| Gene symbol |
Time course plot |
Accession |
Description |
Treatment |
15min |
30min |
1hr |
2hr |
4hr |
8hr |
20hr |
AK050363
| 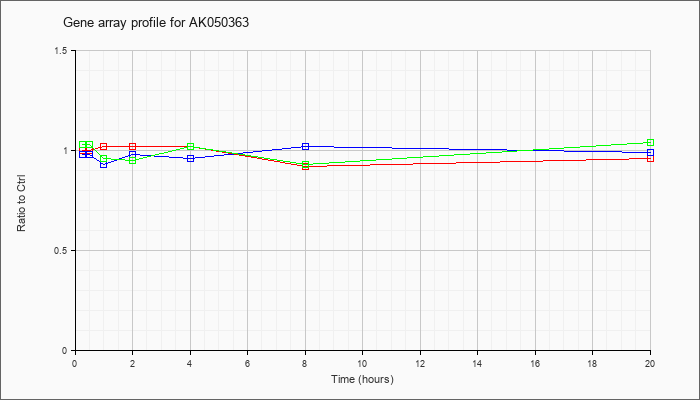 |
AK050363 |
adult male liver tumor cDNA, RIKEN full-length enriched library, clone:C730040N22 product:unclassifiable, full insert sequence. [AK050363] |
KLA | 1.00 |
.98 |
1.02 |
1.02 |
1.02 |
.92 |
.96 |
| ATP | .98 |
1.03 |
.93 |
.98 |
.96 |
1.02 |
.99 |
| KLA/ATP | 1.03 |
.96 |
.96 |
.95 |
1.02 |
.93 |
1.04 |
|
| Akt1 | 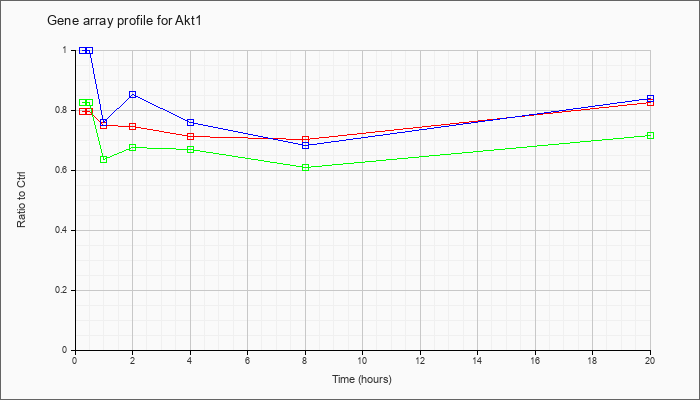 |
NM_009652 |
thymoma viral proto-oncogene 1 (Akt1), mRNA [NM_009652] |
KLA | .80 |
.80 |
.75 |
.75 |
.71 |
.70 |
.82 |
| ATP | 1.00 |
1.02 |
.76 |
.85 |
.76 |
.68 |
.84 |
| KLA/ATP | .82 |
.80 |
.64 |
.68 |
.67 |
.61 |
.72 |
|
| Akt2 | 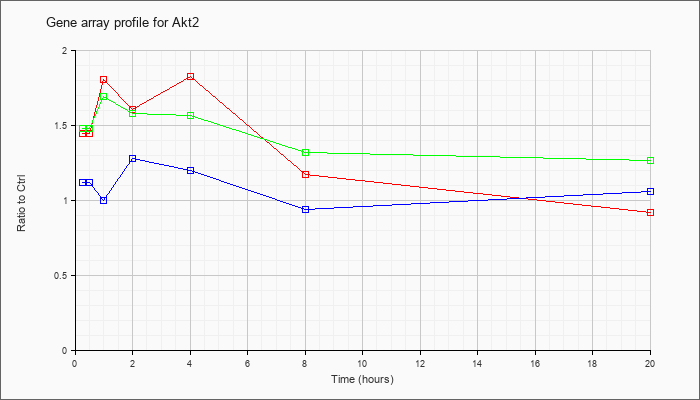 |
NM_007434 |
thymoma viral proto-oncogene 2 (Akt2), transcript variant 2, mRNA [NM_007434] |
KLA | 1.45 |
1.49 |
1.81 |
1.61 |
1.83 |
1.17 |
.92 |
| ATP | 1.12 |
1.12 |
1.00 |
1.28 |
1.20 |
.94 |
1.06 |
| KLA/ATP | 1.48 |
1.70 |
1.69 |
1.58 |
1.57 |
1.32 |
1.27 |
|
| Akt3 | 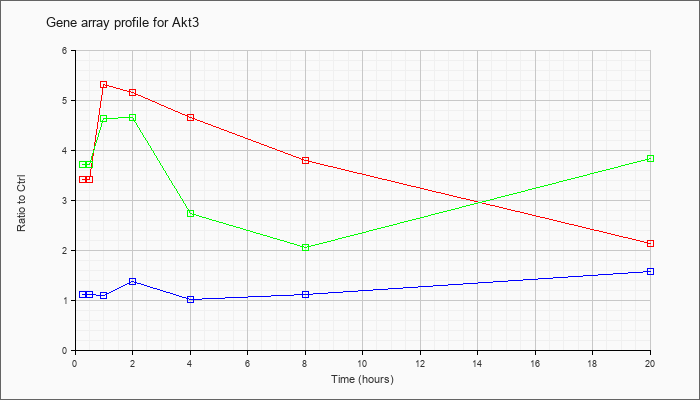 |
NM_011785 |
thymoma viral proto-oncogene 3 (Akt3), mRNA [NM_011785] |
KLA | 3.42 |
3.45 |
5.31 |
5.16 |
4.65 |
3.80 |
2.12 |
| ATP | 1.12 |
1.22 |
1.10 |
1.37 |
1.02 |
1.11 |
1.56 |
| KLA/ATP | 3.72 |
4.17 |
4.62 |
4.64 |
2.74 |
2.05 |
3.82 |
|
| Araf | 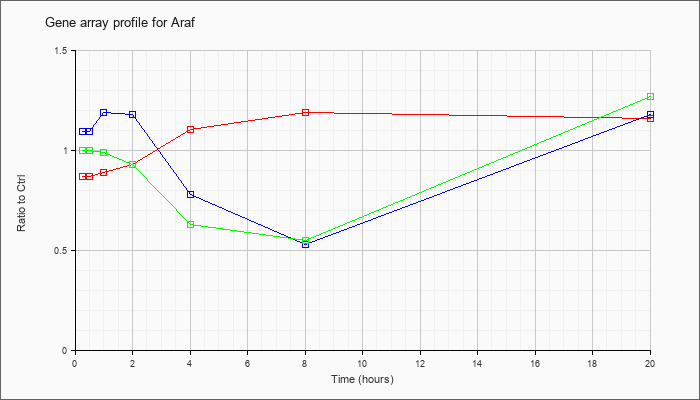 |
NM_009703 |
v-raf murine sarcoma 3611 viral oncogene homolog (Araf), mRNA [NM_009703] |
KLA | .87 |
.80 |
.89 |
.93 |
1.10 |
1.19 |
1.16 |
| ATP | 1.09 |
1.11 |
1.19 |
1.18 |
.78 |
.53 |
1.18 |
| KLA/ATP | 1.00 |
.95 |
.99 |
.93 |
.63 |
.55 |
1.27 |
|
| Arnt | 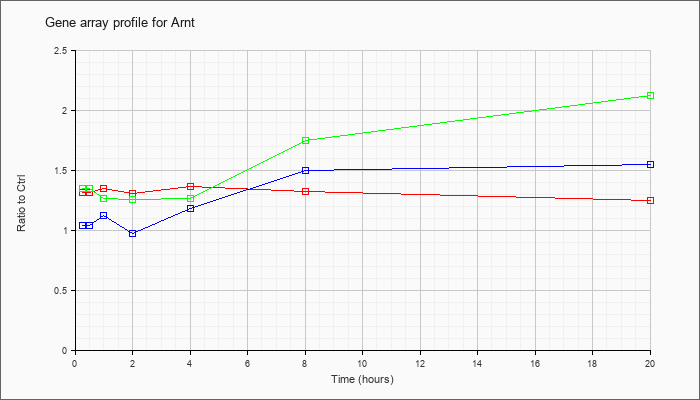 |
AK037762 |
16 days neonate thymus cDNA, RIKEN full-length enriched library, clone:A130047M16 product:aryl hydrocarbon receptor nuclear translocator, full insert sequence. [AK037762] |
KLA | 1.01 |
.87 |
.97 |
.92 |
1.13 |
.91 |
1.00 |
| ATP | 1.16 |
1.22 |
1.22 |
.84 |
1.57 |
1.19 |
1.05 |
| KLA/ATP | 1.06 |
1.06 |
1.08 |
.95 |
1.40 |
1.18 |
1.06 |
|
| Arnt |  |
NM_001037737 |
aryl hydrocarbon receptor nuclear translocator (Arnt), transcript variant 1, mRNA [NM_001037737] |
KLA | 1.23 |
1.27 |
1.32 |
1.23 |
1.34 |
1.33 |
1.30 |
| ATP | 1.02 |
1.09 |
.96 |
.90 |
.99 |
1.33 |
1.38 |
| KLA/ATP | 1.30 |
1.28 |
1.05 |
1.12 |
1.07 |
1.64 |
1.89 |
|
| Arnt |  |
U14333 |
aromatic hydrocarbon receptor nuclear translocator Arnt (ARNT) mRNA, complete cds. [U14333] |
KLA | 1.78 |
1.78 |
1.78 |
1.85 |
1.64 |
1.71 |
1.40 |
| ATP | .96 |
1.05 |
1.37 |
1.24 |
1.19 |
2.13 |
2.37 |
| KLA/ATP | 1.72 |
1.87 |
1.87 |
1.83 |
1.53 |
2.53 |
3.65 |
|
| Arnt2 | 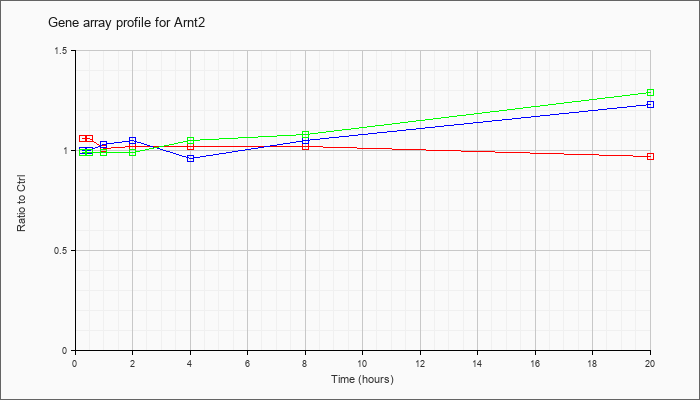 |
NM_007488 |
aryl hydrocarbon receptor nuclear translocator 2 (Arnt2), mRNA [NM_007488] |
KLA | 1.06 |
1.02 |
1.01 |
1.02 |
1.02 |
1.02 |
.97 |
| ATP | 1.00 |
1.00 |
1.03 |
1.05 |
.96 |
1.05 |
1.23 |
| KLA/ATP | .99 |
.97 |
.99 |
.99 |
1.05 |
1.08 |
1.29 |
|
BG242006
| 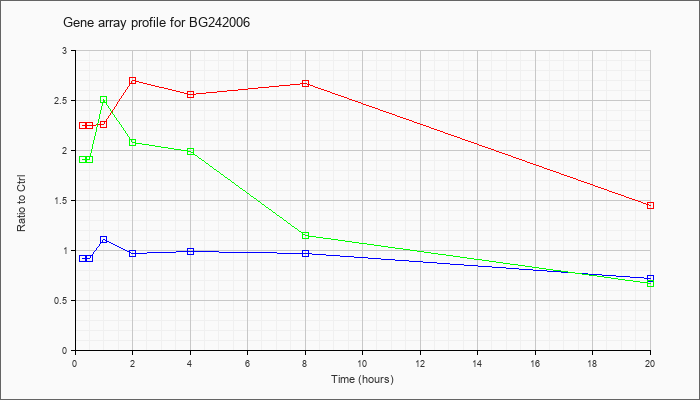 |
BG242006 |
gb|602354740F1 NCI_CGAP_Mam1 Mus musculus cDNA clone IMAGE:4483064 5. [BG242006] |
KLA | 2.25 |
2.23 |
2.26 |
2.70 |
2.56 |
2.67 |
1.45 |
| ATP | .92 |
.87 |
1.11 |
.97 |
.99 |
.97 |
.72 |
| KLA/ATP | 1.91 |
2.04 |
2.51 |
2.08 |
1.99 |
1.15 |
.67 |
|
| Braf |  |
NM_139294 |
Braf transforming gene (Braf), mRNA [NM_139294] |
KLA | 1.52 |
1.57 |
1.55 |
1.33 |
1.83 |
1.52 |
1.60 |
| ATP | 1.08 |
1.17 |
1.02 |
1.37 |
.95 |
1.15 |
.95 |
