| Gene symbol |
Time course plot |
Accession |
Description |
Treatment |
15min |
30min |
1hr |
2hr |
4hr |
8hr |
20hr |
| Abcc8 | 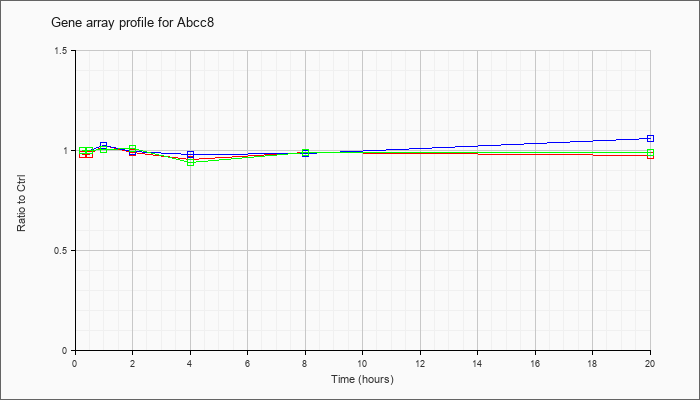 |
NM_011510 |
ATP-binding cassette, sub-family C (CFTR/MRP), member 8 (Abcc8), mRNA [NM_011510] |
KLA | .98 |
1.00 |
1.03 |
.99 |
.96 |
.99 |
.98 |
| ATP | 1.00 |
.97 |
1.03 |
1.00 |
.98 |
.99 |
1.06 |
| KLA/ATP | 1.00 |
1.01 |
1.01 |
1.01 |
.94 |
.99 |
.99 |
|
| Adipoq | 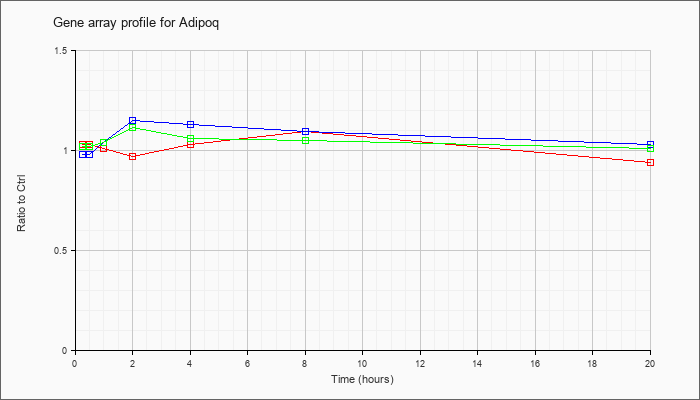 |
NM_009605 |
adiponectin, C1Q and collagen domain containing (Adipoq), mRNA [NM_009605] |
KLA | 1.03 |
.99 |
1.01 |
.97 |
1.03 |
1.09 |
.94 |
| ATP | .98 |
.98 |
1.04 |
1.15 |
1.13 |
1.09 |
1.03 |
| KLA/ATP | 1.02 |
.99 |
1.04 |
1.11 |
1.06 |
1.05 |
1.01 |
|
| Cacna1a | 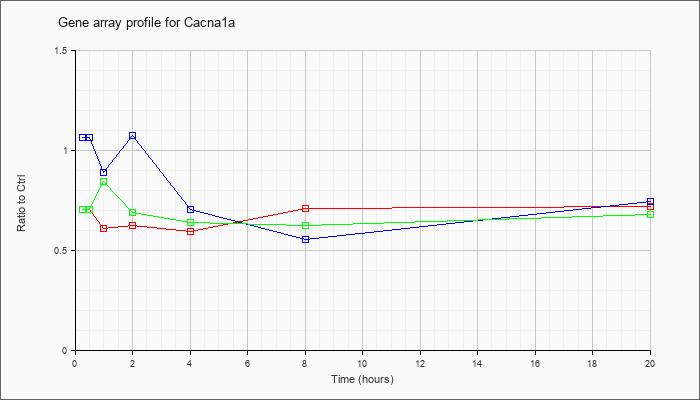 |
NM_007578 |
calcium channel, voltage-dependent, P/Q type, alpha 1A subunit (Cacna1a), mRNA [NM_007578] |
KLA | .71 |
.66 |
.61 |
.63 |
.60 |
.71 |
.72 |
| ATP | 1.07 |
1.04 |
.89 |
1.08 |
.71 |
.56 |
.75 |
| KLA/ATP | .71 |
.72 |
.85 |
.69 |
.64 |
.63 |
.68 |
|
| Cacna1b | 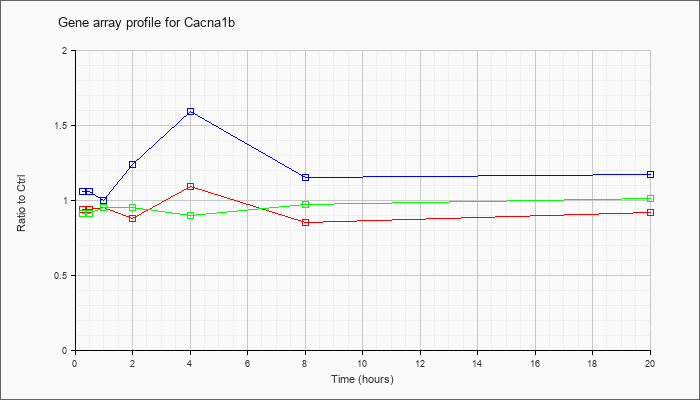 |
U04999 |
N-type calcium channel alpha1 subunit mRNA, complete cds [U04999] |
KLA | .94 |
.94 |
.95 |
.88 |
1.09 |
.85 |
.92 |
| ATP | 1.06 |
1.17 |
1.00 |
1.24 |
1.59 |
1.15 |
1.17 |
| KLA/ATP | .91 |
.91 |
.95 |
.95 |
.90 |
.97 |
1.01 |
|
| Cacna1d | 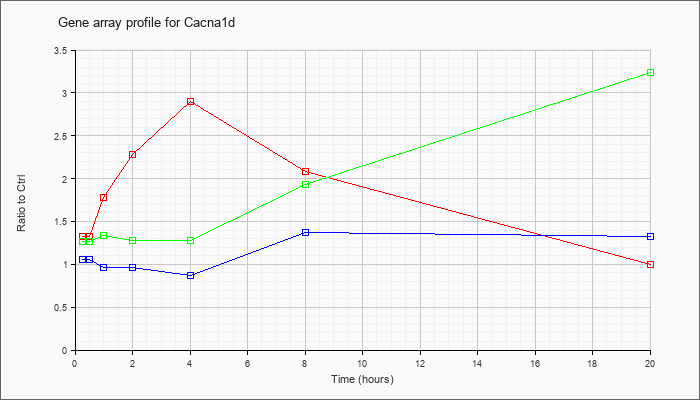 |
NM_001083616 |
calcium channel, voltage-dependent, L type, alpha 1D subunit (Cacna1d), transcript variant 2, mRNA [NM_001083616] |
KLA | 1.57 |
1.58 |
2.50 |
3.62 |
4.55 |
2.93 |
.99 |
| ATP | 1.12 |
.99 |
.97 |
.94 |
.75 |
1.71 |
1.48 |
| KLA/ATP | 1.47 |
1.51 |
1.82 |
1.56 |
1.67 |
2.93 |
4.60 |
|
| Cacna1d |  |
NM_028981 |
calcium channel, voltage-dependent, L type, alpha 1D subunit (Cacna1d), transcript variant 1, mRNA [NM_028981] |
KLA | 1.20 |
1.26 |
1.43 |
1.62 |
2.07 |
1.66 |
1.00 |
| ATP | 1.03 |
1.03 |
.97 |
.97 |
.93 |
1.20 |
1.26 |
| KLA/ATP | 1.16 |
1.12 |
1.10 |
1.14 |
1.08 |
1.44 |
2.56 |
|
| Cacna1e | 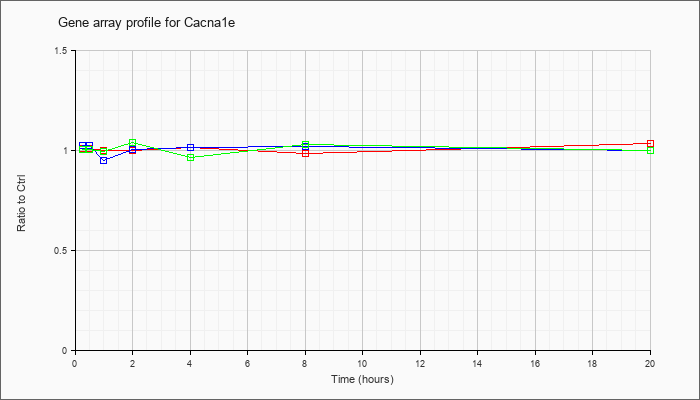 |
NM_009782 |
calcium channel, voltage-dependent, R type, alpha 1E subunit (Cacna1e), mRNA [NM_009782] |
KLA | 1.01 |
.96 |
1.00 |
1.00 |
1.02 |
.99 |
1.04 |
| ATP | 1.03 |
.96 |
.95 |
1.01 |
1.02 |
1.02 |
1.00 |
| KLA/ATP | 1.01 |
1.01 |
1.00 |
1.04 |
.97 |
1.03 |
1.00 |
|
| Cacna1g | 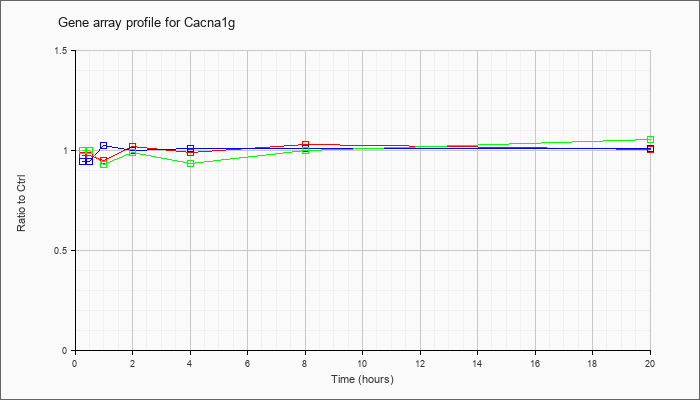 |
NM_009783 |
calcium channel, voltage-dependent, T type, alpha 1G subunit (Cacna1g), transcript variant 1, mRNA [NM_009783] |
