| Gene symbol |
Time course plot |
Accession |
Description |
Treatment |
15min |
30min |
1hr |
2hr |
4hr |
8hr |
20hr |
14927508
3 | 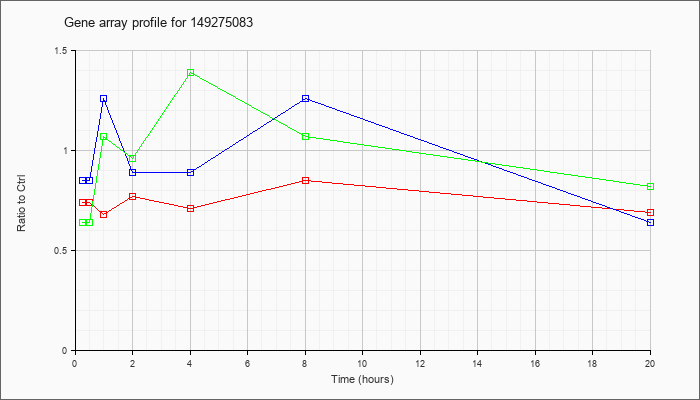 |
149275083 |
Unknown |
KLA | .74 |
.68 |
.68 |
.77 |
.71 |
.85 |
.69 |
| ATP | .85 |
.69 |
1.26 |
.89 |
.89 |
1.26 |
.64 |
| KLA/ATP | .64 |
.63 |
1.07 |
.96 |
1.39 |
1.07 |
.82 |
|
| Cd28 | 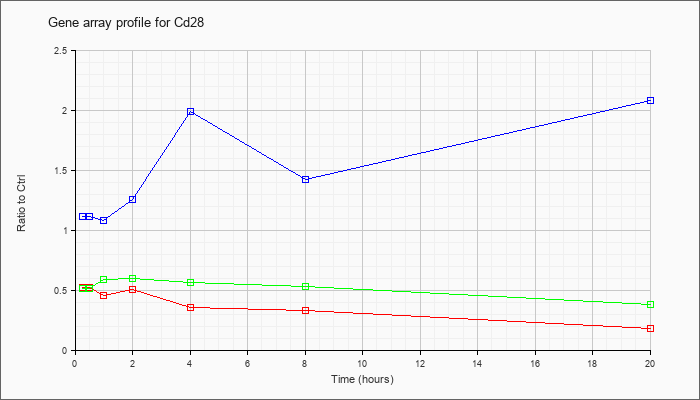 |
NM_007642 |
CD28 antigen (Cd28), mRNA [NM_007642] |
KLA | .52 |
.51 |
.45 |
.50 |
.36 |
.33 |
.18 |
| ATP | 1.11 |
1.26 |
1.08 |
1.25 |
1.99 |
1.42 |
2.08 |
| KLA/ATP | .51 |
.51 |
.58 |
.60 |
.56 |
.53 |
.38 |
|
| Cd80 | 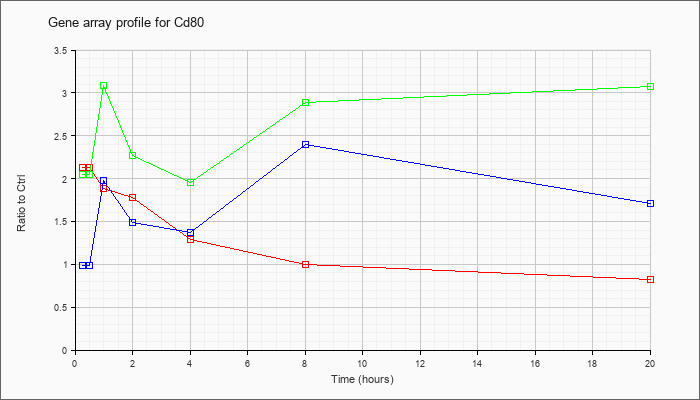 |
NM_009855 |
CD80 antigen (Cd80), mRNA [NM_009855] |
KLA | 2.13 |
1.91 |
1.89 |
1.78 |
1.29 |
1.00 |
.82 |
| ATP | .99 |
1.20 |
1.98 |
1.49 |
1.37 |
2.40 |
1.71 |
| KLA/ATP | 2.05 |
2.32 |
3.09 |
2.27 |
1.95 |
2.89 |
3.07 |
|
| Cd86 | 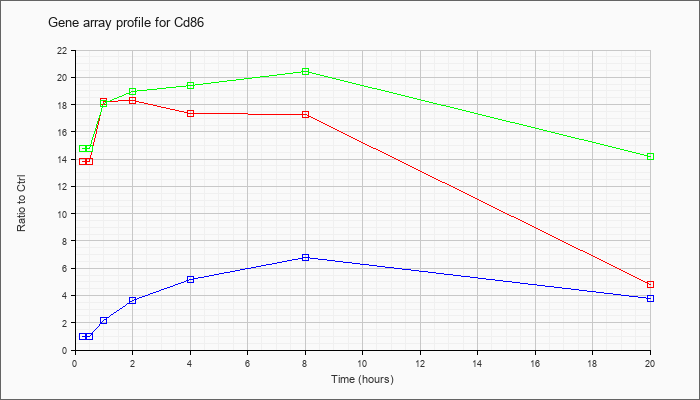 |
NM_019388 |
