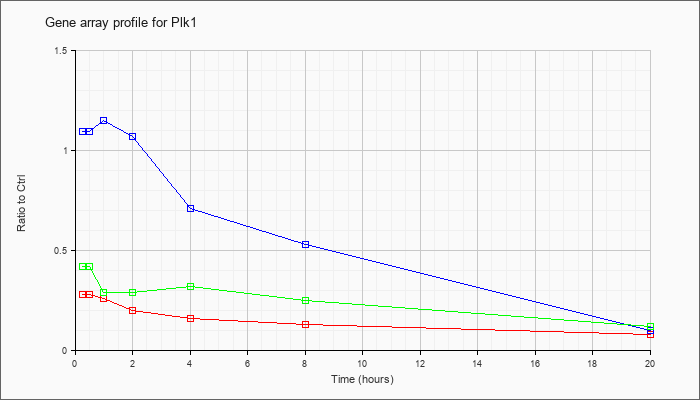 |
NM_011121 |
polo-like kinase 1 (Drosophila) (Plk1), mRNA [NM_011121] |
KLA | .28 |
.23 |
.26 |
.20 |
.16 |
.13 |
.08 |
| ATP | 1.09 |
1.06 |
1.15 |
1.07 |
.71 |
.53 |
.10 |
| KLA/ATP | .42 |
.26 |
.29 |
.29 |
.32 |
.25 |
.12 |
|
| Prkaca | 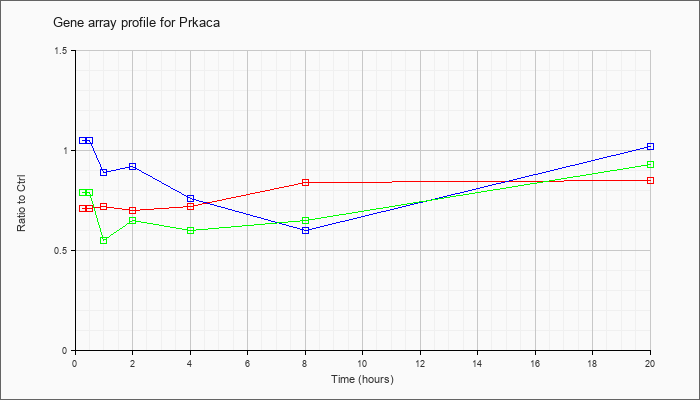 |
NM_008854 |
protein kinase, cAMP dependent, catalytic, alpha (Prkaca), mRNA [NM_008854] |
KLA | .71 |
.74 |
.72 |
.70 |
.72 |
.84 |
.85 |
| ATP | 1.05 |
.98 |
.89 |
.92 |
.76 |
.60 |
1.02 |
| KLA/ATP | .79 |
.74 |
.55 |
.65 |
.60 |
.65 |
.93 |
|
| Prkacb | 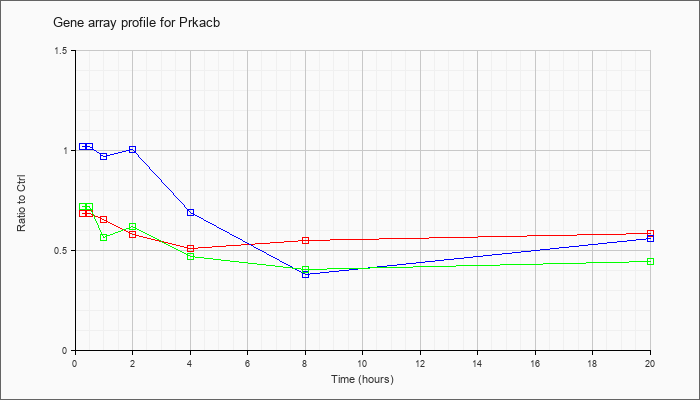 |
NM_011100 |
protein kinase, cAMP dependent, catalytic, beta (Prkacb), mRNA [NM_011100] |
KLA | .69 |
.70 |
.66 |
.58 |
.51 |
.55 |
.59 |
| ATP | 1.02 |
1.19 |
.97 |
1.01 |
.69 |
.38 |
.56 |
| KLA/ATP | .72 |
.73 |
.57 |
.62 |
.47 |
.41 |
.45 |
|
| Prkx | 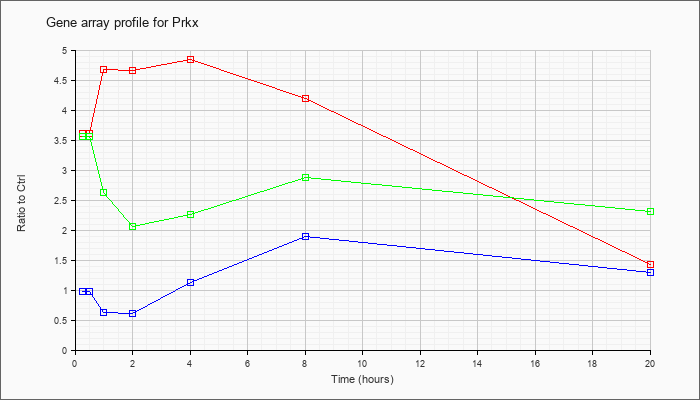 |
NM_016979 |
protein kinase, X-linked (Prkx), mRNA [NM_016979] |
KLA | 3.61 |
3.79 |
4.67 |
4.66 |
4.85 |
4.19 |
1.43 |
| ATP | .98 |
.90 |
.63 |
.61 |
1.12 |
1.90 |
1.29 |
