| Gene symbol |
Time course plot |
Accession |
Description |
Treatment |
15min |
30min |
1hr |
2hr |
4hr |
8hr |
20hr |
| Akt1 | 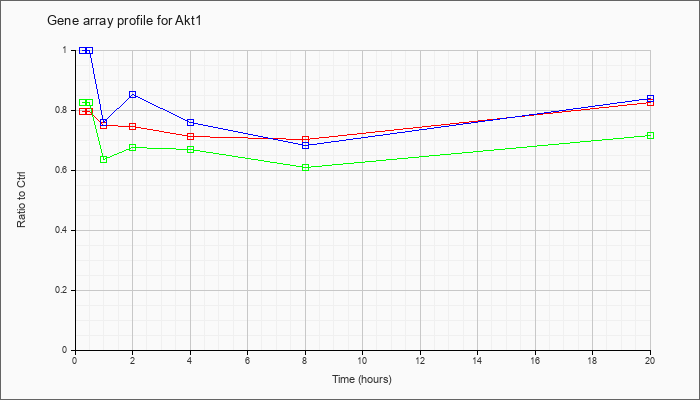 |
NM_009652 |
thymoma viral proto-oncogene 1 (Akt1), mRNA [NM_009652] |
KLA | .80 |
.80 |
.75 |
.75 |
.71 |
.70 |
.82 |
| ATP | 1.00 |
1.02 |
.76 |
.85 |
.76 |
.68 |
.84 |
| KLA/ATP | .82 |
.80 |
.64 |
.68 |
.67 |
.61 |
.72 |
|
| Akt2 | 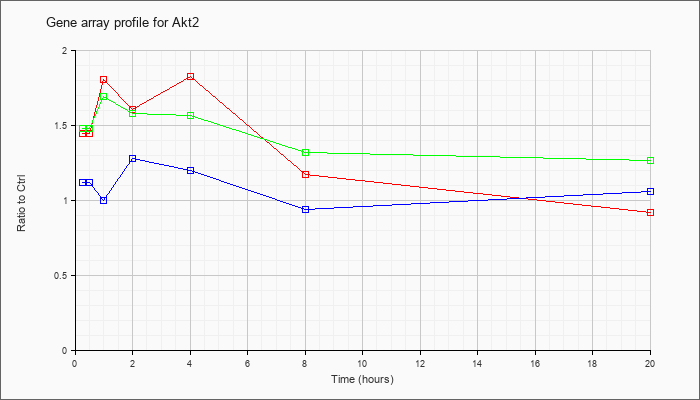 |
NM_007434 |
thymoma viral proto-oncogene 2 (Akt2), transcript variant 2, mRNA [NM_007434] |
KLA | 1.45 |
1.49 |
1.81 |
1.61 |
1.83 |
1.17 |
.92 |
| ATP | 1.12 |
1.12 |
1.00 |
1.28 |
1.20 |
.94 |
1.06 |
| KLA/ATP | 1.48 |
1.70 |
1.69 |
1.58 |
1.57 |
1.32 |
1.27 |
|
| Akt3 | 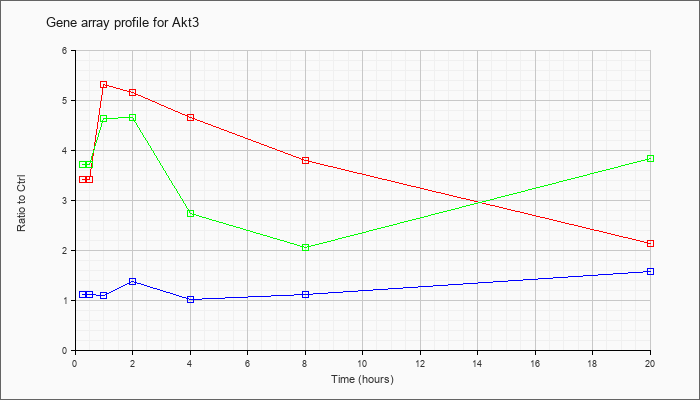 |
NM_011785 |
thymoma viral proto-oncogene 3 (Akt3), mRNA [NM_011785] |
KLA | 3.42 |
3.45 |
5.31 |
5.16 |
4.65 |
3.80 |
2.12 |
| ATP | 1.12 |
1.22 |
1.10 |
1.37 |
1.02 |
1.11 |
1.56 |
| KLA/ATP | 3.72 |
4.17 |
4.62 |
4.64 |
2.74 |
2.05 |
3.82 |
|
BB229853
|  |
BB229853 |
gb|BB229853 RIKEN full-length enriched, 3 days neonate thymus Mus musculus cDNA clone A630023O21 3. [BB229853] |
KLA | 6.32 |
5.94 |
6.13 |
6.41 |
7.39 |
12.33 |
6.08 |
| ATP | .39 |
.24 |
.33 |
.38 |
1.18 |
5.75 |
2.85 |
| KLA/ATP | 2.27 |
2.02 |
3.25 |
4.04 |
7.11 |
10.22 |
13.93 |
|
| Ccnd1 | 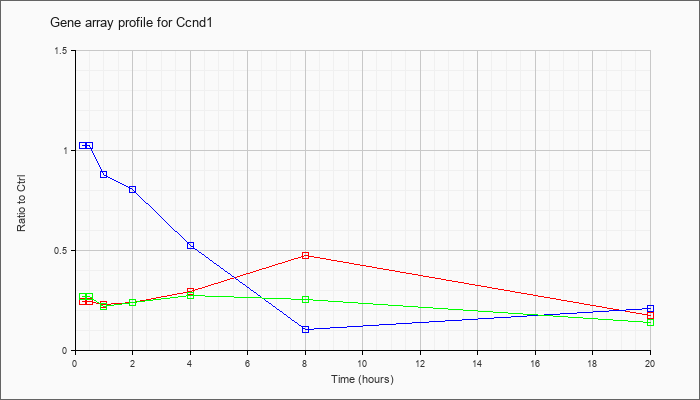 |
S78355 |
gb|Cyl-1=cyclin D1 [mice, BALB/c, brain, mRNA, 3737 nt]. [S78355] |
KLA | .24 |
.23 |
.23 |
.24 |
.29 |
.47 |
.17 |
| ATP | 1.02 |
1.02 |
.88 |
.80 |
.52 |
.10 |
.21 |
| KLA/ATP | .27 |
.23 |
.22 |
.24 |
.27 |
.25 |
.14 |
|
| Ccnd2 | 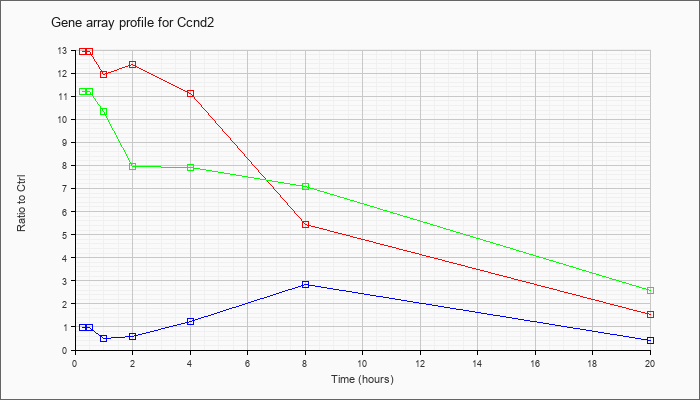 |
NM_009829 |
cyclin D2 (Ccnd2), mRNA [NM_009829] |
KLA | 12.92 |
13.66 |
11.94 |
12.37 |
11.11 |
5.44 |
1.53 |
| ATP | .98 |
.75 |
.50 |
.57 |
1.24 |
2.83 |
.40 |
| KLA/ATP | 11.19 |
11.62 |
10.34 |
7.96 |
7.92 |
7.10 |
2.57 |
|
| Cga | 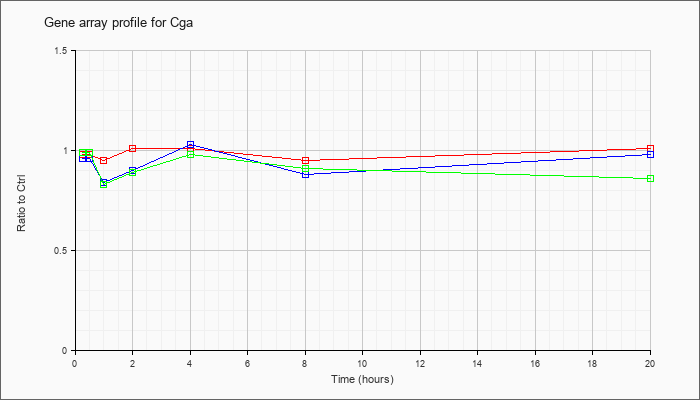 |
NM_009889 |
glycoprotein hormones, alpha subunit (Cga), mRNA [NM_009889] |
KLA | .98 |
.95 |
.95 |
1.01 |
1.01 |
.95 |
1.01 |
| ATP | .96 |
.89 |
.84 |
.90 |
1.03 |
.88 |
.98 |
| KLA/ATP | .99 |
.98 |
.83 |
.89 |
.98 |
.91 |
.86 |
|
| Cish | 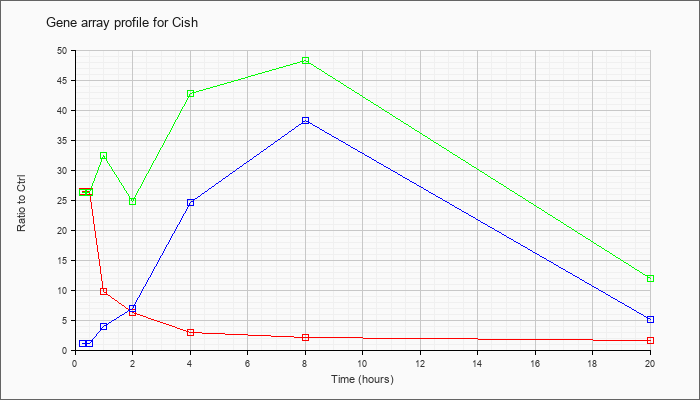 |
NM_009895 |
cytokine inducible SH2-containing protein (Cish), mRNA [NM_009895] |
KLA | 26.44 |
22.18 |
9.67 |
6.27 |
2.93 |
2.06 |
1.53 |
| ATP | 1.03 |
1.35 |
3.93 |
7.00 |
24.56 |
38.19 |
5.08 |
| KLA/ATP | 26.32 |
33.26 |
32.35 |
24.73 |
42.75 |
48.19 |
11.96 |
|
| Csn2 | 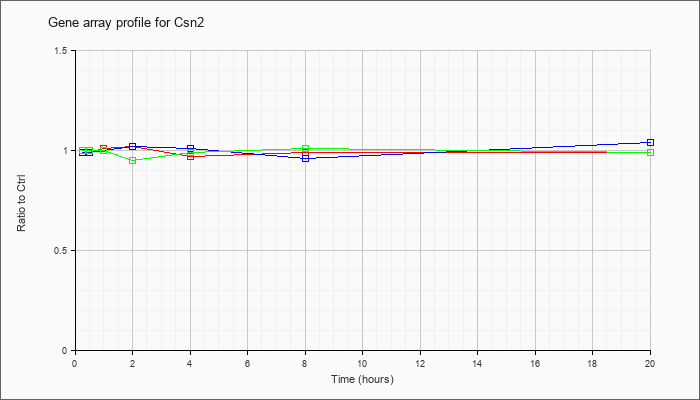 |
NM_009972 |
casein beta (Csn2), mRNA [NM_009972] |
KLA | .99 |
1.07 |
1.01 |
1.02 |
.97 |
.99 |
.99 |
| ATP | .99 |
.94 |
1.00 |
1.02 |
1.01 |
.96 |
1.04 |
| KLA/ATP | 1.00 |
.98 |
1.00 |
.95 |
.99 |
1.01 |
.99 |
|
| Cyp17a1 | 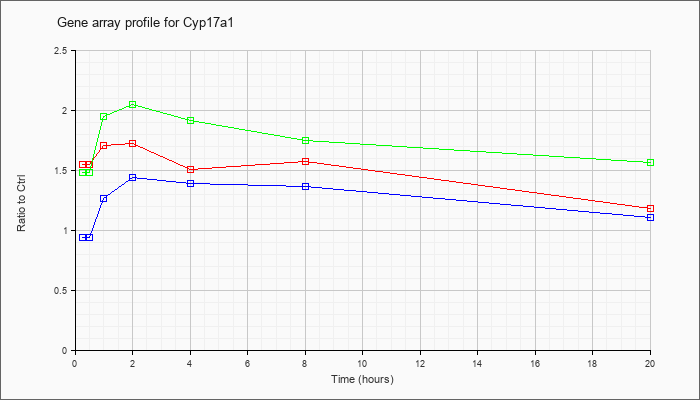 |
NM_007809 |
cytochrome P450, family 17, subfamily a, polypeptide 1 (Cyp17a1), mRNA [NM_007809] |
KLA | 1.55 |
1.60 |
1.71 |
1.73 |
1.51 |
1.58 |
1.18 |
| ATP | .94 |
1.01 |
1.27 |
1.44 |
1.39 |
1.36 |
1.11 |
| KLA/ATP | 1.48 |
1.79 |
1.95 |
2.05 |
1.92 |
1.75 |
1.56 |
|
| Elf5 | 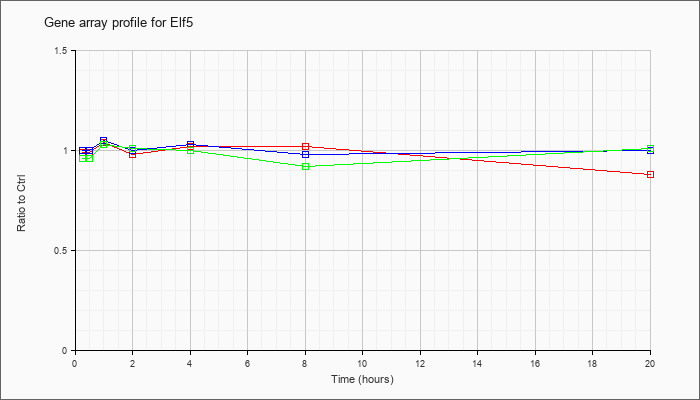 |
NM_010125 |
E74-like factor 5 (Elf5), mRNA [NM_010125] |
KLA | .99 |
.99 |
1.04 |
.98 |
1.02 |
1.02 |
.88 |
| ATP | 1.00 |
1.02 |
1.05 |
1.00 |
1.03 |
.98 |
1.00 |
| KLA/ATP | .96 |
1.05 |
1.03 |
1.01 |
1.00 |
.92 |
1.01 |
|
| Esr1 | 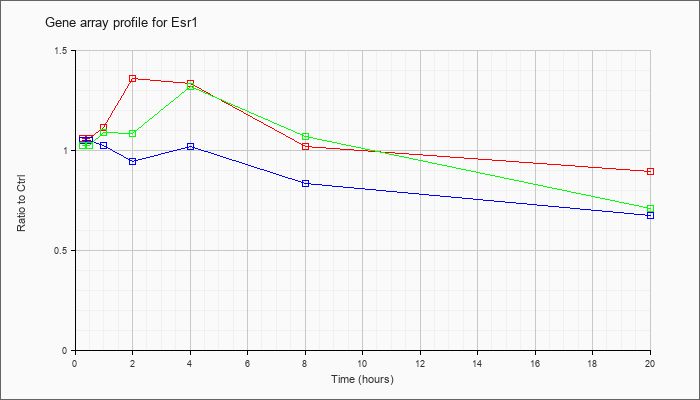 |
NM_007956 |
estrogen receptor 1 (alpha) (Esr1), mRNA [NM_007956] |
KLA | 1.06 |
1.05 |
1.11 |
1.36 |
1.33 |
1.02 |
.90 |
| ATP | 1.05 |
.92 |
1.02 |
.94 |
1.02 |
.83 |
.67 |
| KLA/ATP | 1.02 |
.98 |
1.09 |
1.08 |
1.32 |
1.07 |
.71 |
|
| Esr2 | 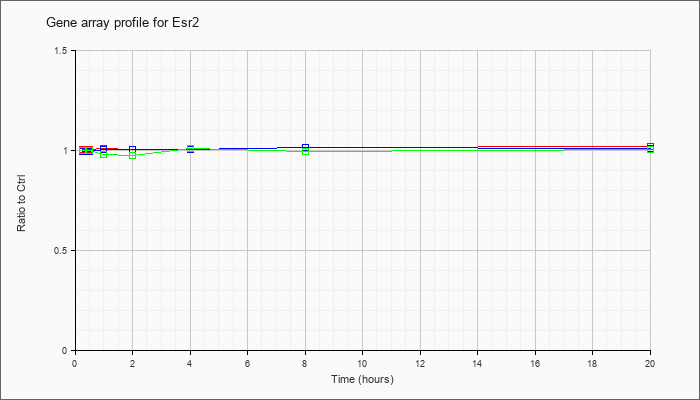 |
NM_207707 |
