| Gene symbol |
Time course plot |
Accession |
Description |
Treatment |
15min |
30min |
1hr |
2hr |
4hr |
8hr |
20hr |
BB229853
|  |
BB229853 |
gb|BB229853 RIKEN full-length enriched, 3 days neonate thymus Mus musculus cDNA clone A630023O21 3. [BB229853] |
KLA | 6.32 |
5.94 |
6.13 |
6.41 |
7.39 |
12.33 |
6.08 |
| ATP | .39 |
.24 |
.33 |
.38 |
1.18 |
5.75 |
2.85 |
| KLA/ATP | 2.27 |
2.02 |
3.25 |
4.04 |
7.11 |
10.22 |
13.93 |
|
| C3 | 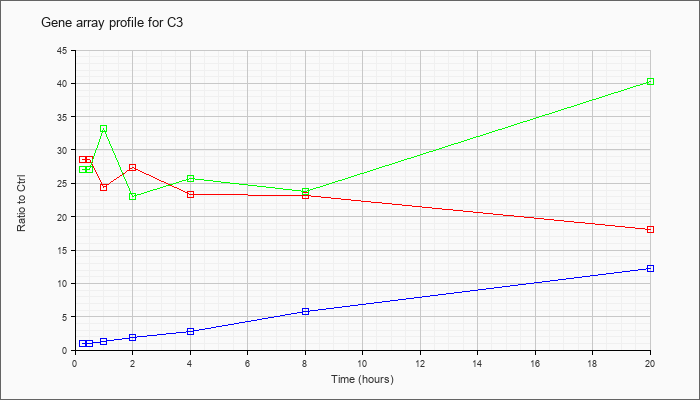 |
NM_009778 |
complement component 3 (C3), mRNA [NM_009778] |
KLA | 28.58 |
25.57 |
24.39 |
27.34 |
23.38 |
23.21 |
18.10 |
| ATP | 1.04 |
.93 |
1.21 |
1.90 |
2.80 |
5.74 |
12.19 |
| KLA/ATP | 27.04 |
26.10 |
33.29 |
23.09 |
25.65 |
23.77 |
40.22 |
|
| Cyba | 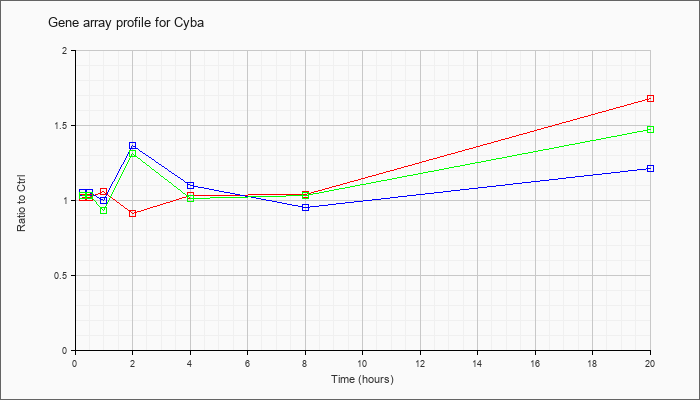 |
NM_007806 |
cytochrome b-245, alpha polypeptide (Cyba), mRNA [NM_007806] |
KLA | 1.02 |
1.05 |
1.06 |
.91 |
1.03 |
1.04 |
1.68 |
| ATP | 1.05 |
1.11 |
1.00 |
1.36 |
1.10 |
.95 |
1.21 |
| KLA/ATP | 1.03 |
1.11 |
.93 |
1.31 |
1.01 |
1.03 |
1.47 |
|
EG629860
| 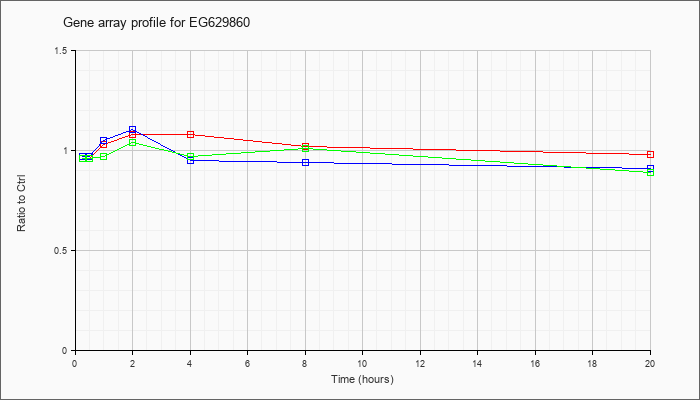 |
M17953 |
gb|Mouse Ig rearranged H-chain V-region mRNA VJ1 [M17953] |
KLA | .96 |
1.05 |
1.03 |
1.08 |
1.08 |
1.02 |
.98 |
| ATP | .97 |
.99 |
1.05 |
1.10 |
.95 |
.94 |
.91 |
| KLA/ATP | .96 |
.94 |
.97 |
1.04 |
.97 |
1.01 |
.89 |
|
| Elk1 | 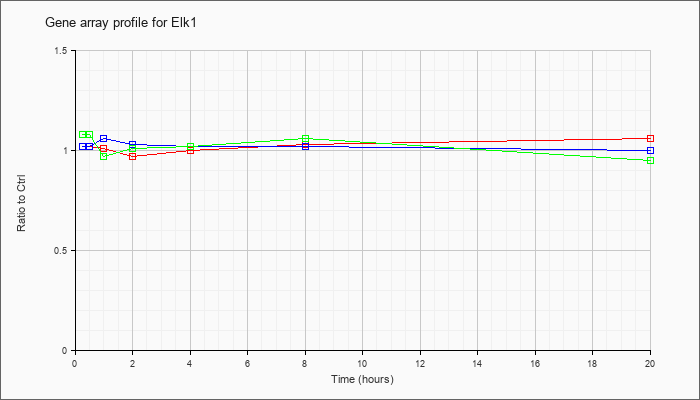 |
AK082260 |
0 day neonate cerebellum cDNA, RIKEN full-length enriched library, clone:C230030F12 product:ELK1, member of ETS oncogene family, full insert sequence. [AK082260] |
KLA | 1.02 |
1.01 |
1.01 |
.97 |
1.00 |
1.03 |
1.06 |
| ATP | 1.02 |
1.07 |
1.06 |
1.03 |
1.02 |
1.02 |
1.00 |
| KLA/ATP | 1.08 |
.99 |
.97 |
1.01 |
1.02 |
1.06 |
.95 |
|
| Fcgr1 | 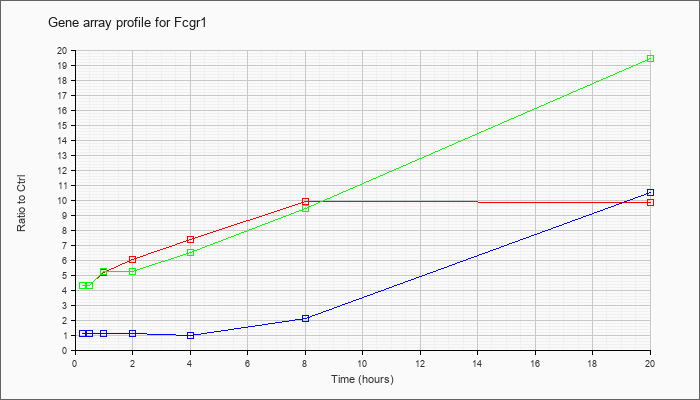 |
AF143181 |
strain AB/H (Biozzi) high affinity immunoglobulin gamma Fc receptor I (Fcgr1) mRNA, Fcgr1-d allele, complete cds. [AF143181] |
KLA | 4.27 |
4.21 |
5.59 |
6.14 |
8.13 |
9.76 |
10.98 |
| ATP | 1.23 |
1.12 |
1.08 |
1.36 |
1.09 |
2.09 |
11.81 |
| KLA/ATP | 4.56 |
4.93 |
4.95 |
5.88 |
6.66 |
9.64 |
23.09 |
|
| Fcgr1 |  |
NM_010186 |
Fc receptor, IgG, high affinity I (Fcgr1), mRNA [NM_010186] |
KLA | 4.31 |
4.37 |
4.70 |
5.97 |
6.59 |
10.06 |
8.67 |
| ATP | 1.01 |
.88 |
1.16 |
.89 |
.90 |
2.15 |
9.23 |
| KLA/ATP | 4.01 |
4.10 |
5.52 |
4.54 |
6.31 |
9.24 |
15.83 |
|
| Fcgr3 | 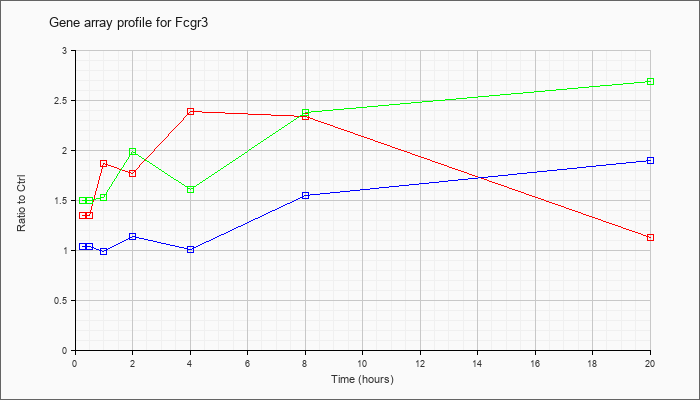 |
NM_010188 |
Fc receptor, IgG, low affinity III (Fcgr3), mRNA [NM_010188] |
KLA | 1.35 |
1.47 |
1.87 |
1.77 |
2.39 |
2.34 |
1.13 |
| ATP | 1.04 |
1.10 |
.99 |
1.14 |
1.01 |
1.55 |
1.90 |
| KLA/ATP | 1.50 |
1.53 |
1.53 |
1.99 |
1.61 |
2.38 |
2.69 |
|
| Fcgr4 | 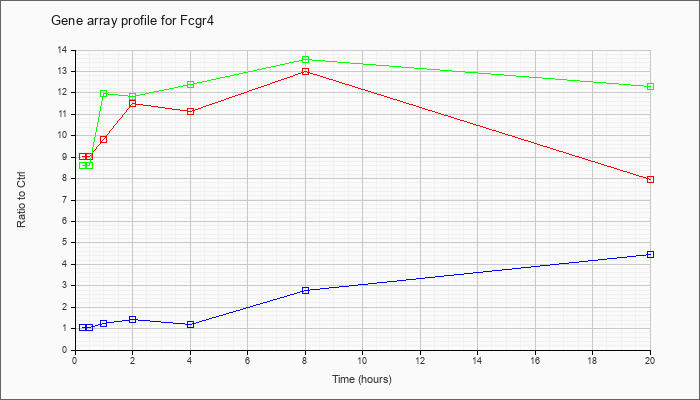 |
NM_144559 |
Fc receptor, IgG, low affinity IV (Fcgr4), mRNA [NM_144559] |
KLA | 9.03 |
8.68 |
9.82 |
11.51 |
11.12 |
13.01 |
7.95 |
| ATP | 1.04 |
.95 |
1.22 |
1.43 |
1.17 |
2.80 |
4.48 |
| KLA/ATP | 8.63 |
8.84 |
11.99 |
11.84 |
12.40 |
13.56 |
12.31 |
|
| Fos | 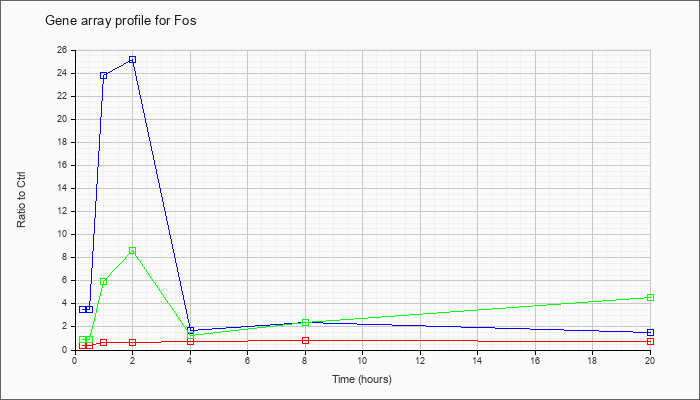 |
NM_010234 |
FBJ osteosarcoma oncogene (Fos), mRNA [NM_010234] |
KLA | .42 |
.33 |
.64 |
.61 |
.71 |
.85 |
.71 |
| ATP | 3.48 |
5.46 |
23.78 |
25.22 |
1.71 |
2.36 |
1.50 |
| KLA/ATP | .90 |
1.74 |
5.97 |
8.62 |
1.28 |
2.42 |
4.52 |
|
| H2-Aa | 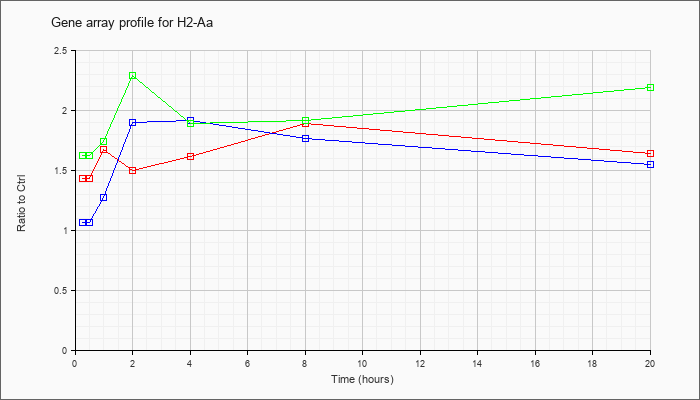 |