KLA | .98 |
.96 |
.95 |
1.02 |
.99 |
1.03 |
1.01 |
| ATP | .95 |
.98 |
1.03 |
1.00 |
1.01 |
1.01 |
1.01 |
| KLA/ATP | 1.00 |
.97 |
.93 |
.99 |
.94 |
1.00 |
1.06 |
|
| Frap1 | 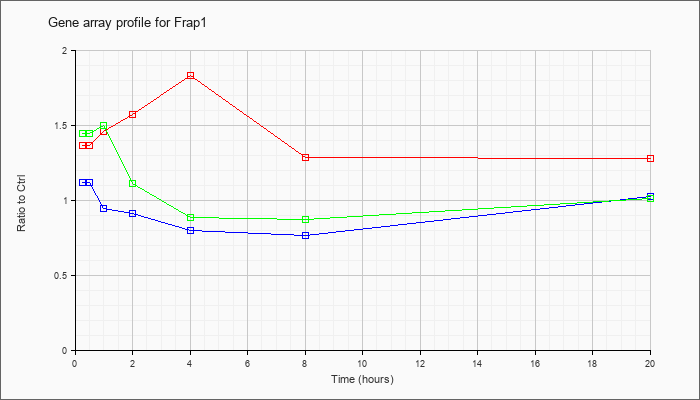 |
NM_020009 |
FK506 binding protein 12-rapamycin associated protein 1 (Frap1), mRNA [NM_020009] |
KLA | 1.37 |
1.32 |
1.46 |
1.57 |
1.83 |
1.29 |
1.28 |
| ATP | 1.12 |
1.07 |
.95 |
.91 |
.80 |
.77 |
1.03 |
| KLA/ATP | 1.45 |
1.51 |
1.50 |
1.11 |
.89 |
.87 |
1.01 |
|
| Gck | 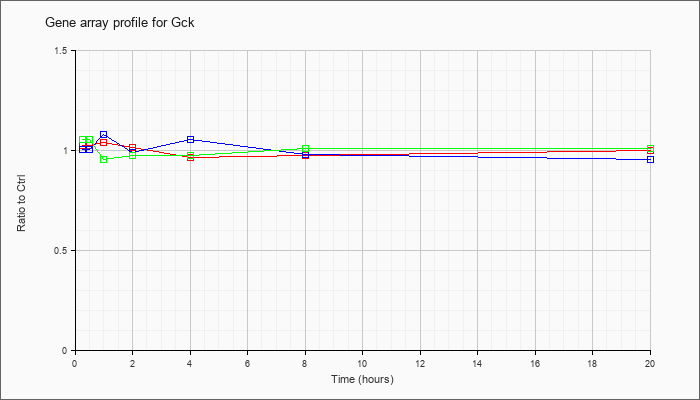 |
NM_010292 |
glucokinase (Gck), mRNA [NM_010292] |
KLA | 1.03 |
.99 |
1.04 |
1.02 |
.97 |
.98 |
1.00 |
| ATP | 1.01 |
.97 |
1.08 |
.99 |
1.06 |
.98 |
.96 |
| KLA/ATP | 1.06 |
1.01 |
.96 |
.98 |
.98 |
1.01 |
1.01 |
|
| Hk1 | 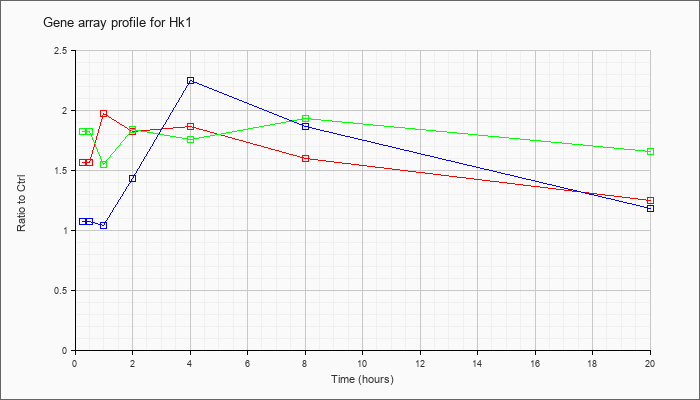 |
J05277 |
gb|Mouse hexokinase mRNA, complete cds [J05277] |
KLA | 1.49 |
1.45 |
1.66 |
1.78 |
1.51 |
1.29 |
.99 |
| ATP | 1.11 |
1.05 |
1.15 |
1.20 |
2.31 |
2.17 |
1.09 |
| KLA/ATP | 1.63 |
1.51 |
1.70 |
1.47 |
1.94 |
1.92 |
1.40 |
|
| Hk1 |  |
NM_010438 |
hexokinase 1 (Hk1), nuclear gene encoding mitochondrial protein, mRNA [NM_010438] |
KLA | 1.58 |
1.75 |
2.05 |
1.84 |
1.96 |
1.67 |
1.31 |
| ATP | 1.06 |
1.25 |
1.01 |
1.49 |
2.23 |
1.79 |
1.20 |
| KLA/ATP | 1.87 |
1.96 |
1.51 |
1.93 |
1.71 |
1.94 |
1.72 |
|
| Hk2 | 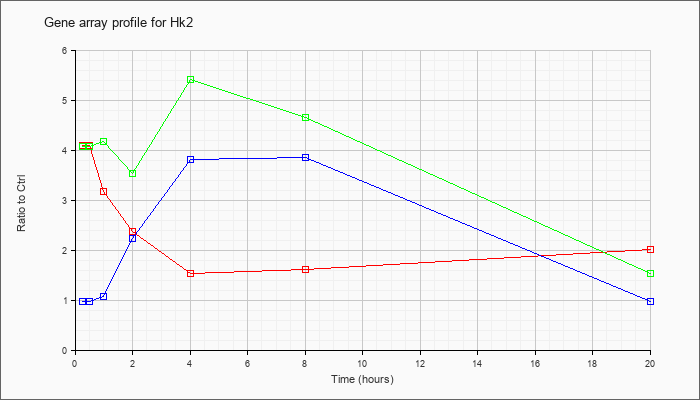 |
NM_013820 |
hexokinase 2 (Hk2), mRNA [NM_013820] |
KLA | 4.09 |
4.09 |
3.17 |
2.37 |
1.53 |
1.62 |
2.02 |
| ATP | .98 |
.92 |
1.08 |
2.24 |
3.82 |
3.86 |
.97 |
| KLA/ATP | 4.07 |
3.94 |
4.17 |
3.53 |
5.42 |
4.65 |
1.54 |
|
| Hk3 | 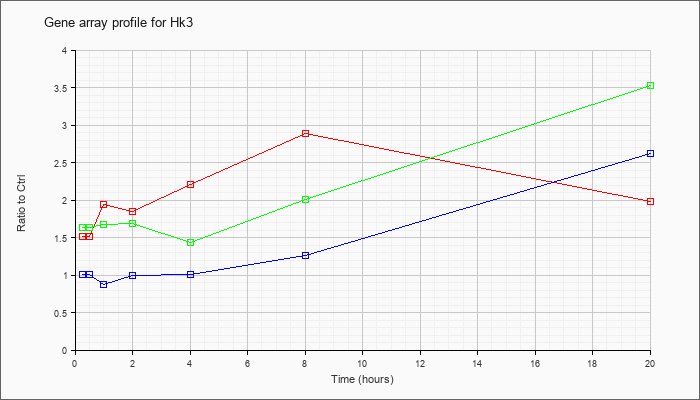 |
NM_001033245 |
hexokinase 3 (Hk3), nuclear gene encoding mitochondrial protein, mRNA [NM_001033245] |
KLA | 1.52 |
1.68 |
1.94 |
1.85 |
2.21 |
2.89 |
1.98 |
| ATP | 1.01 |
1.07 |
.87 |
.99 |
1.01 |
1.26 |
2.62 |
| KLA/ATP | 1.63 |
1.74 |
1.67 |
1.69 |
1.44 |
2.01 |
3.53 |
|
| Hkdc1 | 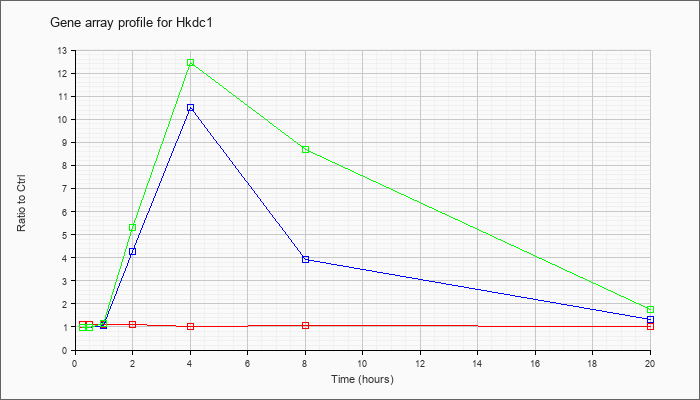 |
NM_145419 |
hexokinase domain containing 1 (Hkdc1), mRNA [NM_145419] |
KLA | 1.10 |
1.08 |
1.12 |
1.12 |
1.02 |
1.07 |
1.04 |
| ATP | .96 |
1.04 |
1.06 |
4.28 |
10.50 |
3.91 |
1.34 |
| KLA/ATP | .98 |
1.04 |
1.14 |
5.33 |
12.44 |
8.68 |
1.77 |
|
| Ikbkb | 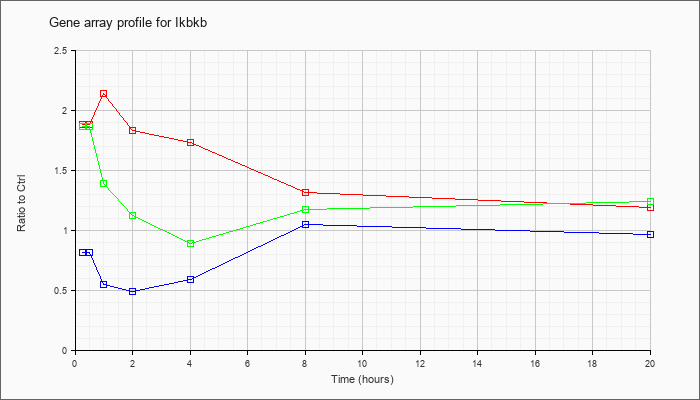 |
NM_010546 |
inhibitor of kappaB kinase beta (Ikbkb), mRNA [NM_010546] |
KLA | 1.88 |
2.10 |
2.14 |
1.83 |
1.73 |
1.31 |
1.19 |
| ATP | .81 |
.74 |
.55 |