| KLA/ATP | 1.64 |
1.69 |
1.39 |
1.51 |
1.04 |
1.31 |
1.26 |
|
| Cdc42 | 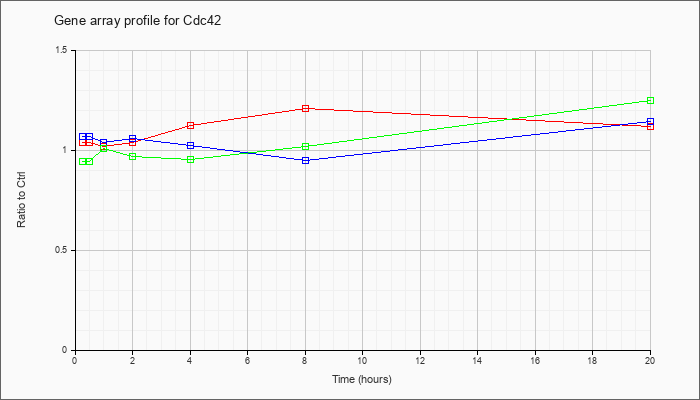 |
NM_009861 |
cell division cycle 42 homolog (S. cerevisiae) (Cdc42), mRNA [NM_009861] |
KLA | 1.04 |
1.01 |
1.02 |
1.04 |
1.12 |
1.21 |
1.12 |
| ATP | 1.07 |
1.10 |
1.04 |
1.06 |
1.02 |
.95 |
1.14 |
| KLA/ATP | .94 |
1.04 |
1.01 |
.97 |
.95 |
1.02 |
1.25 |
|
| Crebbp | 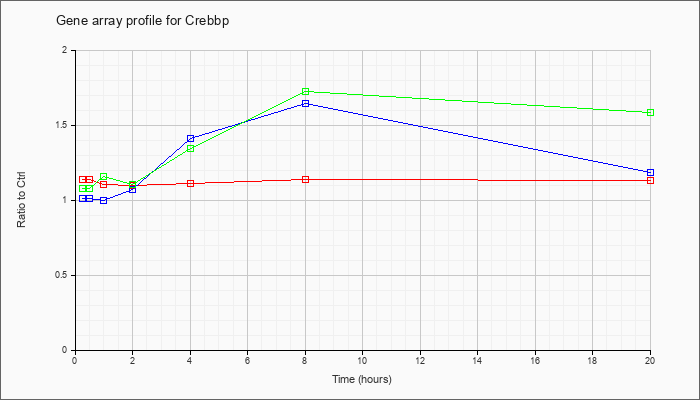 |
AK048818 |
0 day neonate cerebellum cDNA, RIKEN full-length enriched library, clone:C230072N21 product:hypothetical protein, full insert sequence [AK048818] |
KLA | 1.44 |
1.19 |
1.26 |
1.25 |
1.25 |
1.36 |
1.24 |
| ATP | .98 |
.90 |
1.04 |
1.11 |
1.95 |
2.72 |
1.51 |
| KLA/ATP | 1.17 |
1.27 |
1.45 |
1.28 |
1.98 |
2.80 |
2.52 |
|
| Crebbp |  |
NM_001025432 |
CREB binding protein (Crebbp), mRNA [NM_001025432] |
KLA | .97 |
1.01 |
.98 |
1.06 |
1.05 |
.99 |
1.07 |
| ATP | 1.07 |
1.05 |
.96 |
1.01 |
1.15 |
1.14 |
1.00 |
| KLA/ATP | 1.01 |
.97 |
.97 |
1.00 |
1.03 |
1.23 |
1.13 |
|
| Crebbp |  |
S66385 |
gb|CREB-binding protein [mice, brain, mRNA Partial, 7326 nt]. [S66385] |
KLA | 1.00 |
.98 |
1.08 |
.98 |
1.04 |
1.07 |
1.08 |
| ATP | .99 |
.98 |
.99 |
1.09 |
1.13 |
1.08 |
1.05 |
| KLA/ATP | 1.06 |
1.00 |
1.05 |
1.03 |
1.02 |
1.14 |
1.11 |
|
| Crk | 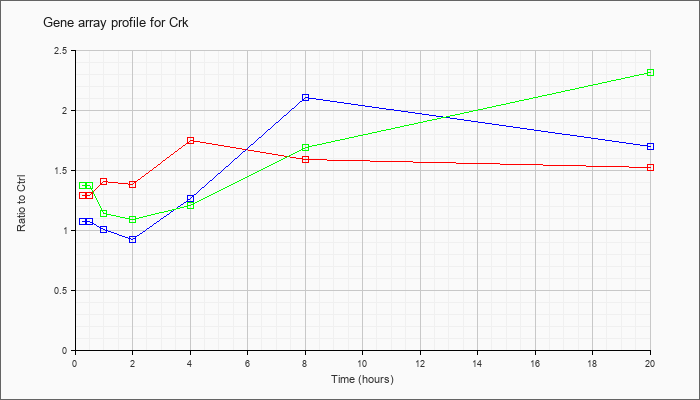 |
NM_133656 |
v-crk sarcoma virus CT10 oncogene homolog (avian) (Crk), mRNA [NM_133656] |
KLA | 1.29 |
1.32 |
1.41 |
1.38 |
1.75 |
1.59 |
1.52 |
| ATP | 1.08 |
1.17 |
1.01 |
.93 |
1.27 |
2.11 |
1.70 |
| KLA/ATP | 1.37 |
1.49 |
1.14 |
1.09 |
1.21 |
1.69 |
2.32 |
|
| Crkl | 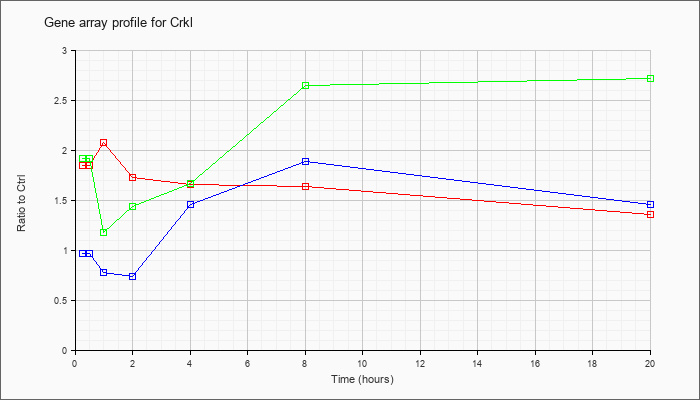 |
NM_007764 |
v-crk sarcoma virus CT10 oncogene homolog (avian)-like (Crkl), mRNA [NM_007764] |
KLA | 1.85 |
1.97 |
2.08 |
1.73 |
1.66 |
1.64 |
1.36 |
| ATP | .97 |
.94 |
.78 |
.74 |
1.46 |
1.89 |
1.46 |
| KLA/ATP | 1.92 |
1.88 |
1.18 |
1.44 |
1.67 |
2.65 |
2.72 |
|
| Cul2 | 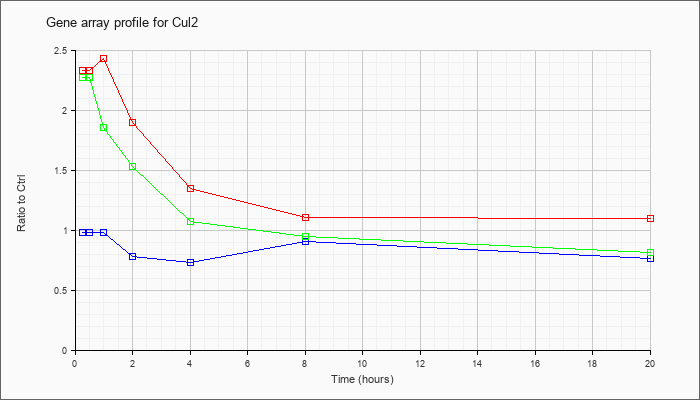 |
NM_029402 |
cullin 2 (Cul2), mRNA [NM_029402] |
KLA | 2.33 |
2.37 |
2.43 |
1.90 |
1.34 |
1.10 |
1.09 |
| ATP | .98 |
.92 |
.98 |
.78 |
.73 |
.90 |
.76 |
| KLA/ATP | 2.27 |
2.19 |
1.86 |
1.53 |
1.07 |
.95 |
.81 |
|
| Egln1 | 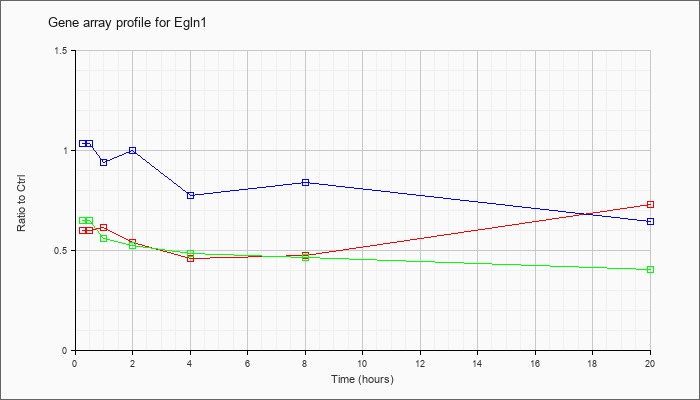 |
NM_053207 |
EGL nine homolog 1 (C. elegans) (Egln1), mRNA [NM_053207] |
KLA | .60 |
.60 |
.62 |
.54 |
.46 |
.48 |
.73 |
| ATP | 1.04 |
1.10 |
.94 |
1.00 |
.78 |
.84 |
.65 |
| KLA/ATP | .65 |
.64 |
.56 |
.53 |
.49 |
.47 |
.41 |
|
| Egln2 | 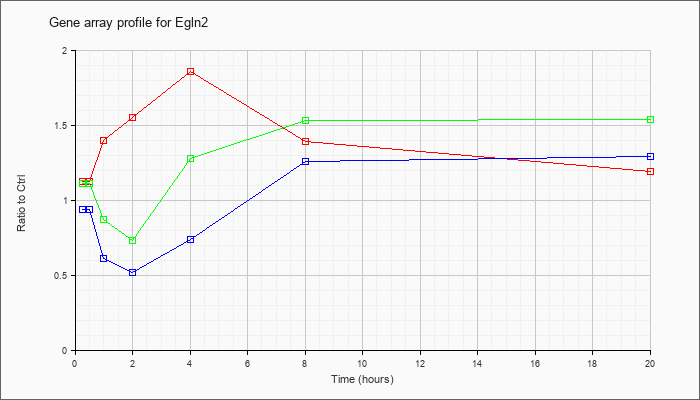 |
NM_053208 |
EGL nine homolog 2 (C. elegans) (Egln2), mRNA [NM_053208] |
KLA | 1.12 |
1.22 |
1.40 |
1.55 |
1.86 |
1.39 |
1.19 |
| ATP | .94 |
.82 |
.61 |
.52 |
.74 |
1.26 |
1.29 |
| KLA/ATP | 1.11 |
1.04 |
.87 |
.73 |
1.28 |
1.53 |
1.54 |
|
| Egln3 | 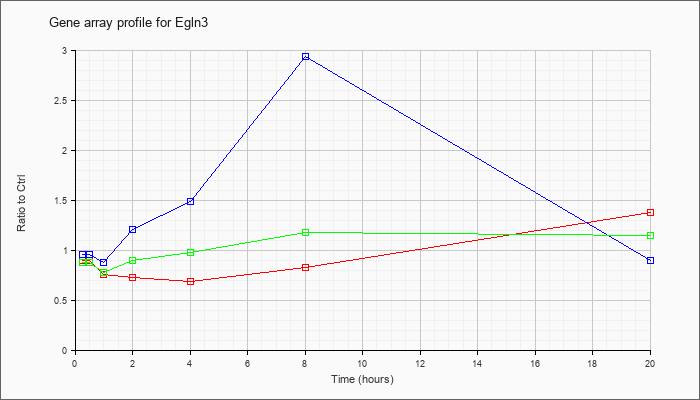 |
NM_028133 |
EGL nine homolog 3 (C. elegans) (Egln3), mRNA [NM_028133] |
KLA | .90 |
.90 |
.76 |
.73 |
.69 |
.83 |
1.38 |
| ATP | .96 |
1.03 |
.88 |
1.21 |
1.49 |
2.94 |
.90 |
| KLA/ATP | .88 |
.96 |
.78 |
.90 |
.98 |
1.18 |
1.15 |
|
| Ep300 | 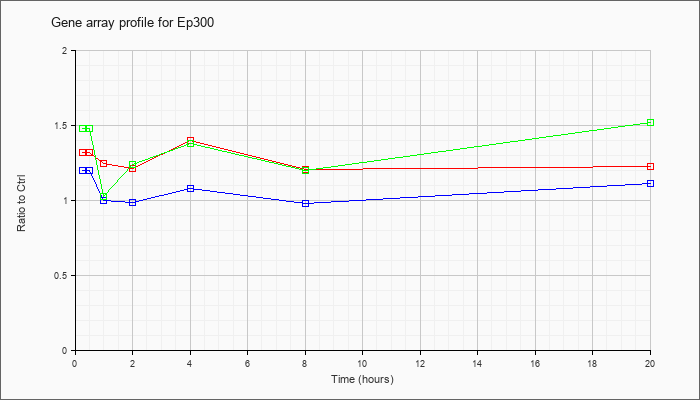 |
AK042627 |
7 days neonate cerebellum cDNA, RIKEN full-length enriched library, clone:A730011L11 product:P300 TRANSCRIPTION COACTIVATOR (FRAGMENT) homolog [Mus sp], full insert sequence. [AK042627] |
KLA | 1.02 |
.95 |
.92 |
1.04 |
1.01 |
1.00 |
1.05 |
| ATP | 1.06 |
.89 |
.88 |
.89 |
1.00 |
.90 |
1.07 |
| KLA/ATP | 1.10 |
1.02 |
.87 |
.91 |
1.04 |
.97 |
.98 |
|
| Ep300 |  |
NM_177821 |