NM_010378 |
histocompatibility 2, class II antigen A, alpha (H2-Aa), mRNA [NM_010378] |
KLA | 1.43 |
1.59 |
1.68 |
1.50 |
1.62 |
1.89 |
1.64 |
| ATP | 1.07 |
1.15 |
1.27 |
1.90 |
1.92 |
1.77 |
1.55 |
| KLA/ATP | 1.62 |
1.81 |
1.74 |
2.29 |
1.89 |
1.92 |
2.19 |
|
| H2-Ab1 |  |
BC010322 |
histocompatibility 2, class II antigen A, beta 1, mRNA (cDNA clone MGC:6297 IMAGE:2651058), complete cds [BC010322] |
KLA | 1.73 |
1.76 |
1.77 |
1.55 |
1.95 |
2.10 |
2.27 |
| ATP | 1.03 |
1.18 |
1.32 |
1.99 |
1.79 |
1.56 |
1.61 |
| KLA/ATP | 1.70 |
2.01 |
1.74 |
2.52 |
1.58 |
2.01 |
2.75 |
|
| H2-Ab1 |  |
NM_207105 |
histocompatibility 2, class II antigen A, beta 1 (H2-Ab1), mRNA [NM_207105] |
KLA | 2.02 |
2.15 |
2.07 |
2.09 |
2.32 |
2.78 |
3.08 |
| ATP | 1.11 |
1.20 |
1.42 |
2.33 |
2.33 |
2.25 |
1.87 |
| KLA/ATP | 2.11 |
2.32 |
2.52 |
3.62 |
2.67 |
2.96 |
5.04 |
|
| H2-DMa | 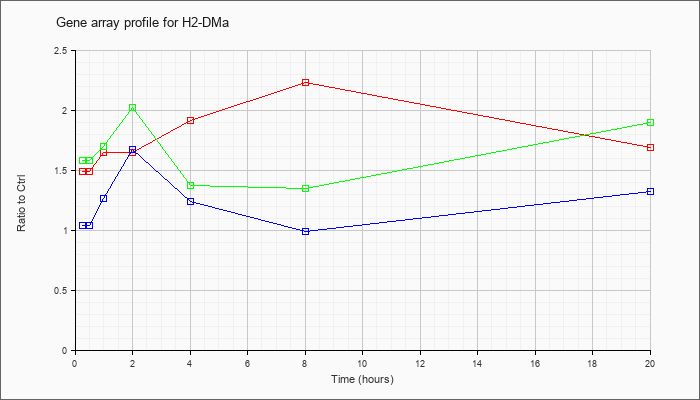 |
NM_010386 |
histocompatibility 2, class II, locus DMa (H2-DMa), mRNA [NM_010386] |
KLA | 1.49 |
1.50 |
1.65 |
1.65 |
1.91 |
2.23 |
1.69 |
| ATP | 1.04 |
1.12 |
1.26 |
1.67 |
1.24 |
.99 |
1.32 |
| KLA/ATP | 1.58 |
1.68 |
1.70 |
2.02 |
1.37 |
1.35 |
1.90 |
|
| H2-DMb1 | 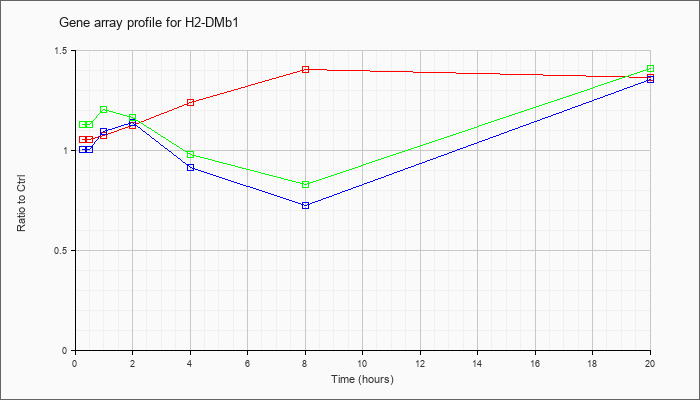 |
NM_010387 |
histocompatibility 2, class II, locus Mb1 (H2-DMb1), mRNA [NM_010387] |
KLA | 1.06 |
1.14 |
1.08 |
1.13 |
1.24 |
1.41 |
1.37 |
| ATP | 1.01 |
1.02 |
1.09 |
1.14 |
.92 |
.73 |
1.36 |
| KLA/ATP | 1.13 |
1.05 |
1.21 |
1.17 |
.98 |
.83 |
1.41 |
|
| H2-DMb2 | 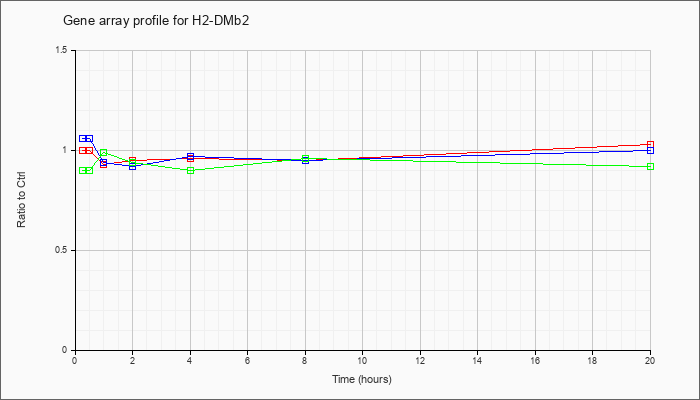 |
NM_010388 |
histocompatibility 2, class II, locus Mb2 (H2-DMb2), mRNA [NM_010388] |
KLA | 1.00 |
1.04 |
.93 |
.95 |
.96 |
.95 |
1.03 |
| ATP | 1.06 |
1.04 |
.94 |
.92 |
.97 |
.95 |
1.00 |
| KLA/ATP | .90 |
.94 |
.99 |
.94 |
.90 |
.96 |
.92 |
|
| H2-Eb1 | 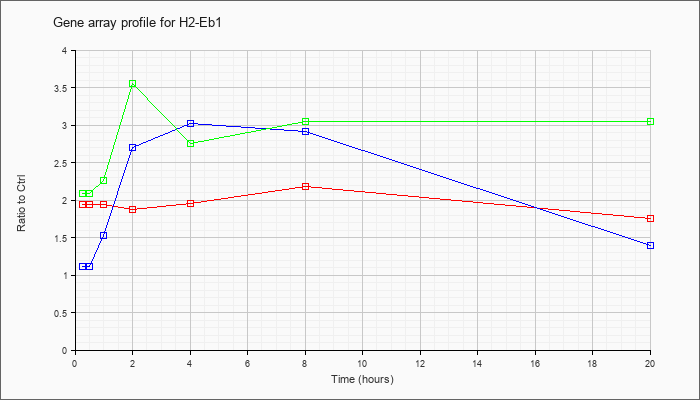 |
X52641 |
gb|Mouse mRNA for I-A beta NOD. [X52641] |
KLA | 1.94 |
2.06 |
1.94 |
1.87 |
1.96 |
2.18 |
1.76 |
| ATP | 1.11 |
1.20 |
1.53 |
2.70 |
3.02 |
2.91 |
1.40 |
| KLA/ATP | 2.09 |
2.11 |
2.26 |
3.55 |
2.76 |
3.05 |
3.05 |
|
| H2-Oa | 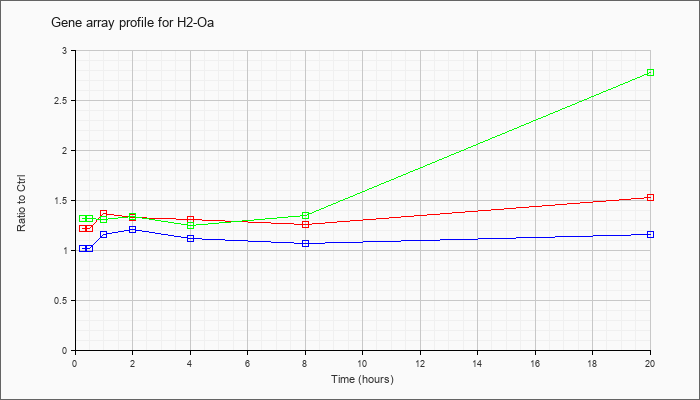 |
NM_008206 |
histocompatibility 2, O region alpha locus (H2-Oa), mRNA [NM_008206] |
KLA | 1.22 |
1.32 |
1.37 |
1.33 |
1.31 |
1.26 |
1.53 |
| ATP | 1.02 |
1.05 |
1.16 |
1.21 |
1.12 |
1.07 |
1.16 |
| KLA/ATP | 1.32 |
1.34 |
1.31 |
1.34 |
1.25 |
1.35 |
2.78 |
|
| H2-Ob | 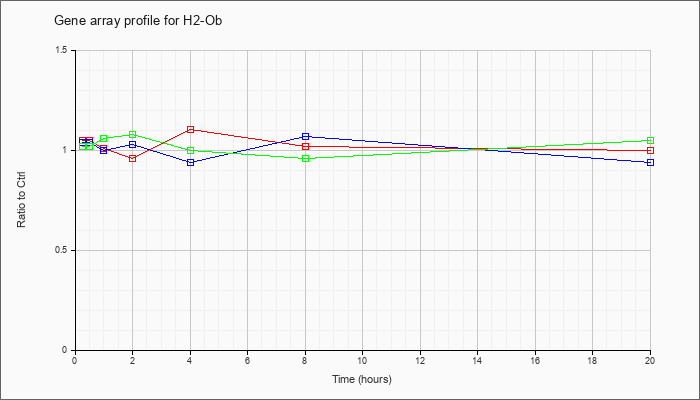 |
NM_010389 |
histocompatibility 2, O region beta locus (H2-Ob), mRNA [NM_010389] |
KLA | 1.05 |
1.07 |
1.01 |
.96 |
1.10 |
1.02 |
1.00 |
| ATP | 1.04 |
.99 |
1.00 |
1.03 |
.94 |
1.07 |
.94 |
| KLA/ATP | 1.02 |
.97 |
1.06 |
1.08 |
1.00 |
.96 |
1.05 |
|
| Ifng | 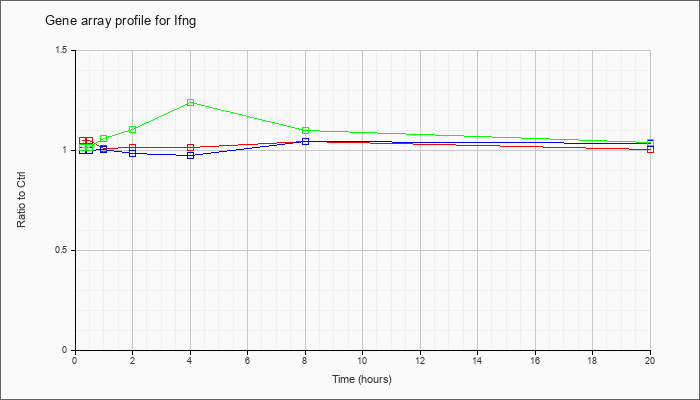 |
NM_008337 |
interferon gamma (Ifng), mRNA [NM_008337] |
KLA | 1.05 |
1.02 |
1.01 |
1.01 |
1.01 |
1.04 |
1.00 |
| ATP | 1.00 |
1.01 |
1.01 |
.98 |
.97 |
1.04 |
1.03 |
| KLA/ATP | 1.01 |
1.01 |
1.06 |
1.10 |
1.24 |
1.10 |
1.04 |
|
| Ifngr1 | 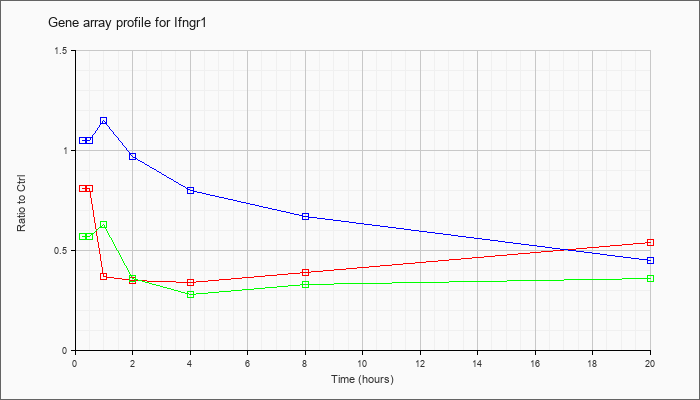 |
NM_010511 |
interferon gamma receptor 1 (Ifngr1), mRNA [NM_010511] |
KLA | .81 |
.60 |
.37 |
.35 |
.34 |
.39 |
.54 |
| ATP | 1.05 |
.96 |
1.15 |
.97 |
.80 |
.67 |
.45 |
| KLA/ATP | .57 |
.57 |
.63 |
.36 |
.28 |
.33 |
.36 |
|
| Ifngr2 | 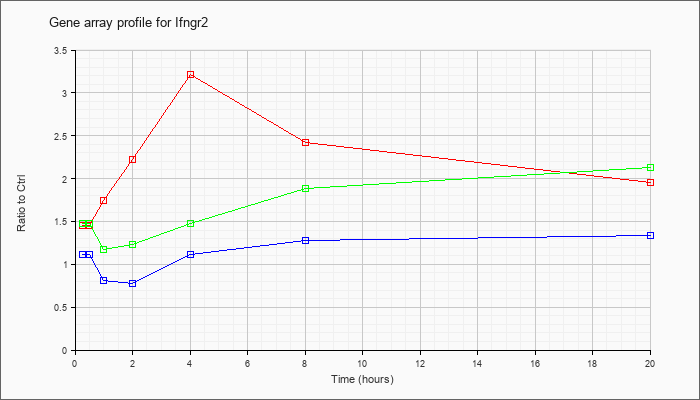 |