.49 |
.59 |
1.05 |
.96 |
| KLA/ATP | 1.86 |
1.73 |
1.39 |
1.12 |
.89 |
1.17 |
1.24 |
|
| Ins1 | 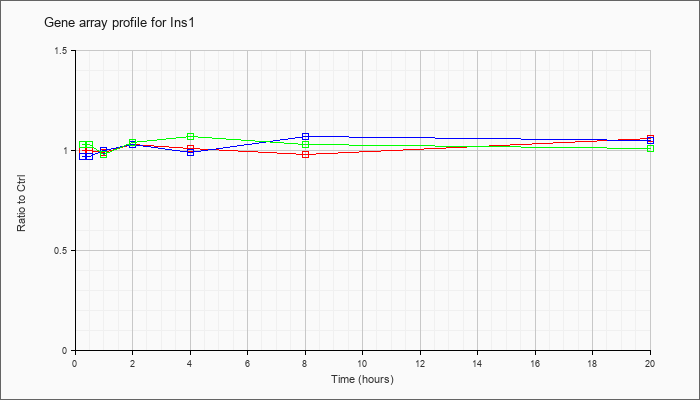 |
NM_008386 |
insulin I (Ins1), mRNA [NM_008386] |
KLA | 1.00 |
.99 |
.99 |
1.03 |
1.01 |
.98 |
1.06 |
| ATP | .97 |
1.03 |
1.00 |
1.03 |
.99 |
1.07 |
1.05 |
| KLA/ATP | 1.03 |
.93 |
.98 |
1.04 |
1.07 |
1.03 |
1.01 |
|
| Ins2 | 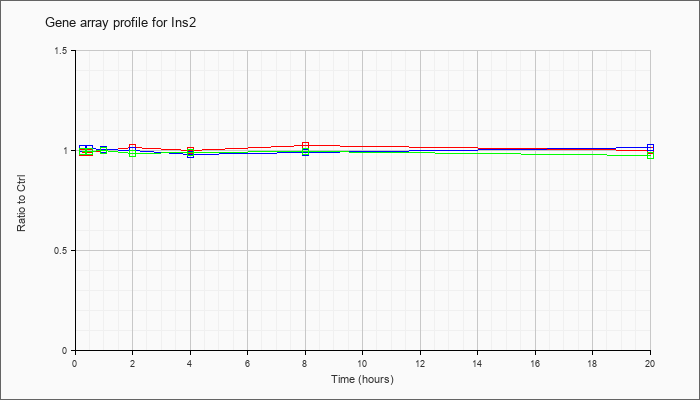 |
NM_008387 |
insulin II (Ins2), mRNA [NM_008387] |
KLA | .99 |
1.00 |
1.00 |
1.01 |
1.00 |
1.02 |
1.00 |
| ATP | 1.01 |
.99 |
1.00 |
1.00 |
.98 |
.99 |
1.01 |
| KLA/ATP | .99 |
1.01 |
1.00 |
.98 |
.99 |
.99 |
.98 |
|
| Insr | 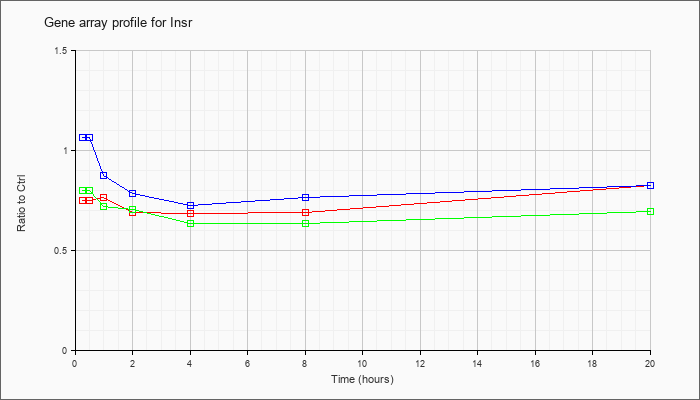 |
NM_010568 |
insulin receptor (Insr), mRNA [NM_010568] |
KLA | .75 |
.81 |
.77 |
.69 |
.69 |
.69 |
.83 |
| ATP | 1.07 |
1.06 |
.88 |
.79 |
.73 |
.77 |
.83 |
| KLA/ATP | .80 |
.80 |
.72 |
.71 |
.64 |
.64 |
.70 |
|
| Irs1 | 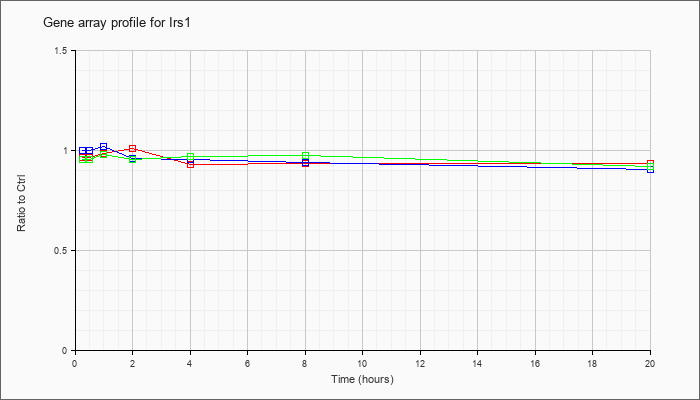 |
NM_010570 |
insulin receptor substrate 1 (Irs1), mRNA [NM_010570] |
KLA | .97 |
.96 |
.99 |
1.01 |
.93 |
.94 |
.94 |
| ATP | 1.00 |
.99 |
1.02 |
.96 |
.96 |
.94 |
.91 |
| KLA/ATP | .96 |
.96 |
.98 |
.96 |
.97 |
.98 |
.92 |
|
| Irs3 | 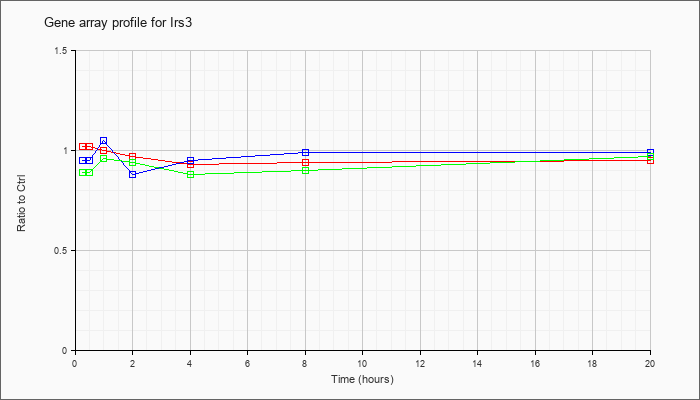 |
NM_010571 |
insulin receptor substrate 3 (Irs3), mRNA [NM_010571] |
KLA | 1.02 |
.92 |
1.00 |
.97 |
.93 |
.94 |
.95 |
| ATP | .95 |
1.06 |
1.05 |
.88 |
.95 |
.99 |
.99 |
| KLA/ATP | .89 |
.88 |
.96 |
.94 |
.88 |
.90 |
.97 |
|
| Irs4 | 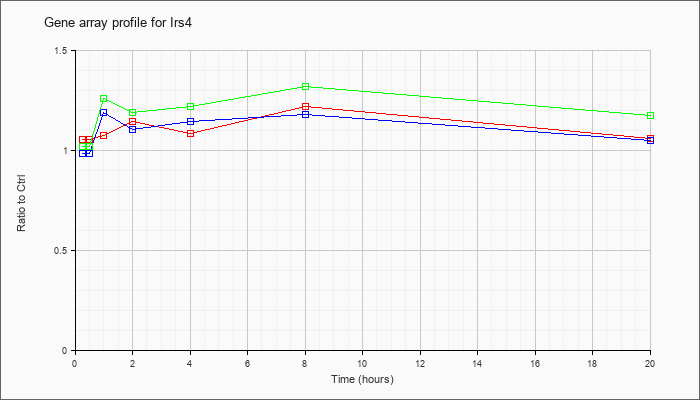 |
NM_010572 |
insulin receptor substrate 4 (Irs4), mRNA [NM_010572] |
KLA | 1.06 |
1.14 |
1.08 |
1.15 |
1.09 |
1.22 |
1.06 |
| ATP | .99 |
.98 |
1.19 |
1.10 |
1.15 |
1.18 |
1.05 |
| KLA/ATP | 1.02 |
1.10 |
1.26 |
1.19 |
1.22 |
1.32 |
1.18 |
|
| Kcnj11 | 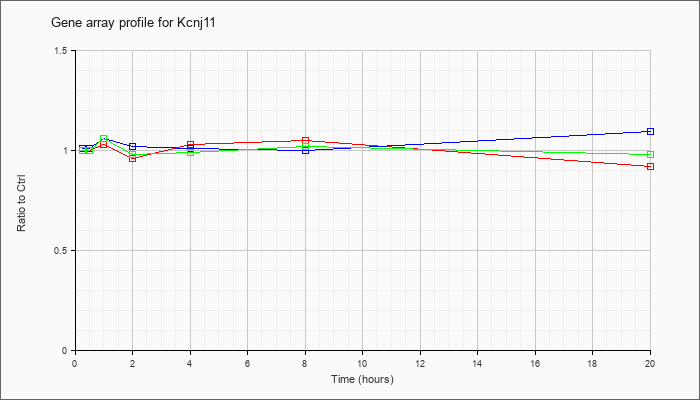 |
NM_010602 |
potassium inwardly rectifying channel, subfamily J, member 11 (Kcnj11), mRNA [NM_010602] |
KLA | 1.00 |
1.02 |
1.03 |
.96 |
1.03 |
1.05 |
.92 |
| ATP | 1.01 |
1.03 |
1.06 |
1.02 |
1.01 |
1.00 |
1.09 |
| KLA/ATP | 1.00 |
1.01 |
1.06 |
.98 |
.99 |
1.02 |
.98 |
|
| Mafa | 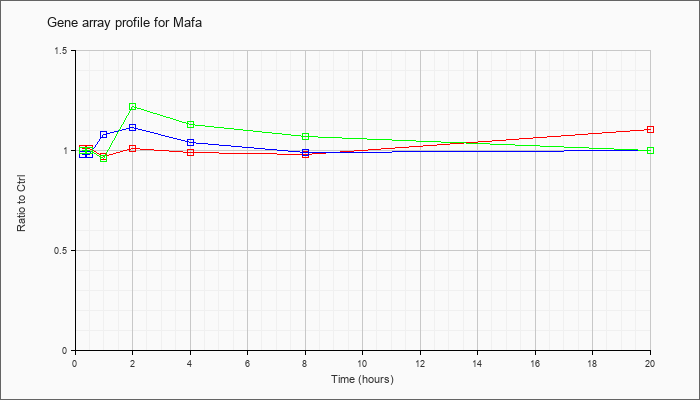 |
NM_194350 |
v-maf musculoaponeurotic fibrosarcoma oncogene family, protein A (avian) (Mafa), mRNA [NM_194350] |
KLA | 1.01 |
1.00 |
.97 |
1.01 |
.99 |
.98 |
1.10 |
| ATP | .98 |
1.08 |
1.08 |
1.11 |
1.04 |
.99 |
1.00 |
| KLA/ATP | 1.00 |
1.02 |
.96 |
1.22 |
1.13 |
1.07 |
1.00 |
|
| Mapk1 | 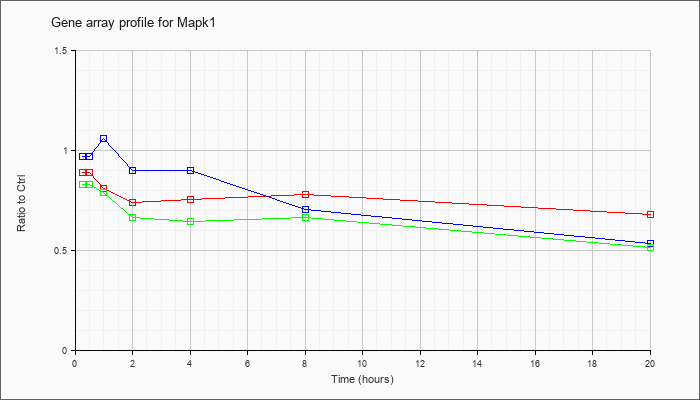 |
NM_001038663 |