E1A binding protein p300 (Ep300), mRNA [NM_177821] |
KLA | 1.61 |
1.26 |
1.57 |
1.38 |
1.78 |
1.41 |
1.40 |
| ATP | 1.34 |
1.60 |
1.11 |
1.08 |
1.16 |
1.05 |
1.15 |
| KLA/ATP | 1.86 |
1.94 |
1.18 |
1.56 |
1.71 |
1.42 |
2.05 |
|
| Epas1 | 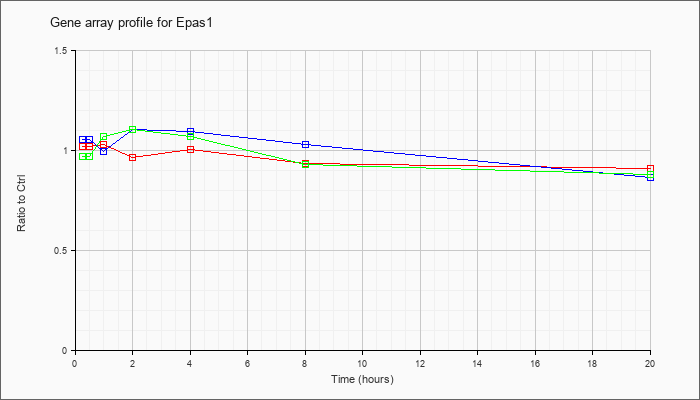 |
AK087208 |
0 day neonate lung cDNA, RIKEN full-length enriched library, clone:E030034H04 product:endothelial PAS domain protein 1, full insert sequence. [AK087208] |
KLA | .92 |
.91 |
.87 |
.93 |
1.01 |
.85 |
.83 |
| ATP | 1.02 |
1.12 |
1.03 |
1.29 |
1.23 |
1.12 |
.88 |
| KLA/ATP | .85 |
.85 |
1.05 |
1.23 |
1.14 |
.93 |
.81 |
|
| Epas1 |  |
NM_010137 |
endothelial PAS domain protein 1 (Epas1), mRNA [NM_010137] |
KLA | 1.12 |
1.03 |
1.19 |
1.00 |
1.00 |
1.02 |
.99 |
| ATP | 1.09 |
1.03 |
.96 |
.91 |
.95 |
.94 |
.85 |
| KLA/ATP | 1.09 |
1.21 |
1.09 |
.97 |
1.00 |
.93 |
.95 |
|
| Ets1 | 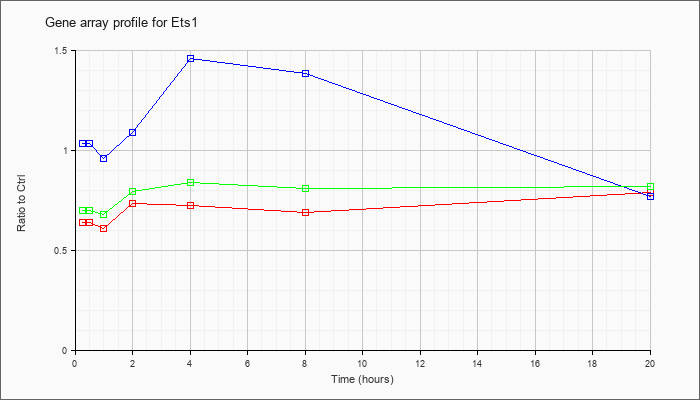 |
NM_011808 |
E26 avian leukemia oncogene 1, 5 domain (Ets1), transcript variant 1, mRNA [NM_011808] |
KLA | .55 |
.54 |
.49 |
.66 |
.62 |
.58 |
.67 |
| ATP | 1.01 |
1.14 |
.98 |
1.11 |
1.59 |
1.48 |
.67 |
| KLA/ATP | .58 |
.53 |
.58 |
.70 |
.80 |
.76 |
.76 |
|
| Ets1 |  |
X55787 |
gb|M.musculus c-ets-1 gene. [X55787] |
KLA | .83 |
.93 |
.85 |
.89 |
.94 |
.90 |
1.03 |
| ATP | 1.08 |
1.11 |
.92 |
1.04 |
1.19 |
1.19 |
.98 |
| KLA/ATP | .94 |
.90 |
.88 |
.98 |
.91 |
.92 |
.95 |
|
| Fh1 | 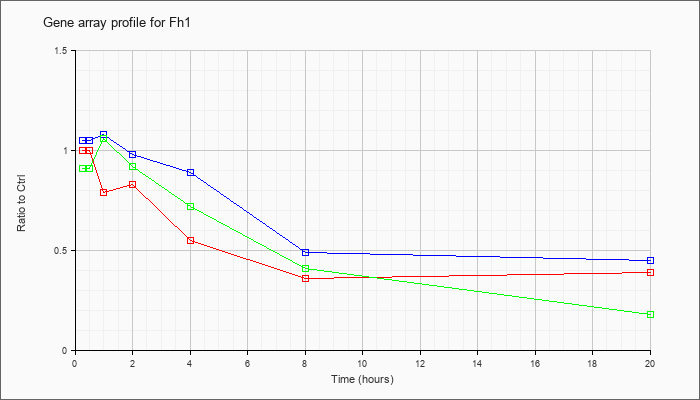 |
NM_010209 |
fumarate hydratase 1 (Fh1), mRNA [NM_010209] |
KLA | 1.00 |
.88 |
.79 |
.83 |
.55 |
.36 |
.39 |
| ATP | 1.05 |
1.05 |
1.08 |
.98 |
.89 |
.49 |
.45 |
| KLA/ATP | .91 |
.95 |
1.06 |
.92 |
.72 |
.41 |
.18 |
|
| Flcn | 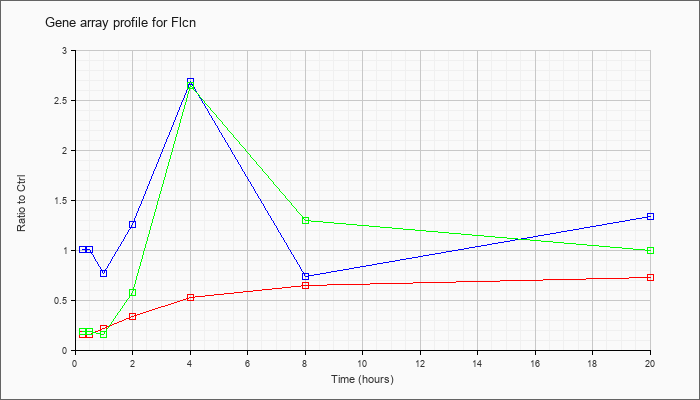 |
NM_146018 |
folliculin (Flcn), mRNA [NM_146018] |
KLA | .16 |
.18 |
.22 |
.34 |
.53 |
.65 |
.73 |
| ATP | 1.01 |
.98 |
.77 |
1.26 |
2.69 |
.74 |
1.34 |
| KLA/ATP | .19 |
.18 |
.16 |
.58 |
2.65 |
1.30 |
1.00 |
|
| Gab1 | 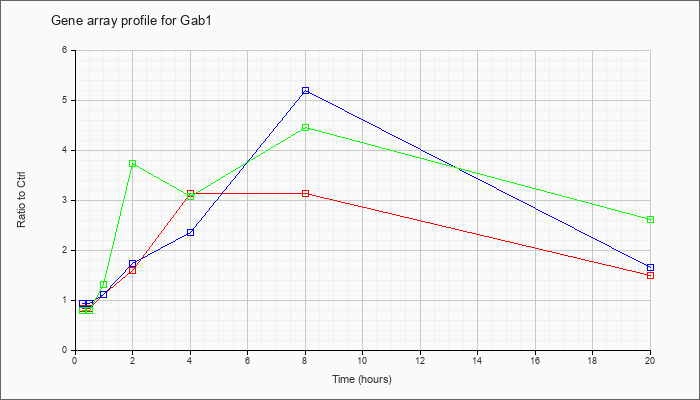 |
NM_021356 |
growth factor receptor bound protein 2-associated protein 1 (Gab1), mRNA [NM_021356] |
KLA | .83 |
.95 |
1.12 |
1.59 |
3.13 |
3.14 |
1.50 |
| ATP | .94 |
.96 |
1.12 |
1.74 |
2.35 |
5.20 |
1.66 |
| KLA/ATP | .79 |
.78 |
1.32 |
3.74 |
3.08 |
4.44 |
2.62 |
|
| Grb2 | 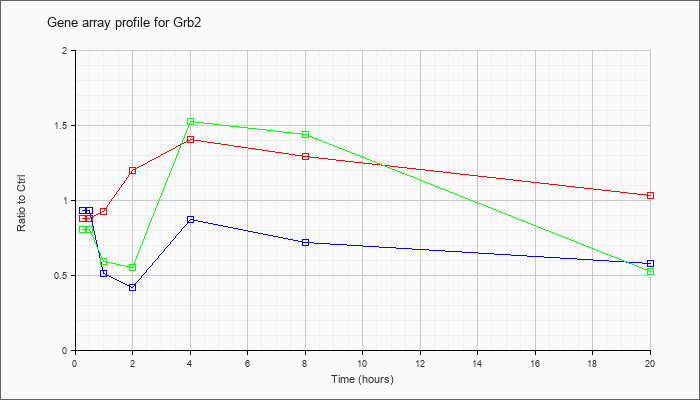 |
NM_008163 |
growth factor receptor bound protein 2 (Grb2), mRNA [NM_008163] |
KLA | .92 |
.88 |
.90 |
1.27 |
1.43 |
1.22 |
1.00 |
| ATP | 1.02 |
.83 |
.51 |
.42 |
.92 |
.70 |
.57 |
| KLA/ATP | .84 |
.77 |
.61 |
.56 |
1.56 |
1.49 |
.52 |
|
| Grb2 |  |
U07617 |
Grb2 adaptor protein (grb2) mRNA, complete cds. [U07617] |
KLA | .83 |
.80 |
.95 |
1.13 |
1.38 |
1.36 |
1.06 |
| ATP | .84 |
.74 |
.51 |
.41 |
.82 |
.74 |
.59 |
| KLA/ATP | .77 |
.70 |
.57 |
.54 |
1.49 |
1.39 |
.53 |
|
| Hgf | 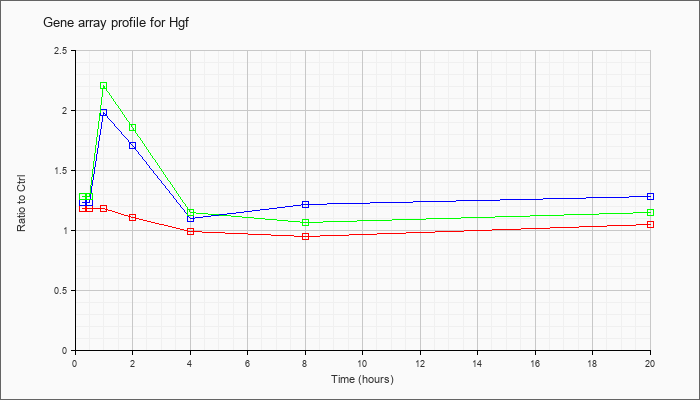 |
AK033274 |
15 days embryo male testis cDNA, RIKEN full-length enriched library, clone:8030481P22 product:inferred: hepatocyte growth factor [AK033274] |
KLA | 1.34 |
1.22 |
.96 |
.93 |
.82 |
.93 |
.87 |
| ATP | 1.64 |
2.45 |
3.30 |
1.89 |
.97 |
1.12 |
.90 |
| KLA/ATP | 1.49 |
2.50 |
3.96 |
1.88 |
1.07 |
1.00 |
.83 |
|
| Hgf |  |
NM_010427 |
hepatocyte growth factor (Hgf), mRNA [NM_010427] |
KLA | 1.11 |
1.18 |
1.30 |
1.20 |
1.08 |
.96 |
1.14 |
| ATP | 1.02 |
1.14 |
1.32 |
1.62 |
1.17 |
1.26 |
1.47 |
| KLA/ATP | 1.18 |
1.24 |
1.33 |
1.84 |
1.19 |
1.09 |
1.31 |
|
| Hif1a | 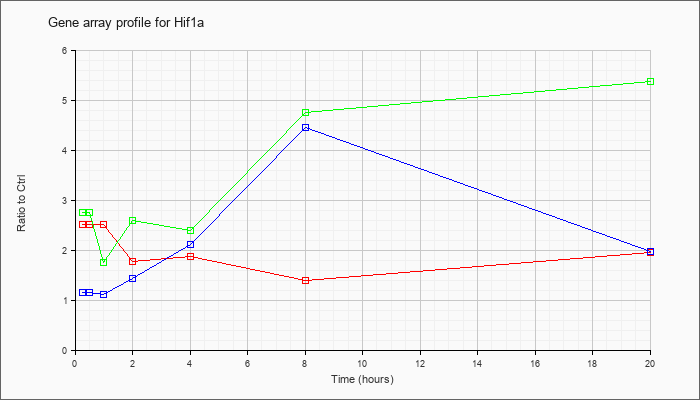 |
NM_010431 |
hypoxia inducible factor 1, alpha subunit (Hif1a), mRNA [NM_010431] |