NM_008338 |
interferon gamma receptor 2 (Ifngr2), mRNA [NM_008338] |
KLA | 1.45 |
1.49 |
1.74 |
2.22 |
3.22 |
2.42 |
1.96 |
| ATP | 1.11 |
1.03 |
.81 |
.78 |
1.12 |
1.28 |
1.34 |
| KLA/ATP | 1.48 |
1.41 |
1.17 |
1.23 |
1.48 |
1.89 |
2.13 |
|
Igh-VJ55
8 | 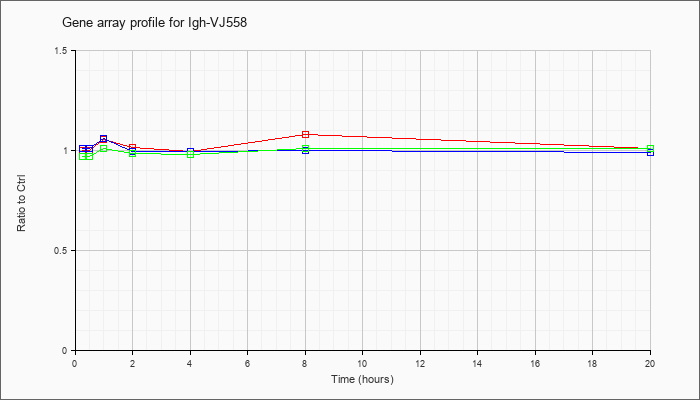 |
AY547436 |
immunoglobulin heavy chain VDJ region mRNA, partial cds [AY547436] |
KLA | 1.06 |
1.05 |
1.01 |
.93 |
1.03 |
.96 |
.93 |
| ATP | .99 |
1.00 |
1.06 |
.95 |
.96 |
1.06 |
1.05 |
| KLA/ATP | .98 |
.94 |
1.07 |
.95 |
.97 |
.99 |
1.02 |
|
Igh-VJ55
8 |  |
CA578712 |
gb|K0727F04-5N NIA Mouse Hematopoietic Stem Cell (Lin- [CA578712] |
KLA | .99 |
.94 |
1.01 |
1.10 |
.93 |
.97 |
.92 |
| ATP | .93 |
1.04 |
1.01 |
.98 |
1.05 |
.96 |
.95 |
| KLA/ATP | .99 |
.95 |
.95 |
.98 |
1.02 |
.98 |
.95 |
|
Igh-VJ55
8 |  |
ENSMUST00000103412 |
ens|Immunoglobulin heavy chain C gene segment [Source:IMGT/GENE_DB;Acc:IGHA] [ENSMUST00000103412] |
KLA | 1.01 |
.97 |
1.20 |
1.01 |
.98 |
1.44 |
1.24 |
| ATP | .99 |
.98 |
1.07 |
1.03 |
1.00 |
1.01 |
1.21 |
| KLA/ATP | .99 |
1.02 |
.96 |
1.03 |
.97 |
1.13 |
1.30 |
|
Igh-VJ55
8 |  |
ENSMUST00000103413 |
ens|Immunoglobulin heavy chain C gene segment [Source:IMGT/GENE_DB;Acc:IGHE] [ENSMUST00000103413] |
KLA | 1.00 |
.99 |
1.06 |
1.06 |
.99 |
1.03 |
1.02 |
| ATP | 1.08 |
1.06 |
1.06 |
1.06 |
1.02 |
1.06 |
.95 |
| KLA/ATP | 1.00 |
1.04 |
1.09 |
1.02 |
1.06 |
1.02 |
.95 |
|
Igh-VJ55
8 |  |
U39781 |
J558+ IgM heavy chain mRNA, hybridoma clone ME2B7, partial cds. [U39781] |
KLA | .93 |
.89 |
.98 |
.97 |
1.03 |
1.00 |
.94 |
| ATP | 1.04 |
1.02 |
1.08 |
.94 |
.93 |
.90 |
.79 |
| KLA/ATP | .88 |
.89 |
.97 |
.93 |
.88 |
.92 |
.82 |
|
| Il10 | 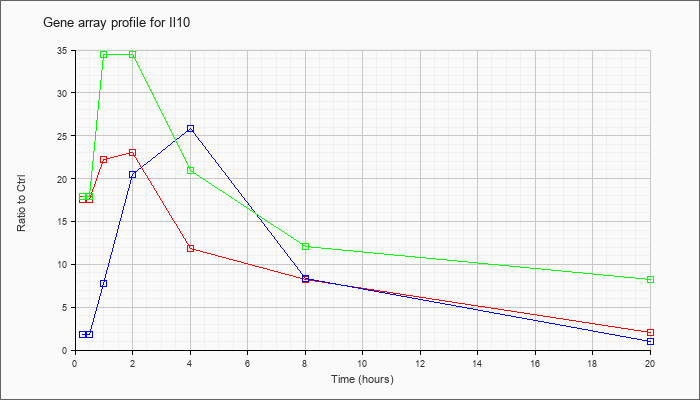 |
NM_010548 |
interleukin 10 (Il10), mRNA [NM_010548] |
KLA | 17.59 |
18.64 |
22.22 |
23.06 |
11.83 |
8.27 |
2.07 |
| ATP | 1.86 |
3.22 |
7.72 |
20.51 |
25.86 |
8.38 |
1.01 |
| KLA/ATP | 17.91 |
23.49 |
34.53 |
34.48 |
20.98 |
12.03 |
8.17 |
|
| Il12a | 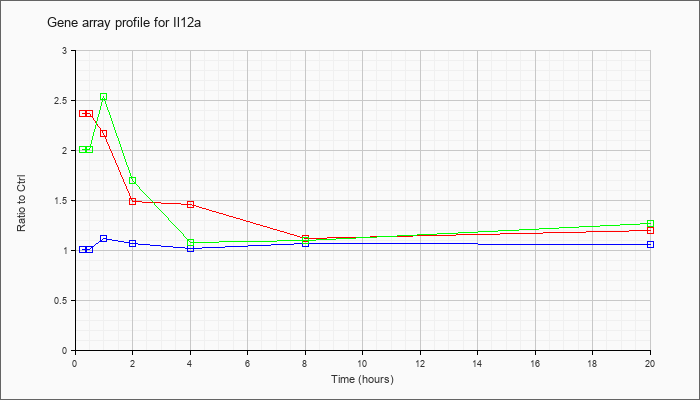 |
NM_008351 |
interleukin 12a (Il12a), mRNA [NM_008351] |
KLA | 2.37 |
2.47 |
2.17 |
1.49 |
1.46 |
1.12 |
1.20 |
| ATP | 1.01 |
.97 |
1.12 |
1.07 |
1.02 |
1.07 |
1.06 |
| KLA/ATP | 2.01 |
2.76 |
2.54 |
1.70 |
1.08 |
1.10 |
1.27 |
|
| Il12b | 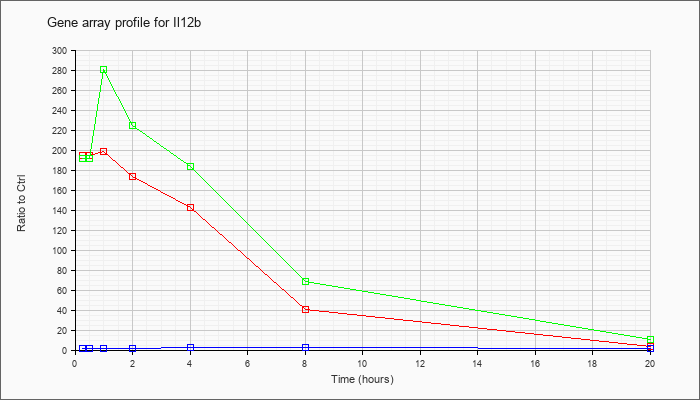 |
NM_008352 |
interleukin 12b (Il12b), mRNA [NM_008352] |
KLA | 194.79 |
211.32 |
198.97 |
173.01 |
142.24 |
40.28 |
3.10 |
| ATP | 1.05 |
1.10 |
1.59 |
1.99 |
2.71 |
2.72 |
1.52 |
| KLA/ATP | 191.27 |
223.23 |
280.10 |
224.29 |
183.03 |
68.43 |
10.69 |
|
| Il1a | 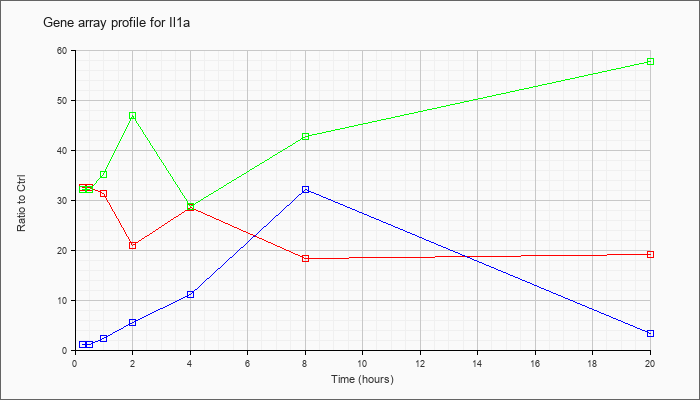 |
NM_010554 |
interleukin 1 alpha (Il1a), mRNA [NM_010554] |
KLA | 32.45 |
35.22 |
31.30 |
20.95 |
28.57 |
18.36 |
19.02 |
| ATP | 1.04 |
1.02 |
2.21 |
5.46 |
11.06 |
32.07 |
3.40 |
| KLA/ATP | 32.02 |
41.20 |
35.12 |
46.91 |
28.76 |
42.72 |
57.80 |
|
| Il1b | 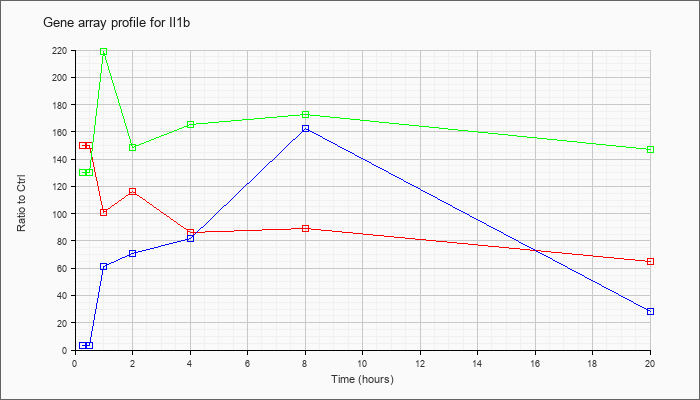 |
NM_008361 |
interleukin 1 beta (Il1b), mRNA [NM_008361] |
KLA | 149.78 |
133.59 |
100.91 |
116.58 |
86.48 |
89.37 |
64.72 |
| ATP | 3.49 |
13.96 |
61.44 |
70.91 |
82.12 |
162.79 |
28.18 |
| KLA/ATP | 130.32 |
139.00 |
219.12 |
148.17 |
165.44 |
172.57 |
146.98 |
|
| Il4 | 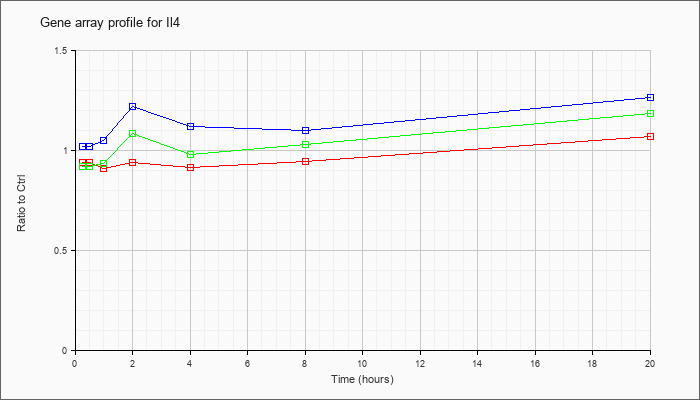 |
NM_021283 |
interleukin 4 (Il4), mRNA [NM_021283] |