estrogen receptor 2 (beta) (Esr2), transcript variant 1, mRNA [NM_207707] |
KLA | 1.00 |
1.00 |
1.01 |
1.00 |
1.00 |
1.01 |
1.02 |
| ATP | .99 |
1.00 |
1.00 |
1.00 |
1.00 |
1.01 |
1.01 |
| KLA/ATP | 1.00 |
1.02 |
.98 |
.97 |
1.01 |
.99 |
1.00 |
|
| Fos | 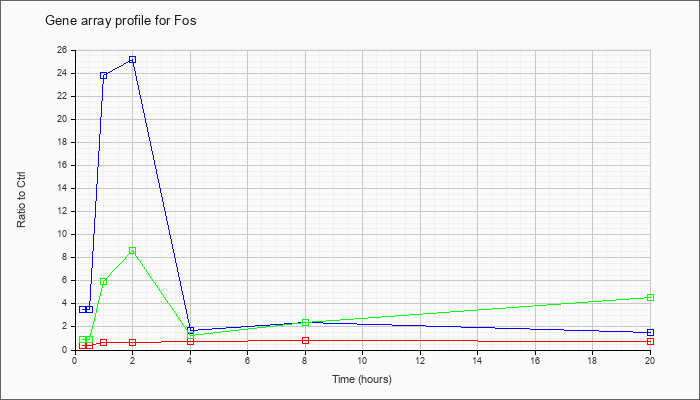 |
NM_010234 |
FBJ osteosarcoma oncogene (Fos), mRNA [NM_010234] |
KLA | .42 |
.33 |
.64 |
.61 |
.71 |
.85 |
.71 |
| ATP | 3.48 |
5.46 |
23.78 |
25.22 |
1.71 |
2.36 |
1.50 |
| KLA/ATP | .90 |
1.74 |
5.97 |
8.62 |
1.28 |
2.42 |
4.52 |
|
| Foxo3a | 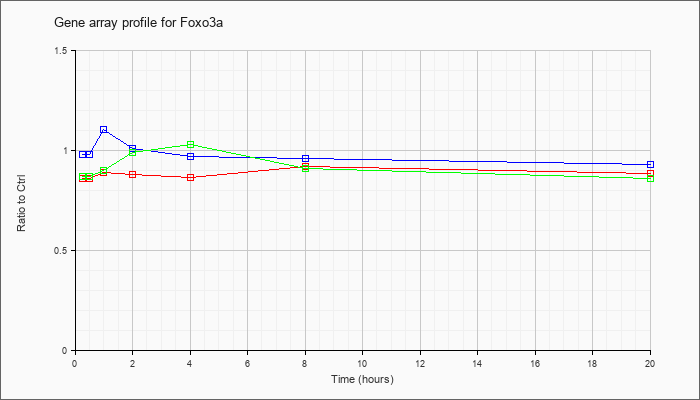 |
AK079567 |
adult male hypothalamus cDNA, RIKEN full-length enriched library, clone:A230087I01 product:unclassifiable, full insert sequence. [AK079567] |
KLA | .82 |
.81 |
.91 |
.83 |
.86 |
.86 |
.81 |
| ATP | .98 |
1.00 |
1.13 |
1.03 |
.97 |
.90 |
.82 |
| KLA/ATP | .86 |
.80 |
.90 |
.99 |
1.07 |
.84 |
.73 |
|
| Foxo3a |  |
NM_019740 |
forkhead box O3a (Foxo3a), mRNA [NM_019740] |
KLA | .94 |
.92 |
.86 |
.97 |
.87 |
1.04 |
1.04 |
| ATP | .99 |
1.09 |
1.06 |
.97 |
.96 |
1.07 |
1.15 |
| KLA/ATP | .90 |
.79 |
.90 |
.99 |
.95 |
1.04 |
1.11 |
|
| Galt | 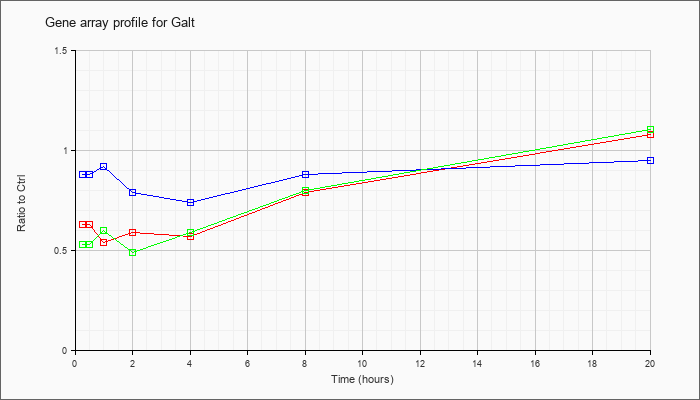 |
NM_016658 |
galactose-1-phosphate uridyl transferase (Galt), mRNA [NM_016658] |
KLA | .63 |
.59 |
.54 |
.59 |
.57 |
.79 |
1.08 |
| ATP | .88 |
.77 |
.92 |
.79 |
.74 |
.88 |
.95 |
| KLA/ATP | .53 |
.53 |
.60 |
.49 |
.59 |
.80 |
1.10 |
|
| Gck | 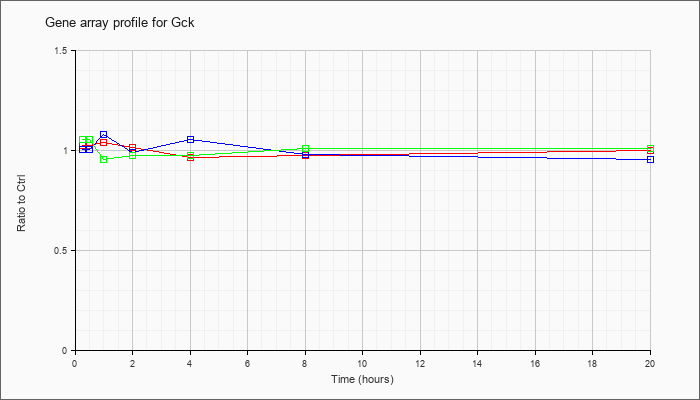 |
NM_010292 |
glucokinase (Gck), mRNA [NM_010292] |
KLA | 1.03 |
.99 |
1.04 |
1.02 |
.97 |
.98 |
1.00 |
| ATP | 1.01 |
.97 |
1.08 |
.99 |
1.06 |
.98 |
.96 |
| KLA/ATP | 1.06 |
1.01 |
.96 |
.98 |
.98 |
1.01 |
1.01 |
|
| Grb2 | 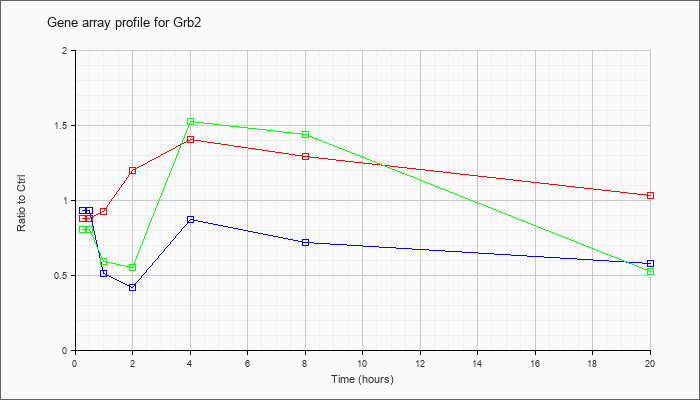 |
NM_008163 |
growth factor receptor bound protein 2 (Grb2), mRNA [NM_008163] |
KLA | .92 |
.88 |
.90 |
1.27 |
1.43 |
1.22 |
1.00 |
| ATP | 1.02 |
.83 |
.51 |
.42 |
.92 |
.70 |
.57 |
| KLA/ATP | .84 |
.77 |
.61 |
.56 |
1.56 |
1.49 |
.52 |
|
| Grb2 |  |
U07617 |
Grb2 adaptor protein (grb2) mRNA, complete cds. [U07617] |
KLA | .83 |
.80 |
.95 |
1.13 |
1.38 |
1.36 |
1.06 |
| ATP | .84 |
.74 |
.51 |
.41 |
.82 |
.74 |
.59 |
| KLA/ATP | .77 |
.70 |
.57 |
.54 |
1.49 |
1.39 |
.53 |
|
| Gsk3b | 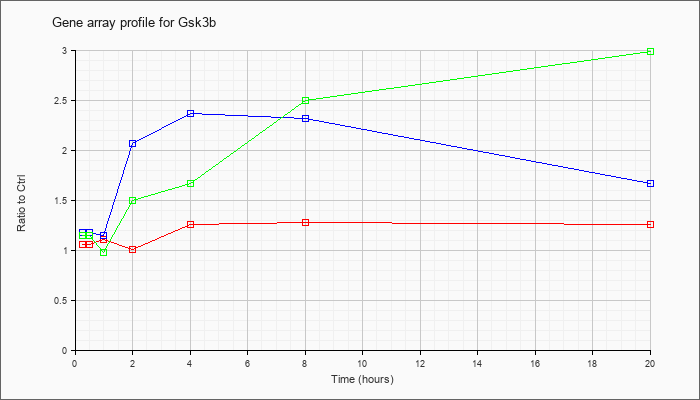 |
NM_019827 |
glycogen synthase kinase 3 beta (Gsk3b), mRNA [NM_019827] |
KLA | 1.06 |
1.03 |
1.11 |
1.01 |
1.26 |
1.28 |
1.26 |
| ATP | 1.18 |
1.27 |
1.15 |
2.07 |
2.37 |
2.32 |
1.67 |
| KLA/ATP | 1.15 |
1.21 |
.98 |
1.50 |
1.67 |
2.50 |
2.99 |
|
| Hras1 | 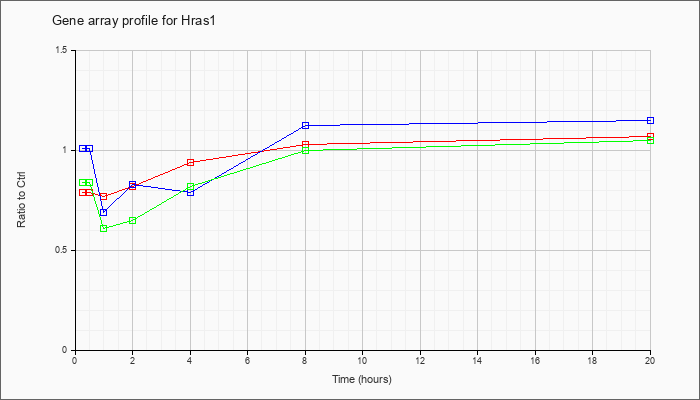 |
NM_008284 |
Harvey rat sarcoma virus oncogene 1 (Hras1), mRNA [NM_008284] |
KLA | .79 |
.82 |
.77 |
.82 |
.94 |
1.03 |
1.07 |
| ATP | 1.01 |
.95 |
.69 |
.83 |
.79 |
1.12 |
1.15 |
| KLA/ATP | .84 |
.77 |
.61 |
.65 |
.82 |
1.00 |
1.05 |
|
| Ins1 | 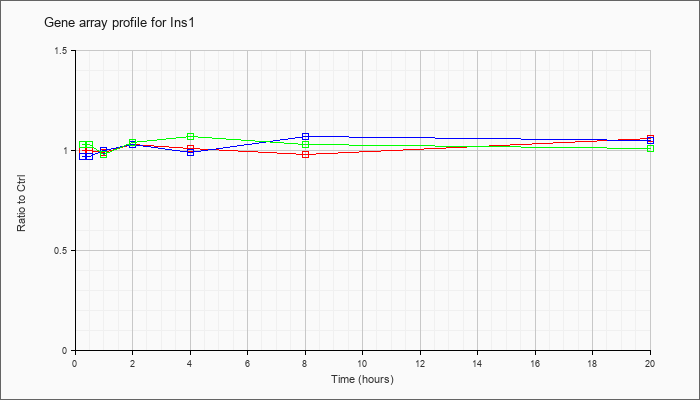 |
NM_008386 |
insulin I (Ins1), mRNA [NM_008386] |
KLA | 1.00 |
.99 |
.99 |
1.03 |
1.01 |
.98 |
1.06 |
| ATP | .97 |
1.03 |
1.00 |
1.03 |
.99 |
1.07 |
1.05 |
| KLA/ATP | 1.03 |
.93 |
.98 |
1.04 |
1.07 |
1.03 |
1.01 |
|
| Ins2 | 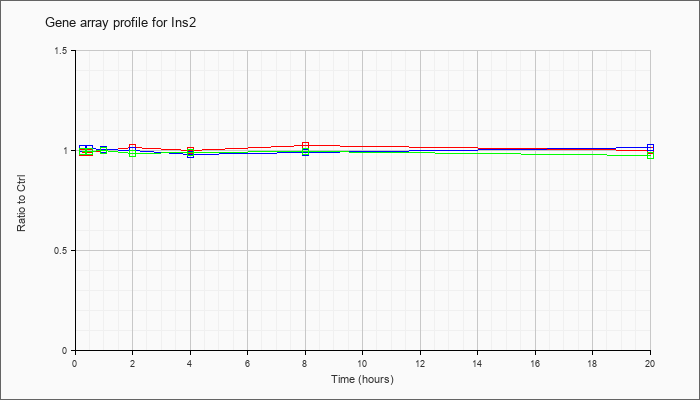 |
NM_008387 |
insulin II (Ins2), mRNA [NM_008387] |
KLA | .99 |
1.00 |
1.00 |
1.01 |
1.00 |
1.02 |
1.00 |
| ATP | 1.01 |
.99 |
1.00 |
1.00 |
.98 |
.99 |
1.01 |
| KLA/ATP | .99 |
1.01 |
1.00 |
.98 |
.99 |
.99 |
.98 |
|
| Irf1 | 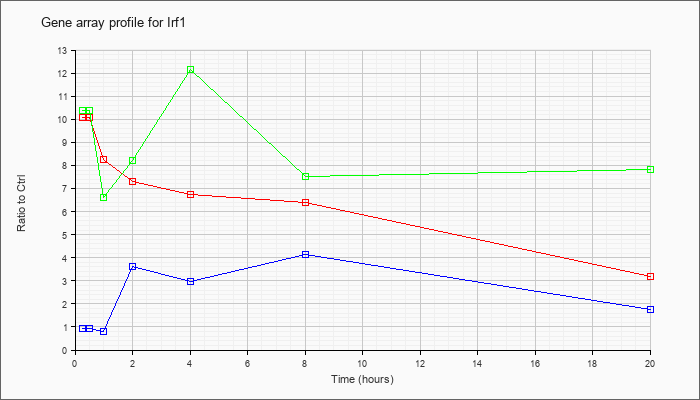 |
NM_008390 |
interferon regulatory factor 1 (Irf1), mRNA [NM_008390] |
KLA | 10.08 |
9.47 |
8.27 |
7.31 |
6.74 |
6.39 |
3.17 |
| ATP | .95 |
.81 |
.80 |
3.61 |
2.99 |
4.15 |
1.77 |
| KLA/ATP | 10.40 |
10.08 |
6.59 |
8.23 |
12.18 |
7.53 |
7.82 |
|
| Jak2 | 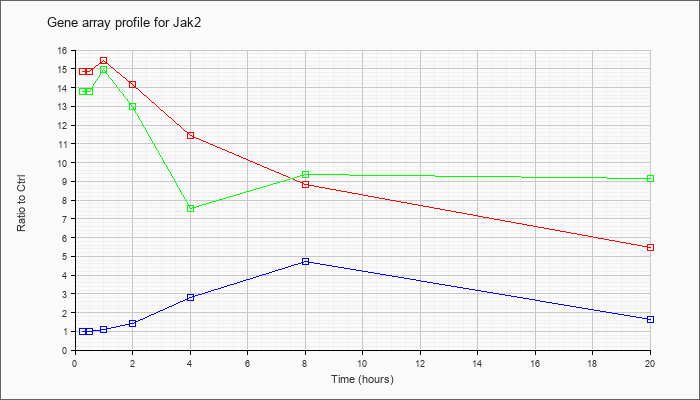 |
NM_008413 |
Janus kinase 2 (Jak2), transcript variant 1, mRNA [NM_008413] |