KLA | 2.52 |
2.50 |
2.52 |
1.78 |
1.87 |
1.39 |
1.96 |
| ATP | 1.16 |
1.37 |
1.11 |
1.44 |
2.11 |
4.46 |
1.97 |
| KLA/ATP | 2.76 |
2.92 |
1.75 |
2.60 |
2.39 |
4.75 |
5.37 |
|
| Hras1 | 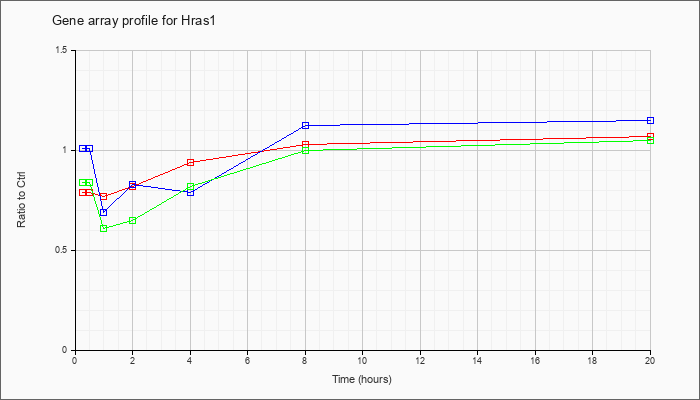 |
NM_008284 |
Harvey rat sarcoma virus oncogene 1 (Hras1), mRNA [NM_008284] |
KLA | .79 |
.82 |
.77 |
.82 |
.94 |
1.03 |
1.07 |
| ATP | 1.01 |
.95 |
.69 |
.83 |
.79 |
1.12 |
1.15 |
| KLA/ATP | .84 |
.77 |
.61 |
.65 |
.82 |
1.00 |
1.05 |
|
| Jun | 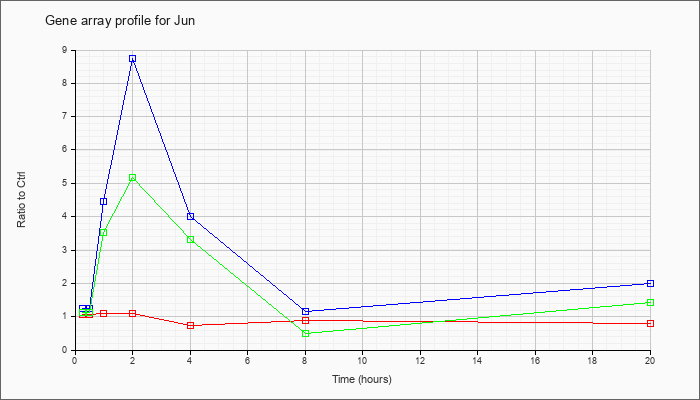 |
NM_010591 |
Jun oncogene (Jun), mRNA [NM_010591] |
KLA | 1.08 |
.96 |
1.09 |
1.09 |
.75 |
.88 |
.81 |
| ATP | 1.26 |
1.48 |
4.45 |
8.73 |
4.02 |
1.16 |
1.99 |
| KLA/ATP | 1.12 |
1.38 |
3.52 |
5.17 |
3.32 |
.49 |
1.42 |
|
| Kras | 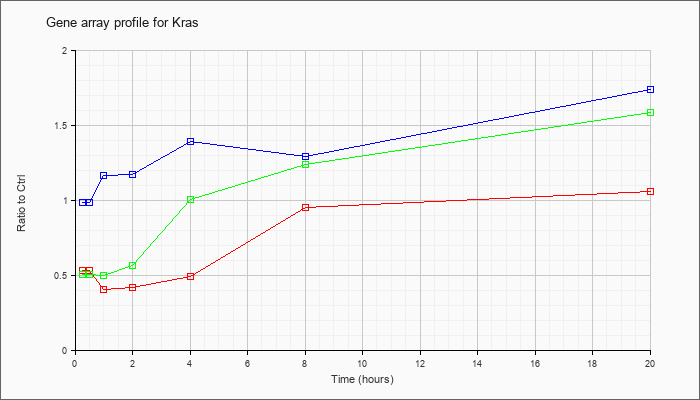 |
NM_021284 |
v-Ki-ras2 Kirsten rat sarcoma viral oncogene homolog (Kras), mRNA [NM_021284] |
KLA | .53 |
.49 |
.40 |
.42 |
.49 |
.95 |
1.05 |
| ATP | .98 |
1.01 |
1.16 |
1.17 |
1.39 |
1.29 |
1.74 |
| KLA/ATP | .50 |
.47 |
.50 |
.56 |
1.00 |
1.24 |
1.58 |
|
| Map2k1 | 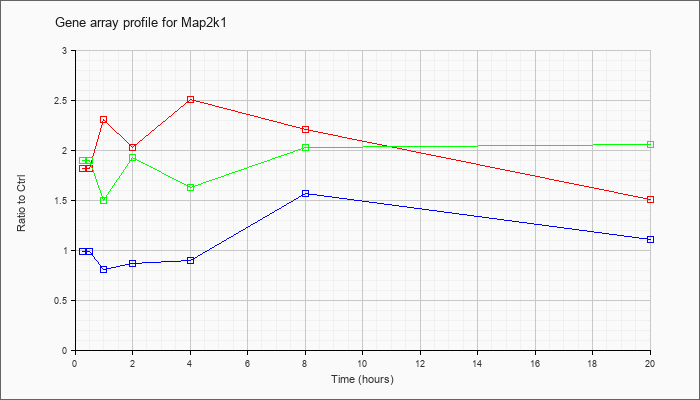 |
NM_008927 |
mitogen-activated protein kinase kinase 1 (Map2k1), mRNA [NM_008927] |
KLA | 1.82 |
2.00 |
2.31 |
2.03 |
2.51 |
2.20 |
1.51 |
| ATP | .99 |
1.14 |
.81 |
.87 |
.90 |
1.57 |
1.11 |
| KLA/ATP | 1.90 |
2.03 |
1.50 |
1.93 |
1.63 |
2.03 |
2.06 |
|
| Map2k2 | 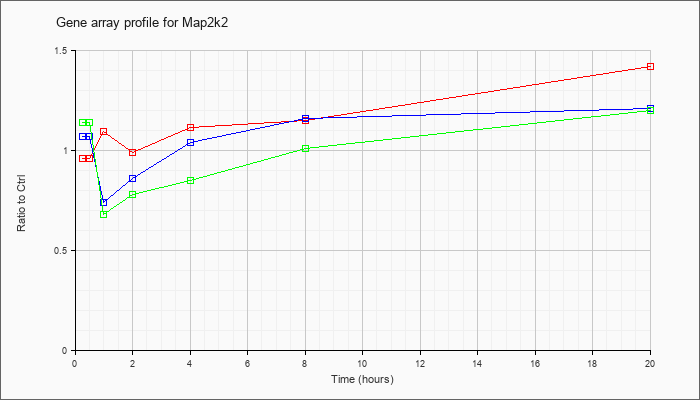 |
NM_023138 |
mitogen-activated protein kinase kinase 2 (Map2k2), mRNA [NM_023138] |
KLA | .96 |
1.01 |
1.09 |
.99 |
1.11 |
1.15 |
1.42 |
| ATP | 1.07 |
1.05 |
.74 |
.86 |
1.04 |
1.16 |
1.21 |
| KLA/ATP | 1.14 |
1.03 |
.68 |
.78 |
.85 |
1.01 |
1.20 |
|
| Mapk1 | 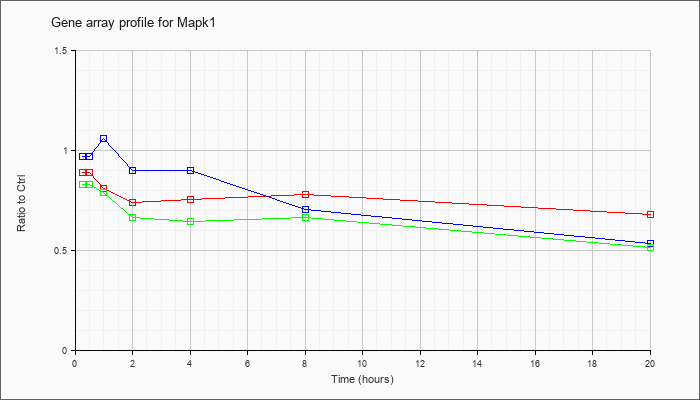 |
NM_001038663 |
mitogen-activated protein kinase 1 (Mapk1), transcript variant 2, mRNA [NM_001038663] |
KLA | .97 |
1.03 |
.93 |
.86 |
.79 |
1.01 |
1.04 |
| ATP | .96 |
1.18 |
1.01 |
1.13 |
.84 |
1.04 |
1.16 |
| KLA/ATP | .99 |
.99 |
.84 |
.87 |
.63 |
.96 |
1.26 |
|
| Mapk1 |  |
NM_011949 |
mitogen-activated protein kinase 1 (Mapk1), transcript variant 1, mRNA [NM_011949] |
KLA | .88 |
.80 |
.80 |
.73 |
.75 |
.76 |
.65 |
| ATP | .97 |
.96 |
1.06 |
.88 |
.90 |
.68 |
.48 |
| KLA/ATP | .82 |
.81 |
.78 |
.65 |
.64 |
.64 |
.45 |
|
| Mapk3 | 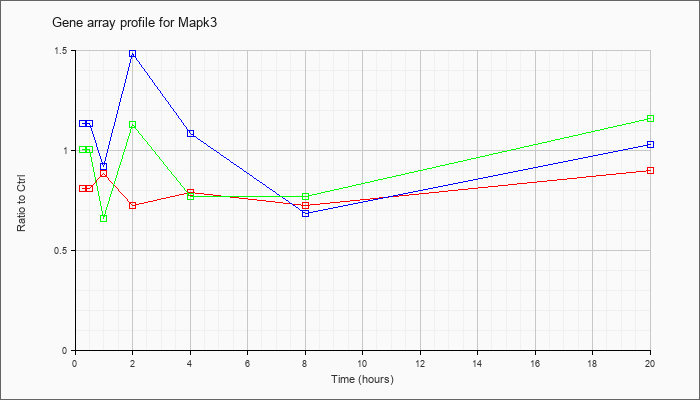 |
NM_011952 |
mitogen-activated protein kinase 3 (Mapk3), mRNA [NM_011952] |
KLA | .81 |
.85 |
.88 |
.72 |
.79 |
.73 |
.90 |
| ATP | 1.13 |
1.30 |
.92 |
1.48 |
1.08 |
.68 |
1.03 |
| KLA/ATP | 1.01 |
1.01 |
.66 |
1.13 |
.77 |
.77 |
1.16 |
|
| Met | 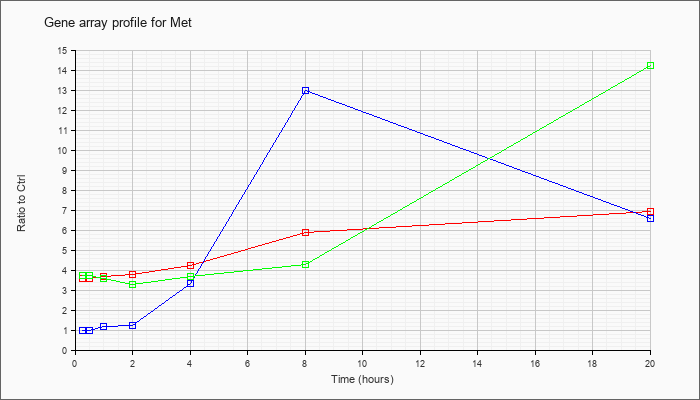 |
NM_008591 |
met proto-oncogene (Met), mRNA [NM_008591] |
KLA | 3.56 |
3.71 |
3.68 |
3.79 |
4.24 |
5.89 |
6.95 |
| ATP | .98 |
.96 |
1.16 |
1.21 |
3.32 |
12.98 |
6.60 |
| KLA/ATP | 3.75 |
3.80 |
3.58 |
3.28 |
3.69 |
4.27 |
14.22 |
|
Mm.13157
2 | 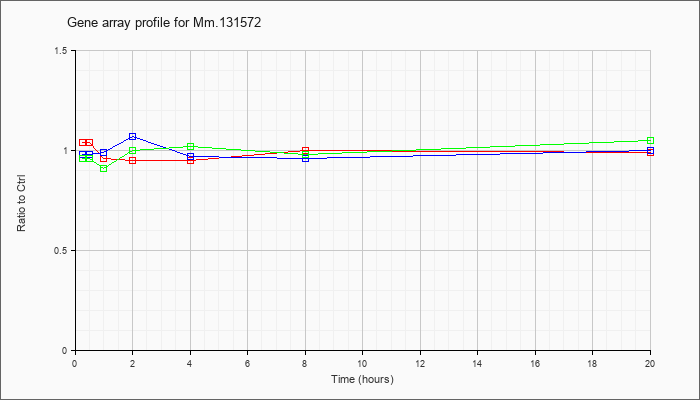 |
134949023 |
Unknown |
KLA | 1.04 |
1.01 |
.96 |
.95 |
.95 |
1.00 |
.99 |
| ATP | .98 |
1.02 |
.99 |
1.07 |
.97 |
.96 |
1.00 |
| KLA/ATP | .96 |
.88 |
.91 |
1.00 |
1.02 |
.98 |
1.05 |
|
Mm.21312
8 | 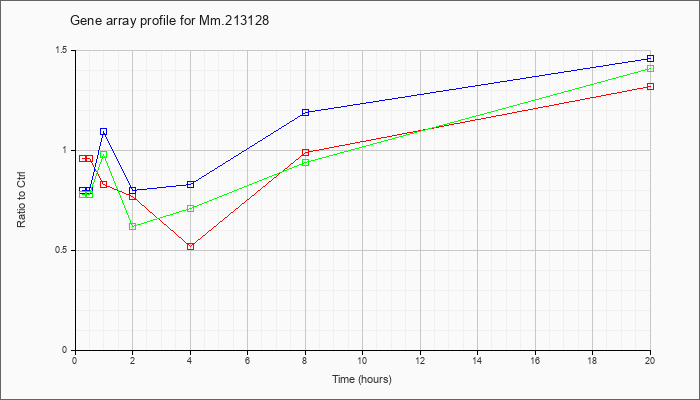 |
141801344 |
Unknown |
KLA | .96 |
.95 |
.83 |
.77 |
.52 |
.99 |
1.32 |
| ATP | .80 |
.82 |
1.09 |
.80 |
.83 |
1.19 |
1.46 |
| KLA/ATP | .78 |
.92 |
.98 |
.62 |
.71 |
.94 |
1.41 |
|
Mm.22094
6 | 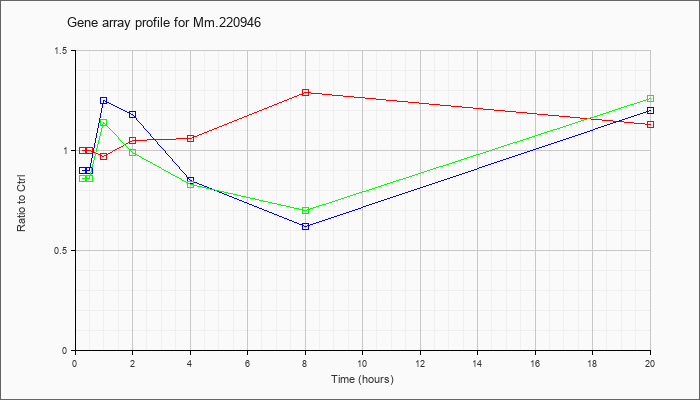 |
27545180 |
Unknown |
KLA | 1.00 |
.93 |
.97 |
1.05 |
1.06 |
1.29 |
1.13 |
| ATP | .90 |
.94 |
1.25 |
1.18 |
.85 |
.62 |
1.20 |