KLA | .94 |
.95 |
.91 |
.94 |
.91 |
.94 |
1.07 |
| ATP | 1.02 |
1.06 |
1.05 |
1.22 |
1.12 |
1.10 |
1.26 |
| KLA/ATP | .92 |
.96 |
.93 |
1.08 |
.98 |
1.03 |
1.18 |
|
| Irak1 | 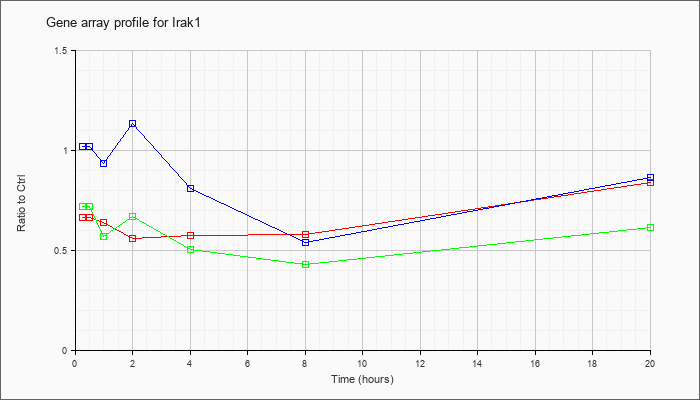 |
NM_008363 |
interleukin-1 receptor-associated kinase 1 (Irak1), mRNA [NM_008363] |
KLA | .67 |
.67 |
.64 |
.56 |
.58 |
.58 |
.84 |
| ATP | 1.02 |
1.10 |
.94 |
1.14 |
.81 |
.54 |
.87 |
| KLA/ATP | .72 |
.72 |
.57 |
.67 |
.51 |
.43 |
.62 |
|
| Irak4 | 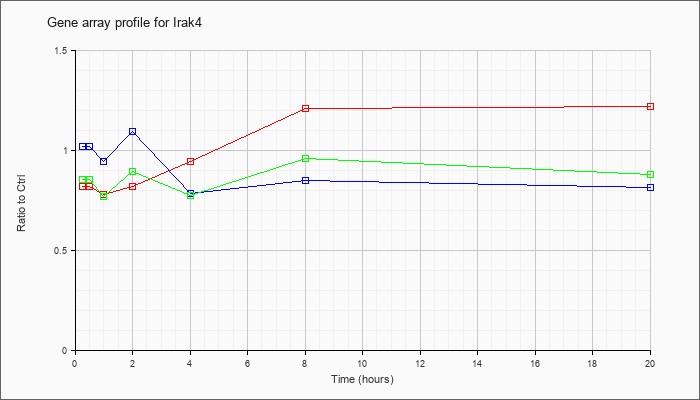 |
NM_029926 |
interleukin-1 receptor-associated kinase 4 (Irak4), mRNA [NM_029926] |
KLA | .82 |
.86 |
.78 |
.82 |
.95 |
1.21 |
1.22 |
| ATP | 1.02 |
1.18 |
.95 |
1.10 |
.79 |
.85 |
.82 |
| KLA/ATP | .86 |
.89 |
.77 |
.90 |
.78 |
.96 |
.88 |
|
| Itga4 | 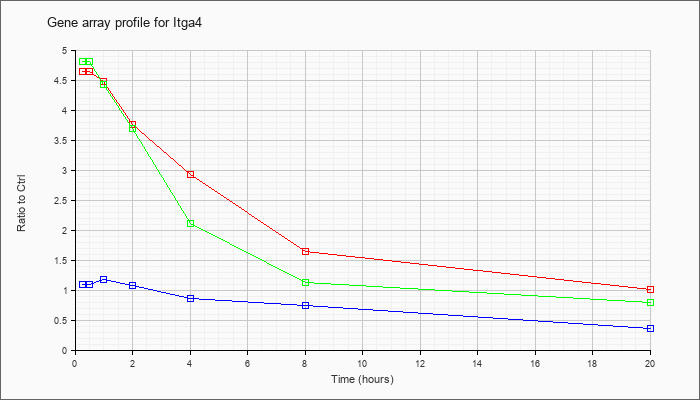 |
NM_010576 |
integrin alpha 4 (Itga4), mRNA [NM_010576] |
KLA | 4.65 |
4.83 |
4.47 |
3.76 |
2.93 |
1.65 |
1.01 |
| ATP | 1.10 |
1.12 |
1.17 |
1.08 |
.86 |
.75 |
.36 |
| KLA/ATP | 4.82 |
5.02 |
4.43 |
3.69 |
2.11 |
1.13 |
.79 |
|
| Itgam | 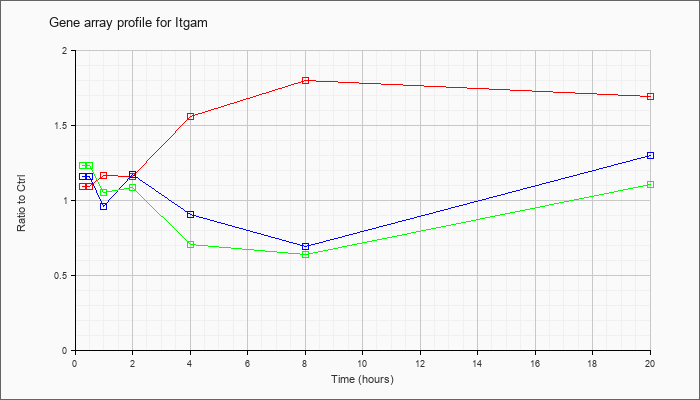 |
NM_001082960 |
integrin alpha M (Itgam), transcript variant 1, mRNA [NM_001082960] |
KLA | 1.04 |
1.12 |
1.14 |
1.07 |
1.54 |
1.80 |
1.72 |
| ATP | 1.17 |
1.19 |
.97 |
1.16 |
.86 |
.65 |
1.33 |
| KLA/ATP | 1.19 |
1.19 |
.94 |
1.06 |
.64 |
.59 |
1.08 |
|
| Itgam |  |
NM_008401 |
integrin alpha M (Itgam), transcript variant 2, mRNA [NM_008401] |
KLA | 1.20 |
1.16 |
1.21 |
1.34 |
1.59 |
1.79 |
1.64 |
| ATP | 1.15 |
1.16 |
.94 |
1.20 |
1.00 |
.78 |
1.25 |
| KLA/ATP | 1.32 |
1.37 |
1.29 |
1.14 |
.84 |
.75 |
1.16 |
|
| Itgb1 | 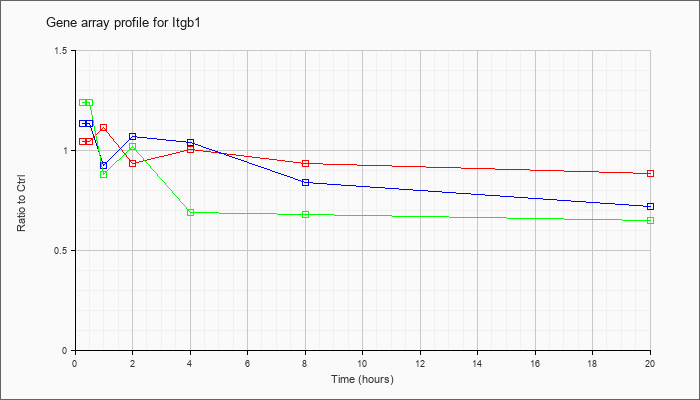 |
NM_010578 |
integrin beta 1 (fibronectin receptor beta) (Itgb1), mRNA [NM_010578] |
KLA | 1.05 |
1.09 |
1.13 |
.92 |
1.01 |
.93 |
.88 |
| ATP | 1.14 |
1.30 |
.92 |
1.07 |
1.04 |
.83 |
.70 |
| KLA/ATP | 1.26 |
1.30 |
.87 |
1.02 |
.66 |
.65 |
.62 |
|
| Itgb1 |  |
U37029 |
integrin beta1D mRNA, partial cds. [U37029] |
KLA | .93 |
.95 |
.96 |
1.01 |
.97 |
1.01 |
.98 |
| ATP | 1.05 |
.99 |
.99 |
1.03 |
1.03 |
.97 |
.94 |
| KLA/ATP | .99 |
.98 |
1.04 |
.97 |
1.02 |
1.01 |
.99 |
|
| Itgb2 | 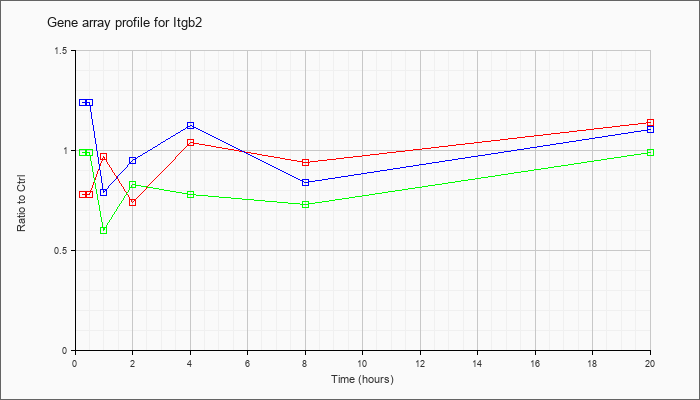 |
NM_008404 |
integrin beta 2 (Itgb2), mRNA [NM_008404] |
KLA | .78 |
.78 |
.97 |
.74 |
1.04 |
.94 |
1.14 |
| ATP | 1.24 |
1.23 |
.79 |
.95 |
1.12 |
.84 |
1.10 |
| KLA/ATP | .99 |
1.01 |
.60 |
.83 |
.78 |
.73 |
.99 |
|
| Itgb2l | 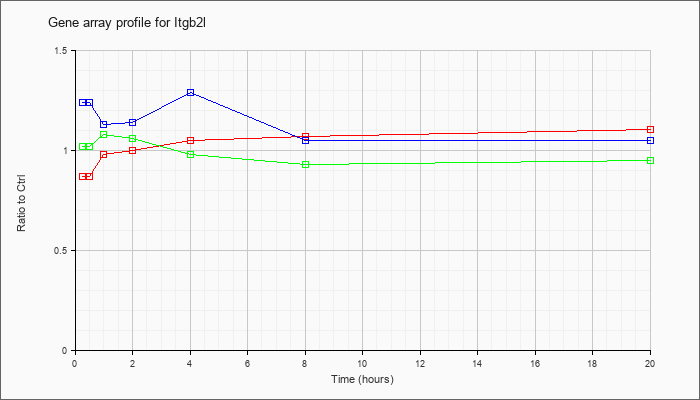 |
NM_008405 |
integrin beta 2-like (Itgb2l), mRNA [NM_008405] |
KLA | .87 |
.98 |
.98 |
1.00 |
1.05 |
1.07 |
1.10 |
| ATP | 1.24 |
1.23 |
1.13 |
1.14 |
1.29 |
1.05 |
1.05 |
| KLA/ATP | 1.02 |
1.08 |
1.08 |
1.06 |
.98 |
.93 |
.95 |
|
| Jak1 | 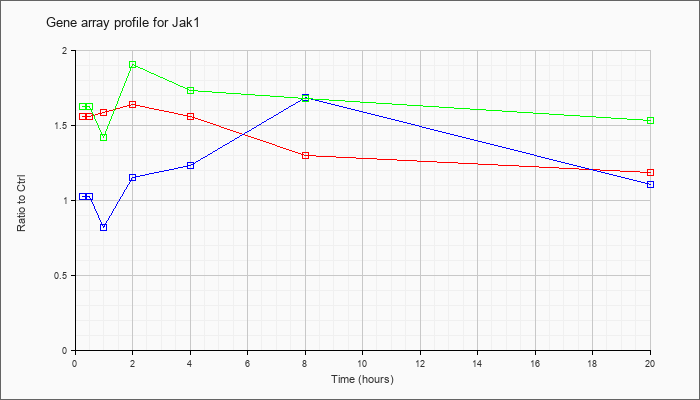 |
AK029232 |
0 day neonate head cDNA, RIKEN full-length enriched library, clone:4831433J01 product:Janus kinase 1, full insert sequence. [AK029232] |
KLA | 1.25 |
1.13 |
1.19 |
1.31 |
1.24 |
1.17 |
1.14 |
| ATP | 1.01 |
1.11 |
.96 |
1.38 |
1.06 |
1.26 |
1.05 |
| KLA/ATP | 1.19 |
1.29 |
1.27 |
1.59 |
1.18 |
1.38 |
1.24 |
|
| Jak1 |  |
NM_146145 |
Janus kinase 1 (Jak1), mRNA [NM_146145] |
KLA | 1.71 |
1.75 |
1.79 |
1.81 |
1.72 |
1.37 |
1.21 |
| ATP | 1.03 |
1.01 |
.75 |
1.04 |
1.32 |
1.90 |
1.14 |
| KLA/ATP | 1.84 |
1.73 |