CD86 antigen (Cd86), mRNA [NM_019388] |
KLA | 13.80 |
14.41 |
18.21 |
18.33 |
17.37 |
17.26 |
4.82 |
| ATP | 1.02 |
1.00 |
2.20 |
3.64 |
5.16 |
6.79 |
3.81 |
| KLA/ATP | 14.80 |
14.66 |
18.10 |
18.96 |
19.38 |
20.42 |
14.21 |
|
| Fas | 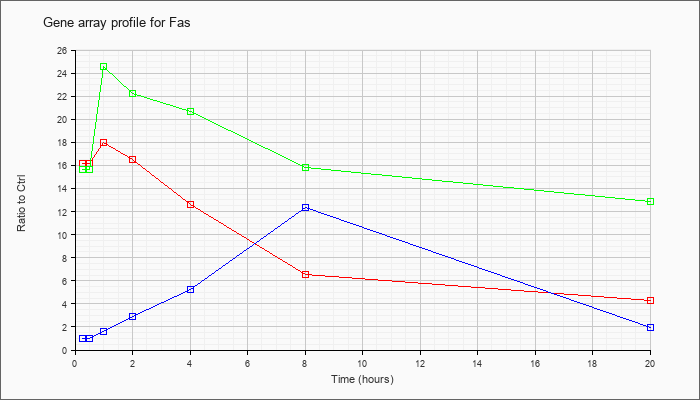 |
AK086933 |
0 day neonate lung cDNA, RIKEN full-length enriched library, clone:E030012O08 product:tumor necrosis factor receptor superfamily, member 6, full insert sequence. [AK086933] |
KLA | 6.31 |
6.56 |
6.33 |
6.70 |
4.93 |
3.74 |
2.48 |
| ATP | .95 |
1.04 |
1.33 |
1.55 |
1.85 |
6.00 |
2.77 |
| KLA/ATP | 5.67 |
7.16 |
7.71 |
7.73 |
6.02 |
5.79 |
7.85 |
|
| Fas |  |
NM_007987 |
Fas (TNF receptor superfamily member 6) (Fas), mRNA [NM_007987] |
KLA | 17.08 |
17.45 |
19.08 |
17.38 |
13.33 |
6.78 |
4.46 |
| ATP | .98 |
.96 |
1.64 |
3.05 |
5.58 |
12.91 |
1.89 |
| KLA/ATP | 16.57 |
17.25 |
26.06 |
23.57 |
22.03 |
16.74 |
13.29 |
|
| Fasl | 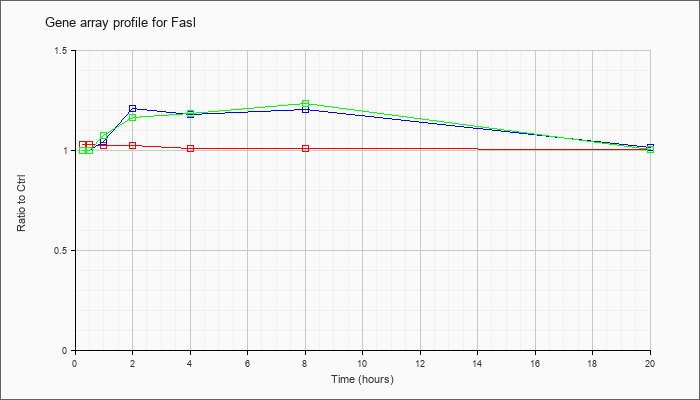 |
NM_010177 |