KLA | 14.86 |
15.05 |
15.42 |
14.15 |
11.46 |
8.81 |
5.47 |
| ATP | .99 |
1.06 |
1.09 |
1.43 |
2.83 |
4.71 |
1.62 |
| KLA/ATP | 13.77 |
14.74 |
14.94 |
12.97 |
7.53 |
9.34 |
9.17 |
|
| Kras | 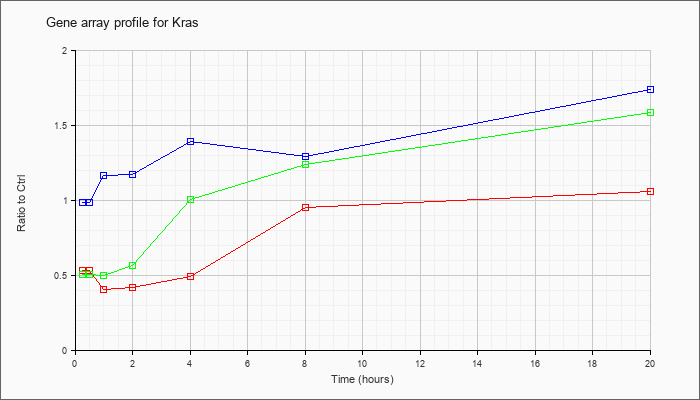 |
NM_021284 |
v-Ki-ras2 Kirsten rat sarcoma viral oncogene homolog (Kras), mRNA [NM_021284] |
KLA | .53 |
.49 |
.40 |
.42 |
.49 |
.95 |
1.05 |
| ATP | .98 |
1.01 |
1.16 |
1.17 |
1.39 |
1.29 |
1.74 |
| KLA/ATP | .50 |
.47 |
.50 |
.56 |
1.00 |
1.24 |
1.58 |
|
| Lhb | 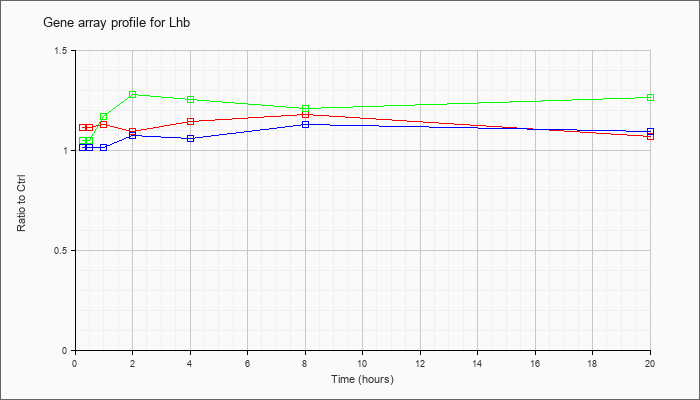 |
AK138425 |
adult male hypothalamus cDNA, RIKEN full-length enriched library, clone:A230101A12 product:luteinizing hormone beta, full insert sequence [AK138425] |
KLA | 1.06 |
1.01 |
1.02 |
.98 |
1.00 |
1.07 |
.91 |
| ATP | 1.02 |
1.04 |
.91 |
.96 |
.90 |
1.05 |
.95 |
| KLA/ATP | .98 |
.96 |
1.01 |
1.03 |
1.04 |
1.00 |
1.09 |
|
| Lhb |  |
NM_008497 |
luteinizing hormone beta (Lhb), mRNA [NM_008497] |
KLA | 1.16 |
1.26 |
1.24 |
1.20 |
1.29 |
1.29 |
1.23 |
| ATP | 1.01 |
1.01 |
1.12 |
1.19 |
1.22 |
1.21 |
1.23 |
| KLA/ATP | 1.12 |
1.12 |
1.33 |
1.53 |
1.47 |
1.42 |
1.44 |
|
| Lhcgr | 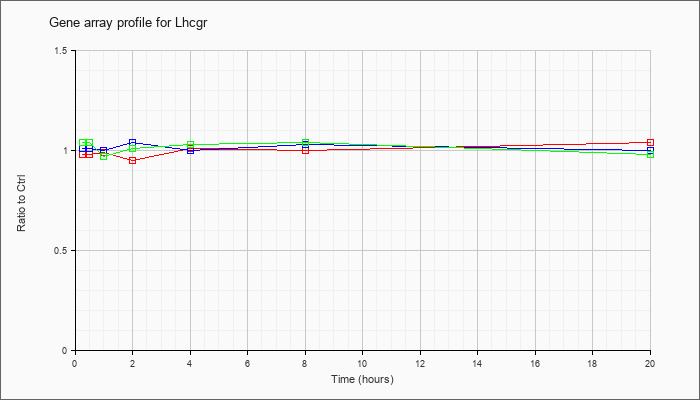 |
NM_013582 |
luteinizing hormone/choriogonadotropin receptor (Lhcgr), mRNA [NM_013582] |
KLA | .98 |
1.06 |
.99 |
.95 |
1.01 |
1.00 |
1.04 |
| ATP | 1.01 |
.95 |
1.00 |
1.04 |
1.00 |
1.03 |
1.00 |
| KLA/ATP | 1.04 |
1.02 |
.97 |
1.01 |
1.03 |
1.04 |
.98 |
|
| Map2k1 | 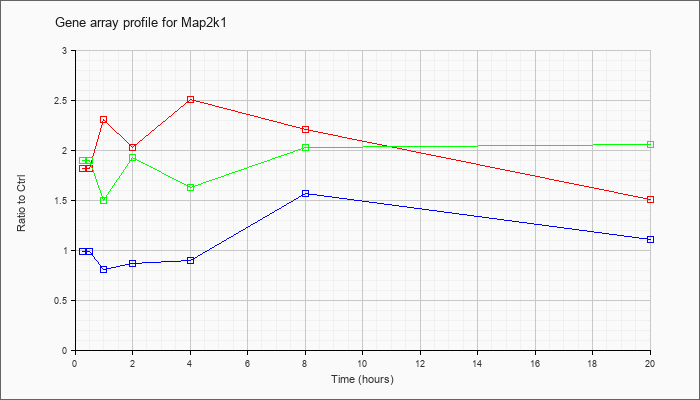 |
NM_008927 |
mitogen-activated protein kinase kinase 1 (Map2k1), mRNA [NM_008927] |
KLA | 1.82 |
2.00 |
2.31 |
2.03 |
2.51 |
2.20 |
1.51 |
| ATP | .99 |
1.14 |
.81 |
.87 |
.90 |
1.57 |
1.11 |
| KLA/ATP | 1.90 |
2.03 |
1.50 |
1.93 |
1.63 |
2.03 |
2.06 |
|
| Map2k2 | 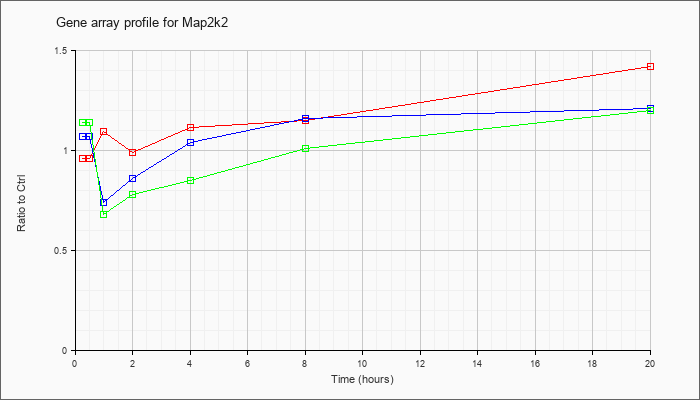 |
NM_023138 |
mitogen-activated protein kinase kinase 2 (Map2k2), mRNA [NM_023138] |
KLA | .96 |
1.01 |
1.09 |
.99 |
1.11 |
1.15 |
1.42 |
| ATP | 1.07 |
1.05 |
.74 |
.86 |
1.04 |
1.16 |
1.21 |
| KLA/ATP | 1.14 |
1.03 |
.68 |
.78 |
.85 |
1.01 |
1.20 |
|
| Mapk1 | 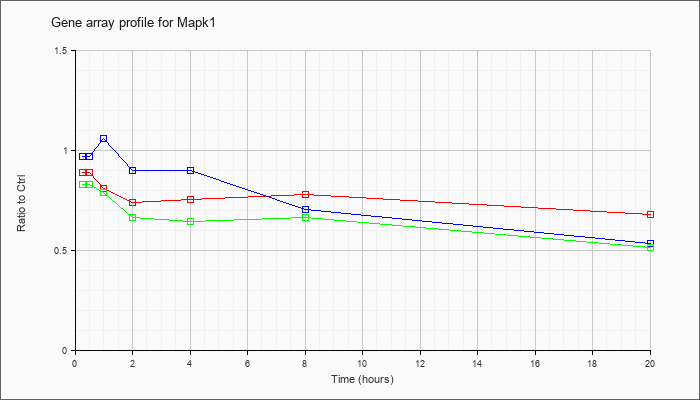 |
NM_001038663 |
mitogen-activated protein kinase 1 (Mapk1), transcript variant 2, mRNA [NM_001038663] |
KLA | .97 |
1.03 |
.93 |
.86 |
.79 |
1.01 |
1.04 |
| ATP | .96 |
1.18 |
1.01 |
1.13 |
.84 |
1.04 |
1.16 |
| KLA/ATP | .99 |
.99 |
.84 |
.87 |
.63 |
.96 |
1.26 |
|
| Mapk1 |  |
NM_011949 |
mitogen-activated protein kinase 1 (Mapk1), transcript variant 1, mRNA [NM_011949] |
KLA | .88 |
.80 |
.80 |
.73 |
.75 |
.76 |
.65 |
| ATP | .97 |
.96 |
1.06 |
.88 |
.90 |
.68 |
.48 |
| KLA/ATP | .82 |
.81 |
.78 |
.65 |
.64 |
.64 |
.45 |
|
| Mapk10 | 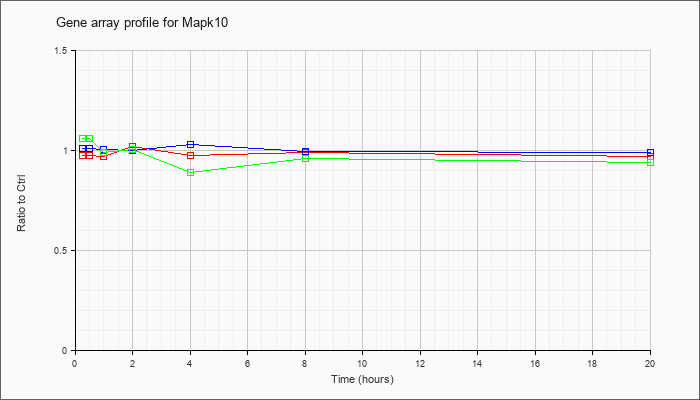 |
AK076990 |
adult male testis cDNA, RIKEN full-length enriched library, clone:4930597C01 product:mitogen activated protein kinase 10, full insert sequence. [AK076990] |
KLA | 1.03 |
1.05 |
.98 |
1.04 |
.94 |
.95 |
1.02 |
| ATP | 1.00 |
.99 |
1.06 |
.97 |
1.01 |
.96 |
.99 |
| KLA/ATP | 1.07 |
1.01 |
.95 |
1.05 |
.82 |
.95 |
.87 |
|
| Mapk10 |  |
NM_009158 |
mitogen-activated protein kinase 10 (Mapk10), transcript variant 1, mRNA [NM_009158] |
KLA | .95 |
1.00 |
.97 |
1.01 |
.99 |
1.01 |
.95 |
| ATP | 1.01 |
1.02 |
.98 |
1.01 |
1.04 |
1.01 |
.99 |
| KLA/ATP | 1.05 |
.99 |
1.01 |
.98 |
.92 |
.97 |
.98 |
|
| Mapk11 | 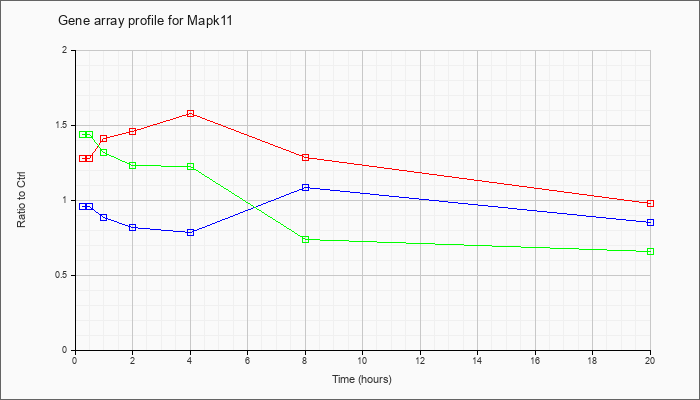 |
NM_011161 |
mitogen-activated protein kinase 11 (Mapk11), mRNA [NM_011161] |
KLA | 1.28 |
1.30 |
1.41 |
1.46 |
1.58 |
1.29 |
.98 |
| ATP | .96 |
.95 |
.89 |
.82 |
.79 |
1.09 |
.85 |
| KLA/ATP | 1.44 |
1.42 |
1.32 |
1.23 |
1.23 |
.74 |
.66 |
|
| Mapk12 | 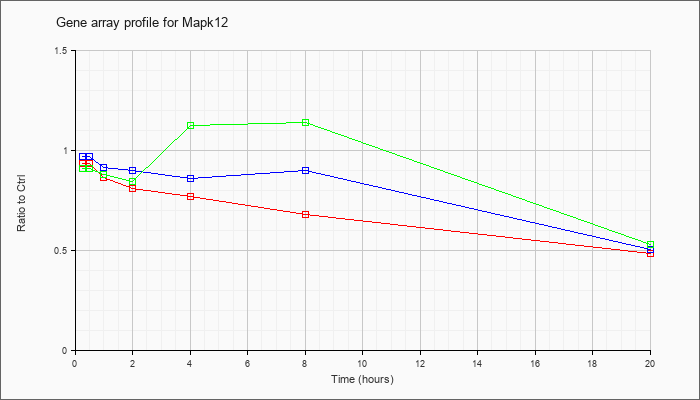 |
NM_013871 |
mitogen-activated protein kinase 12 (Mapk12), mRNA [NM_013871] |
KLA | .94 |
.86 |
.87 |