1.49 |
2.07 |
2.01 |
1.83 |
1.68 |
|
| Jak2 | 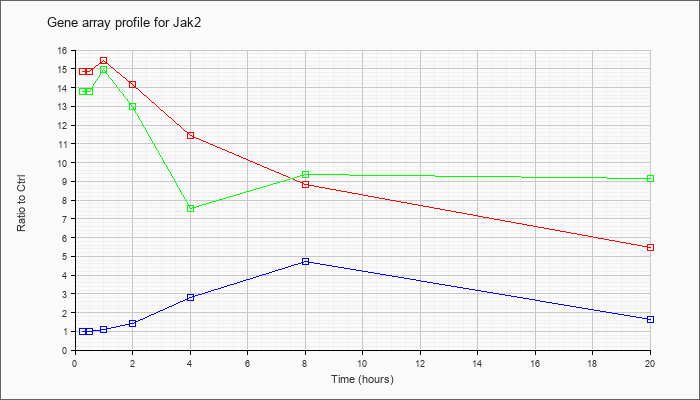 |
NM_008413 |
Janus kinase 2 (Jak2), transcript variant 1, mRNA [NM_008413] |
KLA | 14.86 |
15.05 |
15.42 |
14.15 |
11.46 |
8.81 |
5.47 |
| ATP | .99 |
1.06 |
1.09 |
1.43 |
2.83 |
4.71 |
1.62 |
| KLA/ATP | 13.77 |
14.74 |
14.94 |
12.97 |
7.53 |
9.34 |
9.17 |
|
| Jun | 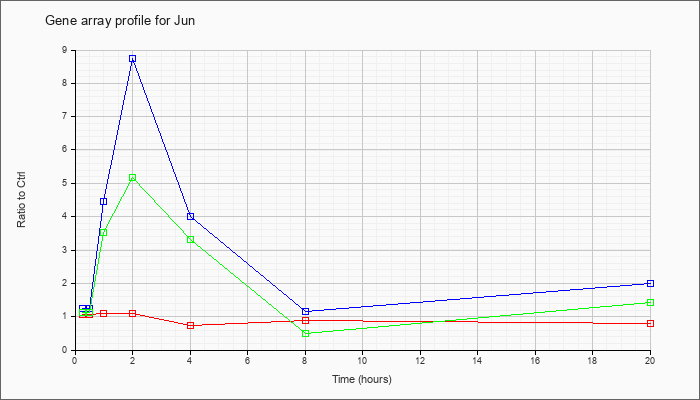 |
NM_010591 |
Jun oncogene (Jun), mRNA [NM_010591] |
KLA | 1.08 |
.96 |
1.09 |
1.09 |
.75 |
.88 |
.81 |
| ATP | 1.26 |
1.48 |
4.45 |
8.73 |
4.02 |
1.16 |
1.99 |
| KLA/ATP | 1.12 |
1.38 |
3.52 |
5.17 |
3.32 |
.49 |
1.42 |
|
| Map3k7 | 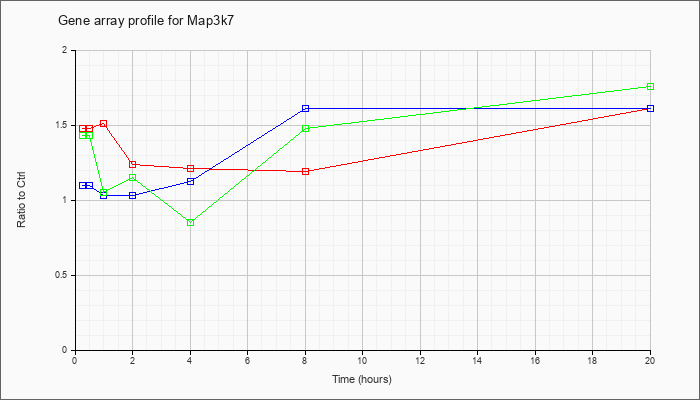 |
NM_172688 |
mitogen-activated protein kinase kinase kinase 7 (Map3k7), mRNA [NM_172688] |
KLA | 1.48 |
1.49 |
1.51 |
1.24 |
1.21 |
1.19 |
1.61 |
| ATP | 1.10 |
1.16 |
1.03 |
1.03 |
1.12 |
1.61 |
1.61 |
| KLA/ATP | 1.43 |
1.56 |
1.05 |
1.15 |
.85 |
1.48 |
1.76 |
|
Map3k7ip
1 | 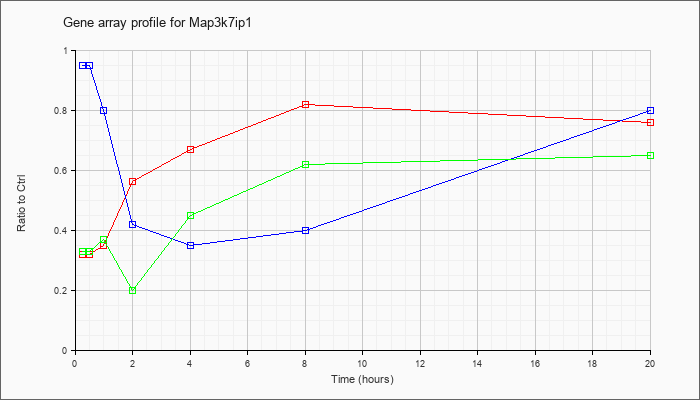 |
NM_025609 |
mitogen-activated protein kinase kinase kinase 7 interacting protein 1 (Map3k7ip1), mRNA [NM_025609] |
KLA | .32 |
.30 |
.35 |
.56 |
.67 |
.82 |
.76 |
| ATP | .95 |
.81 |
.80 |
.42 |
.35 |
.40 |
.80 |
| KLA/ATP | .33 |
.30 |
.37 |
.20 |
.45 |
.62 |
.65 |
|
Map3k7ip
2 | 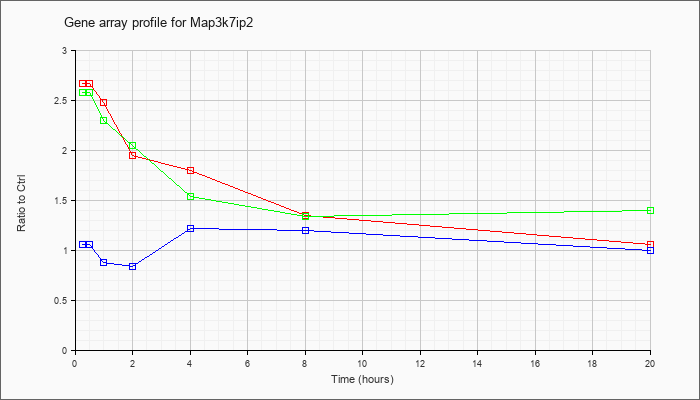 |
AK086230 |
15 days embryo head cDNA, RIKEN full-length enriched library, clone:D930015C19 product:inferred: Mus musculus, Similar to TAK1-binding protein 2; KIAA0733 protein, clone MGC:8010 IMAGE:3586132, mRNA, complete cds, full insert sequence. [A |
KLA | 1.14 |
1.02 |
1.07 |
1.03 |
1.13 |
1.02 |
.96 |
| ATP | 1.00 |
.88 |
.92 |
.90 |
1.06 |
.96 |
.94 |
| KLA/ATP | 1.05 |
1.00 |
.90 |
.98 |
1.02 |
.97 |
.97 |
|
Map3k7ip
2 |  |
NM_138667 |
mitogen-activated protein kinase kinase kinase 7 interacting protein 2 (Map3k7ip2), mRNA [NM_138667] |
KLA | 3.43 |
3.22 |
3.19 |
2.41 |
2.13 |
1.51 |
1.10 |
| ATP | 1.08 |
1.08 |
.86 |
.81 |
1.30 |
1.32 |
1.02 |
| KLA/ATP | 3.34 |
3.66 |
3.00 |
2.58 |
1.80 |
1.52 |
1.61 |
|
| Mapk1 | 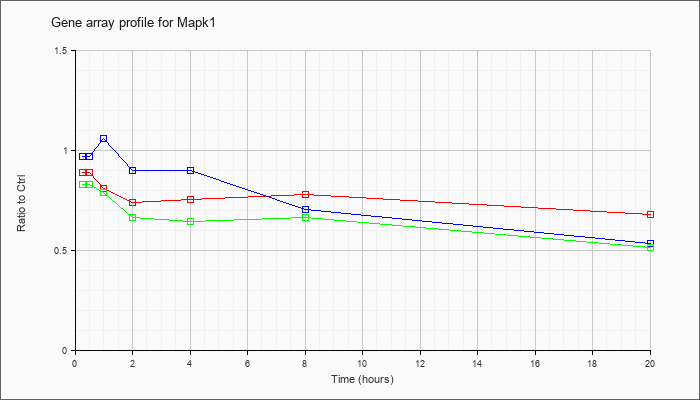 |
NM_001038663 |
mitogen-activated protein kinase 1 (Mapk1), transcript variant 2, mRNA [NM_001038663] |
KLA | .97 |
1.03 |
.93 |
.86 |
.79 |
1.01 |
1.04 |
| ATP | .96 |
1.18 |
1.01 |
1.13 |
.84 |
1.04 |
1.16 |
| KLA/ATP | .99 |
.99 |
.84 |
.87 |
.63 |
.96 |
1.26 |
|
| Mapk1 |  |
NM_011949 |
mitogen-activated protein kinase 1 (Mapk1), transcript variant 1, mRNA [NM_011949] |
KLA | .88 |
.80 |
.80 |
.73 |
.75 |
.76 |
.65 |
| ATP | .97 |
.96 |
1.06 |
.88 |
.90 |
.68 |
.48 |
| KLA/ATP | .82 |
.81 |
.78 |
.65 |
.64 |
.64 |
.45 |
|
| Mapk11 | 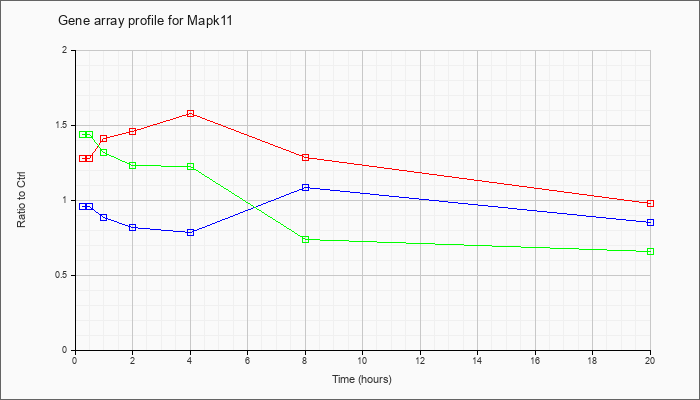 |
NM_011161 |
mitogen-activated protein kinase 11 (Mapk11), mRNA [NM_011161] |
KLA | 1.28 |
1.30 |
1.41 |
1.46 |
1.58 |
1.29 |
.98 |
| ATP | .96 |
.95 |
.89 |
.82 |
.79 |
1.09 |
.85 |
| KLA/ATP | 1.44 |
1.42 |
1.32 |
1.23 |
1.23 |
.74 |
.66 |
|
| Mapk12 | 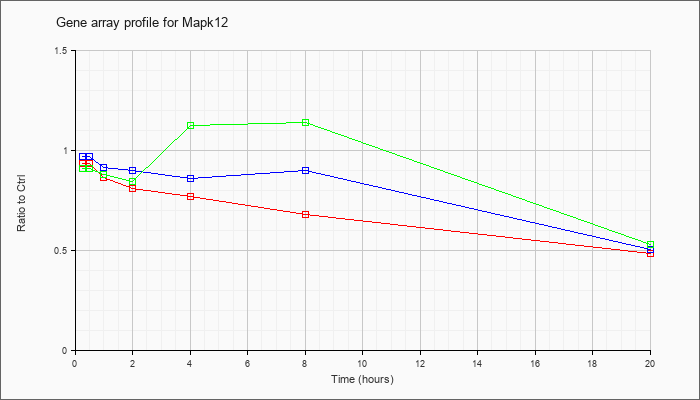 |
NM_013871 |
mitogen-activated protein kinase 12 (Mapk12), mRNA [NM_013871] |
KLA | .94 |
.86 |
.87 |
.81 |
.77 |
.68 |
.49 |
| ATP | .97 |
.83 |
.92 |
.90 |
.86 |
.90 |
.51 |
| KLA/ATP | .91 |
.89 |
.88 |
.85 |
1.12 |
1.14 |
.53 |
|
| Mapk13 | 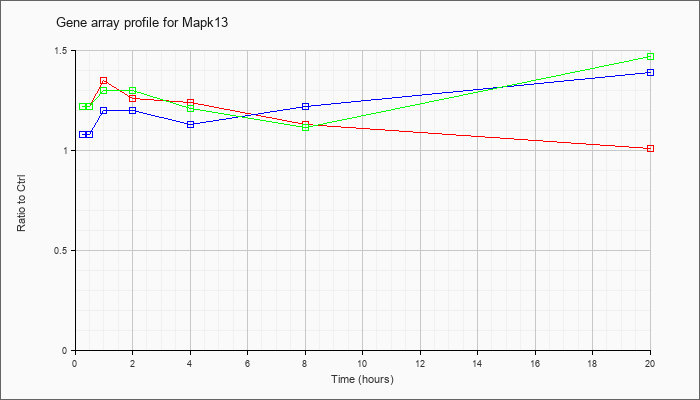 |
NM_011950 |