mitogen-activated protein kinase 1 (Mapk1), transcript variant 2, mRNA [NM_001038663] |
KLA | .97 |
1.03 |
.93 |
.86 |
.79 |
1.01 |
1.04 |
| ATP | .96 |
1.18 |
1.01 |
1.13 |
.84 |
1.04 |
1.16 |
| KLA/ATP | .99 |
.99 |
.84 |
.87 |
.63 |
.96 |
1.26 |
|
| Mapk1 |  |
NM_011949 |
mitogen-activated protein kinase 1 (Mapk1), transcript variant 1, mRNA [NM_011949] |
KLA | .88 |
.80 |
.80 |
.73 |
.75 |
.76 |
.65 |
| ATP | .97 |
.96 |
1.06 |
.88 |
.90 |
.68 |
.48 |
| KLA/ATP | .82 |
.81 |
.78 |
.65 |
.64 |
.64 |
.45 |
|
| Mapk10 | 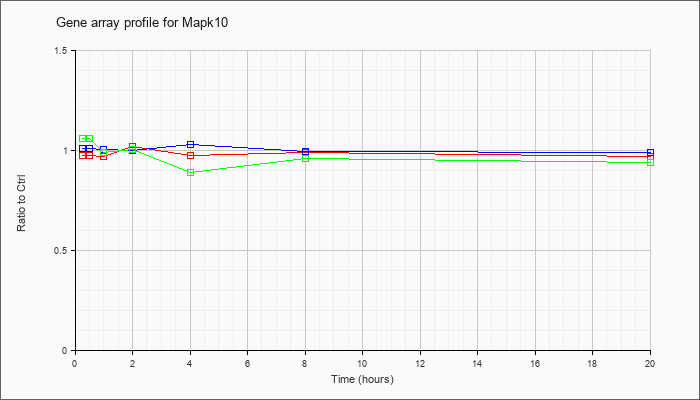 |
AK076990 |
adult male testis cDNA, RIKEN full-length enriched library, clone:4930597C01 product:mitogen activated protein kinase 10, full insert sequence. [AK076990] |
KLA | 1.03 |
1.05 |
.98 |
1.04 |
.94 |
.95 |
1.02 |
| ATP | 1.00 |
.99 |
1.06 |
.97 |
1.01 |
.96 |
.99 |
| KLA/ATP | 1.07 |
1.01 |
.95 |
1.05 |
.82 |
.95 |
.87 |
|
| Mapk10 |  |
NM_009158 |
mitogen-activated protein kinase 10 (Mapk10), transcript variant 1, mRNA [NM_009158] |
KLA | .95 |
1.00 |
.97 |
1.01 |
.99 |
1.01 |
.95 |
| ATP | 1.01 |
1.02 |
.98 |
1.01 |
1.04 |
1.01 |
.99 |
| KLA/ATP | 1.05 |
.99 |
1.01 |
.98 |
.92 |
.97 |
.98 |
|
| Mapk3 | 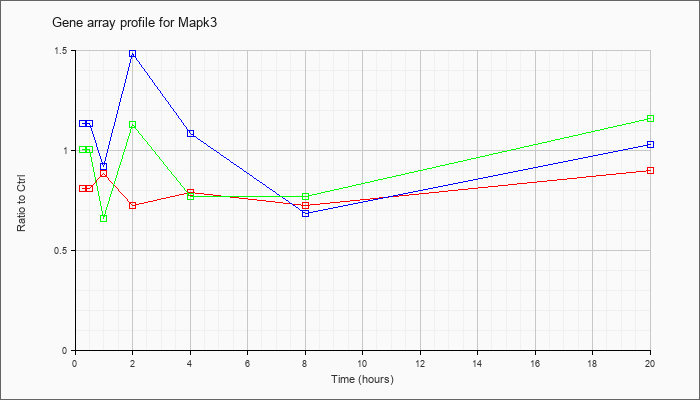 |
NM_011952 |
mitogen-activated protein kinase 3 (Mapk3), mRNA [NM_011952] |
KLA | .81 |
.85 |
.88 |
.72 |
.79 |
.73 |
.90 |
| ATP | 1.13 |
1.30 |
.92 |
1.48 |
1.08 |
.68 |
1.03 |
| KLA/ATP | 1.01 |
1.01 |
.66 |
1.13 |
.77 |
.77 |
1.16 |
|
| Mapk8 |  |
AK162915 |
adult male spinal cord cDNA, RIKEN full-length enriched library, clone:A330031O19 product:mitogen activated protein kinase 8, full insert sequence [AK162915] |
KLA | .94 |
.86 |
.80 |
.83 |
.77 |
.85 |
.83 |
| ATP | 1.08 |
1.12 |
.94 |
.80 |
.83 |
.71 |
.75 |
| KLA/ATP | .83 |
.81 |
.80 |
.74 |
.74 |
.75 |
.67 |
|
| Mapk8 |  |
AK163829 |
16 days embryo head cDNA, RIKEN full-length enriched library, clone:C130070A06 product:mitogen activated protein kinase 8, full insert sequence [AK163829] |
KLA | .65 |
.61 |
.66 |
.70 |
.90 |
1.02 |
1.03 |
| ATP | .99 |
.94 |
1.68 |
1.44 |
1.75 |
1.84 |
1.10 |
| KLA/ATP | .63 |
.67 |
1.11 |
1.00 |
2.01 |
1.68 |
1.15 |
|
| Mapk8 |  |
NM_016700 |
mitogen-activated protein kinase 8 (Mapk8), mRNA [NM_016700] |
KLA | .96 |
1.00 |
.94 |
.96 |
.94 |
.99 |
1.04 |
| ATP | 1.01 |
1.05 |
1.01 |
.99 |
1.03 |
.98 |
1.02 |
| KLA/ATP | .96 |
.97 |
.96 |
.97 |
.93 |
.94 |
.96 |
|
| Mapk9 | 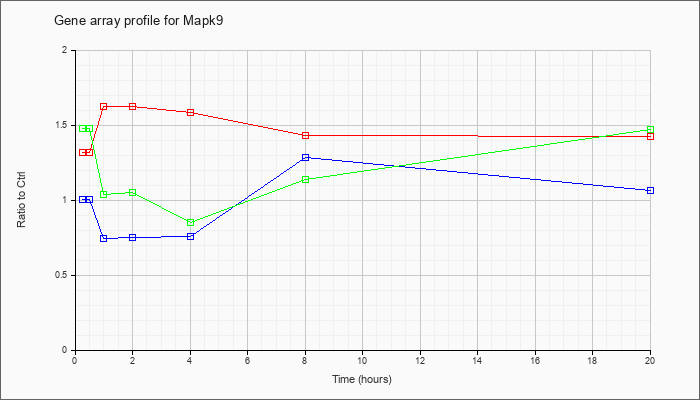 |
NM_016961 |