| KLA/ATP | 3.56 |
3.36 |
2.62 |
2.06 |
2.26 |
2.87 |
2.31 |
|
| Raf1 | 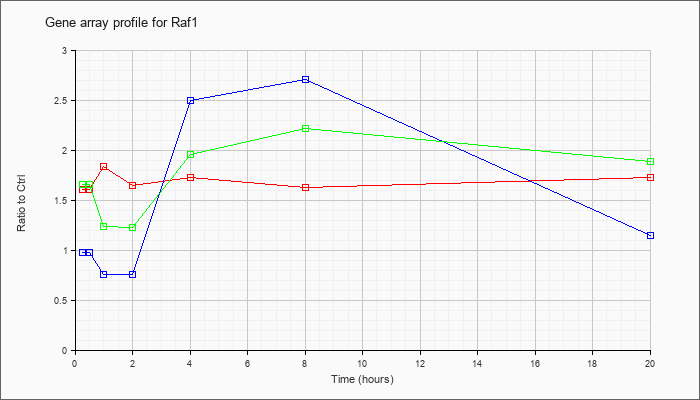 |
AK036317 |
16 days neonate cerebellum cDNA, RIKEN full-length enriched library, clone:9630056E02 product:v-raf-1 leukemia viral oncogene 1, full insert sequence. [AK036317] |
KLA | 1.47 |
1.34 |
1.55 |
1.29 |
1.59 |
1.32 |
1.46 |
| ATP | 1.05 |
.98 |
.54 |
.63 |
3.32 |
2.07 |
1.01 |
| KLA/ATP | 1.57 |
1.33 |
.79 |
.96 |
2.26 |
1.38 |
1.35 |
|
| Raf1 |  |
NM_029780 |
v-raf-leukemia viral oncogene 1 (Raf1), mRNA [NM_029780] |
KLA | 1.65 |
1.64 |
1.93 |
1.77 |
1.77 |
1.72 |
1.81 |
| ATP | .95 |
.96 |
.83 |
.80 |
2.22 |
2.92 |
1.19 |
| KLA/ATP | 1.68 |
1.64 |
1.39 |
1.32 |
1.85 |
2.50 |
2.06 |
|
| Rps6ka1 | 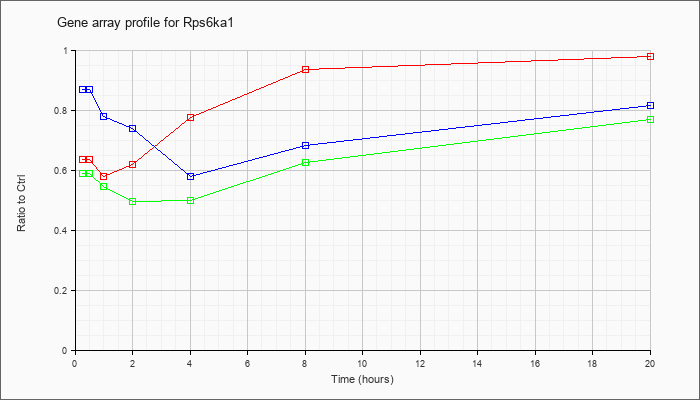 |
NM_009097 |
ribosomal protein S6 kinase polypeptide 1 (Rps6ka1), mRNA [NM_009097] |
KLA | .64 |
.65 |
.58 |
.62 |
.78 |
.94 |
.98 |
| ATP | .87 |
.85 |
.78 |
.74 |
.58 |
.68 |
.82 |
| KLA/ATP | .59 |
.52 |
.55 |
.50 |
.50 |
.63 |
.77 |
|
| Rps6ka2 | 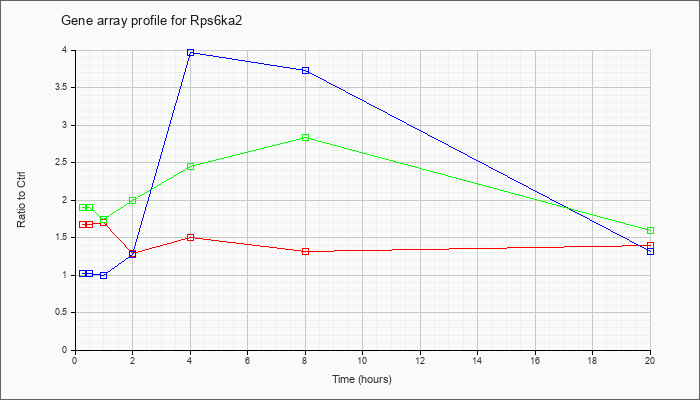 |
NM_011299 |