mitogen-activated protein kinase 13 (Mapk13), mRNA [NM_011950] |
KLA | 1.22 |
1.21 |
1.35 |
1.26 |
1.24 |
1.13 |
1.01 |
| ATP | 1.08 |
1.00 |
1.20 |
1.20 |
1.13 |
1.22 |
1.39 |
| KLA/ATP | 1.22 |
1.27 |
1.30 |
1.30 |
1.21 |
1.11 |
1.47 |
|
| Mapk14 | 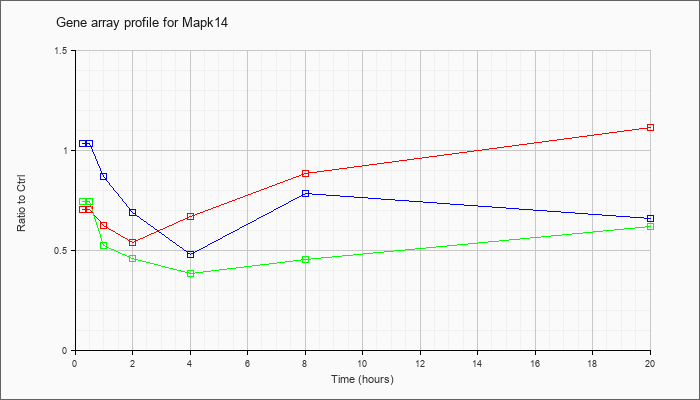 |
NM_011951 |
mitogen-activated protein kinase 14 (Mapk14), mRNA [NM_011951] |
KLA | .70 |
.70 |
.62 |
.54 |
.67 |
.88 |
1.11 |
| ATP | 1.03 |
.98 |
.87 |
.69 |
.48 |
.78 |
.66 |
| KLA/ATP | .74 |
.68 |
.52 |
.46 |
.38 |
.45 |
.62 |
|
| Mapk3 | 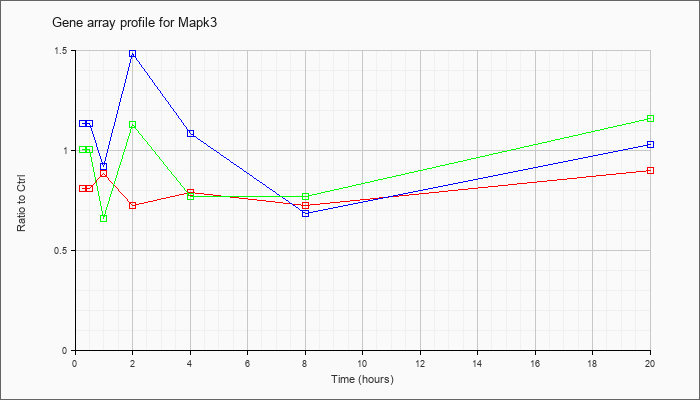 |
NM_011952 |
mitogen-activated protein kinase 3 (Mapk3), mRNA [NM_011952] |
KLA | .81 |
.85 |
.88 |
.72 |
.79 |
.73 |
.90 |
| ATP | 1.13 |
1.30 |
.92 |
1.48 |
1.08 |
.68 |
1.03 |
| KLA/ATP | 1.01 |
1.01 |
.66 |
1.13 |
.77 |
.77 |
1.16 |
|
Marcksl1
| 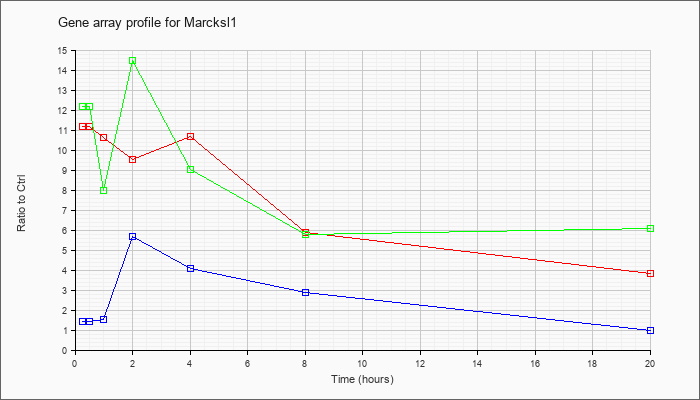 |
NM_010807 |
MARCKS-like 1 (Marcksl1), mRNA [NM_010807] |
KLA | 11.17 |
9.95 |
10.63 |
9.53 |
10.67 |
5.90 |
3.84 |
| ATP | 1.42 |
1.91 |
1.55 |
5.70 |
4.10 |
2.89 |
.97 |
| KLA/ATP | 12.20 |
13.11 |
7.97 |
14.47 |
9.03 |
5.78 |
6.06 |
|
Mm.24936
4 | 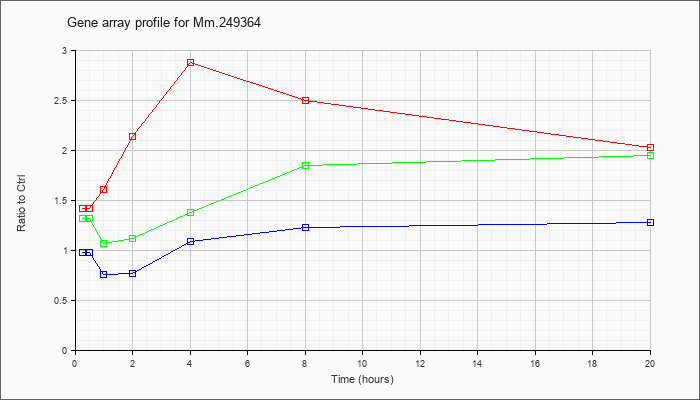 |
133892283 |
Unknown |
KLA | 1.42 |
1.41 |
1.61 |
2.14 |
2.88 |
2.49 |
2.03 |
| ATP | .98 |
.98 |
.76 |
.77 |
1.09 |
1.23 |
1.28 |
| KLA/ATP | 1.32 |
1.32 |
1.07 |
1.12 |
1.38 |
1.85 |
1.95 |
|
Mm.27970
| 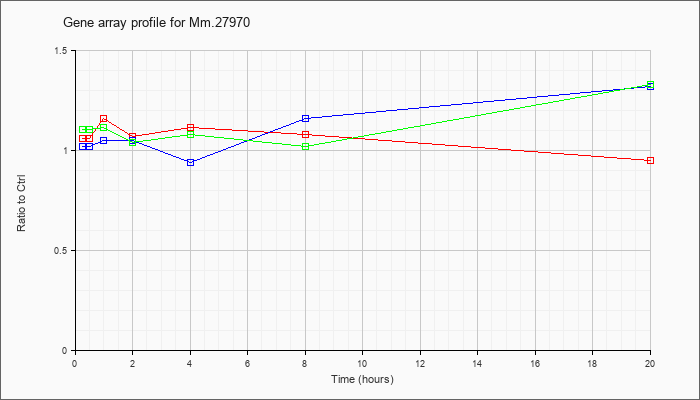 |
13994135 |
Unknown |
KLA | 1.06 |
1.11 |
1.16 |
1.07 |
1.11 |
1.08 |
.95 |
| ATP | 1.02 |
1.09 |
1.05 |
1.05 |
.94 |
1.16 |
1.32 |
| KLA/ATP | 1.10 |
1.19 |
1.11 |
1.04 |
1.08 |
1.02 |
1.33 |
|
Mm.38049
| 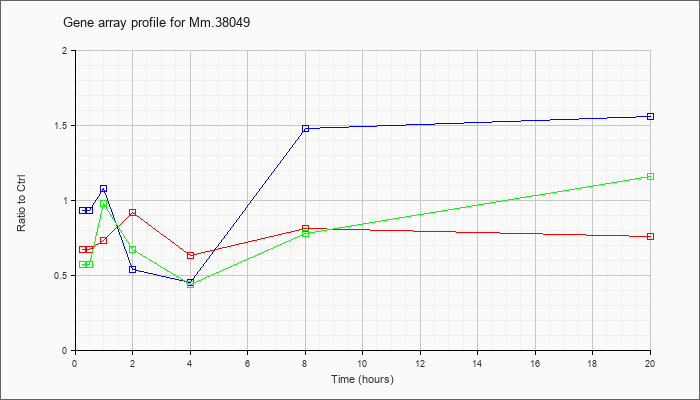 |
118130391 |
Unknown |
KLA | .67 |
.65 |
.73 |
.92 |
.63 |
.81 |
.76 |
| ATP | .93 |
.92 |
1.08 |
.54 |
.45 |
1.48 |
1.56 |
| KLA/ATP | .57 |
.65 |
.98 |
.67 |
.44 |
.78 |
1.16 |
|
Mm.42529
6 | 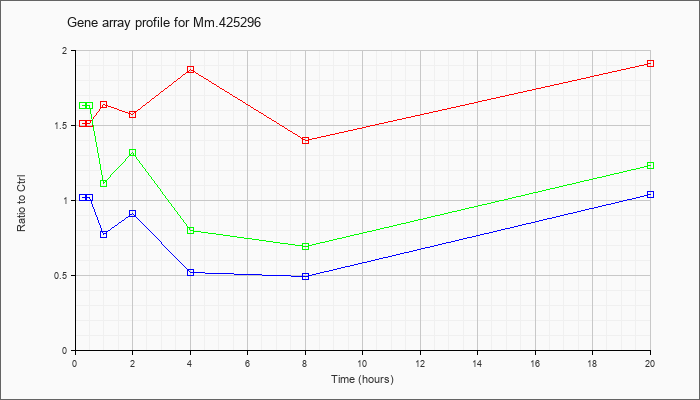 |
170172552 |
Unknown |
KLA | 1.51 |
1.59 |
1.64 |
1.57 |
1.87 |
1.40 |
1.91 |
| ATP | 1.02 |
1.04 |
.77 |
.91 |
.52 |
.49 |
1.04 |
| KLA/ATP | 1.63 |
1.53 |
1.11 |
1.32 |
.80 |
.69 |
1.23 |
|
Mm.45825
9 | 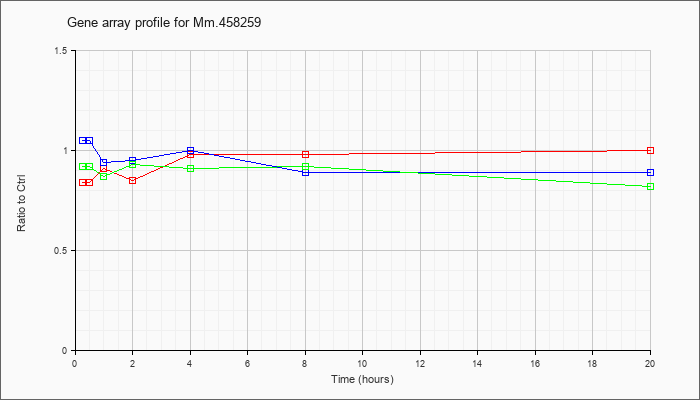 |
116292181 |
Unknown |
KLA | .84 |
.86 |
.91 |
.85 |
.98 |
.98 |
1.00 |
| ATP | 1.05 |
1.06 |
.94 |
.95 |
1.00 |
.89 |
.89 |
| KLA/ATP | .92 |
.84 |
.87 |
.93 |
.91 |
.92 |
.82 |
|
Mm.46002
4 | 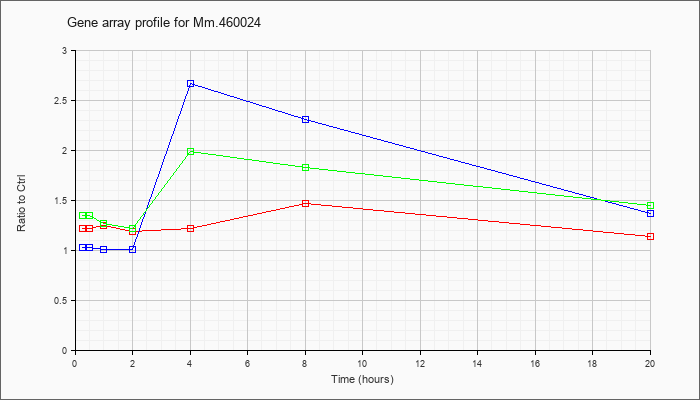 |
38348245 |
Unknown |
KLA | 1.22 |
1.25 |
1.25 |
1.19 |
1.22 |
1.47 |
1.14 |
| ATP | 1.03 |
.94 |
1.01 |
1.01 |
2.67 |
2.31 |
1.37 |
| KLA/ATP | 1.35 |
1.19 |
1.27 |
1.22 |
1.99 |
1.83 |
1.45 |
|
| Mm.874 | 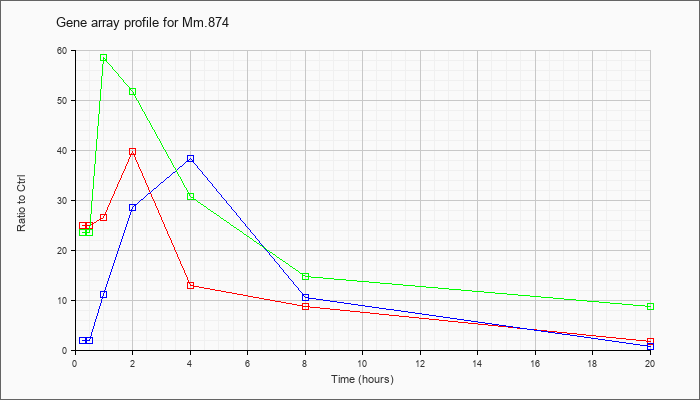 |
6754317 |
Unknown |
KLA | 24.98 |
24.87 |
26.43 |
39.61 |
12.88 |
8.64 |
1.63 |
| ATP | 1.97 |
3.50 |
11.05 |
28.47 |
38.30 |
10.44 |
.65 |
| KLA/ATP | 23.45 |
29.37 |
58.52 |
51.77 |
30.64 |
14.61 |
8.63 |
|
| Myd88 | 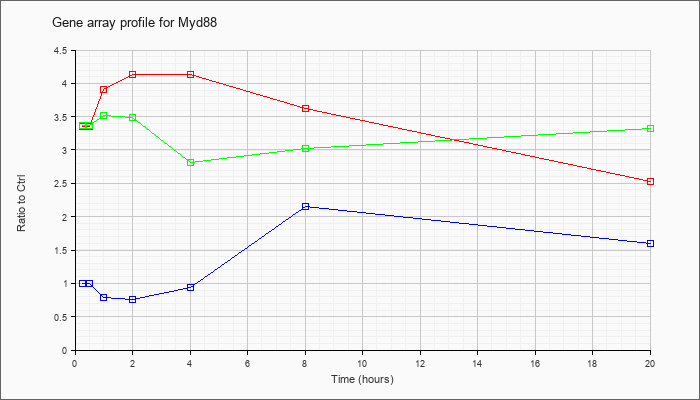 |
NM_010851 |
myeloid differentiation primary response gene 88 (Myd88), mRNA [NM_010851] |
KLA | 3.35 |
3.51 |
3.91 |
4.13 |
4.13 |
3.62 |
2.53 |
| ATP | .99 |
.96 |
.79 |
.75 |
.93 |
2.15 |
1.60 |
| KLA/ATP | 3.37 |
3.52 |
3.52 |
3.48 |
2.81 |
3.03 |
3.33 |
|
| Ncf1 |  |
NM_010876 |
neutrophil cytosolic factor 1 (Ncf1), mRNA [NM_010876] |
KLA | 1.62 |
1.71 |
1.81 |
1.70 |
1.99 |
1.64 |
2.07 |