.81 |
.77 |
.68 |
.49 |
| ATP | .97 |
.83 |
.92 |
.90 |
.86 |
.90 |
.51 |
| KLA/ATP | .91 |
.89 |
.88 |
.85 |
1.12 |
1.14 |
.53 |
|
| Mapk13 | 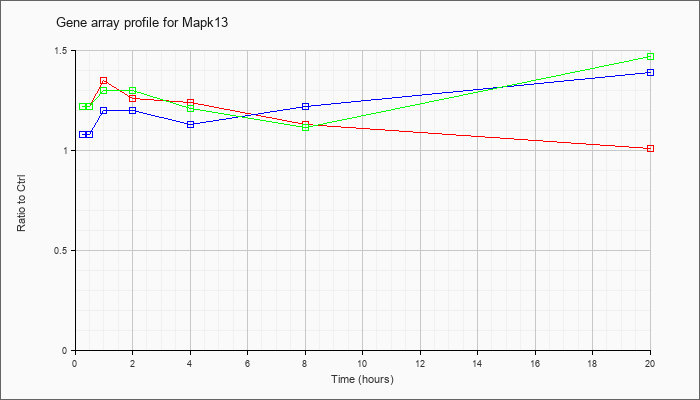 |
NM_011950 |
mitogen-activated protein kinase 13 (Mapk13), mRNA [NM_011950] |
KLA | 1.22 |
1.21 |
1.35 |
1.26 |
1.24 |
1.13 |
1.01 |
| ATP | 1.08 |
1.00 |
1.20 |
1.20 |
1.13 |
1.22 |
1.39 |
| KLA/ATP | 1.22 |
1.27 |
1.30 |
1.30 |
1.21 |
1.11 |
1.47 |
|
| Mapk14 | 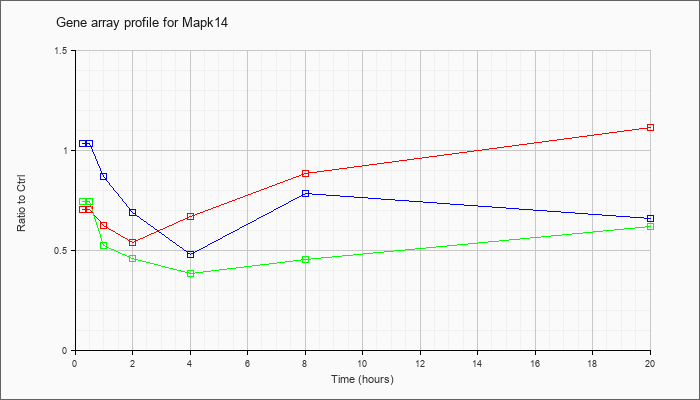 |
NM_011951 |
mitogen-activated protein kinase 14 (Mapk14), mRNA [NM_011951] |
KLA | .70 |
.70 |
.62 |
.54 |
.67 |
.88 |
1.11 |
| ATP | 1.03 |
.98 |
.87 |
.69 |
.48 |
.78 |
.66 |
| KLA/ATP | .74 |
.68 |
.52 |
.46 |
.38 |
.45 |
.62 |
|
| Mapk3 | 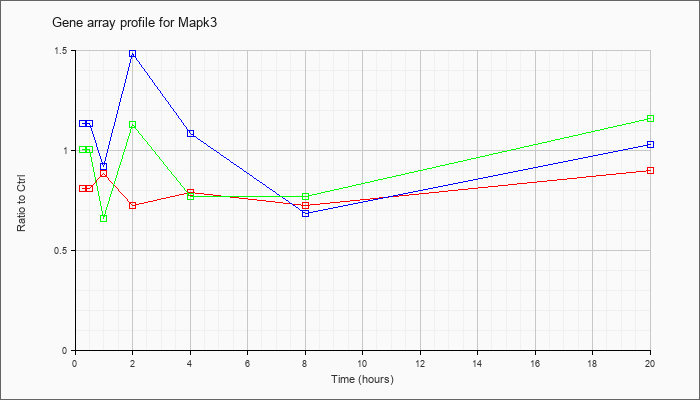 |
NM_011952 |
mitogen-activated protein kinase 3 (Mapk3), mRNA [NM_011952] |
KLA | .81 |
.85 |
.88 |
.72 |
.79 |
.73 |
.90 |
| ATP | 1.13 |
1.30 |
.92 |
1.48 |
1.08 |
.68 |
1.03 |
| KLA/ATP | 1.01 |
1.01 |
.66 |
1.13 |
.77 |
.77 |
1.16 |
|
| Mapk8 |  |
AK162915 |
adult male spinal cord cDNA, RIKEN full-length enriched library, clone:A330031O19 product:mitogen activated protein kinase 8, full insert sequence [AK162915] |
KLA | .94 |
.86 |
.80 |
.83 |
.77 |
.85 |
.83 |
| ATP | 1.08 |
1.12 |
.94 |
.80 |
.83 |
.71 |
.75 |
| KLA/ATP | .83 |
.81 |
.80 |
.74 |
.74 |
.75 |
.67 |
|
| Mapk8 |  |
AK163829 |
16 days embryo head cDNA, RIKEN full-length enriched library, clone:C130070A06 product:mitogen activated protein kinase 8, full insert sequence [AK163829] |
KLA | .65 |
.61 |
.66 |
.70 |
.90 |
1.02 |
1.03 |
| ATP | .99 |
.94 |
1.68 |
1.44 |
1.75 |
1.84 |
1.10 |
| KLA/ATP | .63 |
.67 |
1.11 |
1.00 |
2.01 |
1.68 |
1.15 |
|
| Mapk8 |  |
NM_016700 |
mitogen-activated protein kinase 8 (Mapk8), mRNA [NM_016700] |
KLA | .96 |
1.00 |
.94 |
.96 |
.94 |
.99 |
1.04 |
| ATP | 1.01 |
1.05 |
1.01 |
.99 |
1.03 |
.98 |
1.02 |
| KLA/ATP | .96 |
.97 |
.96 |
.97 |
.93 |
.94 |
.96 |
|
| Mapk9 | 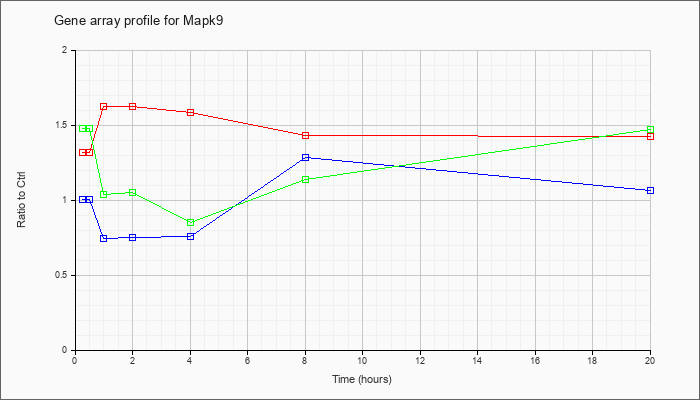 |
NM_016961 |
mitogen-activated protein kinase 9 (Mapk9), transcript variant 2, mRNA [NM_016961] |
KLA | 1.28 |
1.37 |
1.59 |
1.60 |
1.61 |
1.43 |
1.38 |
| ATP | 1.03 |
.99 |
.74 |
.71 |
.79 |
1.20 |
1.01 |
| KLA/ATP | 1.53 |
1.30 |
1.07 |
.99 |
.83 |
1.10 |
1.40 |
|
| Mapk9 |  |
NM_207692 |
mitogen-activated protein kinase 9 (Mapk9), transcript variant 1, mRNA [NM_207692] |
KLA | 1.39 |
1.53 |
1.69 |
1.69 |
1.55 |
1.44 |
1.51 |
| ATP | .95 |
.92 |
.75 |
.85 |
.70 |
1.46 |
1.17 |
| KLA/ATP | 1.37 |
1.35 |
.99 |
1.18 |
.91 |
1.22 |
1.62 |
|
Mm.12688
5 | 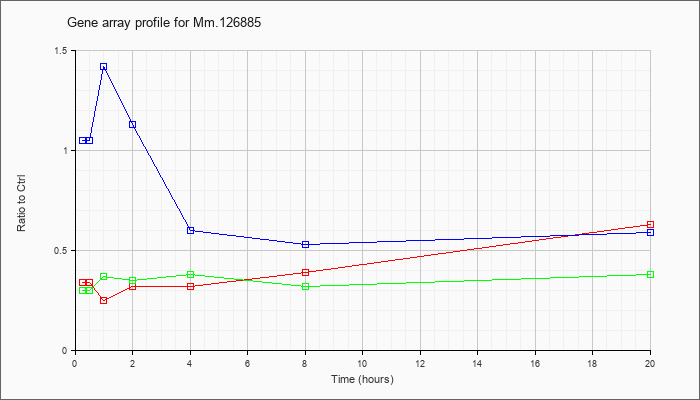 |
34328248 |
Unknown |
KLA | .34 |
.32 |
.25 |
.32 |
.32 |
.39 |
.63 |
| ATP | 1.05 |
1.19 |
1.42 |
1.13 |
.60 |
.53 |
.59 |
| KLA/ATP | .30 |
.27 |
.37 |
.35 |
.38 |
.32 |
.38 |
|
Mm.21312
8 | 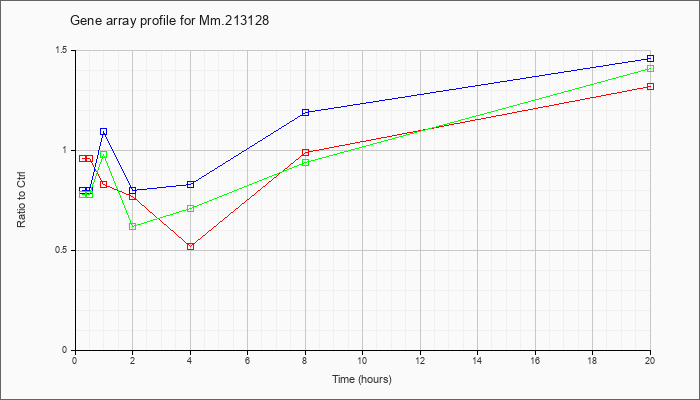 |
141801344 |
Unknown |
KLA | .96 |
.95 |
.83 |
.77 |
.52 |
.99 |
1.32 |
| ATP | .80 |
.82 |
1.09 |
.80 |
.83 |
1.19 |
1.46 |
| KLA/ATP | .78 |
.92 |
.98 |
.62 |
.71 |
.94 |
1.41 |
|
Mm.25381
9 | 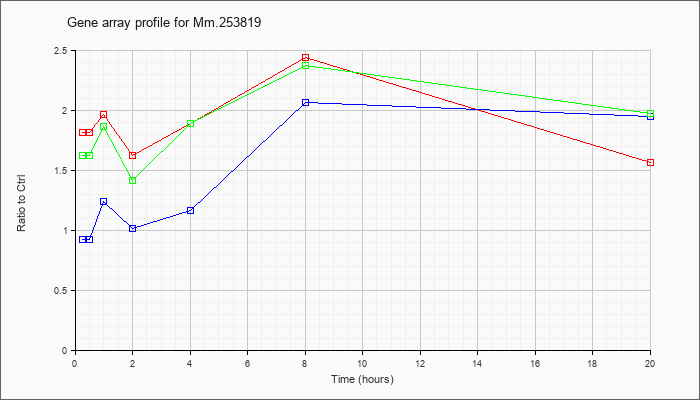 |
118130676 |
Unknown |
KLA | 1.81 |
1.88 |
1.96 |
1.62 |
1.89 |
2.44 |
1.56 |
| ATP | .92 |
.91 |
1.24 |
1.01 |
1.16 |
2.06 |
1.95 |
| KLA/ATP | 1.62 |
1.86 |
1.86 |
1.41 |
1.89 |
2.37 |
1.97 |
|
Mm.27304
9 | 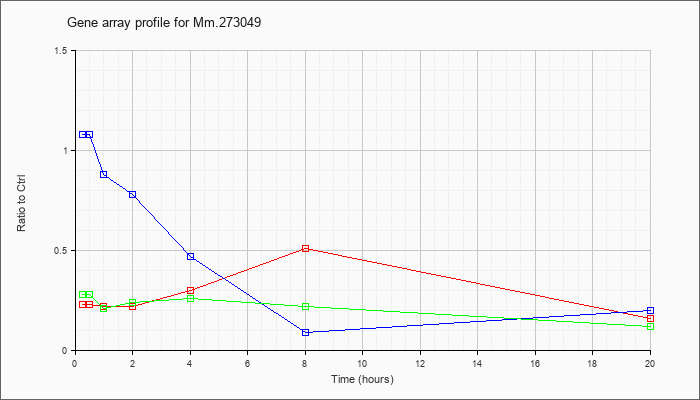 |
119672895 |
Unknown |
KLA | .23 |
.22 |
.22 |
.22 |
.30 |
.51 |
.16 |
| ATP | 1.08 |
1.05 |
.88 |
.78 |
.47 |
.09 |
.20 |
| KLA/ATP | .28 |
.22 |
.21 |
.24 |
.26 |
.22 |
.12 |
|
Mm.27970
| 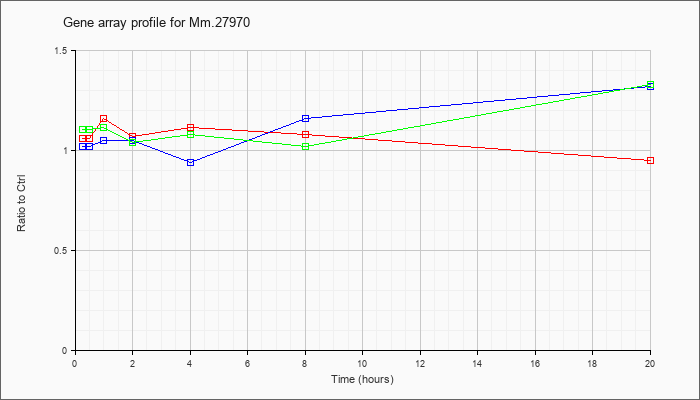 |
13994135 |
Unknown |
KLA | 1.06 |
1.11 |
1.16 |
1.07 |
1.11 |
1.08 |
.95 |
| ATP | 1.02 |
1.09 |
1.05 |
1.05 |
.94 |
1.16 |
1.32 |
| KLA/ATP | 1.10 |
1.19 |
1.11 |
1.04 |
1.08 |