| KLA/ATP | .86 |
.99 |
1.14 |
.99 |
.83 |
.70 |
1.26 |
|
Mm.24185
7 | 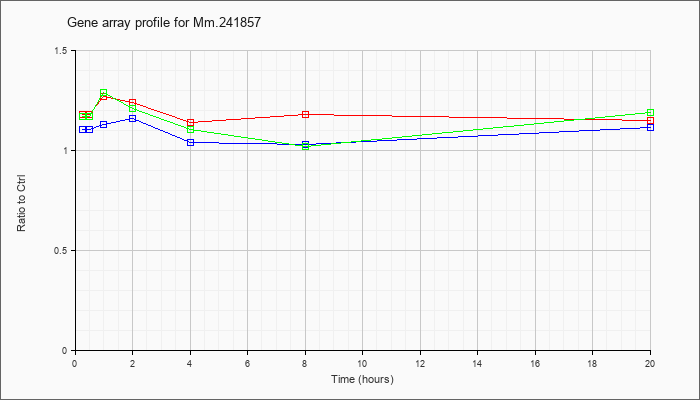 |
142347489 |
Unknown |
KLA | 1.18 |
1.09 |
1.27 |
1.24 |
1.14 |
1.18 |
1.15 |
| ATP | 1.10 |
1.02 |
1.13 |
1.16 |
1.04 |
1.03 |
1.11 |
| KLA/ATP | 1.17 |
1.19 |
1.29 |
1.21 |
1.10 |
1.02 |
1.19 |
|
Mm.25381
9 | 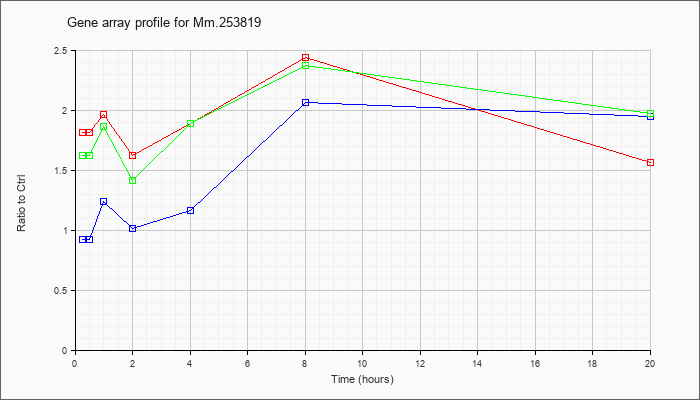 |
118130676 |
Unknown |
KLA | 1.81 |
1.88 |
1.96 |
1.62 |
1.89 |
2.44 |
1.56 |
| ATP | .92 |
.91 |
1.24 |
1.01 |
1.16 |
2.06 |
1.95 |
| KLA/ATP | 1.62 |
1.86 |
1.86 |
1.41 |
1.89 |
2.37 |
1.97 |
|
Mm.28924
8 | 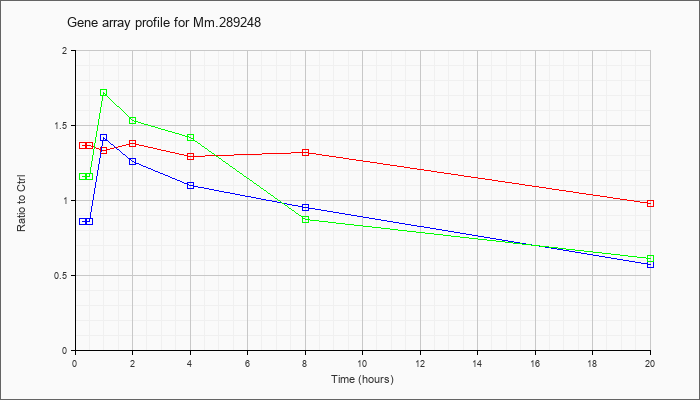 |
148747277 |
Unknown |
KLA | 1.36 |
1.32 |
1.33 |
1.38 |
1.29 |
1.32 |
.98 |
| ATP | .86 |
.92 |
1.42 |
1.26 |
1.10 |
.95 |
.57 |
| KLA/ATP | 1.16 |
1.34 |
1.72 |
1.53 |
1.42 |
.87 |
.61 |
|
| Nras | 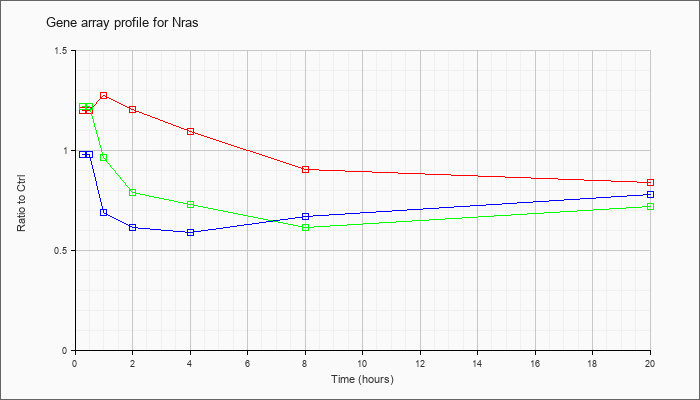 |
NM_010937 |
neuroblastoma ras oncogene (Nras), mRNA [NM_010937] |
KLA | 1.20 |
1.22 |
1.27 |
1.20 |
1.09 |
.91 |
.84 |
| ATP | .98 |
.91 |
.69 |
.61 |
.59 |
.67 |
.78 |
| KLA/ATP | 1.22 |
1.14 |
.97 |
.79 |
.73 |
.61 |
.72 |
|
| Pak1 | 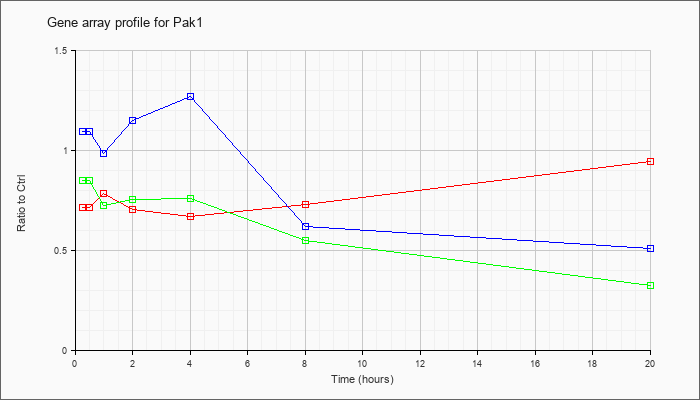 |
NM_011035 |
p21 (CDKN1A)-activated kinase 1 (Pak1), mRNA [NM_011035] |
KLA | .72 |
.77 |
.79 |
.71 |
.67 |
.73 |
.95 |
| ATP | 1.10 |
1.07 |
.99 |
1.15 |
1.27 |
.62 |
.51 |
| KLA/ATP | .85 |
.83 |
.73 |
.76 |
.76 |
.55 |
.33 |
|
| Pak2 | 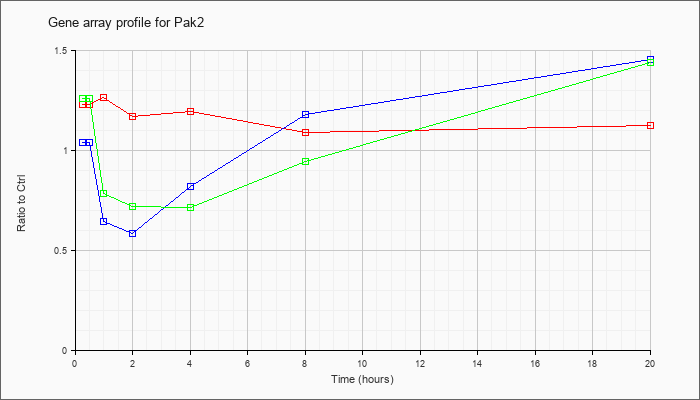 |
NM_177326 |
p21 (CDKN1A)-activated kinase 2 (Pak2), mRNA [NM_177326] |
KLA | 1.23 |
1.24 |
1.26 |
1.17 |
1.19 |
1.09 |
1.13 |
| ATP | 1.04 |
1.01 |
.65 |
.58 |
.82 |
1.18 |
1.45 |
| KLA/ATP | 1.26 |
1.26 |
.78 |
.72 |
.71 |
.95 |
1.44 |
|
| Pak3 | 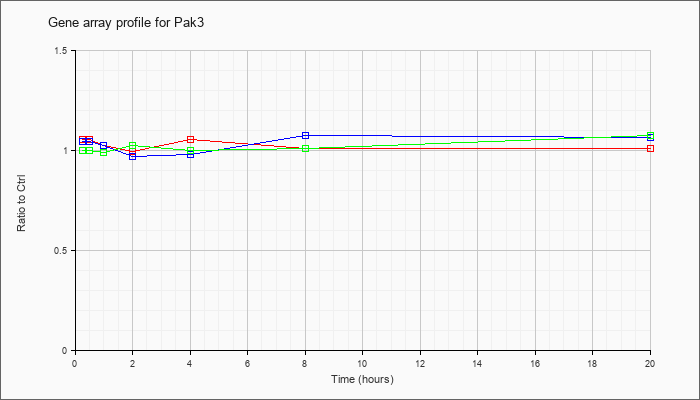 |
NM_008778 |
p21 (CDKN1A)-activated kinase 3 (Pak3), mRNA [NM_008778] |
KLA | 1.06 |
1.02 |
1.03 |
1.00 |
1.06 |
1.01 |
1.01 |
| ATP | 1.05 |
1.01 |
1.03 |
.97 |
.98 |
1.08 |
1.07 |
| KLA/ATP | 1.00 |
.99 |
.99 |
1.03 |
1.00 |
1.01 |
1.08 |
|
| Pak4 | 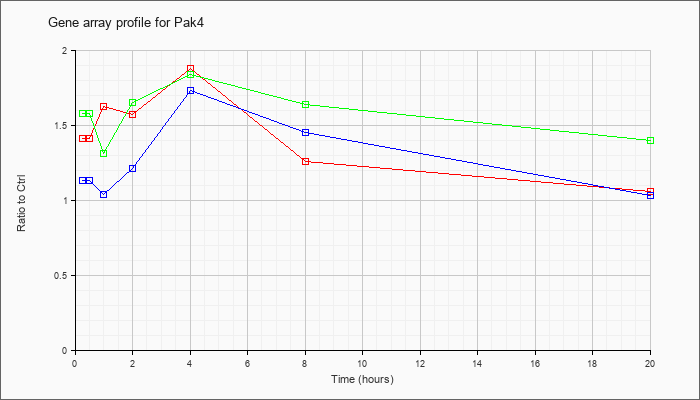 |
NM_027470 |
p21 (CDKN1A)-activated kinase 4 (Pak4), mRNA [NM_027470] |
KLA | 1.41 |
1.51 |
1.62 |
1.57 |
1.88 |
1.26 |
1.06 |
| ATP | 1.13 |
1.15 |
1.04 |
1.21 |
1.73 |
1.45 |
1.03 |
| KLA/ATP | 1.58 |
1.60 |
1.31 |
1.65 |
1.84 |
1.64 |
1.40 |
|
| Pak6 | 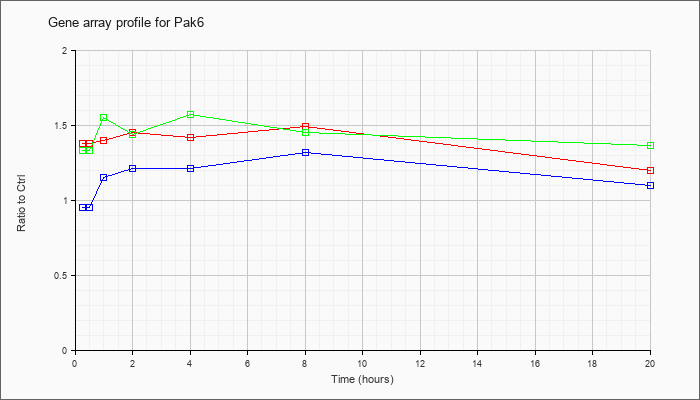 |
AK028788 |
10 days neonate skin cDNA, RIKEN full-length enriched library, clone:4732456M09 product:p21 (CDKN1A)-activated kinase 6, full insert sequence. [AK028788] |
KLA | 1.38 |
1.33 |
1.40 |
1.45 |
1.42 |
1.49 |
1.20 |
| ATP | .95 |
.93 |
1.15 |
1.21 |
1.21 |
1.32 |
1.10 |
| KLA/ATP | 1.33 |
1.38 |
1.55 |
1.44 |
1.57 |
1.45 |
1.36 |
|
| Pak7 | 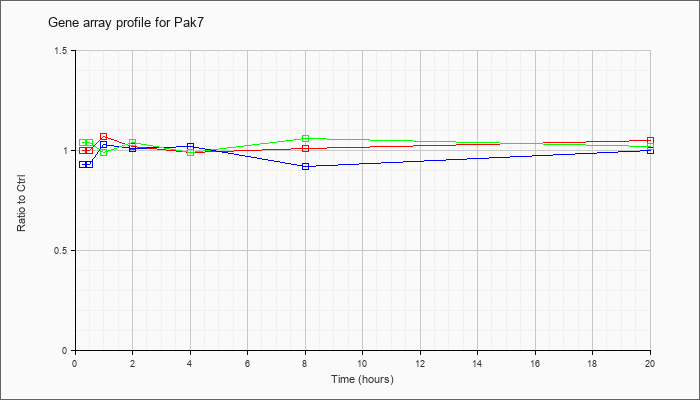 |
AK077967 |
13 days embryo male testis cDNA, RIKEN full-length enriched library, clone:6030481C12 product:hypothetical protein, full insert sequence. [AK077967] |
KLA | 1.00 |
1.01 |
1.07 |
1.02 |
.99 |
1.01 |
1.05 |
| ATP | .93 |
.95 |
1.03 |
1.01 |
1.02 |
.92 |
1.00 |
| KLA/ATP | 1.04 |
1.01 |
.99 |
1.04 |
.99 |
1.06 |
1.02 |
|
| Pdgfa | 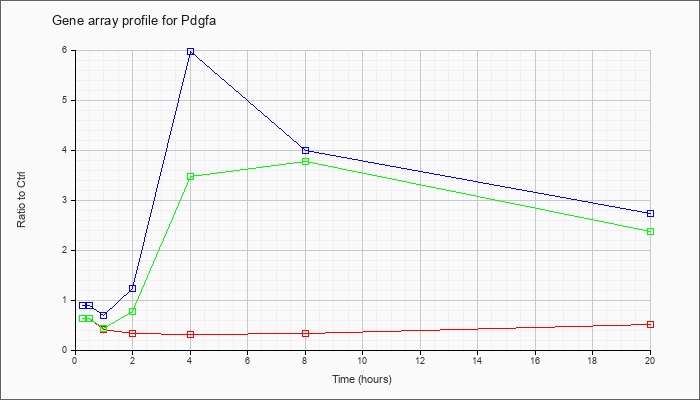 |
NM_008808 |
platelet derived growth factor, alpha (Pdgfa), mRNA [NM_008808] |
KLA | .63 |
.56 |
.42 |
.34 |
.31 |
.33 |
.52 |
| ATP | .89 |
.80 |
.69 |
1.23 |
5.97 |
3.99 |
2.73 |
| KLA/ATP | .64 |
.53 |
.43 |
.77 |
3.48 |
3.78 |
2.37 |
|
| Pdgfb | 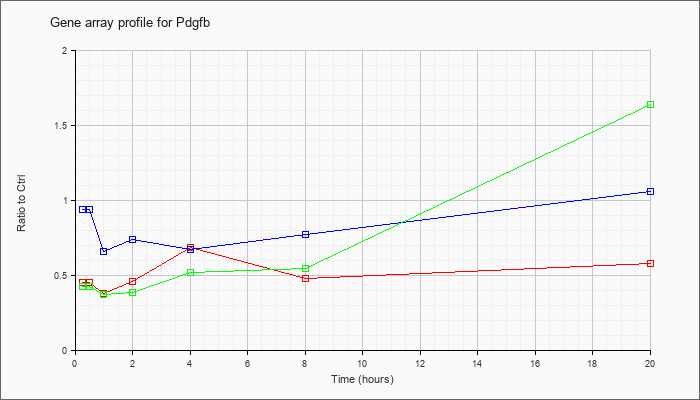 |
NM_011057 |
platelet derived growth factor, B polypeptide (Pdgfb), mRNA [NM_011057] |
KLA | .45 |
.43 |
.38 |
.46 |
.69 |
.48 |
.58 |