Fas ligand (TNF superfamily, member 6) (Fasl), mRNA [NM_010177] |
KLA | 1.03 |
1.01 |
1.03 |
1.02 |
1.01 |
1.01 |
1.00 |
| ATP | 1.00 |
.99 |
1.04 |
1.21 |
1.18 |
1.20 |
1.01 |
| KLA/ATP | 1.00 |
1.02 |
1.07 |
1.16 |
1.18 |
1.23 |
1.00 |
|
| Gzmb | 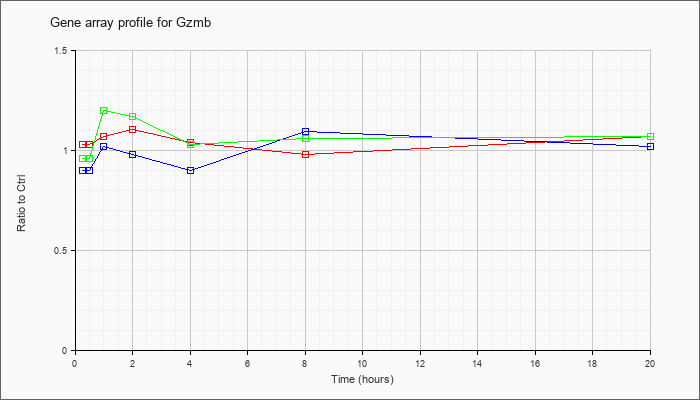 |
NM_013542 |
granzyme B (Gzmb), mRNA [NM_013542] |
KLA | 1.03 |
.96 |
1.07 |
1.10 |
1.04 |
.98 |
1.07 |
| ATP | .90 |
1.05 |
1.02 |
.98 |
.90 |
1.09 |
1.02 |
| KLA/ATP | .96 |
.98 |
1.20 |
1.17 |
1.03 |
1.06 |
1.07 |
|
| H2-Aa | 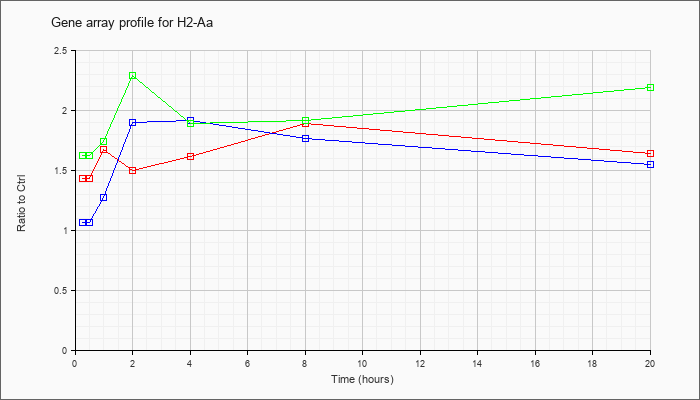 |
NM_010378 |
histocompatibility 2, class II antigen A, alpha (H2-Aa), mRNA [NM_010378] |
KLA | 1.43 |
1.59 |
1.68 |
1.50 |
1.62 |
1.89 |
1.64 |
| ATP | 1.07 |
1.15 |
1.27 |
1.90 |
1.92 |
1.77 |
1.55 |
| KLA/ATP | 1.62 |
1.81 |
1.74 |
2.29 |
1.89 |
1.92 |
2.19 |
|
| H2-Ab1 |  |
BC010322 |
histocompatibility 2, class II antigen A, beta 1, mRNA (cDNA clone MGC:6297 IMAGE:2651058), complete cds [BC010322] |
KLA | 1.73 |
1.76 |
1.77 |