mitogen-activated protein kinase 9 (Mapk9), transcript variant 2, mRNA [NM_016961] |
KLA | 1.28 |
1.37 |
1.59 |
1.60 |
1.61 |
1.43 |
1.38 |
| ATP | 1.03 |
.99 |
.74 |
.71 |
.79 |
1.20 |
1.01 |
| KLA/ATP | 1.53 |
1.30 |
1.07 |
.99 |
.83 |
1.10 |
1.40 |
|
| Mapk9 |  |
NM_207692 |
mitogen-activated protein kinase 9 (Mapk9), transcript variant 1, mRNA [NM_207692] |
KLA | 1.39 |
1.53 |
1.69 |
1.69 |
1.55 |
1.44 |
1.51 |
| ATP | .95 |
.92 |
.75 |
.85 |
.70 |
1.46 |
1.17 |
| KLA/ATP | 1.37 |
1.35 |
.99 |
1.18 |
.91 |
1.22 |
1.62 |
|
Mm.21312
8 | 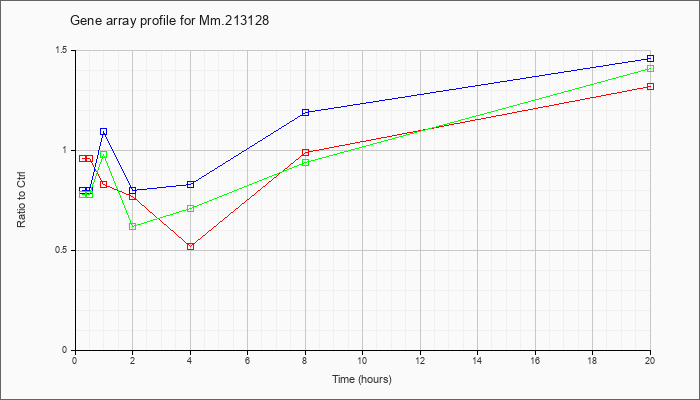 |
141801344 |
Unknown |
KLA | .96 |
.95 |
.83 |
.77 |
.52 |
.99 |
1.32 |
| ATP | .80 |
.82 |
1.09 |
.80 |
.83 |
1.19 |
1.46 |
| KLA/ATP | .78 |
.92 |
.98 |
.62 |
.71 |
.94 |
1.41 |
|
Mm.25381
9 | 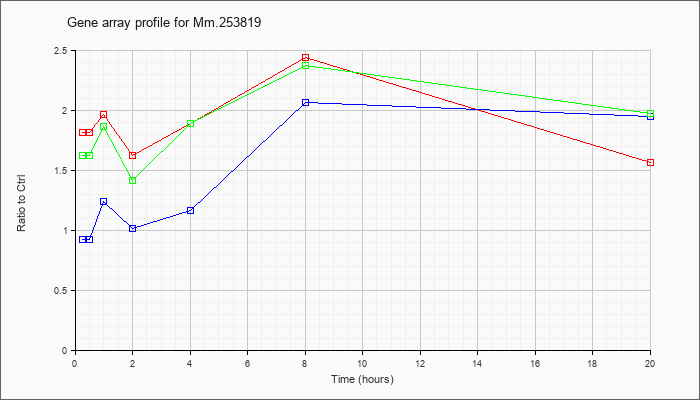 |
118130676 |
Unknown |
KLA | 1.81 |
1.88 |
1.96 |
1.62 |
1.89 |
2.44 |
1.56 |
| ATP | .92 |
.91 |
1.24 |
1.01 |
1.16 |
2.06 |
1.95 |
| KLA/ATP | 1.62 |
1.86 |
1.86 |
1.41 |
1.89 |
2.37 |
1.97 |
|
| Mm.4424 | 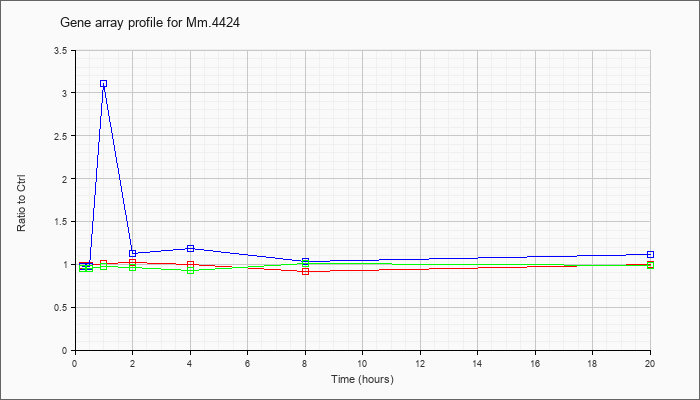 |
110225367 |
Unknown |
KLA | .99 |
.93 |
1.01 |
1.02 |
1.00 |
.92 |
1.00 |
| ATP | .98 |
1.03 |
3.11 |
1.13 |
1.19 |
1.03 |
1.11 |
| KLA/ATP | .95 |
.97 |
.97 |
.96 |
.93 |
1.01 |
.99 |
|
| Pdx1 | 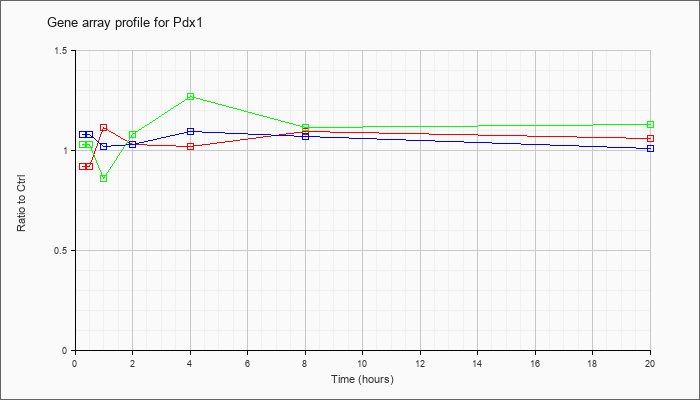 |
NM_008814 |
pancreatic and duodenal homeobox 1 (Pdx1), mRNA [NM_008814] |
KLA | .92 |
1.07 |
1.11 |
1.03 |
1.02 |
1.09 |
1.06 |
| ATP | 1.08 |
1.02 |
1.02 |
1.03 |
1.09 |
1.07 |
1.01 |
| KLA/ATP | 1.03 |
1.12 |
.86 |
1.08 |
1.27 |
1.11 |
1.13 |
|
| Pik3ca | 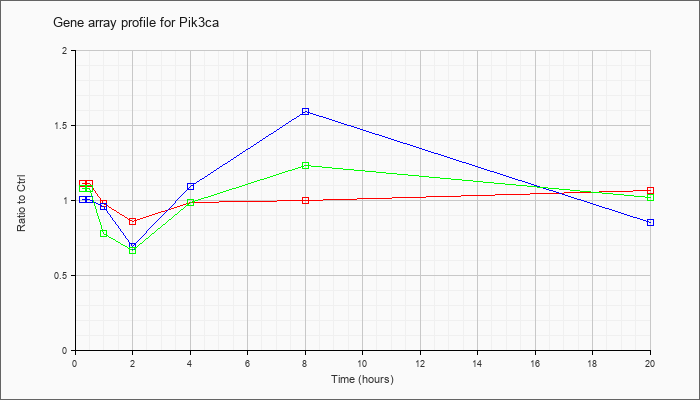 |
ENSMUST00000108243 |
ens|Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha isoform (EC 2.7.1.153) (PI3-kinase p110 subunit alpha) (PtdIns-3- kinase p110) (PI3K). [Source:Uniprot/SWISSPROT;Acc:P42337] [ENSMUST00000108243] |
KLA | 1.07 |
1.01 |
.91 |
.90 |
.98 |
.96 |
.86 |
| ATP | 1.03 |
.87 |
1.12 |
.63 |
1.22 |
1.86 |
.60 |
| KLA/ATP | 1.05 |
.82 |
.91 |
.61 |
1.17 |
1.26 |
.63 |
|
| Pik3ca |  |
NM_008839 |
phosphatidylinositol 3-kinase, catalytic, alpha polypeptide (Pik3ca), mRNA [NM_008839] |
KLA | 1.15 |
1.11 |
1.05 |
.82 |
.99 |
1.03 |
1.27 |
| ATP | .98 |
1.00 |
.79 |
.75 |
.96 |
1.32 |
1.10 |
| KLA/ATP | 1.10 |
.98 |
.64 |
.72 |
.80 |
1.20 |
1.41 |
|
| Pik3cb | 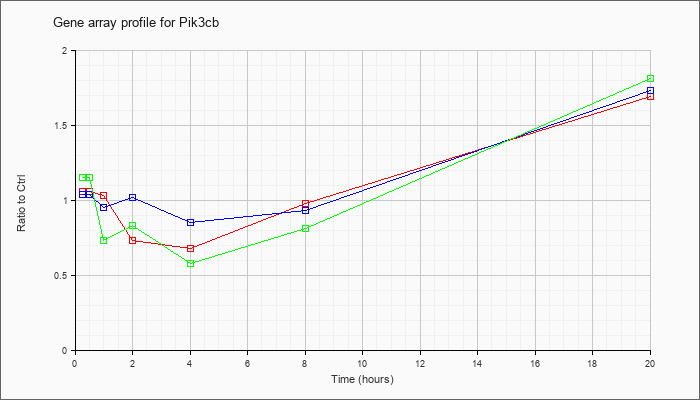 |
NM_029094 |
phosphatidylinositol 3-kinase, catalytic, beta polypeptide (Pik3cb), mRNA [NM_029094] |
KLA | 1.06 |
1.06 |
1.03 |
.73 |
.68 |
.98 |
1.69 |