| ATP | .94 |
.95 |
.66 |
.74 |
.67 |
.77 |
1.06 |
| KLA/ATP | .43 |
.43 |
.37 |
.39 |
.52 |
.55 |
1.64 |
|
| Pik3ca | 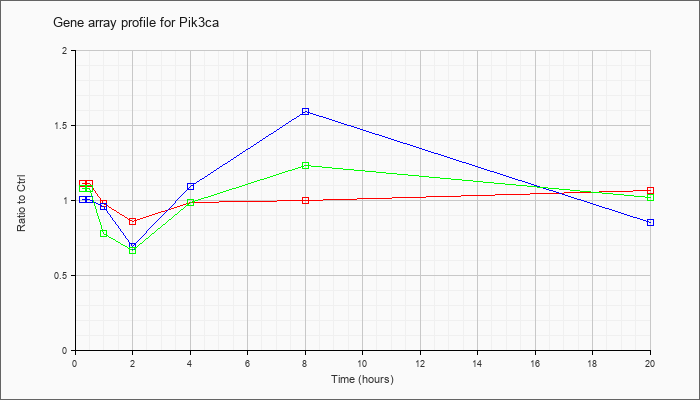 |
ENSMUST00000108243 |
ens|Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha isoform (EC 2.7.1.153) (PI3-kinase p110 subunit alpha) (PtdIns-3- kinase p110) (PI3K). [Source:Uniprot/SWISSPROT;Acc:P42337] [ENSMUST00000108243] |
KLA | 1.07 |
1.01 |
.91 |
.90 |
.98 |
.96 |
.86 |
| ATP | 1.03 |
.87 |
1.12 |
.63 |
1.22 |
1.86 |
.60 |
| KLA/ATP | 1.05 |
.82 |
.91 |
.61 |
1.17 |
1.26 |
.63 |
|
| Pik3ca |  |
NM_008839 |
phosphatidylinositol 3-kinase, catalytic, alpha polypeptide (Pik3ca), mRNA [NM_008839] |
KLA | 1.15 |
1.11 |
1.05 |
.82 |
.99 |
1.03 |
1.27 |
| ATP | .98 |
1.00 |
.79 |
.75 |
.96 |
1.32 |
1.10 |
| KLA/ATP | 1.10 |
.98 |
.64 |
.72 |
.80 |
1.20 |
1.41 |
|
| Pik3cb | 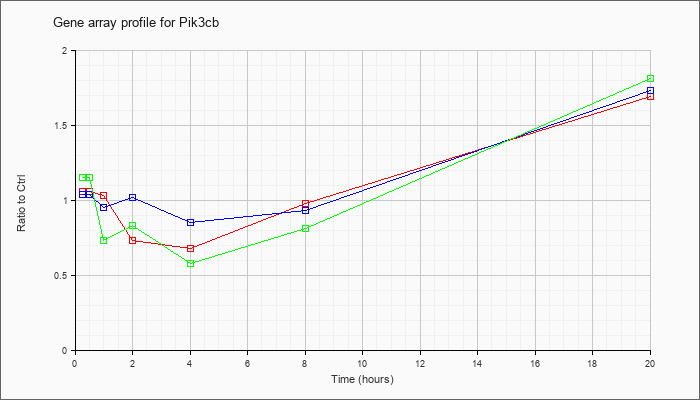 |
NM_029094 |
phosphatidylinositol 3-kinase, catalytic, beta polypeptide (Pik3cb), mRNA [NM_029094] |
KLA | 1.06 |
1.06 |
1.03 |
.73 |
.68 |
.98 |
1.69 |
| ATP | 1.04 |
1.21 |
.95 |
1.02 |
.85 |
.93 |
1.73 |
| KLA/ATP | 1.15 |
1.27 |
.73 |
.83 |
.58 |
.81 |
1.81 |
|
| Pik3cd | 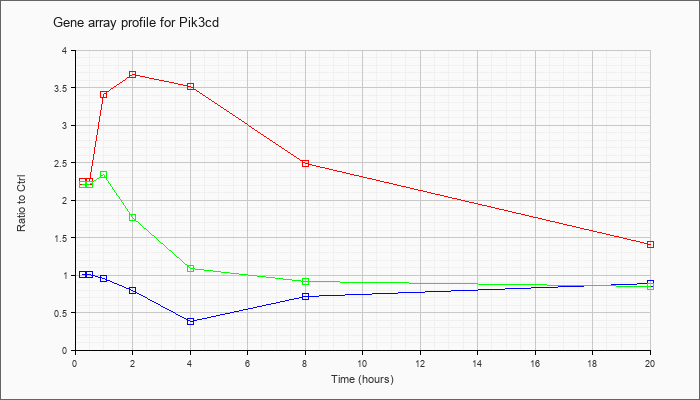 |
NM_008840 |
phosphatidylinositol 3-kinase catalytic delta polypeptide (Pik3cd), transcript variant 1, mRNA [NM_008840] |
KLA | 2.28 |
2.42 |
3.10 |
4.17 |
3.35 |
2.31 |
1.34 |
| ATP | 1.02 |
.91 |
.99 |
.63 |
.30 |
.67 |
.80 |
| KLA/ATP | 2.14 |
2.23 |
2.70 |
1.51 |
1.17 |
.85 |
.72 |
|
| Pik3cd |  |
U86587 |
phosphatidylinositol 3-kinase catalytic subunit p110 delta mRNA, complete cds. [U86587] |
KLA | 2.21 |
2.24 |
3.71 |
3.17 |
3.67 |
2.66 |
1.48 |
| ATP | 1.00 |
1.03 |
.93 |
.96 |
.46 |
.75 |
.97 |
| KLA/ATP | 2.27 |
2.32 |
1.97 |
2.02 |
1.01 |
.99 |
.98 |
|
| Pik3cg | 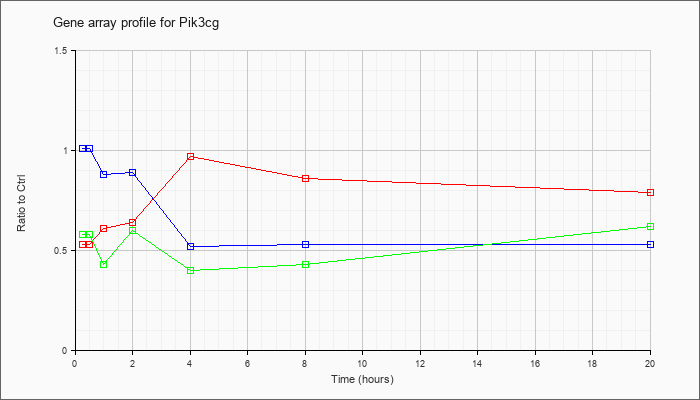 |
NM_020272 |
phosphoinositide-3-kinase, catalytic, gamma polypeptide (Pik3cg), mRNA [NM_020272] |
KLA | .53 |
.53 |
.61 |
.64 |
.97 |
.86 |
.79 |
| ATP | 1.01 |
1.13 |
.88 |
.89 |
.52 |
.53 |
.53 |
| KLA/ATP | .58 |
.53 |
.43 |
.60 |
.40 |
.43 |
.62 |
|
| Pik3r1 | 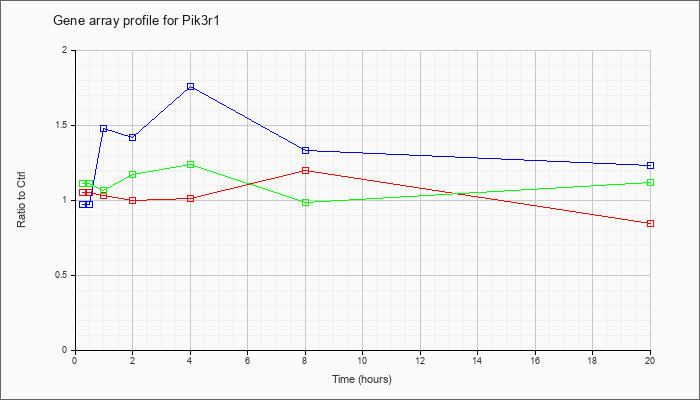 |
NM_001077495 |
phosphatidylinositol 3-kinase, regulatory subunit, polypeptide 1 (p85 alpha) (Pik3r1), transcript variant 2, mRNA [NM_001077495] |
KLA | 1.05 |
1.07 |
1.03 |
1.00 |
1.01 |
1.20 |
.85 |
| ATP | .97 |
1.02 |
1.48 |
1.42 |
1.76 |
1.33 |
1.23 |
| KLA/ATP | 1.11 |
1.12 |
1.06 |
1.17 |
1.24 |
.99 |
1.12 |
|
| Pik3r2 | 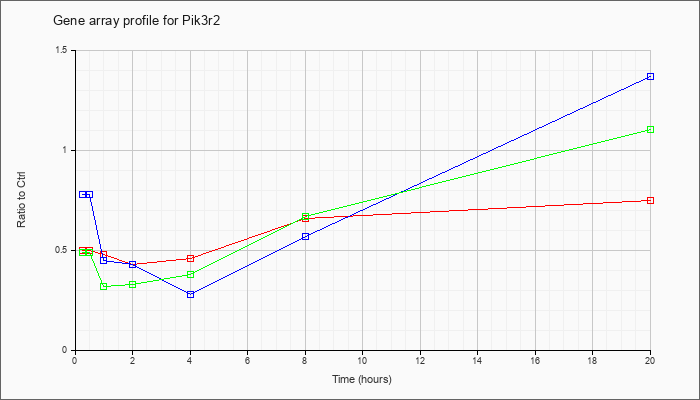 |
NM_008841 |
phosphatidylinositol 3-kinase, regulatory subunit, polypeptide 2 (p85 beta) (Pik3r2), mRNA [NM_008841] |
KLA | .50 |
.50 |
.48 |
.43 |
.46 |
.66 |
.75 |
| ATP | .78 |
.68 |
.45 |
.43 |
.28 |
.57 |
1.37 |
| KLA/ATP | .49 |
.38 |
.32 |
.33 |
.38 |
.67 |
1.10 |
|
| Pik3r3 | 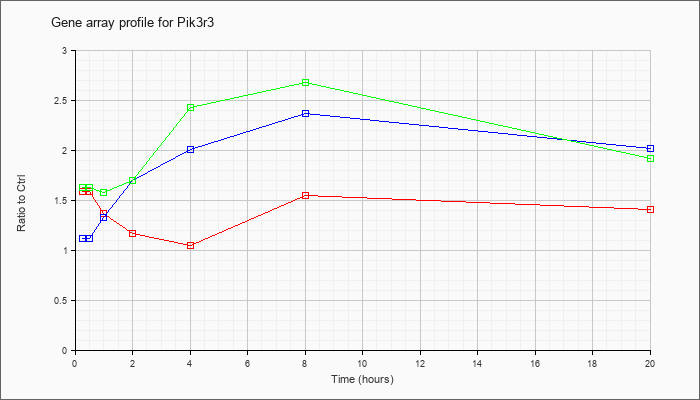 |
NM_181585 |
phosphatidylinositol 3 kinase, regulatory subunit, polypeptide 3 (p55) (Pik3r3), mRNA [NM_181585] |
KLA | 1.59 |
1.49 |
1.37 |
1.17 |
1.05 |
1.55 |
1.41 |
| ATP | 1.12 |
1.00 |
1.33 |
1.70 |
2.01 |
2.37 |
2.02 |
| KLA/ATP | 1.63 |
1.54 |
1.58 |
1.70 |
2.43 |
2.68 |
1.92 |
|
| Pik3r5 | 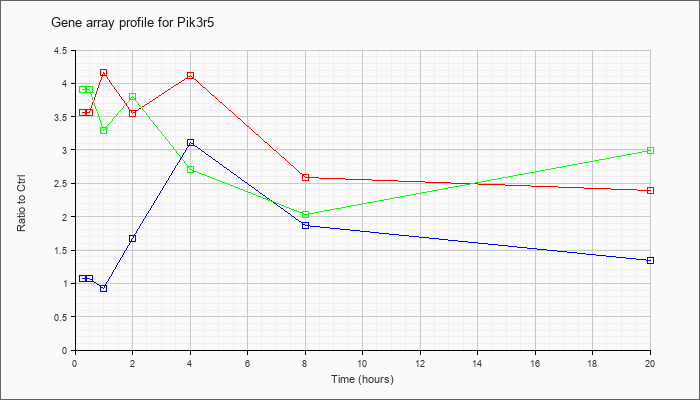 |
NM_177320 |
phosphoinositide-3-kinase, regulatory subunit 5, p101 (Pik3r5), mRNA [NM_177320] |
KLA | 3.57 |
3.58 |
4.16 |
3.54 |
4.11 |
2.59 |
2.39 |
| ATP | 1.08 |
1.08 |
.93 |
1.67 |
3.11 |
1.88 |
1.34 |
| KLA/ATP | 3.91 |
3.70 |
3.30 |
3.80 |
2.70 |
2.04 |
3.00 |
|
| Ptpn11 | 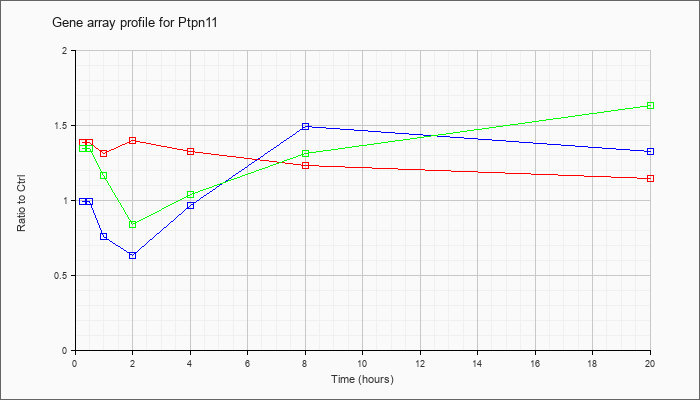 |