ribosomal protein S6 kinase, polypeptide 2 (Rps6ka2), mRNA [NM_011299] |
KLA | 1.67 |
1.69 |
1.70 |
1.29 |
1.50 |
1.32 |
1.40 |
| ATP | 1.02 |
1.02 |
.99 |
1.28 |
3.97 |
3.73 |
1.32 |
| KLA/ATP | 1.90 |
1.63 |
1.74 |
1.99 |
2.45 |
2.84 |
1.59 |
|
| Rps6ka3 | 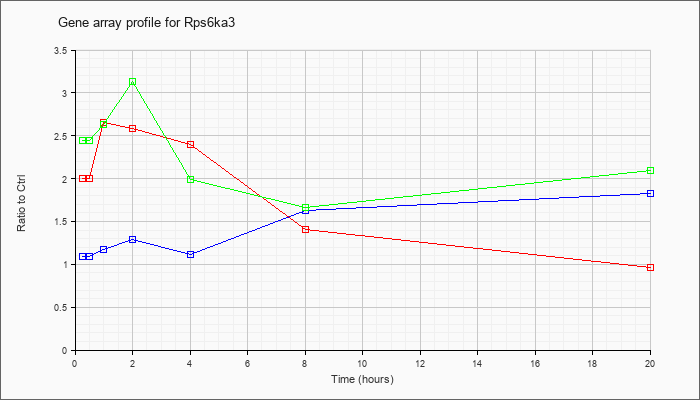 |
NM_148945 |
ribosomal protein S6 kinase polypeptide 3 (Rps6ka3), mRNA [NM_148945] |
KLA | 2.00 |
2.37 |
2.65 |
2.59 |
2.40 |
1.41 |
.96 |
| ATP | 1.09 |
1.09 |
1.17 |
1.29 |
1.11 |
1.63 |
1.83 |
| KLA/ATP | 2.44 |
2.44 |
2.63 |
3.13 |
1.99 |
1.66 |
2.09 |
|
| Rps6ka6 | 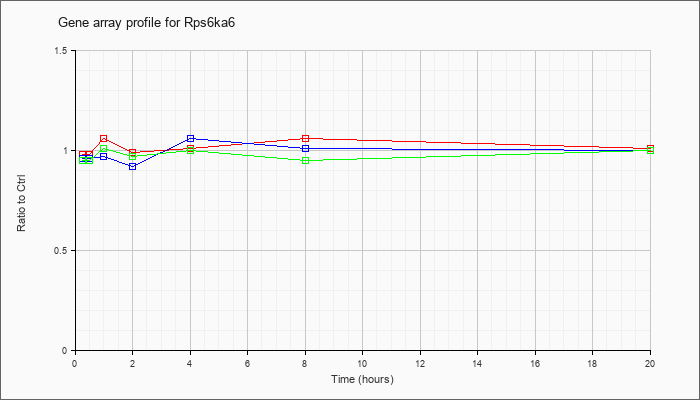 |
NM_025949 |
ribosomal protein S6 kinase polypeptide 6 (Rps6ka6), mRNA [NM_025949] |
KLA | .98 |
1.03 |
1.06 |
.99 |
1.01 |
1.06 |
1.01 |
| ATP | .96 |
1.00 |
.97 |
.92 |
1.06 |
1.01 |
1.00 |
| KLA/ATP | .95 |
1.01 |
1.01 |
.97 |
1.00 |
.95 |
1.00 |
|
| Spdyb |  |
NM_029048 |
speedy homolog B (Drosophila) (Spdyb), mRNA [NM_029048] |
KLA | 1.02 |
1.01 |
1.01 |
.99 |
1.00 |
1.02 |
1.02 |
| ATP | .96 |
1.01 |
1.01 |
1.05 |
1.00 |
1.04 |
1.01 |
| KLA/ATP | 1.01 |
.99 |
.97 |
1.03 |
1.00 |
1.04 |
1.01 |
|