| ATP | 1.04 |
1.21 |
.95 |
1.02 |
.85 |
.93 |
1.73 |
| KLA/ATP | 1.15 |
1.27 |
.73 |
.83 |
.58 |
.81 |
1.81 |
|
| Pik3cd | 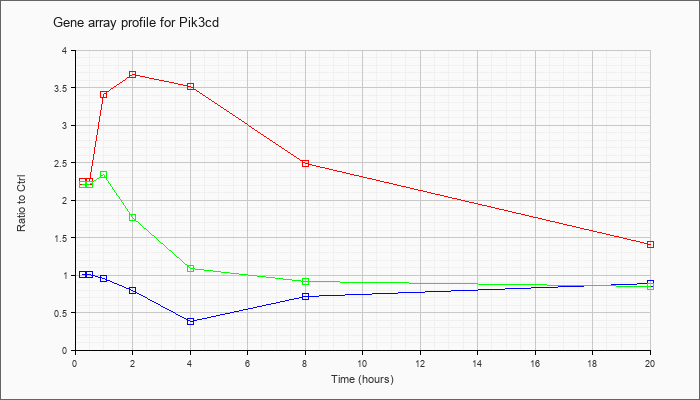 |
NM_008840 |
phosphatidylinositol 3-kinase catalytic delta polypeptide (Pik3cd), transcript variant 1, mRNA [NM_008840] |
KLA | 2.28 |
2.42 |
3.10 |
4.17 |
3.35 |
2.31 |
1.34 |
| ATP | 1.02 |
.91 |
.99 |
.63 |
.30 |
.67 |
.80 |
| KLA/ATP | 2.14 |
2.23 |
2.70 |
1.51 |
1.17 |
.85 |
.72 |
|
| Pik3cd |  |
U86587 |
phosphatidylinositol 3-kinase catalytic subunit p110 delta mRNA, complete cds. [U86587] |
KLA | 2.21 |
2.24 |
3.71 |
3.17 |
3.67 |
2.66 |
1.48 |
| ATP | 1.00 |
1.03 |
.93 |
.96 |
.46 |
.75 |
.97 |
| KLA/ATP | 2.27 |
2.32 |
1.97 |
2.02 |
1.01 |
.99 |
.98 |
|
| Pik3cg | 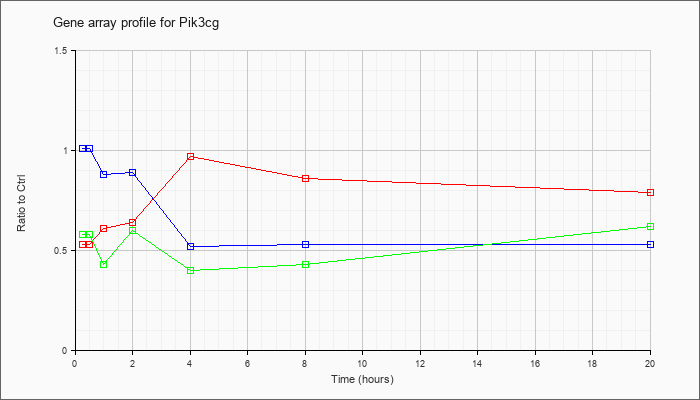 |
NM_020272 |
phosphoinositide-3-kinase, catalytic, gamma polypeptide (Pik3cg), mRNA [NM_020272] |
KLA | .53 |
.53 |
.61 |
.64 |
.97 |
.86 |
.79 |
| ATP | 1.01 |
1.13 |
.88 |
.89 |
.52 |
.53 |
.53 |
| KLA/ATP | .58 |
.53 |
.43 |
.60 |
.40 |
.43 |
.62 |
|
| Pik3r1 | 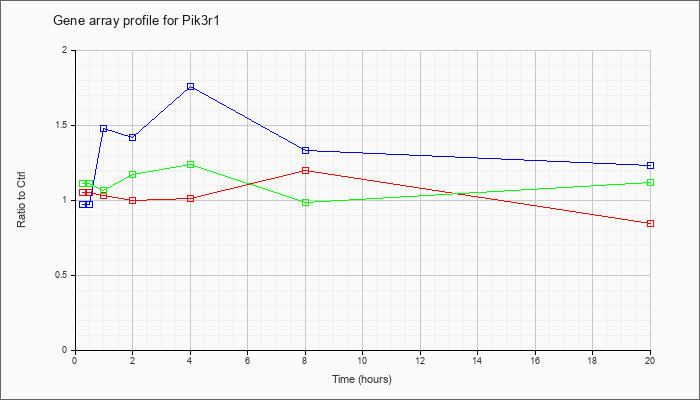 |
NM_001077495 |
phosphatidylinositol 3-kinase, regulatory subunit, polypeptide 1 (p85 alpha) (Pik3r1), transcript variant 2, mRNA [NM_001077495] |
KLA | 1.05 |
1.07 |
1.03 |
1.00 |
1.01 |
1.20 |
.85 |
| ATP | .97 |
1.02 |
1.48 |
1.42 |
1.76 |
1.33 |
1.23 |
| KLA/ATP | 1.11 |
1.12 |
1.06 |
1.17 |
1.24 |
.99 |
1.12 |
|
| Pik3r2 | 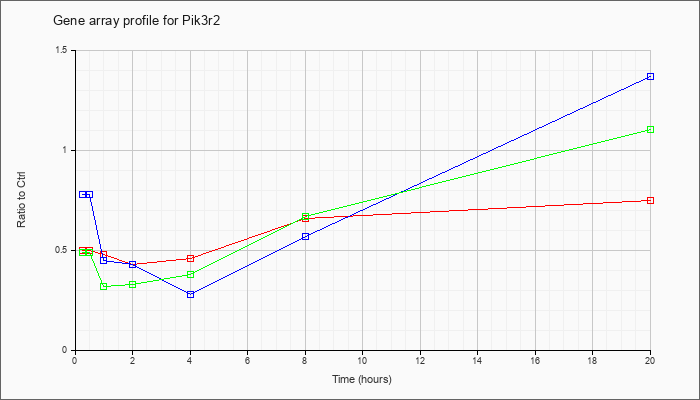 |
NM_008841 |
phosphatidylinositol 3-kinase, regulatory subunit, polypeptide 2 (p85 beta) (Pik3r2), mRNA [NM_008841] |
KLA | .50 |
.50 |
.48 |
.43 |
.46 |
.66 |
.75 |
| ATP | .78 |
.68 |
.45 |
.43 |
.28 |
.57 |
1.37 |
| KLA/ATP | .49 |
.38 |
.32 |
.33 |
.38 |
.67 |
1.10 |
|
| Pik3r3 | 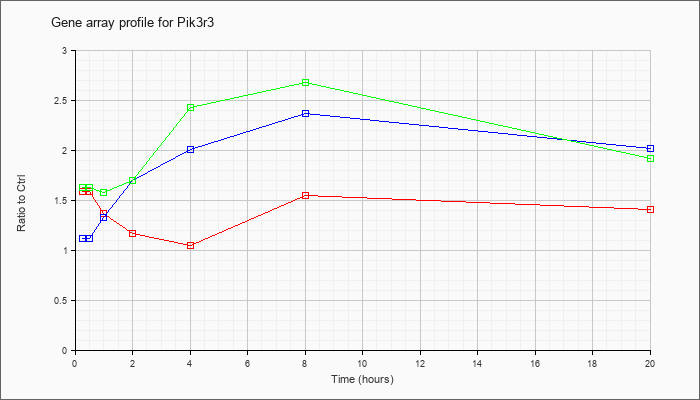 |
NM_181585 |
phosphatidylinositol 3 kinase, regulatory subunit, polypeptide 3 (p55) (Pik3r3), mRNA [NM_181585] |
KLA | 1.59 |
1.49 |
1.37 |
1.17 |
1.05 |
1.55 |
1.41 |
| ATP | 1.12 |
1.00 |
1.33 |
1.70 |
2.01 |
2.37 |
2.02 |
| KLA/ATP | 1.63 |
1.54 |
1.58 |
1.70 |
2.43 |
2.68 |
1.92 |
|
| Pik3r5 | 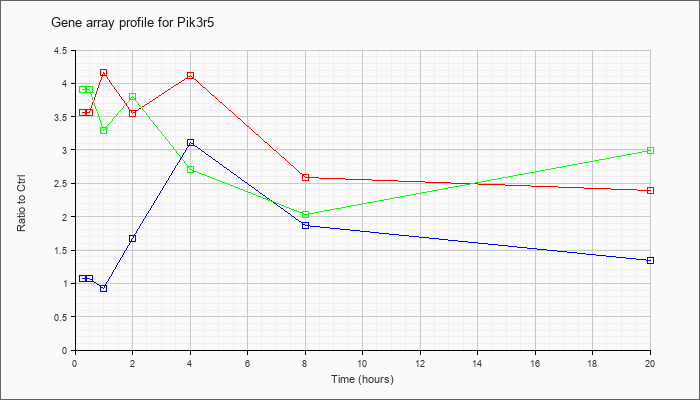 |
NM_177320 |
phosphoinositide-3-kinase, regulatory subunit 5, p101 (Pik3r5), mRNA [NM_177320] |
KLA | 3.57 |
3.58 |
4.16 |
3.54 |
4.11 |
2.59 |
2.39 |
| ATP | 1.08 |
1.08 |
.93 |
1.67 |
3.11 |
1.88 |
1.34 |
| KLA/ATP | 3.91 |
3.70 |
3.30 |
3.80 |
2.70 |
2.04 |
3.00 |
|
| Pklr | 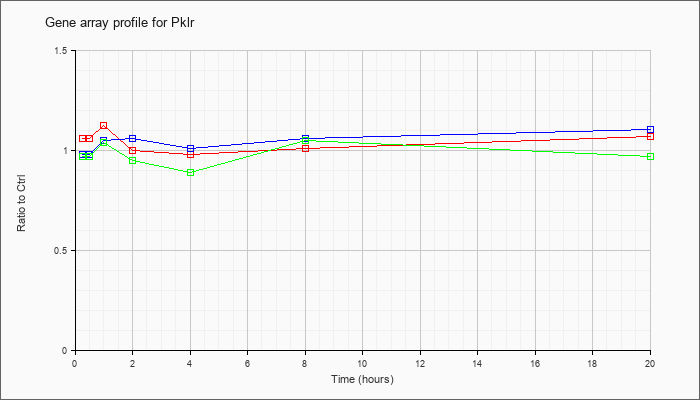 |
NM_013631 |