1.55 |
1.95 |
2.10 |
2.27 |
| ATP | 1.03 |
1.18 |
1.32 |
1.99 |
1.79 |
1.56 |
1.61 |
| KLA/ATP | 1.70 |
2.01 |
1.74 |
2.52 |
1.58 |
2.01 |
2.75 |
|
| H2-Ab1 |  |
NM_207105 |
histocompatibility 2, class II antigen A, beta 1 (H2-Ab1), mRNA [NM_207105] |
KLA | 2.02 |
2.15 |
2.07 |
2.09 |
2.32 |
2.78 |
3.08 |
| ATP | 1.11 |
1.20 |
1.42 |
2.33 |
2.33 |
2.25 |
1.87 |
| KLA/ATP | 2.11 |
2.32 |
2.52 |
3.62 |
2.67 |
2.96 |
5.04 |
|
| H2-Bl | 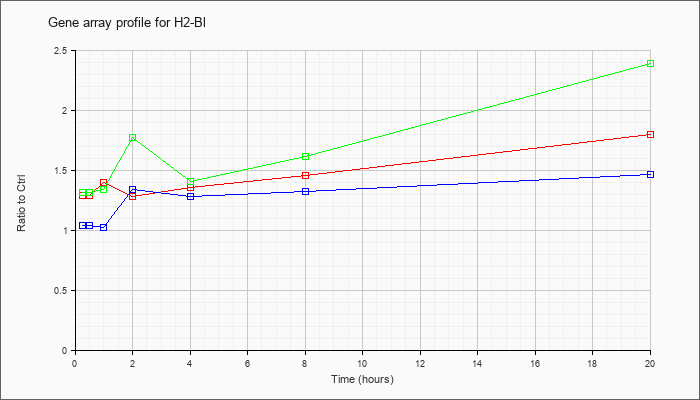 |
NM_008199 |
histocompatibility 2, blastocyst (H2-Bl), mRNA [NM_008199] |
KLA | 1.29 |
1.32 |
1.40 |
1.28 |
1.36 |
1.46 |
1.80 |
| ATP | 1.04 |
1.11 |
1.02 |
1.34 |
1.28 |
1.32 |
1.47 |
| KLA/ATP | 1.32 |
1.39 |
1.34 |
1.77 |
1.41 |
1.61 |
2.39 |
|
| H2-D1 | 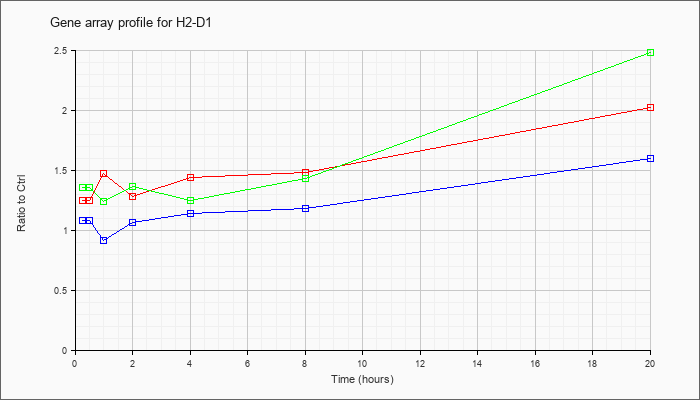 |
AK169750 |
2 days neonate thymus thymic cells cDNA, RIKEN full-length enriched library, clone:E430019E13 product:histocompatibility 2, D region locus 1, full insert sequence [AK169750] |
KLA | .99 |