| ATP | 1.02 |
1.06 |
.78 |
1.02 |
.64 |
.56 |
1.08 |
| KLA/ATP | 1.87 |
1.81 |
1.22 |
1.73 |
1.00 |
.80 |
1.49 |
|
| Ncf2 | 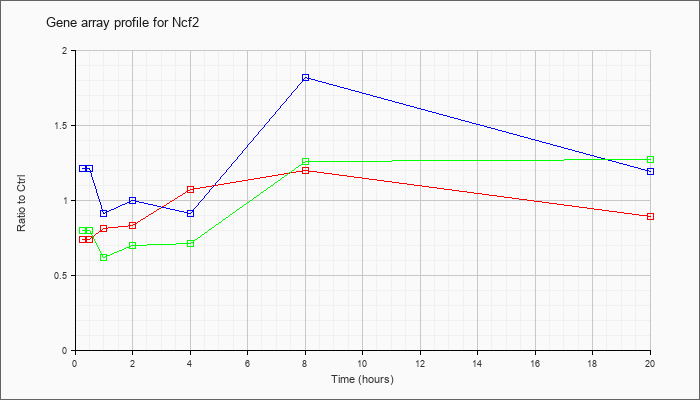 |
NM_010877 |
neutrophil cytosolic factor 2 (Ncf2), mRNA [NM_010877] |
KLA | .74 |
.69 |
.81 |
.83 |
1.07 |
1.20 |
.89 |
| ATP | 1.21 |
1.13 |
.91 |
1.00 |
.91 |
1.82 |
1.19 |
| KLA/ATP | .80 |
.74 |
.62 |
.70 |
.71 |
1.26 |
1.27 |
|
| Ncf4 | 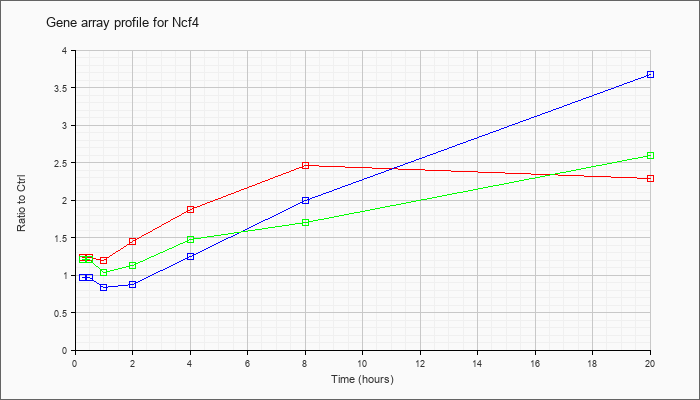 |
NM_008677 |
neutrophil cytosolic factor 4 (Ncf4), mRNA [NM_008677] |
KLA | 1.23 |
1.30 |
1.19 |
1.45 |
1.88 |
2.46 |
2.29 |
| ATP | .97 |
.83 |
.83 |
.87 |
1.25 |
1.99 |
3.67 |
| KLA/ATP | 1.21 |
1.11 |
1.04 |
1.13 |
1.48 |
1.70 |
2.60 |
|
| Nfkb1 | 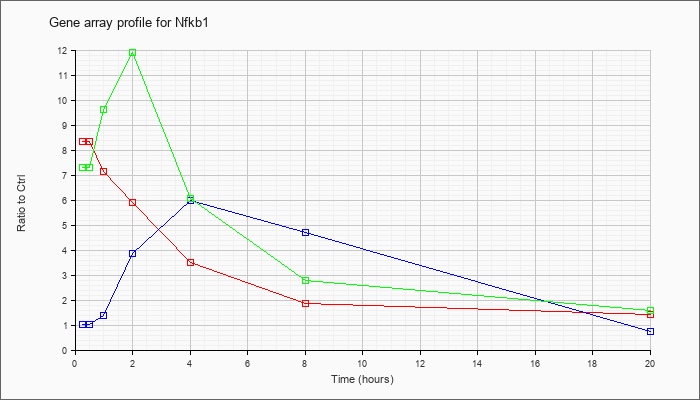 |
BC050841 |
nuclear factor of kappa light chain gene enhancer in B-cells 1, p105, mRNA (cDNA clone IMAGE:6392793), complete cds. [BC050841] |
KLA | 5.81 |
4.96 |
4.77 |
3.21 |
3.37 |
2.00 |
1.71 |
| ATP | 1.39 |
1.52 |
3.25 |
12.63 |
13.64 |
4.80 |
1.08 |
| KLA/ATP | 7.21 |
8.69 |
19.53 |
23.93 |
11.70 |
2.85 |
2.01 |
|
| Nfkb1 |  |
M57999 |
gb|Mouse transcription factor NF-kappa-B DNA binding subunit mRNA, complete cds. [M57999] |
KLA | 8.55 |
7.87 |
7.35 |
6.13 |
3.51 |
1.85 |
1.38 |
| ATP | .98 |
.96 |
1.21 |
3.06 |
5.29 |
4.67 |
.69 |
| KLA/ATP | 7.32 |
7.20 |
8.71 |
10.82 |
5.55 |
2.77 |
1.54 |
|
| Nfkbia | 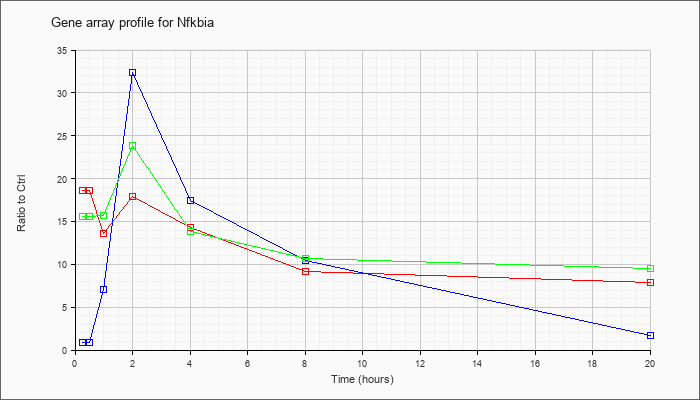 |
NM_010907 |
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha (Nfkbia), mRNA [NM_010907] |
KLA | 18.65 |
18.46 |
13.63 |
17.88 |
14.25 |
9.17 |
7.92 |
| ATP | .91 |
1.09 |
7.11 |
32.39 |
17.44 |
10.49 |
1.72 |
| KLA/ATP | 15.57 |
17.66 |
15.73 |
23.91 |
13.83 |
10.73 |
9.48 |
|
| Nfkbib | 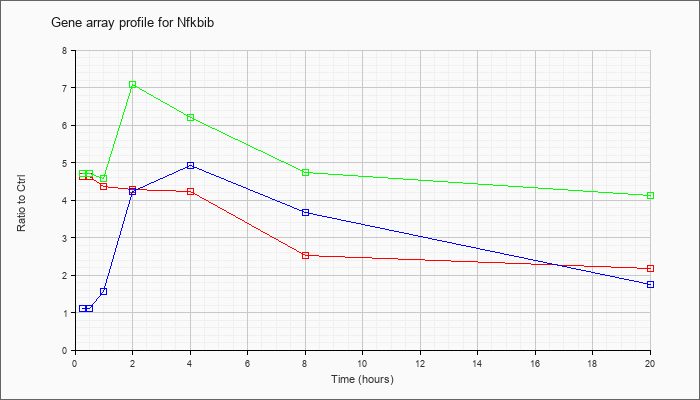 |
NM_010908 |
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, beta (Nfkbib), mRNA [NM_010908] |
KLA | 4.63 |
4.69 |
4.36 |
4.27 |
4.23 |
2.52 |
2.17 |
| ATP | 1.11 |
1.22 |
1.55 |
4.23 |
4.93 |
3.66 |
1.76 |
| KLA/ATP | 4.71 |
5.41 |
4.58 |
7.08 |
6.20 |
4.73 |
4.11 |
|
| Nos2 | 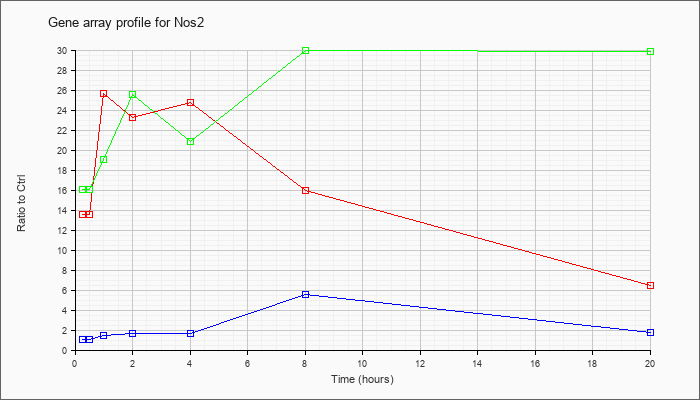 |
NM_010927 |
nitric oxide synthase 2, inducible, macrophage (Nos2), mRNA [NM_010927] |
KLA | 13.50 |
16.36 |
25.65 |
23.25 |
24.78 |
15.99 |
6.48 |
| ATP | 1.03 |
1.03 |
1.48 |
1.66 |
1.65 |
5.55 |
1.70 |
| KLA/ATP | 16.01 |
17.41 |
19.06 |
25.50 |
20.84 |
29.92 |
29.80 |
|
| Prkcb1 | 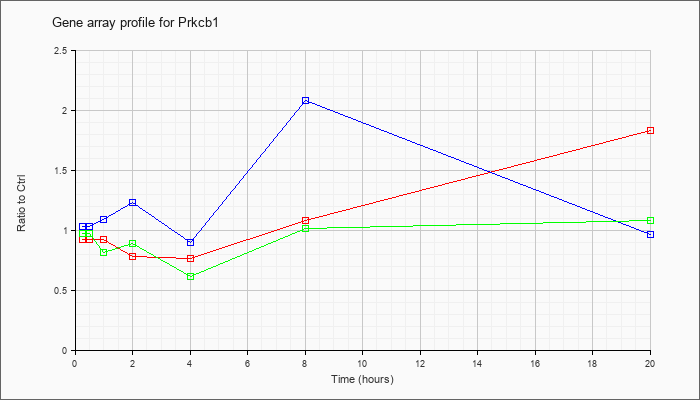 |
NM_008855 |
protein kinase C, beta 1 (Prkcb1), mRNA [NM_008855] |
KLA | .92 |
.95 |
.93 |
.78 |
.77 |
1.08 |
1.83 |
| ATP | 1.03 |
1.12 |
1.09 |
1.23 |
.90 |
2.08 |
.96 |
| KLA/ATP | .98 |
.99 |
.82 |
.89 |
.62 |
1.01 |
1.08 |
|
| Ptgs2 | 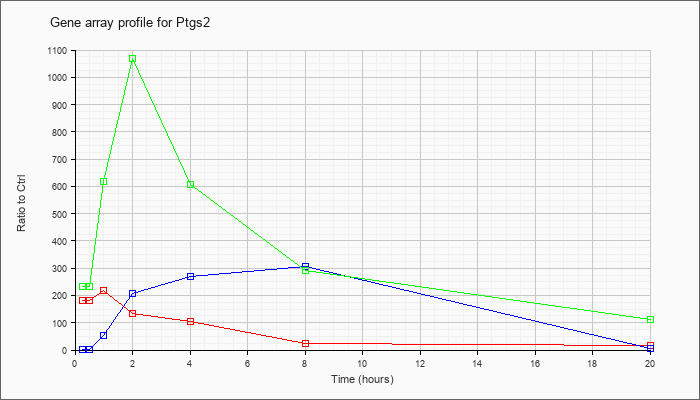 |
NM_011198 |
prostaglandin-endoperoxide synthase 2 (Ptgs2), mRNA [NM_011198] |
KLA | 181.04 |
220.88 |
218.58 |
132.79 |
103.05 |
22.69 |
18.02 |
| ATP | 2.47 |
9.45 |
52.13 |
208.62 |
271.22 |
306.11 |
6.86 |
| KLA/ATP | 233.39 |
387.29 |
617.86 |
###.## |
608.44 |
290.80 |
112.22 |
|
| Ptpn6 | 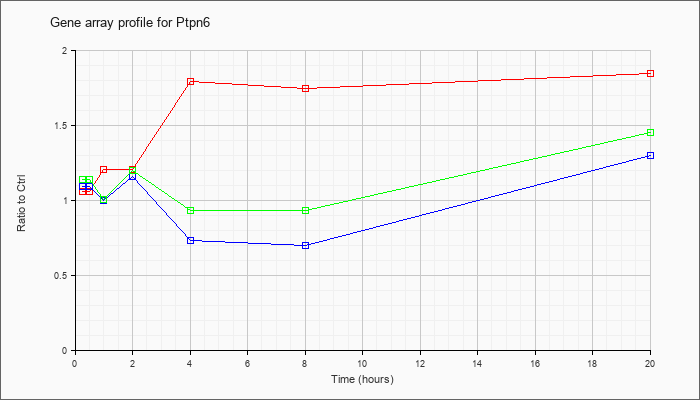 |
NM_001077705 |
protein tyrosine phosphatase, non-receptor type 6 (Ptpn6), transcript variant 2, mRNA [NM_001077705] |
KLA | .87 |
.90 |
1.03 |
1.05 |
1.83 |
1.96 |
2.34 |
| ATP | 1.06 |
1.09 |
.94 |
1.08 |
.51 |
.34 |
1.04 |
| KLA/ATP | .98 |
.93 |
.71 |
.95 |