1.02 |
1.33 |
|
| Nfkb1 | 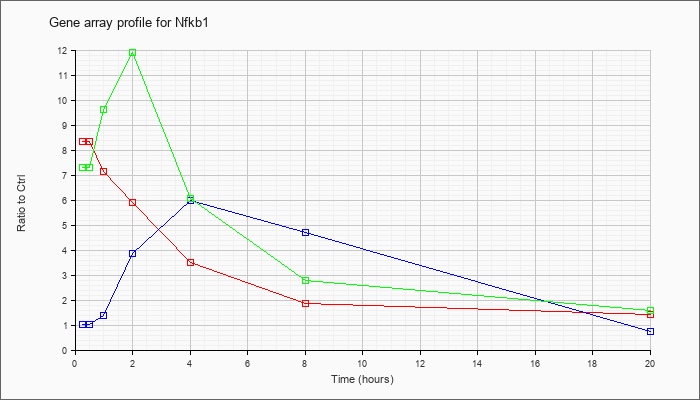 |
BC050841 |
nuclear factor of kappa light chain gene enhancer in B-cells 1, p105, mRNA (cDNA clone IMAGE:6392793), complete cds. [BC050841] |
KLA | 5.81 |
4.96 |
4.77 |
3.21 |
3.37 |
2.00 |
1.71 |
| ATP | 1.39 |
1.52 |
3.25 |
12.63 |
13.64 |
4.80 |
1.08 |
| KLA/ATP | 7.21 |
8.69 |
19.53 |
23.93 |
11.70 |
2.85 |
2.01 |
|
| Nfkb1 |  |
M57999 |
gb|Mouse transcription factor NF-kappa-B DNA binding subunit mRNA, complete cds. [M57999] |
KLA | 8.55 |
7.87 |
7.35 |
6.13 |
3.51 |
1.85 |
1.38 |
| ATP | .98 |
.96 |
1.21 |
3.06 |
5.29 |
4.67 |
.69 |
| KLA/ATP | 7.32 |
7.20 |
8.71 |
10.82 |
5.55 |
2.77 |
1.54 |
|
| Nras | 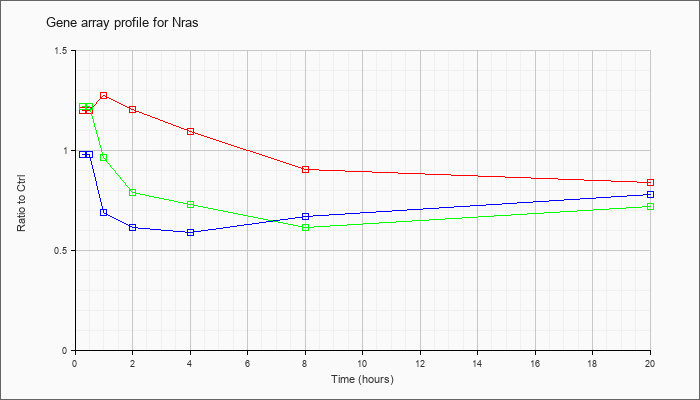 |
NM_010937 |
neuroblastoma ras oncogene (Nras), mRNA [NM_010937] |
KLA | 1.20 |
1.22 |
1.27 |
1.20 |
1.09 |
.91 |
.84 |
| ATP | .98 |
.91 |
.69 |
.61 |
.59 |
.67 |
.78 |
| KLA/ATP | 1.22 |
1.14 |
.97 |
.79 |
.73 |
.61 |
.72 |
|
| Pik3ca | 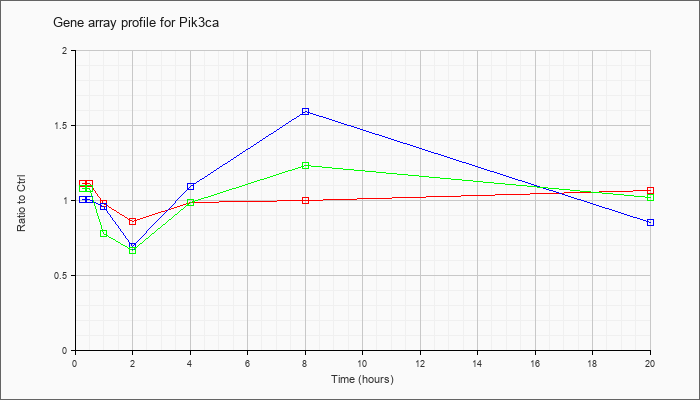 |
ENSMUST00000108243 |
ens|Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha isoform (EC 2.7.1.153) (PI3-kinase p110 subunit alpha) (PtdIns-3- kinase p110) (PI3K). [Source:Uniprot/SWISSPROT;Acc:P42337] [ENSMUST00000108243] |
KLA | 1.07 |
1.01 |
.91 |
.90 |
.98 |
.96 |
.86 |
| ATP | 1.03 |
.87 |
1.12 |
.63 |
1.22 |
1.86 |
.60 |
| KLA/ATP | 1.05 |
.82 |
.91 |
.61 |
1.17 |
1.26 |
.63 |
|
| Pik3ca |  |
NM_008839 |
phosphatidylinositol 3-kinase, catalytic, alpha polypeptide (Pik3ca), mRNA [NM_008839] |
KLA | 1.15 |
1.11 |
1.05 |
.82 |
.99 |
1.03 |
1.27 |
| ATP | .98 |
1.00 |
.79 |
.75 |
.96 |
1.32 |
1.10 |
| KLA/ATP | 1.10 |
.98 |
.64 |
.72 |
.80 |
1.20 |
1.41 |
|
| Pik3cb | 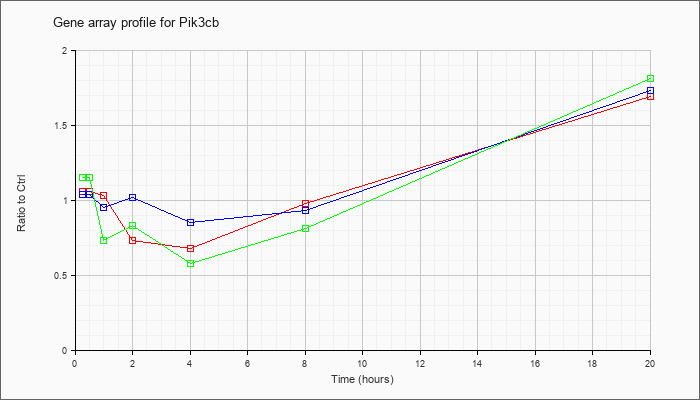 |
NM_029094 |
phosphatidylinositol 3-kinase, catalytic, beta polypeptide (Pik3cb), mRNA [NM_029094] |
KLA | 1.06 |
1.06 |
1.03 |
.73 |
.68 |
.98 |
1.69 |
| ATP | 1.04 |
1.21 |
.95 |
1.02 |
.85 |
.93 |
1.73 |
| KLA/ATP | 1.15 |
1.27 |
.73 |
.83 |
.58 |
.81 |
1.81 |
|
| Pik3cd | 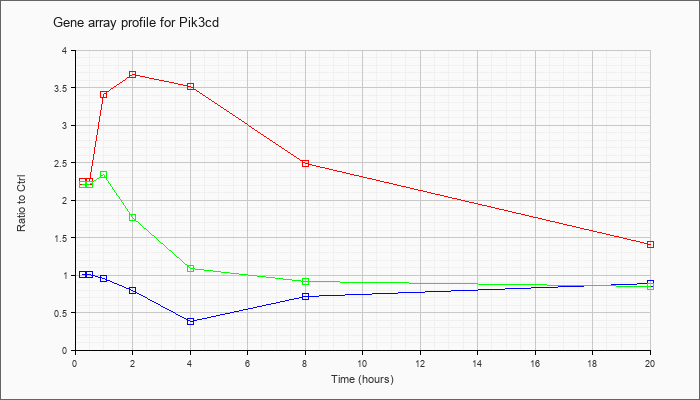 |
NM_008840 |
phosphatidylinositol 3-kinase catalytic delta polypeptide (Pik3cd), transcript variant 1, mRNA [NM_008840] |
KLA | 2.28 |
2.42 |
3.10 |
4.17 |
3.35 |
2.31 |
1.34 |
| ATP | 1.02 |
.91 |
.99 |
.63 |
.30 |
.67 |
.80 |
| KLA/ATP | 2.14 |
2.23 |
2.70 |
1.51 |
1.17 |
.85 |
.72 |
|
| Pik3cd |  |
U86587 |
phosphatidylinositol 3-kinase catalytic subunit p110 delta mRNA, complete cds. [U86587] |
KLA | 2.21 |
2.24 |
3.71 |
3.17 |
3.67 |
2.66 |
1.48 |
| ATP | 1.00 |
1.03 |
.93 |
.96 |
.46 |
.75 |
.97 |
| KLA/ATP | 2.27 |
2.32 |
1.97 |
2.02 |
1.01 |
.99 |
.98 |
|
| Pik3cg | 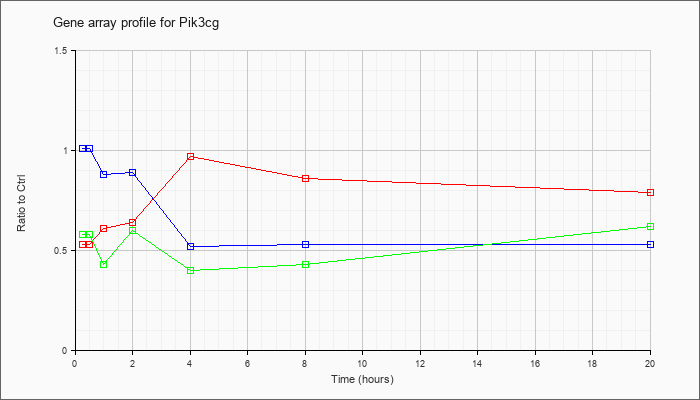 |
NM_020272 |
phosphoinositide-3-kinase, catalytic, gamma polypeptide (Pik3cg), mRNA [NM_020272] |
KLA | .53 |
.53 |
.61 |
.64 |
.97 |
.86 |
.79 |
| ATP | 1.01 |
1.13 |
.88 |
.89 |
.52 |
.53 |
.53 |
| KLA/ATP | .58 |
.53 |
.43 |
.60 |
.40 |
.43 |
.62 |
|
| Pik3r1 | 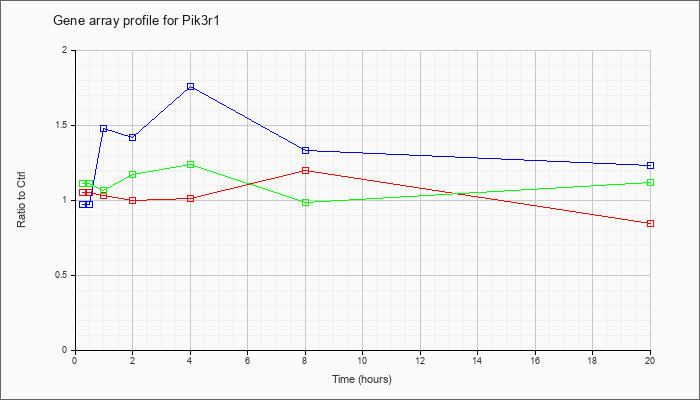 |
NM_001077495 |
phosphatidylinositol 3-kinase, regulatory subunit, polypeptide 1 (p85 alpha) (Pik3r1), transcript variant 2, mRNA [NM_001077495] |
KLA | 1.05 |
1.07 |
1.03 |
1.00 |
1.01 |
1.20 |
.85 |
| ATP | .97 |
1.02 |
1.48 |
1.42 |
1.76 |
1.33 |
1.23 |
| KLA/ATP | 1.11 |
1.12 |
1.06 |
1.17 |
1.24 |
.99 |
1.12 |
|
| Pik3r2 | 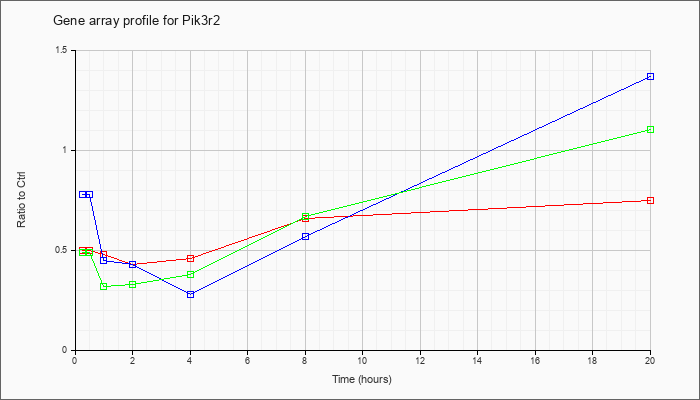 |
NM_008841 |
phosphatidylinositol 3-kinase, regulatory subunit, polypeptide 2 (p85 beta) (Pik3r2), mRNA [NM_008841] |
KLA | .50 |
.50 |
.48 |
.43 |
.46 |
.66 |
.75 |
| ATP | .78 |
.68 |
.45 |
.43 |
.28 |
.57 |
1.37 |
| KLA/ATP | .49 |
.38 |
.32 |
.33 |
.38 |
.67 |
1.10 |
|
| Pik3r3 | 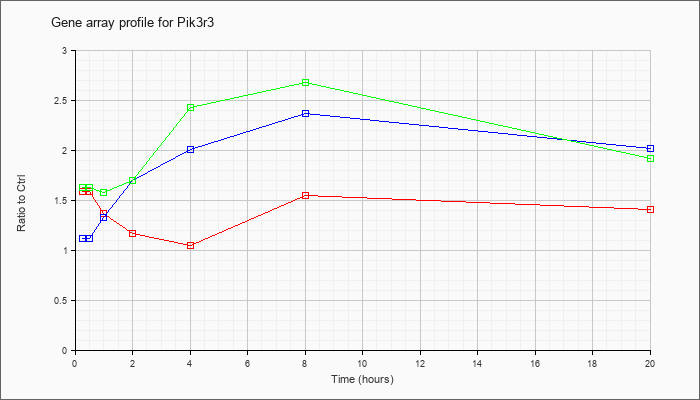 |
NM_181585 |