.98 |
.96 |
.99 |
.93 |
.91 |
.94 |
| ATP | .96 |
1.01 |
1.02 |
.95 |
1.05 |
1.00 |
.98 |
| KLA/ATP | .92 |
.94 |
.97 |
.97 |
.98 |
.92 |
1.03 |
|
| H2-D1 |  |
NM_010380 |
histocompatibility 2, D region locus 1 (H2-D1), mRNA [NM_010380] |
KLA | 1.38 |
1.45 |
1.73 |
1.42 |
1.70 |
1.76 |
2.56 |
| ATP | 1.15 |
1.11 |
.86 |
1.12 |
1.19 |
1.28 |
1.90 |
| KLA/ATP | 1.58 |
1.72 |
1.38 |
1.56 |
1.38 |
1.69 |
3.21 |
|
| H2-DMa | 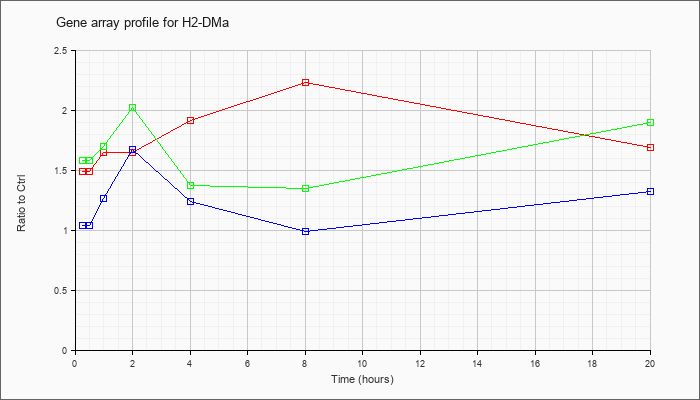 |
NM_010386 |
histocompatibility 2, class II, locus DMa (H2-DMa), mRNA [NM_010386] |
KLA | 1.49 |
1.50 |
1.65 |
1.65 |
1.91 |
2.23 |
1.69 |
| ATP | 1.04 |
1.12 |
1.26 |
1.67 |
1.24 |
.99 |
1.32 |
| KLA/ATP | 1.58 |
1.68 |
1.70 |
2.02 |
1.37 |
1.35 |
1.90 |
|
| H2-DMb1 | 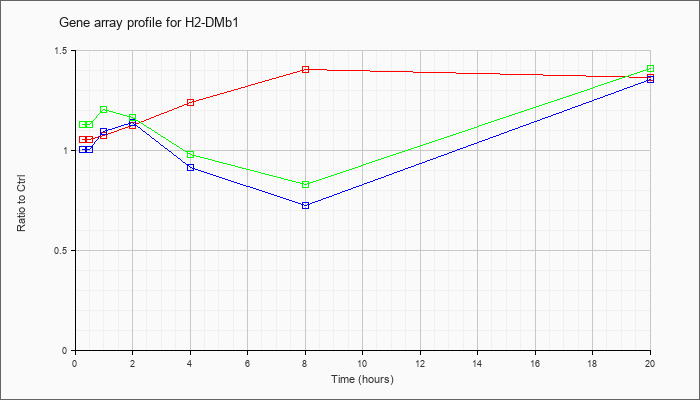 |
NM_010387 |
histocompatibility 2, class II, locus Mb1 (H2-DMb1), mRNA [NM_010387] |
KLA | 1.06 |
1.14 |
1.08 |
1.13 |
1.24 |
1.41 |
1.37 |
| ATP | 1.01 |
1.02 |
1.09 |
1.14 |
.92 |
.73 |
1.36 |