pyruvate kinase liver and red blood cell (Pklr), nuclear gene encoding mitochondrial protein, transcript variant 1, mRNA [NM_013631] |
KLA | 1.06 |
1.01 |
1.12 |
1.00 |
.98 |
1.01 |
1.07 |
| ATP | .98 |
.94 |
1.05 |
1.06 |
1.01 |
1.06 |
1.10 |
| KLA/ATP | .97 |
1.00 |
1.04 |
.95 |
.89 |
1.05 |
.97 |
|
| Pkm2 | 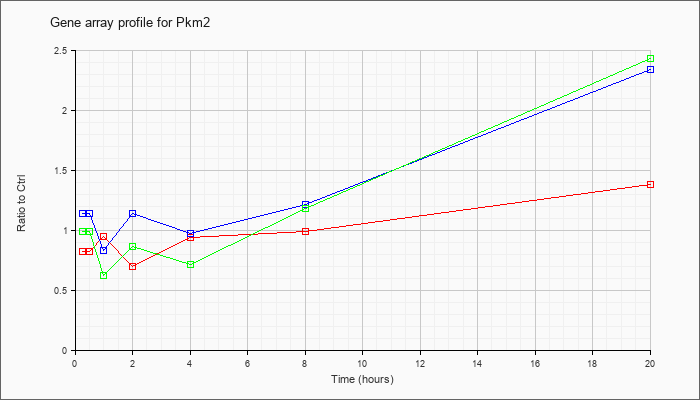 |
D38379 |
mRNA for pyruvate kinase M, complete cds. [D38379] |
KLA | .80 |
.83 |
.94 |
.67 |
.95 |
.95 |
1.42 |
| ATP | 1.22 |
1.30 |
.83 |
1.14 |
.99 |
1.23 |
2.53 |
| KLA/ATP | .99 |
1.03 |
.61 |
.83 |
.70 |
1.16 |
2.54 |
|
| Pkm2 |  |
NM_011099 |
pyruvate kinase, muscle (Pkm2), mRNA [NM_011099] |
KLA | .84 |
.89 |
.95 |
.72 |
.92 |
1.03 |
1.34 |
| ATP | 1.05 |
1.23 |
.83 |
1.13 |
.96 |
1.20 |
2.14 |
| KLA/ATP | .98 |
.97 |
.64 |
.89 |
.73 |
1.20 |
2.32 |
|
| Prkcd | 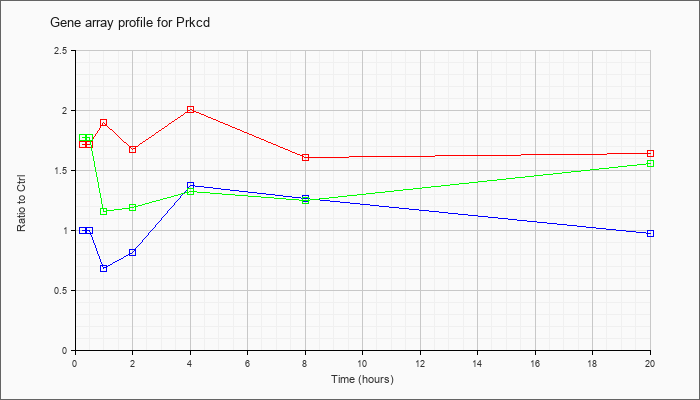 |
NM_011103 |
protein kinase C, delta (Prkcd), mRNA [NM_011103] |
KLA | 1.72 |
1.84 |
1.90 |
1.68 |
2.01 |
1.61 |
1.64 |
| ATP | 1.00 |
1.03 |
.68 |
.81 |
1.38 |
1.26 |
.98 |
| KLA/ATP | 1.77 |
1.79 |
1.16 |
1.19 |
1.33 |
1.25 |
1.56 |
|
| Prkce | 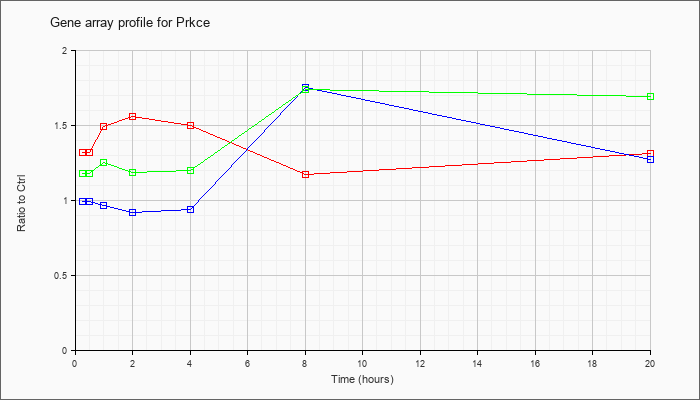 |
AK017901 |
adult male thymus cDNA, RIKEN full-length enriched library, clone:5830406C15 product:unclassifiable, full insert sequence. [AK017901] |
KLA | 1.63 |
1.44 |
1.95 |
2.09 |
1.96 |
1.34 |
1.56 |
| ATP | .99 |
.83 |
.85 |
.78 |
.85 |
2.44 |
1.47 |
| KLA/ATP | 1.36 |
1.45 |
1.44 |
1.23 |
1.30 |
2.39 |
2.29 |
|
| Prkce |  |
AK042994 |
7 days neonate cerebellum cDNA, RIKEN full-length enriched library, clone:A730046G04 product:protein kinase C, epsilon, full insert sequence. [AK042994] |
KLA | 1.01 |
1.07 |
1.03 |
1.03 |
1.04 |
1.00 |
1.06 |
| ATP | .99 |
1.09 |
1.08 |
1.05 |
1.02 |
1.06 |
1.07 |
| KLA/ATP | 1.00 |
.90 |
1.06 |
1.14 |
1.10 |
1.08 |
1.09 |
|
| Prkcz | 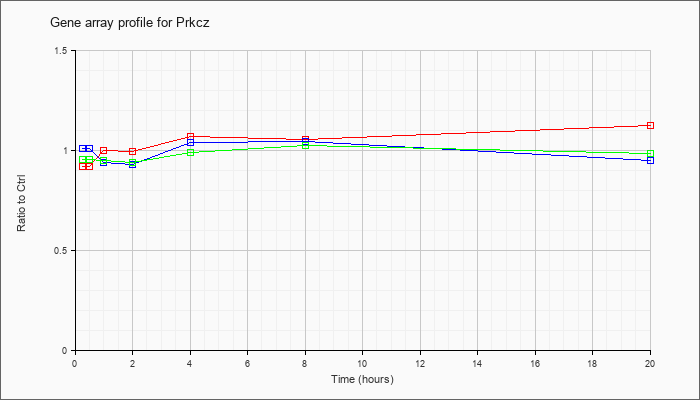 |
NM_008860 |
protein kinase C, zeta (Prkcz), transcript variant 1, mRNA [NM_008860] |
KLA | .92 |
.90 |
1.00 |
.99 |
1.07 |
1.05 |
1.12 |
| ATP | 1.01 |
.95 |
.94 |
.93 |
1.04 |
1.04 |
.95 |
| KLA/ATP | .95 |
.94 |
.95 |
.94 |
.99 |
1.02 |
.98 |
|
| Slc2a2 | 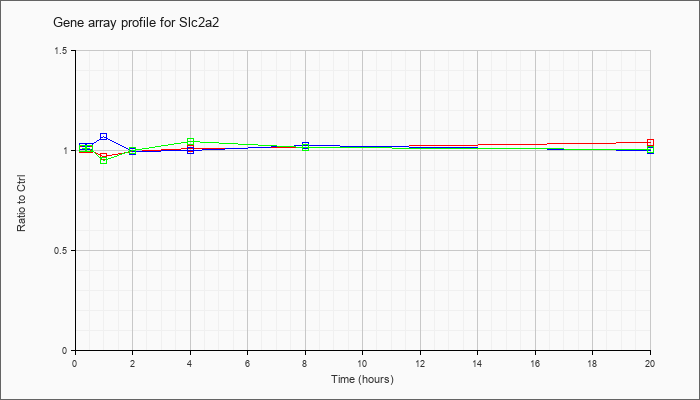 |
NM_031197 |
solute carrier family 2 (facilitated glucose transporter), member 2 (Slc2a2), mRNA [NM_031197] |
KLA | 1.01 |
1.03 |
.97 |
1.00 |
1.01 |
1.02 |
1.04 |
| ATP | 1.02 |
.97 |
1.07 |
1.00 |
1.00 |
1.03 |
1.00 |
| KLA/ATP | 1.01 |
1.01 |
.95 |
1.00 |
1.05 |
1.02 |
1.01 |
|
| Slc2a4 | 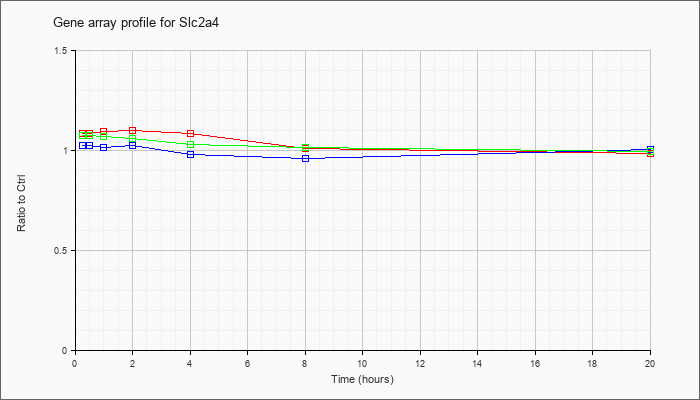 |