.67 |
.64 |
1.44 |
|
| Ptpn6 |  |
NM_013545 |
protein tyrosine phosphatase, non-receptor type 6 (Ptpn6), transcript variant 1, mRNA [NM_013545] |
KLA | 1.24 |
1.20 |
1.46 |
1.42 |
2.43 |
2.27 |
2.10 |
| ATP | 1.18 |
1.20 |
1.10 |
1.49 |
.80 |
.58 |
1.09 |
| KLA/ATP | 1.36 |
1.42 |
1.27 |
1.66 |
1.12 |
.87 |
1.46 |
|
| Ptpn6 |  |
S63763 |
gb|me/Hcph (mev)=viable motheaten/hematopoietic cell protein-tyrosine phosphatase {insertion} [mice, mRNA Partial Mutant, 120 nt]. [S63763] |
KLA | 1.06 |
1.03 |
1.12 |
1.14 |
1.11 |
1.01 |
1.09 |
| ATP | 1.03 |
.97 |
.96 |
.90 |
.89 |
1.17 |
1.76 |
| KLA/ATP | 1.07 |
1.00 |
1.03 |
.98 |
1.00 |
1.29 |
1.44 |
|
| Rela | 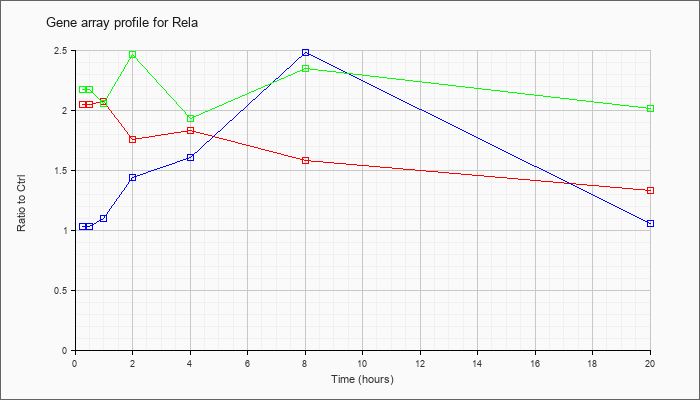 |
NM_009045 |
v-rel reticuloendotheliosis viral oncogene homolog A (avian) (Rela), mRNA [NM_009045] |
KLA | 2.05 |
2.05 |
2.07 |
1.75 |
1.83 |
1.58 |
1.33 |
| ATP | 1.03 |
1.08 |
1.10 |
1.44 |
1.61 |
2.48 |
1.05 |
| KLA/ATP | 2.17 |
2.27 |
2.05 |
2.47 |
1.93 |
2.35 |
2.02 |
|
| Stat1 |  |
NM_009283 |
signal transducer and activator of transcription 1 (Stat1), mRNA [NM_009283] |
KLA | 2.90 |
2.72 |
3.00 |
3.64 |
4.58 |
6.51 |
5.58 |
| ATP | 1.01 |
.92 |
.89 |
.72 |
.51 |
2.22 |
3.07 |
| KLA/ATP | 2.73 |
2.65 |
2.34 |
1.95 |
1.99 |
4.70 |
8.66 |
|
| Tgfb1 | 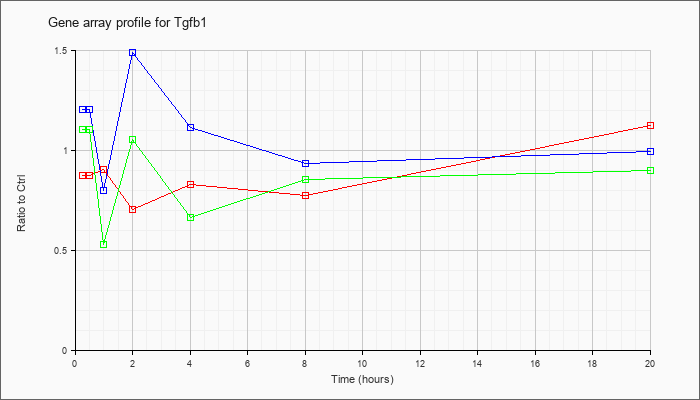 |
NM_011577 |
transforming growth factor, beta 1 (Tgfb1), mRNA [NM_011577] |
KLA | .87 |
.89 |
.90 |
.70 |
.83 |
.77 |
1.12 |
| ATP | 1.20 |
1.41 |
.80 |
1.49 |
1.11 |
.93 |
.99 |
| KLA/ATP | 1.10 |
1.20 |
.53 |
1.05 |
.66 |
.85 |
.90 |
|
| Tgfb2 | 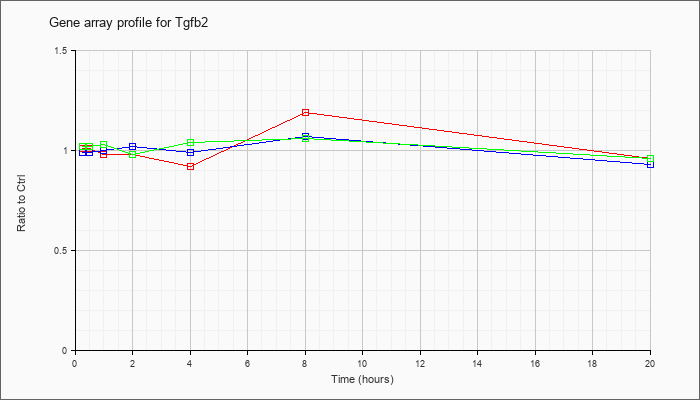 |
NM_009367 |
transforming growth factor, beta 2 (Tgfb2), mRNA [NM_009367] |
KLA | 1.01 |
.91 |
.98 |
.98 |
.92 |
1.19 |
.96 |
| ATP | .99 |
1.03 |
1.00 |
1.02 |
.99 |
1.07 |
.93 |
| KLA/ATP | 1.02 |
.99 |
1.03 |
.98 |
1.04 |
1.06 |
.96 |
|
| Tgfb3 | 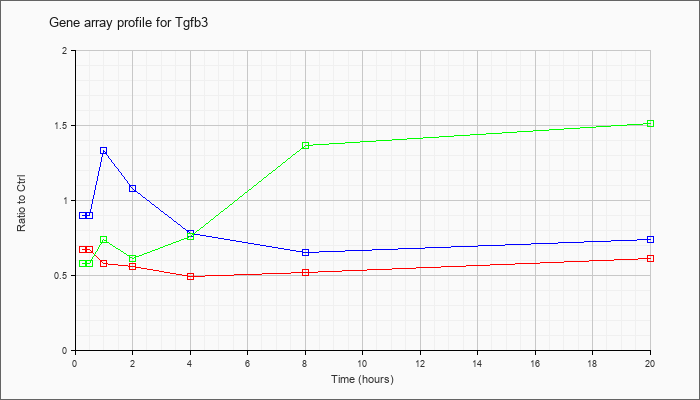 |
NM_009368 |
transforming growth factor, beta 3 (Tgfb3), mRNA [NM_009368] |
KLA | .67 |
.64 |
.58 |
.56 |
.49 |
.52 |
.61 |
| ATP | .90 |
.92 |
1.33 |
1.08 |
.78 |
.65 |
.74 |
| KLA/ATP | .58 |
.60 |
.74 |
.61 |
.76 |
1.36 |
1.51 |
|
| Tlr2 | 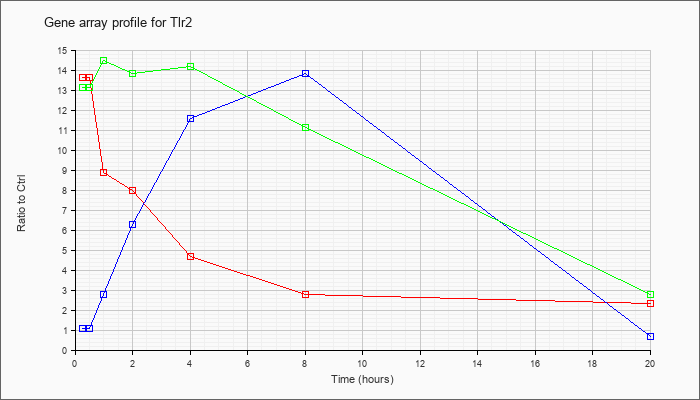 |
NM_011905 |
toll-like receptor 2 (Tlr2), mRNA [NM_011905] |
KLA | 13.65 |
11.82 |
8.87 |
7.95 |
4.67 |
2.78 |
2.33 |
| ATP | 1.06 |
1.24 |
2.76 |
6.28 |
11.55 |
13.82 |
.67 |
| KLA/ATP | 13.11 |
12.91 |
14.48 |
13.84 |
14.18 |
11.14 |
2.78 |
|
| Tlr4 | 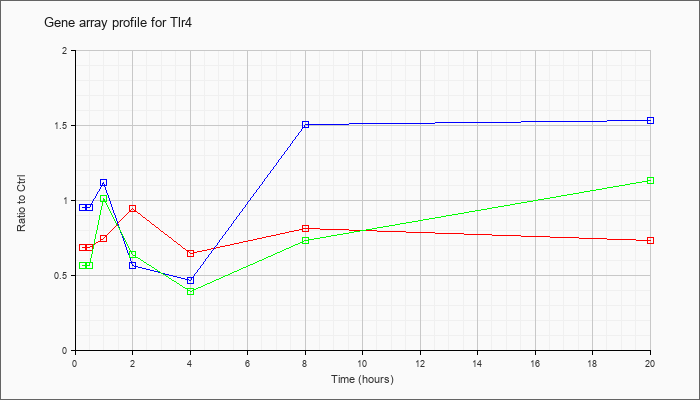 |
NM_021297 |
toll-like receptor 4 (Tlr4), mRNA [NM_021297] |
KLA | .69 |
.61 |
.75 |
.94 |
.64 |
.81 |
.73 |
| ATP | .95 |
.90 |
1.12 |
.56 |
.47 |
1.51 |
1.53 |
| KLA/ATP | .57 |
.62 |
1.01 |
.64 |
.39 |
.73 |
1.13 |
|
| Tnf | 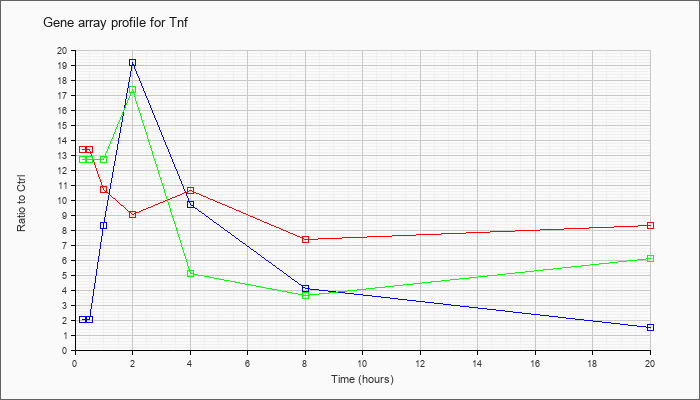 |
NM_013693 |
tumor necrosis factor (Tnf), mRNA [NM_013693] |
KLA | 13.35 |
12.76 |
10.71 |
9.03 |
10.64 |
7.39 |
8.32 |
| ATP | 2.01 |
2.48 |
8.27 |
19.20 |
9.72 |
4.09 |
1.49 |
| KLA/ATP | 12.72 |
15.76 |
12.73 |
17.34 |
5.10 |
3.61 |
6.07 |
|
| Traf6 | 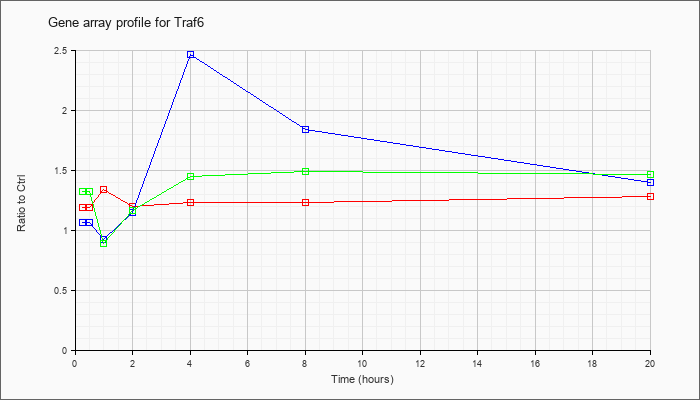 |
NM_009424 |
Tnf receptor-associated factor 6 (Traf6), mRNA [NM_009424] |
KLA | 1.19 |
1.28 |
1.34 |
1.20 |
1.23 |
1.23 |
1.28 |
| ATP | 1.06 |
1.12 |
.92 |
1.15 |
2.46 |
1.84 |
1.40 |
| KLA/ATP | 1.32 |
1.37 |
.89 |
1.16 |
1.45 |
1.49 |
1.46 |
|