| KLA/ATP | 1.13 |
1.05 |
1.21 |
1.17 |
.98 |
.83 |
1.41 |
|
| H2-DMb2 | 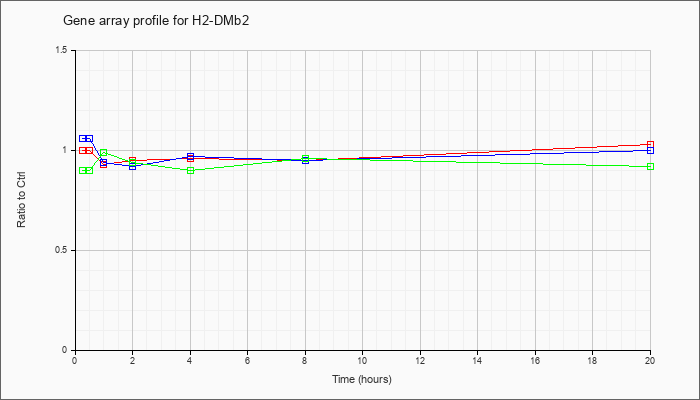 |
NM_010388 |
histocompatibility 2, class II, locus Mb2 (H2-DMb2), mRNA [NM_010388] |
KLA | 1.00 |
1.04 |
.93 |
.95 |
.96 |
.95 |
1.03 |
| ATP | 1.06 |
1.04 |
.94 |
.92 |
.97 |
.95 |
1.00 |
| KLA/ATP | .90 |
.94 |
.99 |
.94 |
.90 |
.96 |
.92 |
|
| H2-Eb1 | 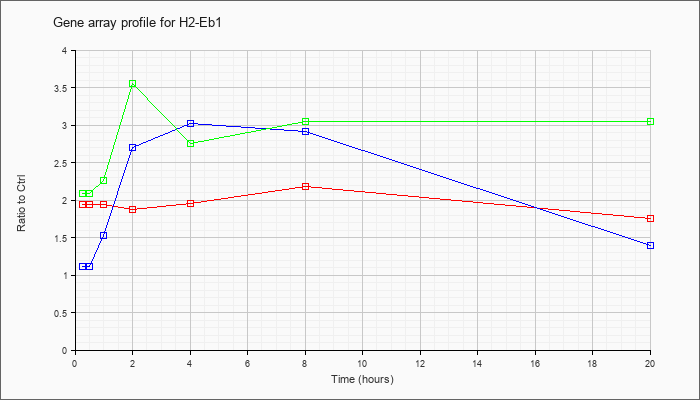 |
X52641 |
gb|Mouse mRNA for I-A beta NOD. [X52641] |
KLA | 1.94 |
2.06 |
1.94 |
1.87 |
1.96 |
2.18 |
1.76 |
| ATP | 1.11 |
1.20 |
1.53 |
2.70 |
3.02 |
2.91 |
1.40 |
| KLA/ATP | 2.09 |
2.11 |
2.26 |
3.55 |
2.76 |
3.05 |
3.05 |
|
| H2-K1 | 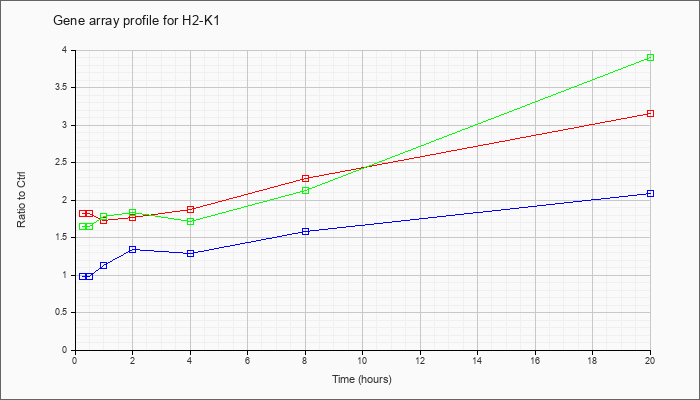 |
M69070 |
mRNA, complete cds. [M69070] |
KLA | 1.76 |
1.66 |
1.78 |
1.78 |
1.87 |
2.23 |
2.99 |
| ATP | .98 |
.97 |
1.19 |
1.45 |
1.36 |
1.68 |
2.17 |
| KLA/ATP | 1.68 |
1.70 |
1.87 |
1.96 |
1.87 |
2.13 |
3.82 |
|
| H2-K1 |  |
NM_001001892 |
histocompatibility 2, K1, K region (H2-K1), transcript variant 1, mRNA [NM_001001892] |