NM_001109992 |
protein tyrosine phosphatase, non-receptor type 11 (Ptpn11), transcript variant 2, mRNA [NM_001109992] |
KLA | 1.52 |
1.71 |
1.59 |
1.78 |
1.62 |
1.57 |
1.35 |
| ATP | 1.04 |
.93 |
.86 |
.79 |
1.09 |
1.95 |
1.90 |
| KLA/ATP | 1.59 |
1.61 |
1.45 |
1.11 |
1.18 |
1.82 |
3.02 |
|
| Ptpn11 |  |
NM_011202 |
protein tyrosine phosphatase, non-receptor type 11 (Ptpn11), transcript variant 1, mRNA [NM_011202] |
KLA | 1.32 |
1.28 |
1.17 |
1.21 |
1.18 |
1.06 |
1.05 |
| ATP | .97 |
.77 |
.71 |
.56 |
.90 |
1.27 |
1.04 |
| KLA/ATP | 1.22 |
1.21 |
1.02 |
.70 |
.96 |
1.06 |
.94 |
|
| Rac1 | 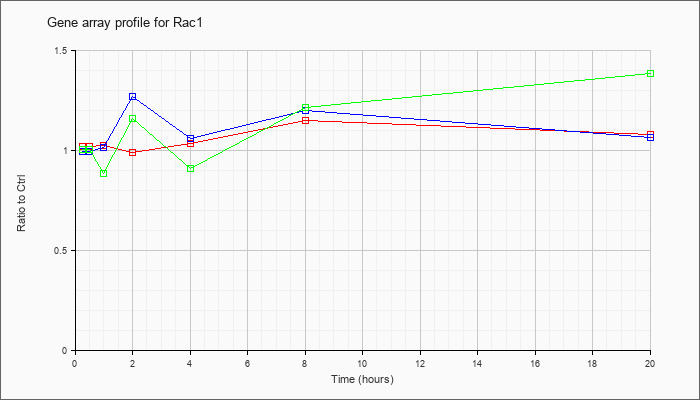 |
NM_009007 |
RAS-related C3 botulinum substrate 1 (Rac1), mRNA [NM_009007] |
KLA | 1.02 |
1.01 |
1.02 |
.99 |
1.03 |
1.15 |
1.08 |
| ATP | .99 |
1.15 |
1.01 |
1.27 |
1.06 |
1.20 |
1.06 |
| KLA/ATP | 1.00 |
1.04 |
.88 |
1.16 |
.91 |
1.21 |
1.38 |
|
| Raf1 | 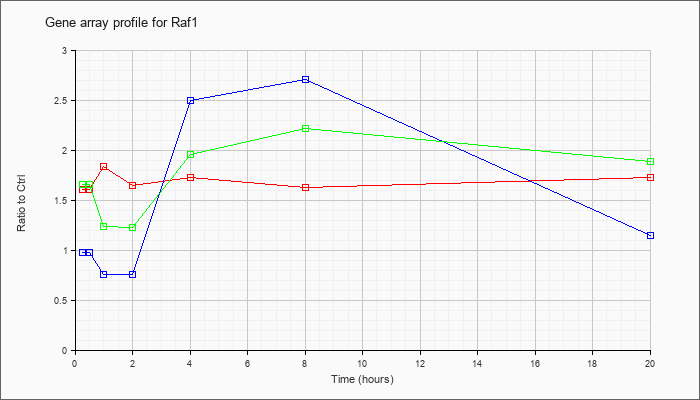 |
AK036317 |
16 days neonate cerebellum cDNA, RIKEN full-length enriched library, clone:9630056E02 product:v-raf-1 leukemia viral oncogene 1, full insert sequence. [AK036317] |
KLA | 1.47 |
1.34 |
1.55 |
1.29 |
1.59 |
1.32 |
1.46 |
| ATP | 1.05 |
.98 |
.54 |
.63 |
3.32 |
2.07 |
1.01 |
| KLA/ATP | 1.57 |
1.33 |
.79 |
.96 |
2.26 |
1.38 |
1.35 |
|
| Raf1 |  |
NM_029780 |
v-raf-leukemia viral oncogene 1 (Raf1), mRNA [NM_029780] |
KLA | 1.65 |
1.64 |
1.93 |
1.77 |
1.77 |
1.72 |
1.81 |
| ATP | .95 |
.96 |
.83 |
.80 |
2.22 |
2.92 |
1.19 |
| KLA/ATP | 1.68 |
1.64 |
1.39 |
1.32 |
1.85 |
2.50 |
2.06 |
|
| Rap1a | 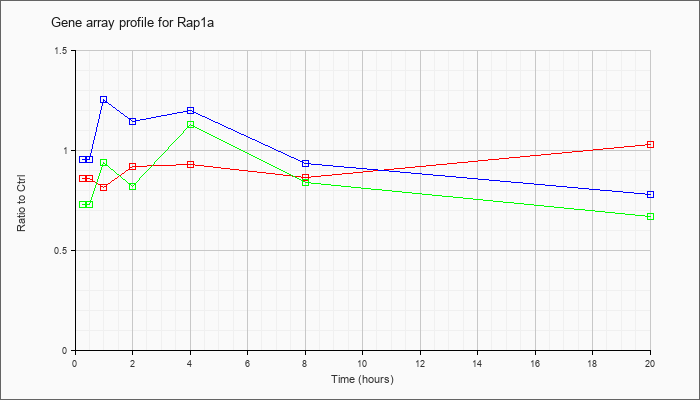 |
AK030897 |
adult male thymus cDNA, RIKEN full-length enriched library, clone:5830450G12 product:RAS-related protein-1a, full insert sequence [AK030897] |
KLA | .83 |
.80 |
.92 |
1.04 |
1.06 |
.91 |
1.06 |
| ATP | 1.04 |
.92 |
1.11 |
1.03 |
1.05 |
.73 |
.72 |
| KLA/ATP | .83 |
.81 |
.97 |
.75 |
1.14 |
.71 |
.53 |
|
| Rap1a |  |
NM_145541 |
RAS-related protein-1a (Rap1a), mRNA [NM_145541] |
KLA | .87 |
.78 |
.76 |
.86 |
.86 |
.84 |
1.01 |
| ATP | .91 |
.95 |
1.33 |
1.20 |
1.27 |
1.04 |
.81 |
| KLA/ATP | .68 |
.75 |
.92 |
.85 |
1.13 |
.90 |
.74 |
|
| Rap1b | 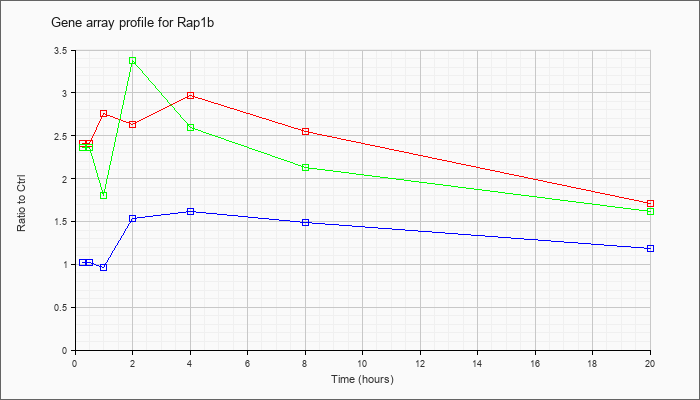 |
M79314 |
GTP-binding protein RAP1b mRNA, partial cds. [M79314] |
KLA | 2.61 |
2.95 |
2.81 |
2.76 |
3.07 |
2.67 |
1.88 |
| ATP | 1.00 |
1.10 |
1.00 |
1.94 |
1.94 |
1.72 |
1.31 |
| KLA/ATP | 2.49 |
2.45 |
2.03 |
4.23 |
3.03 |
2.60 |
1.86 |
|
| Rap1b |  |
NM_024457 |
RAS related protein 1b (Rap1b), mRNA [NM_024457] |
KLA | 2.21 |
2.35 |
2.71 |
2.51 |
2.86 |
2.44 |
1.54 |
| ATP | 1.05 |
.92 |
.93 |
1.14 |
1.30 |
1.26 |
1.07 |
| KLA/ATP | 2.23 |
2.15 |
1.57 |
2.53 |
2.16 |
1.67 |
1.38 |
|
| Rapgef1 | 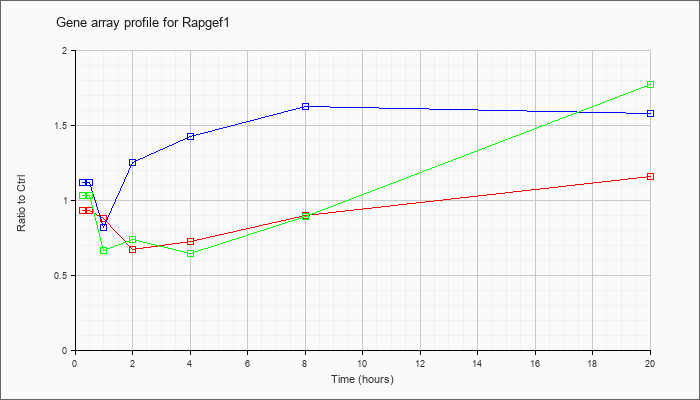 |
NM_001039087 |
Rap guanine nucleotide exchange factor (GEF) 1 (Rapgef1), transcript variant 1, mRNA [NM_001039087] |
KLA | .93 |
.89 |
.88 |
.67 |
.73 |
.90 |
1.16 |
| ATP | 1.12 |
1.09 |
.82 |
1.25 |
1.43 |
1.63 |
1.58 |
| KLA/ATP | 1.03 |
.95 |
.67 |
.74 |
.65 |
.89 |
1.77 |
|
| Rbx1 | 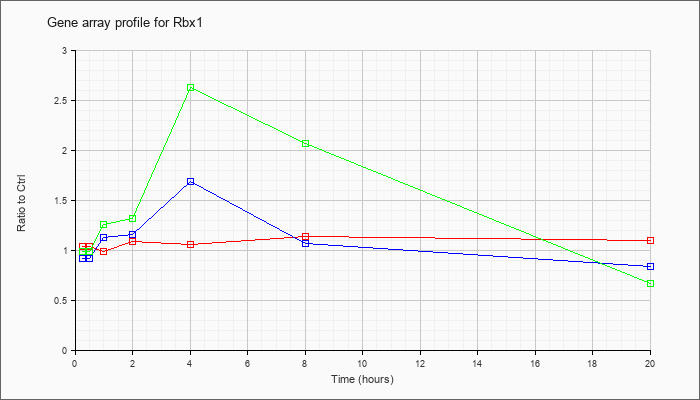 |
NM_019712 |
ring-box 1 (Rbx1), mRNA [NM_019712] |
KLA | 1.04 |
1.01 |
.98 |
1.09 |
1.06 |
1.14 |
1.09 |
| ATP | .92 |
.89 |
1.13 |
1.16 |
1.68 |
1.07 |
.83 |
| KLA/ATP | .99 |
.99 |
1.25 |
1.31 |
2.63 |
2.06 |
.67 |
|
| Slc2a1 | 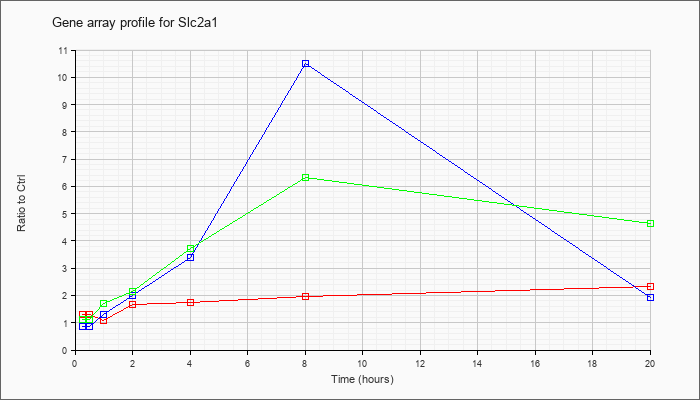 |
NM_011400 |
solute carrier family 2 (facilitated glucose transporter), member 1 (Slc2a1), mRNA [NM_011400] |
KLA | 1.31 |
1.46 |
1.10 |
1.68 |