phosphatidylinositol 3 kinase, regulatory subunit, polypeptide 3 (p55) (Pik3r3), mRNA [NM_181585] |
KLA | 1.59 |
1.49 |
1.37 |
1.17 |
1.05 |
1.55 |
1.41 |
| ATP | 1.12 |
1.00 |
1.33 |
1.70 |
2.01 |
2.37 |
2.02 |
| KLA/ATP | 1.63 |
1.54 |
1.58 |
1.70 |
2.43 |
2.68 |
1.92 |
|
| Pik3r5 | 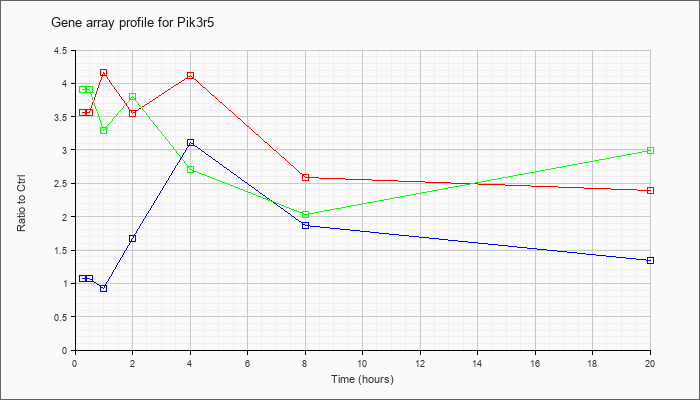 |
NM_177320 |
phosphoinositide-3-kinase, regulatory subunit 5, p101 (Pik3r5), mRNA [NM_177320] |
KLA | 3.57 |
3.58 |
4.16 |
3.54 |
4.11 |
2.59 |
2.39 |
| ATP | 1.08 |
1.08 |
.93 |
1.67 |
3.11 |
1.88 |
1.34 |
| KLA/ATP | 3.91 |
3.70 |
3.30 |
3.80 |
2.70 |
2.04 |
3.00 |
|
| Prl | 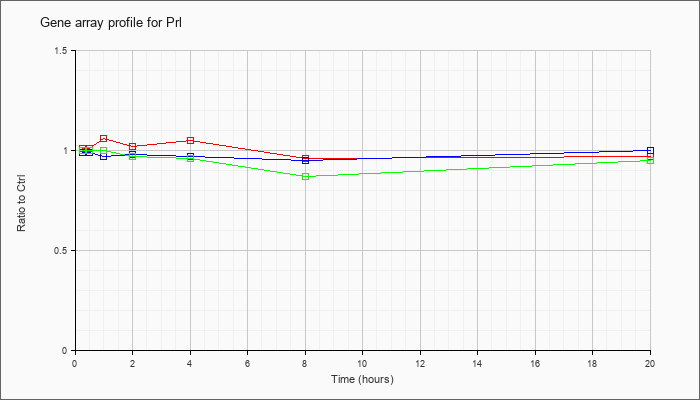 |
NM_011164 |
prolactin (Prl), mRNA [NM_011164] |
KLA | 1.01 |
1.00 |
1.06 |
1.02 |
1.05 |
.96 |
.97 |
| ATP | .99 |
1.02 |
.97 |
.98 |
.97 |
.95 |
1.00 |
| KLA/ATP | 1.00 |
1.01 |
1.00 |
.97 |
.96 |
.87 |
.95 |
|
| Prlr | 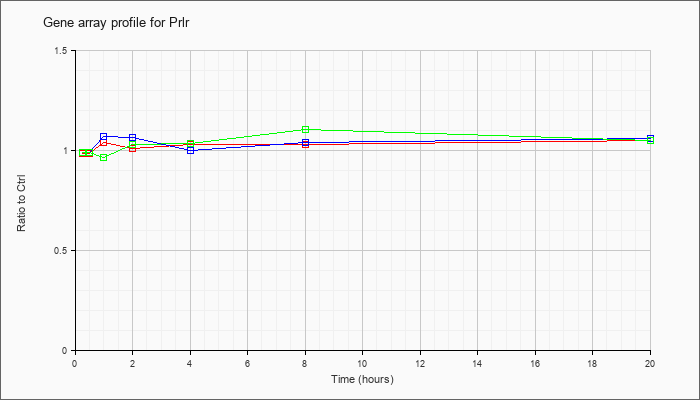 |
AK083198 |
adult male hippocampus cDNA, RIKEN full-length enriched library, clone:C630026G18 product:unclassifiable, full insert sequence [AK083198] |
KLA | .99 |
1.07 |
1.00 |
.95 |
1.01 |
.90 |
1.01 |
| ATP | 1.02 |
.94 |
.97 |
1.00 |
.97 |
.94 |
.96 |
| KLA/ATP | .98 |
.94 |
.96 |
.96 |
.99 |
1.07 |
1.02 |
|
| Prlr |  |
AK087727 |
2 days pregnant adult female ovary cDNA, RIKEN full-length enriched library, clone:E330012C04 product:hypothetical protein, full insert sequence [AK087727] |
KLA | 1.03 |
1.00 |
.99 |
1.01 |
1.02 |
1.04 |
.99 |
| ATP | 1.00 |
.95 |
1.13 |
1.06 |
.97 |
.98 |
1.04 |
| KLA/ATP | .97 |
1.01 |
.88 |
1.02 |
1.01 |
1.03 |
1.05 |
|
| Prlr |  |
NM_011169 |
prolactin receptor (Prlr), mRNA [NM_011169] |
KLA | .93 |
1.03 |
1.12 |
1.06 |
1.05 |
1.14 |
1.15 |
| ATP | .95 |
.99 |
1.11 |
1.13 |
1.06 |
1.19 |
1.18 |
| KLA/ATP | 1.02 |
1.01 |
1.05 |
1.10 |
1.10 |
1.21 |
1.07 |
|
| Raf1 | 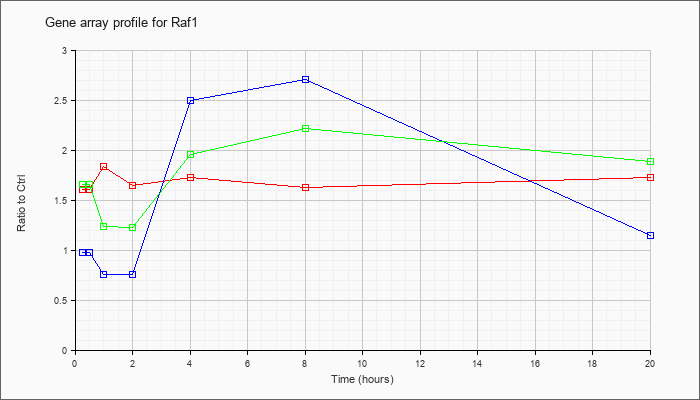 |
AK036317 |
16 days neonate cerebellum cDNA, RIKEN full-length enriched library, clone:9630056E02 product:v-raf-1 leukemia viral oncogene 1, full insert sequence. [AK036317] |
KLA | 1.47 |
1.34 |
1.55 |
1.29 |
1.59 |
1.32 |
1.46 |
| ATP | 1.05 |
.98 |
.54 |
.63 |
3.32 |
2.07 |
1.01 |
| KLA/ATP | 1.57 |
1.33 |
.79 |
.96 |
2.26 |
1.38 |
1.35 |
|
| Raf1 |  |
NM_029780 |
v-raf-leukemia viral oncogene 1 (Raf1), mRNA [NM_029780] |
KLA | 1.65 |
1.64 |
1.93 |
1.77 |
1.77 |
1.72 |
1.81 |
| ATP | .95 |
.96 |
.83 |
.80 |
2.22 |
2.92 |
1.19 |
| KLA/ATP | 1.68 |
1.64 |
1.39 |
1.32 |
1.85 |
2.50 |
2.06 |
|
| Rela | 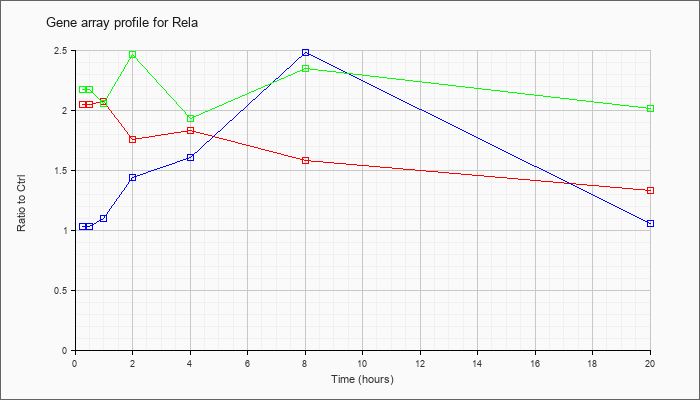 |
NM_009045 |
v-rel reticuloendotheliosis viral oncogene homolog A (avian) (Rela), mRNA [NM_009045] |
KLA | 2.05 |
2.05 |
2.07 |
1.75 |
1.83 |
1.58 |
1.33 |
| ATP | 1.03 |
1.08 |
1.10 |
1.44 |
1.61 |
2.48 |
1.05 |
| KLA/ATP | 2.17 |
2.27 |
2.05 |
2.47 |
1.93 |
2.35 |
2.02 |
|
| Shc1 | 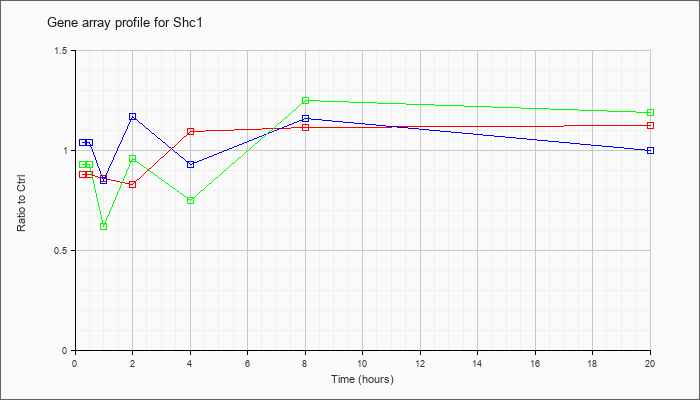 |
NM_011368 |
src homology 2 domain-containing transforming protein C1 (Shc1), transcript variant 2, mRNA [NM_011368] |
KLA | .88 |
.87 |
.86 |
.83 |
1.09 |
1.11 |
1.12 |
| ATP | 1.04 |
1.25 |
.85 |
1.17 |
.93 |
1.16 |
1.00 |
| KLA/ATP | .93 |
.96 |
.62 |
.96 |
.75 |
1.25 |
1.19 |
|
| Shc2 | 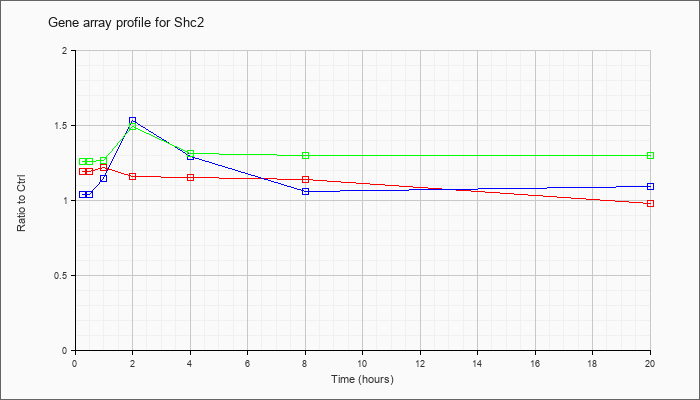 |
NM_001024539 |
src homology 2 domain-containing transforming protein C2 (Shc2), mRNA [NM_001024539] |
KLA | 1.19 |
1.25 |
1.22 |
1.16 |
1.15 |
1.14 |
.98 |
| ATP | 1.04 |
1.14 |
1.15 |
1.53 |
1.29 |
1.06 |
1.09 |
| KLA/ATP | 1.26 |
1.24 |
1.26 |
1.49 |
1.31 |
1.30 |
1.30 |
|
| Shc3 | 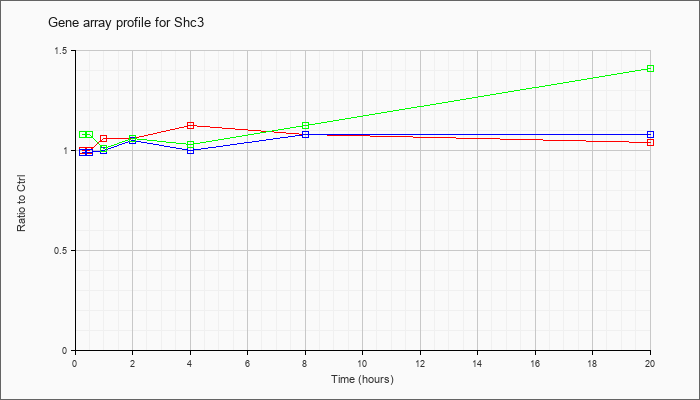 |
NM_009167 |
src homology 2 domain-containing transforming protein C3 (Shc3), mRNA [NM_009167] |
KLA | 1.00 |
.98 |
1.06 |
1.06 |
1.12 |
1.08 |
1.04 |
| ATP | .99 |
1.02 |
1.00 |
1.05 |
1.00 |
1.08 |
1.08 |
| KLA/ATP | 1.08 |
1.06 |
1.01 |
1.06 |
1.03 |
1.12 |
1.41 |
|