KLA | 1.82 |
1.66 |
1.73 |
1.76 |
1.87 |
2.29 |
3.17 |
| ATP | .98 |
.97 |
1.13 |
1.32 |
1.29 |
1.57 |
2.09 |
| KLA/ATP | 1.64 |
1.71 |
1.77 |
1.82 |
1.71 |
2.12 |
3.91 |
|
| H2-M1 | 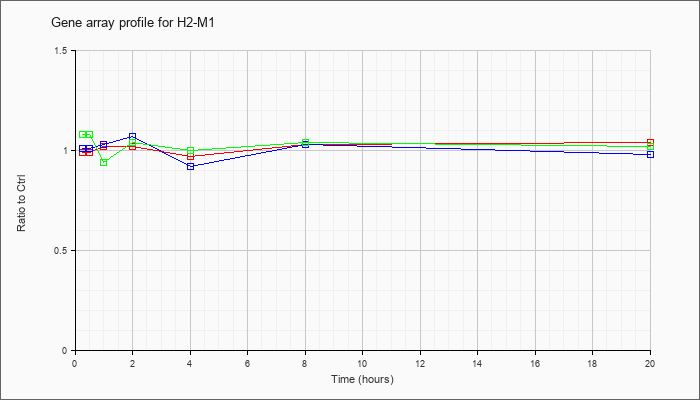 |
NM_177636 |
histocompatibility 2, M region locus 1 (H2-M1), mRNA [NM_177636] |
KLA | .99 |
1.01 |
1.02 |
1.02 |
.97 |
1.03 |
1.04 |
| ATP | 1.01 |
.99 |
1.03 |
1.07 |
.92 |
1.03 |
.98 |
| KLA/ATP | 1.08 |
.97 |
.94 |
1.04 |
1.00 |
1.04 |
1.02 |
|
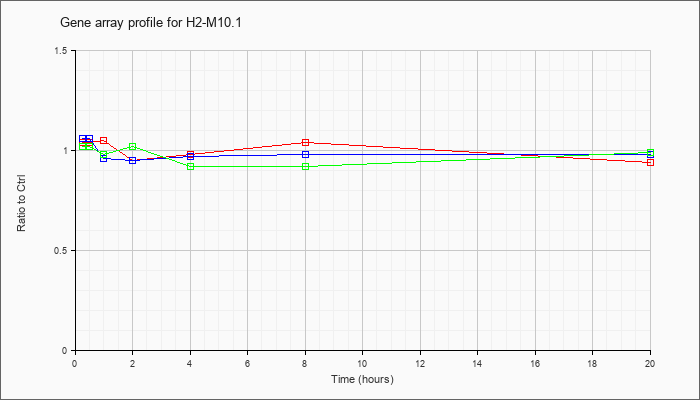
H2-M10.1
|  |
NM_013544 |
histocompatibility 2, M region locus 10.1 (H2-M10.1), mRNA [NM_013544] |
KLA | 1.04 |
1.00 |
1.05 |
.95 |
.98 |
1.04 |
.94 |
| ATP | 1.06 |
1.04 |
.96 |
.95 |
.97 |
.98 |
.98 |
| KLA/ATP | 1.02 |
1.07 |
.98 |
1.02 |
.92 |
.92 |
.99 |
|
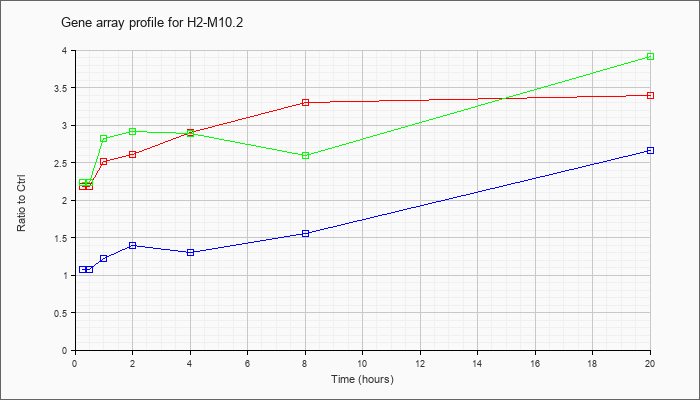
H2-M10.2
|  |
NM_177923 |
histocompatibility 2, M region locus 10.2 (H2-M10.2), mRNA [NM_177923] |
KLA | 2.18 |
2.23 |
2.51 |