NM_009204 |
solute carrier family 2 (facilitated glucose transporter), member 4 (Slc2a4), mRNA [NM_009204] |
KLA | 1.08 |
1.10 |
1.09 |
1.10 |
1.08 |
1.01 |
.98 |
| ATP | 1.02 |
.98 |
1.01 |
1.02 |
.98 |
.96 |
1.00 |
| KLA/ATP | 1.07 |
1.08 |
1.07 |
1.06 |
1.03 |
1.01 |
.99 |
|
| Socs1 | 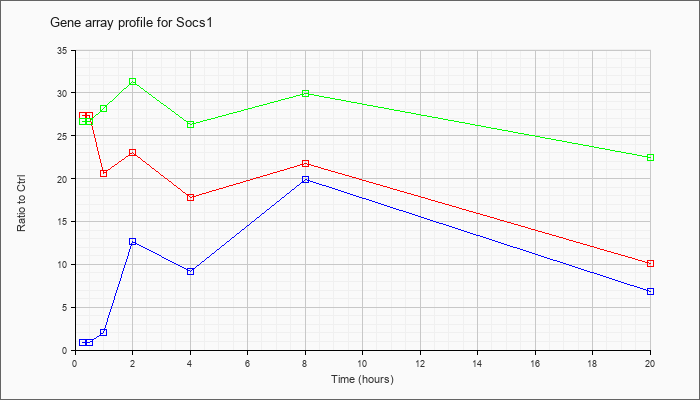 |
NM_009896 |
suppressor of cytokine signaling 1 (Socs1), mRNA [NM_009896] |
KLA | 27.36 |
22.16 |
20.64 |
23.08 |
17.74 |
21.71 |
10.12 |
| ATP | .91 |
.72 |
2.03 |
12.69 |
9.16 |
19.87 |
6.80 |
| KLA/ATP | 26.70 |
24.56 |
28.15 |
31.28 |
26.32 |
29.95 |
22.51 |
|
| Socs2 | 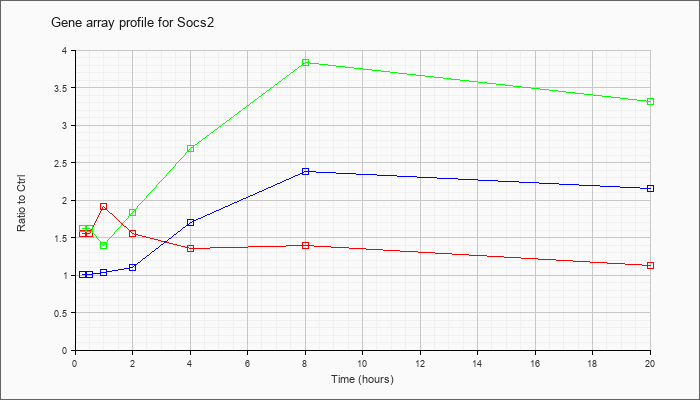 |
AK033206 |
15 days embryo male testis cDNA, RIKEN full-length enriched library, clone:8030460M17 product:hypothetical SH2 motif [AK033206] |
KLA | 1.08 |
1.11 |
1.22 |
1.13 |
1.11 |
1.08 |
.97 |
| ATP | .98 |
.97 |
1.03 |
1.03 |
1.29 |
1.30 |
1.29 |
| KLA/ATP | 1.06 |
1.09 |
1.07 |
1.33 |
2.01 |
1.85 |
1.98 |
|
| Socs2 |  |
NM_007706 |
suppressor of cytokine signaling 2 (Socs2), mRNA [NM_007706] |
KLA | 2.03 |
2.28 |
2.61 |
1.98 |
1.59 |
1.72 |
1.28 |
| ATP | 1.04 |
1.04 |
1.05 |
1.17 |
2.11 |
3.47 |
3.02 |
| KLA/ATP | 2.18 |
1.92 |
1.71 |
2.33 |
3.37 |
5.81 |
4.64 |
|
| Socs3 | 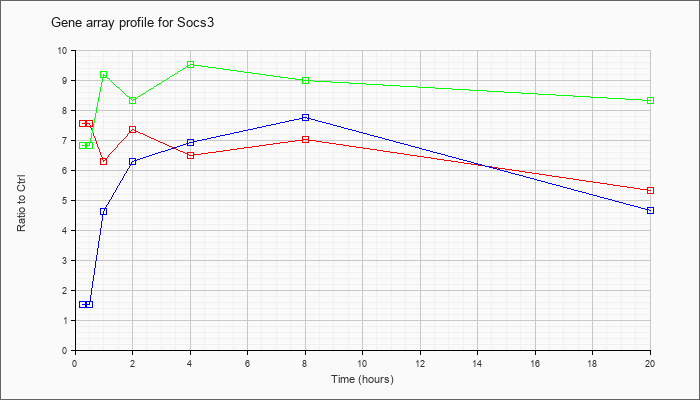 |
NM_007707 |
suppressor of cytokine signaling 3 (Socs3), mRNA [NM_007707] |
KLA | 7.56 |
6.68 |
6.29 |
7.36 |
6.47 |
7.03 |
5.33 |
| ATP | 1.51 |
1.75 |
4.63 |
6.27 |
6.92 |
7.74 |
4.64 |
| KLA/ATP | 6.82 |
6.77 |
9.19 |
8.32 |
9.51 |
8.98 |
8.33 |
|
| Socs4 | 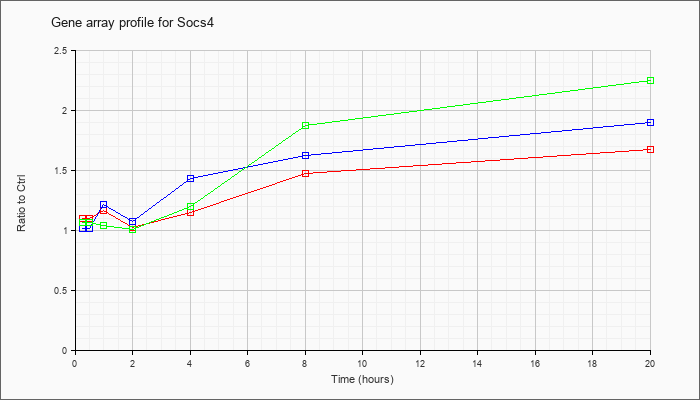 |
AK080408 |
7 days neonate cerebellum cDNA, RIKEN full-length enriched library, clone:A730004F22 product:unclassifiable, full insert sequence [AK080408] |
KLA | .89 |
.90 |
.87 |
.74 |
.87 |
1.05 |
.98 |
| ATP | 1.01 |
.97 |
1.11 |
.76 |
1.00 |
.82 |
1.11 |
| KLA/ATP | .80 |
.82 |
.82 |
.70 |
1.00 |
.80 |
.76 |
|
| Socs4 |  |
NM_080843 |
suppressor of cytokine signaling 4 (Socs4), mRNA [NM_080843] |
KLA | 1.31 |
1.29 |
1.45 |
1.31 |
1.42 |
1.90 |
2.37 |
| ATP | 1.02 |
1.16 |
1.31 |
1.39 |
1.86 |
2.43 |
2.69 |
| KLA/ATP | 1.33 |
1.24 |
1.26 |
1.31 |
1.39 |
2.94 |
3.73 |
|
| Tnf | 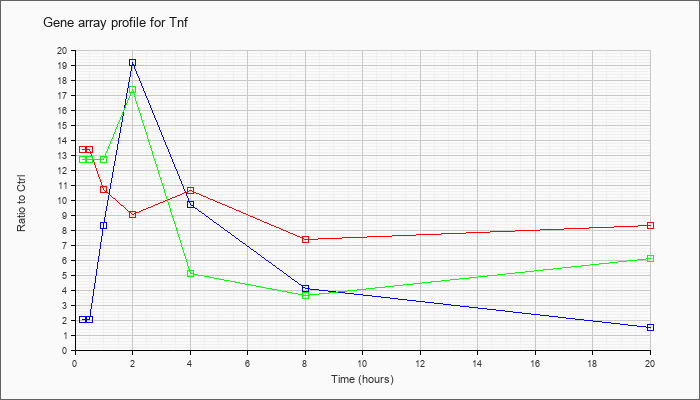 |
NM_013693 |
tumor necrosis factor (Tnf), mRNA [NM_013693] |
KLA | 13.35 |
12.76 |
10.71 |
9.03 |
10.64 |
7.39 |
8.32 |
| ATP | 2.01 |
2.48 |
8.27 |
19.20 |
9.72 |
4.09 |
1.49 |
| KLA/ATP | 12.72 |
15.76 |
12.73 |
17.34 |
5.10 |
3.61 |
6.07 |
|