| Shc4 | 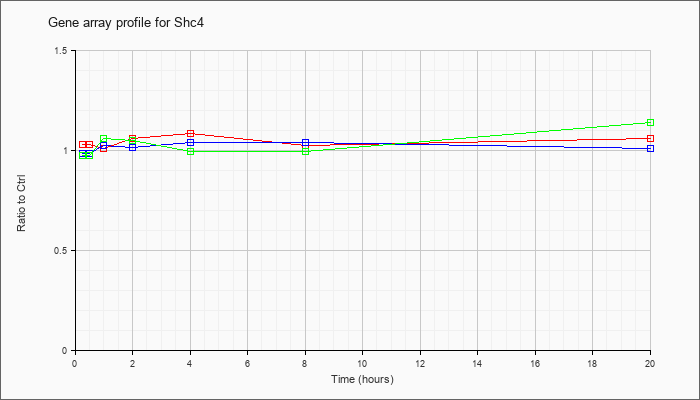 |
NM_199022 |
SHC (Src homology 2 domain containing) family, member 4 (Shc4), mRNA [NM_199022] |
KLA | 1.03 |
1.03 |
1.01 |
1.06 |
1.09 |
1.03 |
1.06 |
| ATP | .99 |
1.00 |
1.03 |
1.02 |
1.04 |
1.04 |
1.01 |
| KLA/ATP | .98 |
.97 |
1.06 |
1.05 |
1.00 |
1.00 |
1.14 |
|
| Slc2a2 | 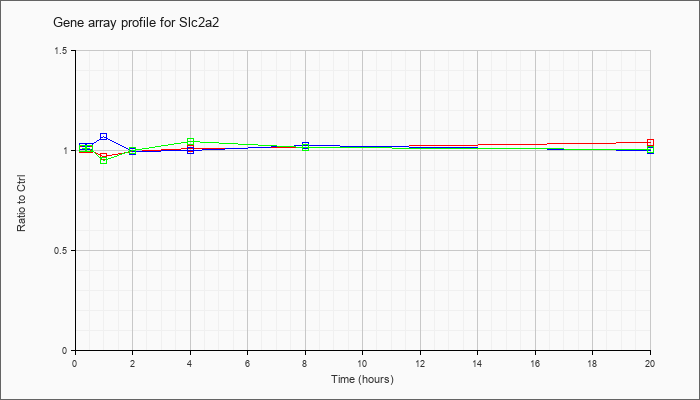 |
NM_031197 |
solute carrier family 2 (facilitated glucose transporter), member 2 (Slc2a2), mRNA [NM_031197] |
KLA | 1.01 |
1.03 |
.97 |
1.00 |
1.01 |
1.02 |
1.04 |
| ATP | 1.02 |
.97 |
1.07 |
1.00 |
1.00 |
1.03 |
1.00 |
| KLA/ATP | 1.01 |
1.01 |
.95 |
1.00 |
1.05 |
1.02 |
1.01 |
|
| Socs1 | 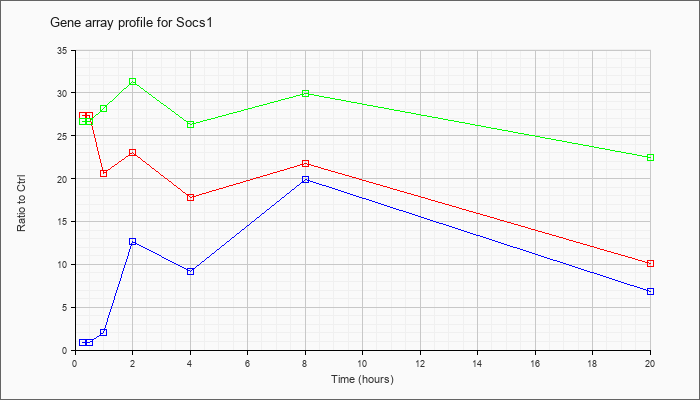 |
NM_009896 |
suppressor of cytokine signaling 1 (Socs1), mRNA [NM_009896] |
KLA | 27.36 |
22.16 |
20.64 |
23.08 |
17.74 |
21.71 |
10.12 |
| ATP | .91 |
.72 |
2.03 |
12.69 |
9.16 |
19.87 |
6.80 |
| KLA/ATP | 26.70 |
24.56 |
28.15 |
31.28 |
26.32 |
29.95 |
22.51 |
|
| Socs2 | 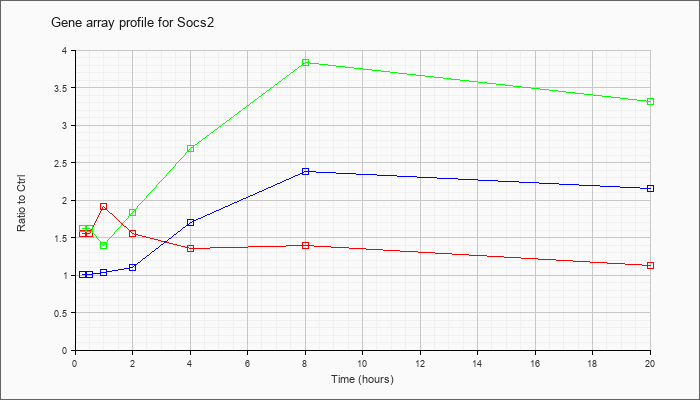 |
AK033206 |
15 days embryo male testis cDNA, RIKEN full-length enriched library, clone:8030460M17 product:hypothetical SH2 motif [AK033206] |
KLA | 1.08 |
1.11 |
1.22 |
1.13 |
1.11 |
1.08 |
.97 |
| ATP | .98 |
.97 |
1.03 |
1.03 |
1.29 |
1.30 |
1.29 |
| KLA/ATP | 1.06 |
1.09 |
1.07 |
1.33 |
2.01 |
1.85 |
1.98 |
|
| Socs2 |  |
NM_007706 |
suppressor of cytokine signaling 2 (Socs2), mRNA [NM_007706] |
KLA | 2.03 |
2.28 |
2.61 |
1.98 |
1.59 |
1.72 |
1.28 |
| ATP | 1.04 |
1.04 |
1.05 |
1.17 |
2.11 |
3.47 |
3.02 |
| KLA/ATP | 2.18 |
1.92 |
1.71 |
2.33 |
3.37 |
5.81 |
4.64 |
|
| Socs3 | 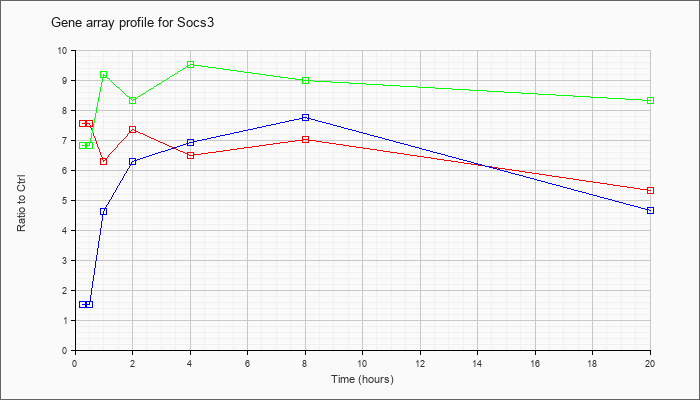 |
NM_007707 |
suppressor of cytokine signaling 3 (Socs3), mRNA [NM_007707] |
KLA | 7.56 |
6.68 |
6.29 |
7.36 |
6.47 |
7.03 |
5.33 |
| ATP | 1.51 |
1.75 |
4.63 |
6.27 |
6.92 |
7.74 |
4.64 |
| KLA/ATP | 6.82 |
6.77 |
9.19 |
8.32 |
9.51 |
8.98 |
8.33 |
|
| Socs4 | 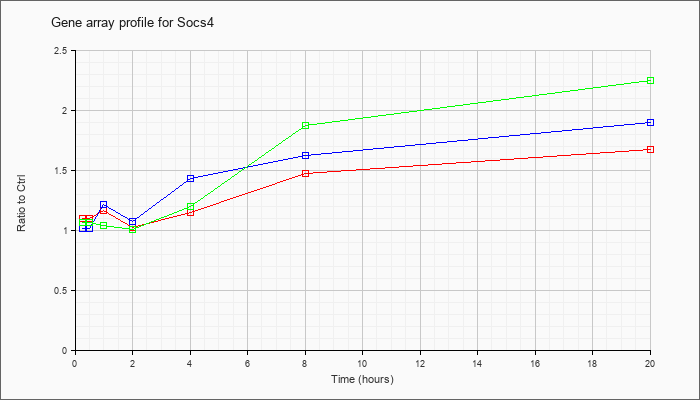 |
AK080408 |
7 days neonate cerebellum cDNA, RIKEN full-length enriched library, clone:A730004F22 product:unclassifiable, full insert sequence [AK080408] |
KLA | .89 |
.90 |
.87 |
.74 |
.87 |
1.05 |
.98 |
| ATP | 1.01 |
.97 |
1.11 |
.76 |
1.00 |
.82 |
1.11 |
| KLA/ATP | .80 |
.82 |
.82 |
.70 |
1.00 |
.80 |
.76 |
|
| Socs4 |  |
NM_080843 |
suppressor of cytokine signaling 4 (Socs4), mRNA [NM_080843] |
KLA | 1.31 |
1.29 |
1.45 |
1.31 |
1.42 |
1.90 |
2.37 |
| ATP | 1.02 |
1.16 |
1.31 |
1.39 |
1.86 |
2.43 |
2.69 |
| KLA/ATP | 1.33 |
1.24 |
1.26 |
1.31 |
1.39 |
2.94 |
3.73 |
|
| Socs5 | 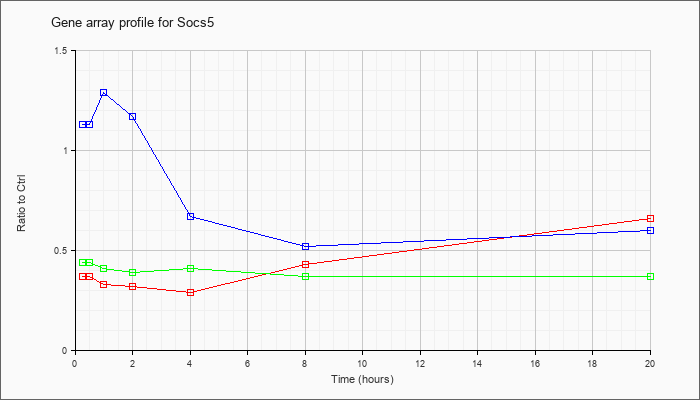 |
NM_019654 |
suppressor of cytokine signaling 5 (Socs5), mRNA [NM_019654] |
KLA | .37 |
.34 |
.33 |
.32 |
.29 |
.43 |
.66 |
| ATP | 1.13 |
1.24 |
1.29 |
1.17 |
.67 |
.52 |
.60 |
| KLA/ATP | .44 |
.37 |
.41 |
.39 |
.41 |
.37 |
.37 |
|
| Socs6 | 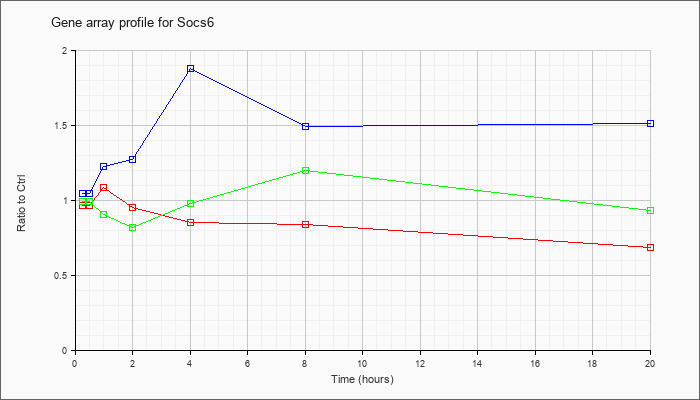 |
AK142069 |
12 days embryo eyeball cDNA, RIKEN full-length enriched library, clone:D230020G23 product:suppressor of cytokine signaling 6, full insert sequence [AK142069] |
KLA | .89 |
.96 |
.95 |
.91 |
.82 |
.89 |
.83 |
| ATP | 1.11 |
1.04 |
.91 |
.88 |
1.06 |
.84 |
1.24 |
| KLA/ATP | 1.08 |
.94 |
.71 |
.80 |
.76 |
.75 |
.74 |
|
| Socs6 |  |
NM_018821 |
suppressor of cytokine signaling 6 (Socs6), mRNA [NM_018821] |
KLA | 1.00 |
.99 |
1.16 |
.98 |
.87 |
.81 |
.61 |
| ATP | 1.02 |
1.10 |
1.38 |
1.47 |
2.29 |
1.81 |
1.65 |
| KLA/ATP | .95 |
.96 |
1.00 |
.83 |
1.09 |
1.42 |
1.03 |
|
| Socs7 | 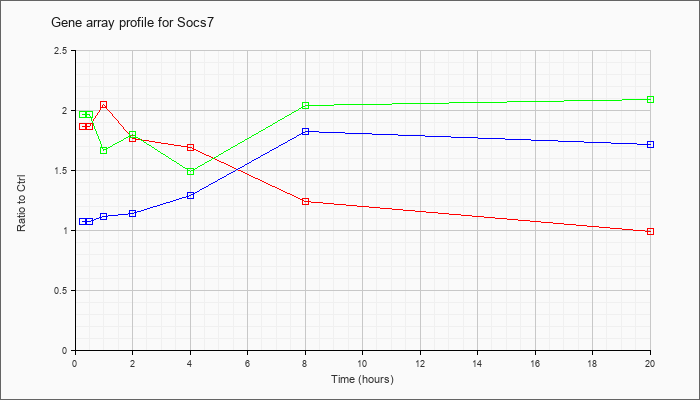 |
NM_138657 |
suppressor of cytokine signaling 7 (Socs7), mRNA [NM_138657] |
KLA | 1.86 |
1.79 |
2.05 |
1.76 |
1.69 |
1.24 |
.99 |
| ATP | 1.07 |
1.08 |
1.11 |
1.14 |
1.29 |
1.82 |
1.71 |
| KLA/ATP | 1.96 |
1.99 |
1.66 |
1.80 |
1.49 |
2.04 |