1.75 |
1.95 |
2.34 |
| ATP | .88 |
.90 |
1.32 |
1.99 |
3.40 |
10.49 |
1.93 |
| KLA/ATP | 1.13 |
1.25 |
1.71 |
2.14 |
3.71 |
6.31 |
4.63 |
|
| Sos1 | 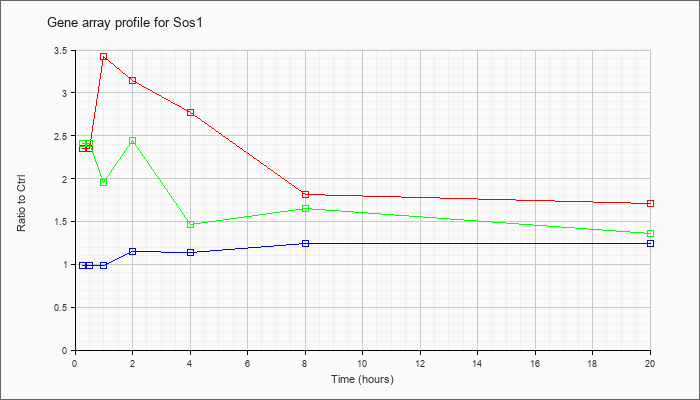 |
NM_009231 |
Son of sevenless homolog 1 (Drosophila) (Sos1), mRNA [NM_009231] |
KLA | 2.36 |
2.40 |
3.42 |
3.15 |
2.78 |
1.81 |
1.71 |
| ATP | .99 |
1.00 |
.99 |
1.15 |
1.14 |
1.25 |
1.24 |
| KLA/ATP | 2.41 |
2.26 |
1.95 |
2.44 |
1.47 |
1.66 |
1.37 |
|
| Sos2 | 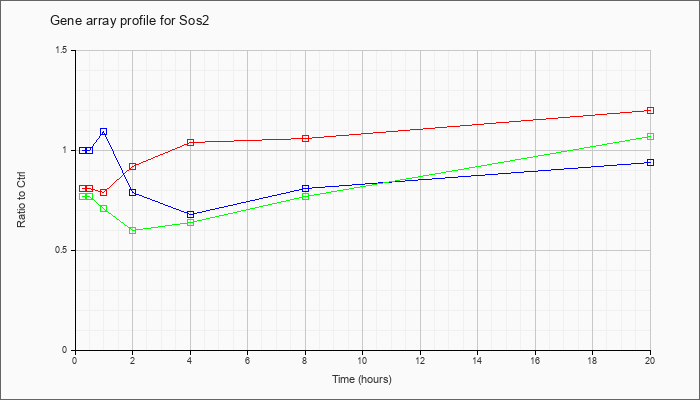 |
XM_127051 |
gb|PREDICTED: Mus musculus Son of sevenless homolog 2 (Drosophila) (Sos2), mRNA [XM_127051] |
KLA | .81 |
.77 |
.79 |
.92 |
1.04 |
1.06 |
1.20 |
| ATP | 1.00 |
1.00 |
1.09 |
.79 |
.68 |
.81 |
.94 |
| KLA/ATP | .77 |
.69 |
.71 |
.60 |
.64 |
.77 |
1.07 |
|
| Tceb1 | 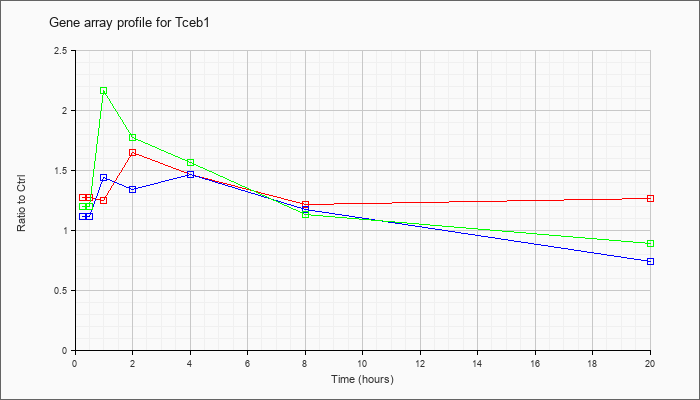 |
NM_026456 |
transcription elongation factor B (SIII), polypeptide 1 (Tceb1), mRNA [NM_026456] |
KLA | 1.27 |
1.21 |
1.25 |
1.65 |
1.46 |
1.21 |
1.26 |
| ATP | 1.11 |
1.01 |
1.44 |
1.34 |
1.46 |
1.17 |
.74 |
| KLA/ATP | 1.20 |
1.48 |
2.16 |
1.77 |
1.56 |
1.13 |
.89 |
|
| Tceb2 | 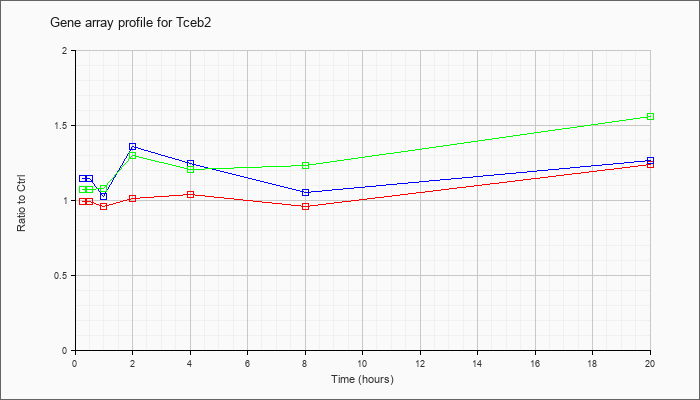 |
NM_026305 |
transcription elongation factor B (SIII), polypeptide 2 (Tceb2), mRNA [NM_026305] |
KLA | .99 |
1.00 |
.96 |
1.01 |
1.04 |
.96 |
1.24 |
| ATP | 1.15 |
1.17 |
1.03 |
1.36 |
1.25 |
1.05 |
1.27 |
| KLA/ATP | 1.07 |
1.13 |
1.08 |
1.30 |
1.21 |
1.23 |
1.56 |
|
| Tgfa | 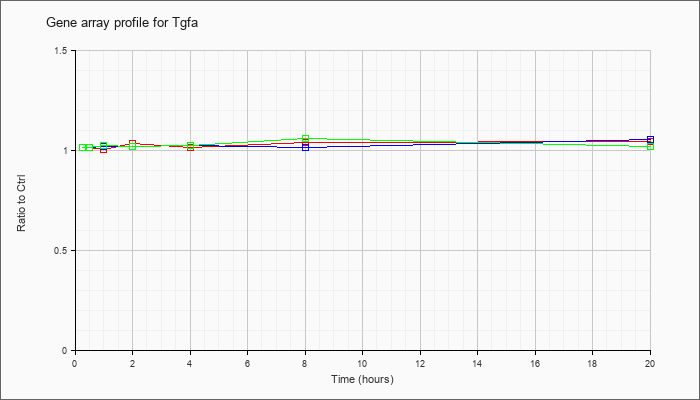 |
NM_031199 |
transforming growth factor alpha (Tgfa), mRNA [NM_031199] |
KLA | 1.01 |
.99 |
1.00 |
1.03 |
1.01 |
1.04 |
1.04 |
| ATP | 1.01 |
1.04 |
1.02 |
1.02 |
1.02 |
1.01 |
1.05 |
| KLA/ATP | 1.01 |
1.02 |
1.02 |
1.02 |
1.02 |
1.06 |
1.02 |
|
| Tgfb1 | 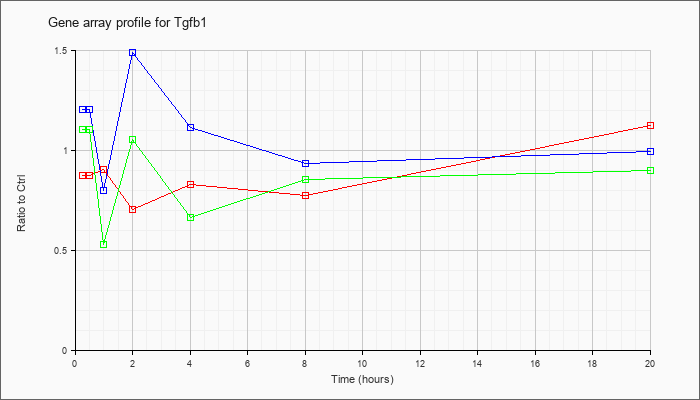 |
NM_011577 |
transforming growth factor, beta 1 (Tgfb1), mRNA [NM_011577] |
KLA | .87 |
.89 |
.90 |
.70 |
.83 |
.77 |
1.12 |
| ATP | 1.20 |
1.41 |
.80 |
1.49 |
1.11 |
.93 |
.99 |
| KLA/ATP | 1.10 |
1.20 |
.53 |
1.05 |
.66 |
.85 |
.90 |
|
| Tgfb2 | 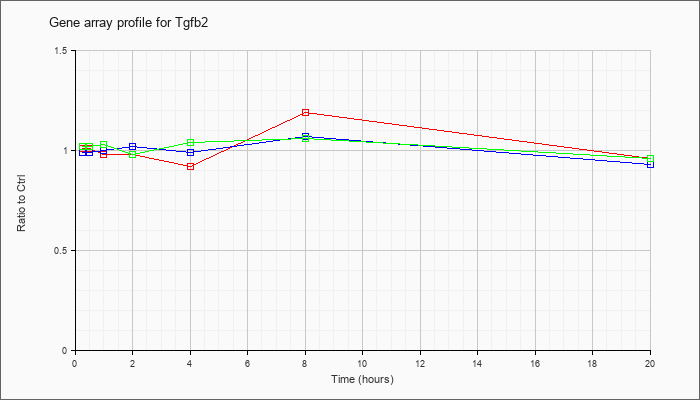 |
NM_009367 |
transforming growth factor, beta 2 (Tgfb2), mRNA [NM_009367] |
KLA | 1.01 |
.91 |
.98 |
.98 |
.92 |
1.19 |
.96 |
| ATP | .99 |
1.03 |
1.00 |
1.02 |
.99 |
1.07 |
.93 |
| KLA/ATP | 1.02 |
.99 |
1.03 |
.98 |
1.04 |
1.06 |
.96 |
|
| Tgfb3 | 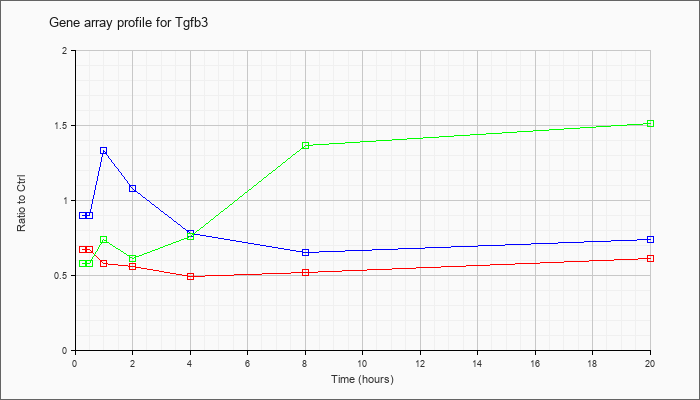 |
NM_009368 |
transforming growth factor, beta 3 (Tgfb3), mRNA [NM_009368] |
KLA | .67 |
.64 |
.58 |
.56 |
.49 |
.52 |
.61 |
| ATP | .90 |
.92 |
1.33 |
1.08 |
.78 |
.65 |
.74 |
| KLA/ATP | .58 |
.60 |
.74 |
.61 |
.76 |
1.36 |
1.51 |
|
| Vegfa | 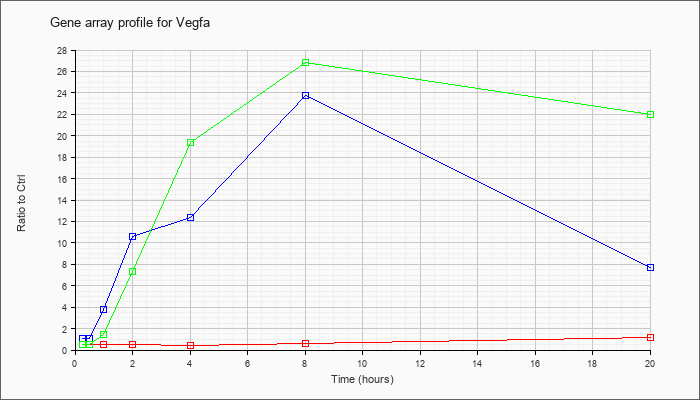 |
NM_001025250 |
vascular endothelial growth factor A (Vegfa), transcript variant 1, mRNA [NM_001025250] |
KLA | .49 |
.48 |
.46 |
.46 |
.39 |
.64 |
1.18 |
| ATP | 1.05 |
1.37 |
3.82 |
10.55 |
12.51 |
24.20 |
7.85 |
| KLA/ATP | .48 |
.59 |
1.45 |
7.42 |
19.75 |
27.42 |
22.35 |
|
| Vegfa |  |
NM_001025257 |
vascular endothelial growth factor A (Vegfa), transcript variant 3, mRNA [NM_001025257] |
KLA | .72 |
.71 |
.66 |
.70 |
.65 |
.80 |
1.22 |
| ATP | 1.01 |
1.21 |
3.24 |
11.12 |
11.09 |
18.59 |
5.43 |
| KLA/ATP | .68 |
.71 |
1.47 |
5.94 |
15.06 |
20.32 |
17.31 |
|
| Vhlh | 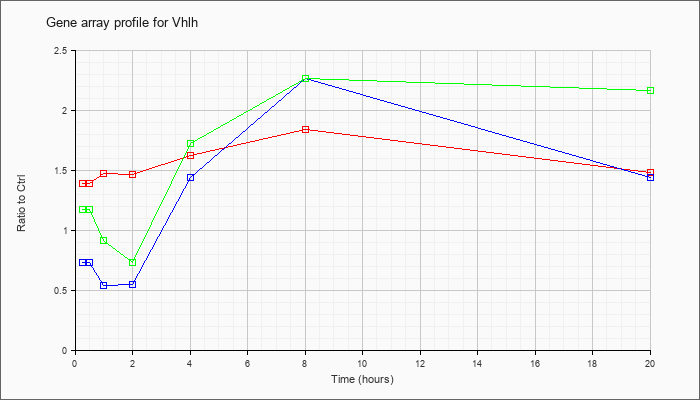 |
S76748 |
gb|mVHLh1=von Hippel-Lindau disease gene [mice, mRNA, 2757 nt]. [S76748] |
KLA | 1.39 |
1.39 |
1.47 |
1.46 |
1.62 |
1.84 |
1.48 |
| ATP | .73 |
.64 |
.54 |
.55 |
1.44 |
2.26 |
1.44 |
| KLA/ATP | 1.17 |
1.13 |
.91 |
.73 |
1.72 |
2.26 |
2.16 |
|