2.09 |
|
| Sos1 | 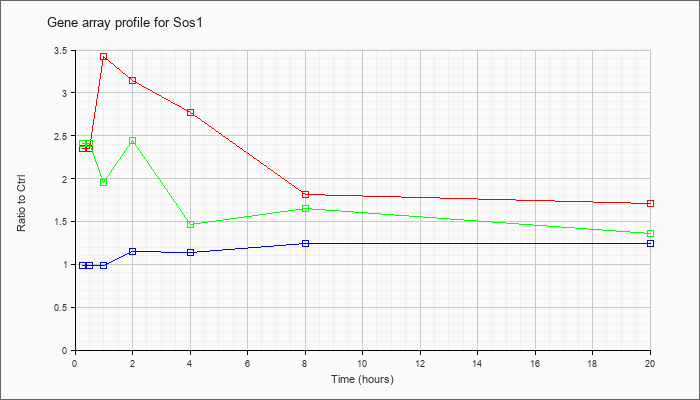 |
NM_009231 |
Son of sevenless homolog 1 (Drosophila) (Sos1), mRNA [NM_009231] |
KLA | 2.36 |
2.40 |
3.42 |
3.15 |
2.78 |
1.81 |
1.71 |
| ATP | .99 |
1.00 |
.99 |
1.15 |
1.14 |
1.25 |
1.24 |
| KLA/ATP | 2.41 |
2.26 |
1.95 |
2.44 |
1.47 |
1.66 |
1.37 |
|
| Sos2 | 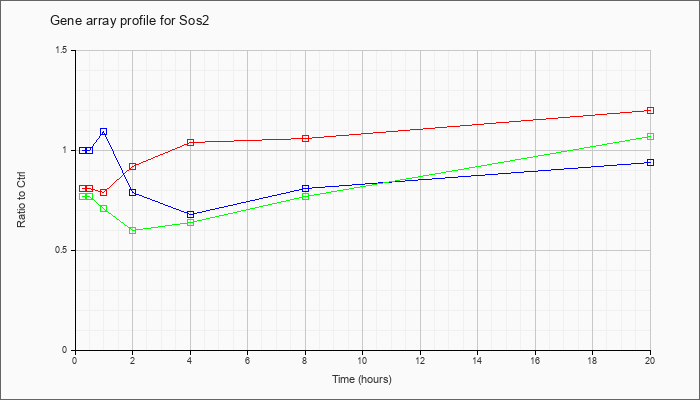 |
XM_127051 |
gb|PREDICTED: Mus musculus Son of sevenless homolog 2 (Drosophila) (Sos2), mRNA [XM_127051] |
KLA | .81 |
.77 |
.79 |
.92 |
1.04 |
1.06 |
1.20 |
| ATP | 1.00 |
1.00 |
1.09 |
.79 |
.68 |
.81 |
.94 |
| KLA/ATP | .77 |
.69 |
.71 |
.60 |
.64 |
.77 |
1.07 |
|
| Src | 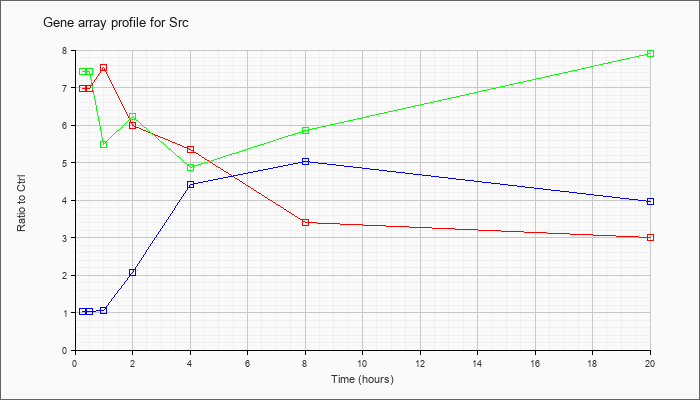 |
NM_009271 |
Rous sarcoma oncogene (Src), transcript variant 1, mRNA [NM_009271] |
KLA | 6.99 |
7.23 |
7.54 |
5.99 |
5.34 |
3.39 |
3.00 |
| ATP | 1.02 |
1.06 |
1.06 |
2.08 |
4.41 |
5.03 |
3.97 |
| KLA/ATP | 7.44 |
7.51 |
5.48 |
6.24 |
4.86 |
5.86 |
7.90 |
|
| Stat1 |  |
NM_009283 |
signal transducer and activator of transcription 1 (Stat1), mRNA [NM_009283] |
KLA | 2.90 |
2.72 |
3.00 |
3.64 |
4.58 |
6.51 |
5.58 |
| ATP | 1.01 |
.92 |
.89 |
.72 |
.51 |
2.22 |
3.07 |
| KLA/ATP | 2.73 |
2.65 |
2.34 |
1.95 |
1.99 |
4.70 |
8.66 |
|
| Stat3 | 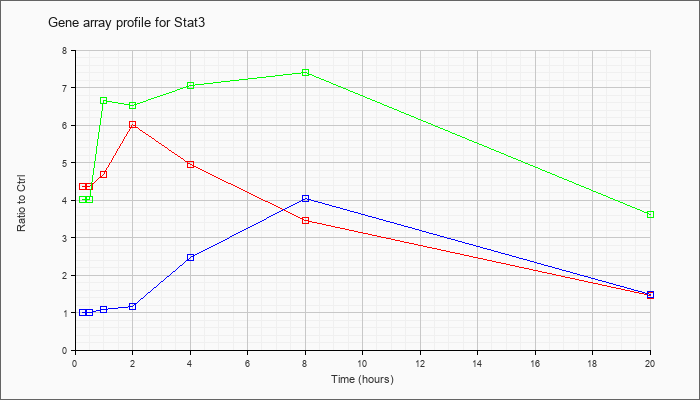 |
NM_213659 |
signal transducer and activator of transcription 3 (Stat3), transcript variant 1, mRNA [NM_213659] |
KLA | 4.36 |
4.42 |
4.67 |
6.02 |
4.95 |
3.45 |
1.46 |
| ATP | 1.01 |
.97 |
1.07 |
1.17 |
2.47 |
4.04 |
1.47 |
| KLA/ATP | 4.02 |
4.57 |
6.65 |
6.51 |
7.06 |
7.39 |
3.61 |
|
| Stat5a | 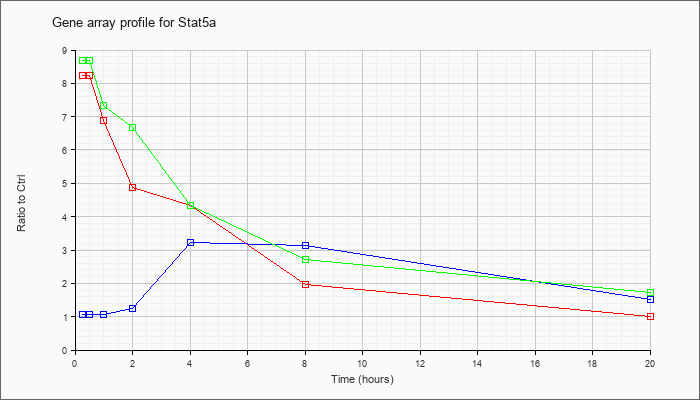 |
NM_011488 |
signal transducer and activator of transcription 5A (Stat5a), mRNA [NM_011488] |
KLA | 8.23 |
8.24 |
6.89 |
4.88 |
4.35 |
1.98 |
1.01 |
| ATP | 1.07 |
1.14 |
1.07 |
1.26 |
3.23 |
3.14 |
1.52 |
| KLA/ATP | 8.70 |
9.53 |
7.33 |
6.68 |
4.32 |
2.73 |
1.72 |
|
| Stat5b | 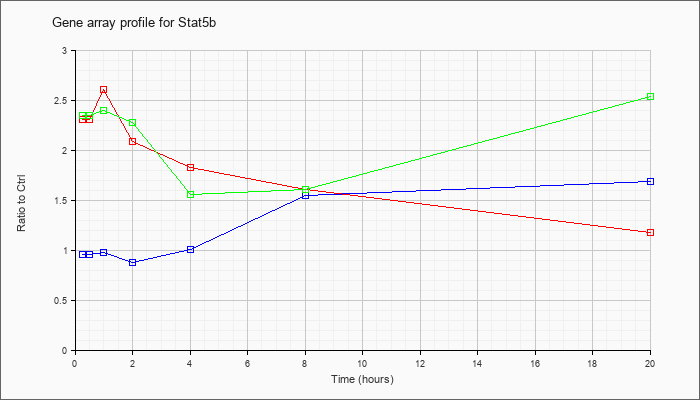 |
AK137889 |
16 days neonate thymus cDNA, RIKEN full-length enriched library, clone:A130035C01 product:unclassifiable, full insert sequence. [AK137889] |
KLA | 1.10 |
1.01 |
.98 |
.90 |
1.03 |
.93 |
.88 |
| ATP | .90 |
.87 |
.61 |
.69 |
1.01 |
.97 |
.89 |
| KLA/ATP | 1.12 |
1.00 |
.87 |
.85 |
.95 |
1.07 |
.83 |
|
| Stat5b |  |
NM_011489 |
signal transducer and activator of transcription 5B (Stat5b), transcript variant 1, mRNA [NM_011489] |
KLA | 2.91 |
2.81 |
3.42 |
2.69 |
2.23 |
1.95 |
1.32 |
| ATP | .99 |
1.04 |
1.16 |
.97 |
1.00 |
1.84 |
2.09 |
| KLA/ATP | 2.96 |
3.05 |
3.17 |
2.99 |
1.86 |
1.87 |
3.40 |
|
| Th | 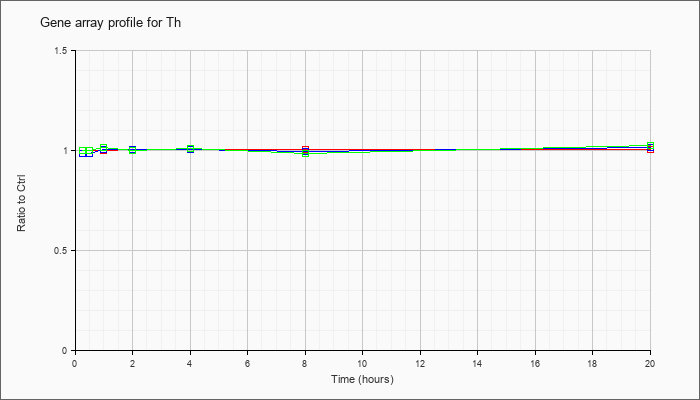 |
NM_009377 |
tyrosine hydroxylase (Th), mRNA [NM_009377] |
KLA | 1.00 |
1.01 |
1.00 |
1.00 |
1.00 |
1.00 |
1.00 |
| ATP | .98 |
1.01 |
1.00 |
1.00 |
1.00 |
.99 |
1.01 |
| KLA/ATP | 1.00 |
1.01 |
1.01 |
1.00 |
1.01 |
.98 |
1.02 |
|
Tnfrsf11
a | 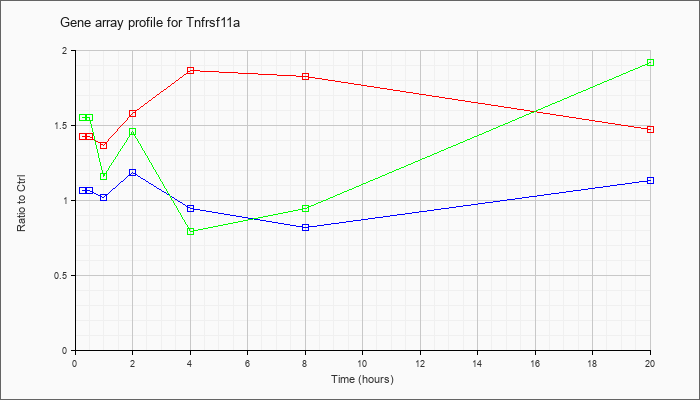 |
NM_009399 |
tumor necrosis factor receptor superfamily, member 11a (Tnfrsf11a), mRNA [NM_009399] |
KLA | 1.43 |
1.33 |
1.36 |
1.58 |
1.87 |
1.83 |
1.47 |
| ATP | 1.07 |
1.17 |
1.02 |
1.19 |
.95 |
.82 |
1.13 |
| KLA/ATP | 1.55 |
1.46 |
1.16 |
1.46 |
.79 |
.95 |
1.92 |
|
| Tnfsf11 | 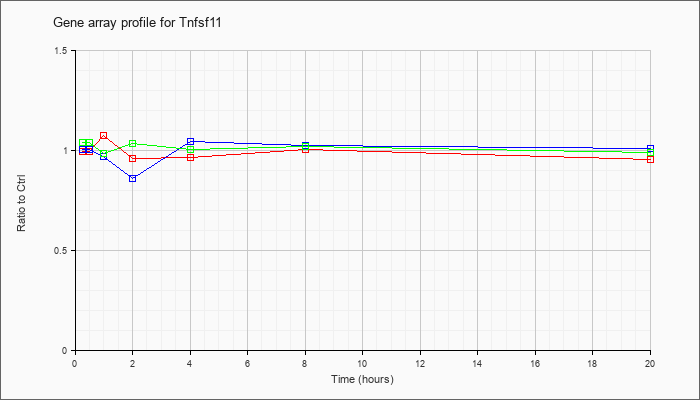 |
NM_011613 |
tumor necrosis factor (ligand) superfamily, member 11 (Tnfsf11), mRNA [NM_011613] |
KLA | 1.00 |
1.00 |
1.08 |
.96 |
.97 |
1.01 |
.96 |
| ATP | 1.01 |
1.05 |
.97 |
.86 |
1.05 |
1.03 |
1.01 |
| KLA/ATP | 1.04 |
1.05 |
.99 |
1.04 |
1.01 |
1.02 |
.99 |
|
| Wap | 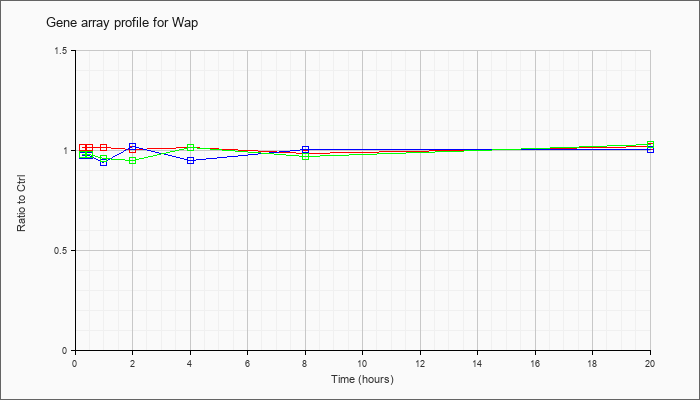 |
NM_011709 |
whey acidic protein (Wap), mRNA [NM_011709] |
KLA | 1.02 |
.97 |
1.02 |
1.01 |
1.02 |
.99 |
1.02 |
| ATP | .98 |
.98 |
.94 |
1.02 |
.95 |
1.01 |
1.01 |
| KLA/ATP | .98 |
.98 |
.96 |
.95 |
1.02 |
.97 |
1.03 |
|