2.61 |
2.90 |
3.30 |
3.40 |
| ATP | 1.08 |
1.02 |
1.22 |
1.39 |
1.30 |
1.56 |
2.66 |
| KLA/ATP | 2.23 |
2.53 |
2.82 |
2.91 |
2.89 |
2.60 |
3.92 |
|
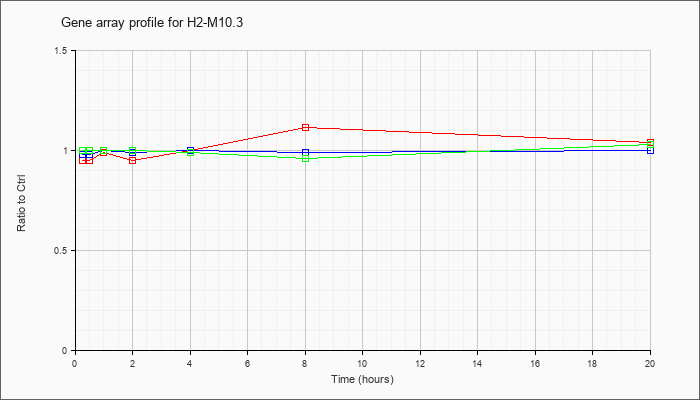
H2-M10.3
|  |
AY211080 |
major histocompatibility complex class Ib M10.3 mRNA, complete cds [AY211080] |
KLA | .95 |
1.08 |
.99 |
.95 |
1.00 |
1.11 |
1.04 |
| ATP | .98 |
.96 |
1.00 |
.99 |
1.00 |
.99 |
1.00 |
| KLA/ATP | 1.00 |
.99 |
1.00 |
1.00 |
.99 |
.96 |
1.03 |
|
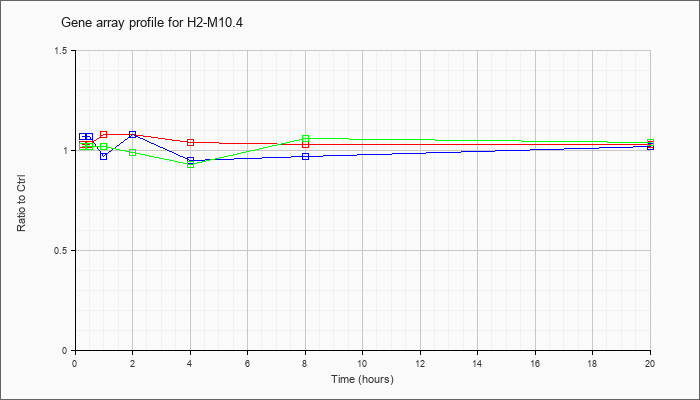
H2-M10.4
|  |
NM_177634 |
histocompatibility 2, M region locus 10.4 (H2-M10.4), mRNA [NM_177634] |
KLA | 1.03 |
1.04 |
1.08 |
1.08 |
1.04 |
1.03 |
1.03 |
| ATP | 1.07 |
1.05 |
.97 |
1.08 |
.95 |
.97 |
1.02 |
| KLA/ATP | 1.02 |
.97 |
1.02 |
.99 |
.93 |
1.06 |
1.04 |
|
H2-M10.5
| 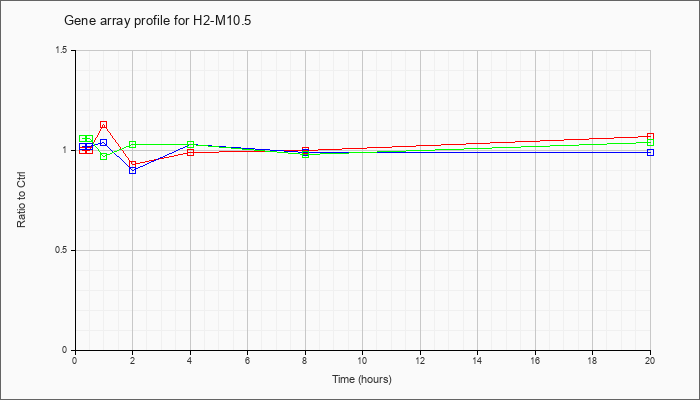 |
NM_177637 |
histocompatibility 2, M region locus 10.5 (H2-M10.5), mRNA [NM_177637] |
KLA | 1.00 |
1.04 |
1.13 |
.93 |
.99 |
1.00 |
1.07 |
| ATP | 1.02 |
1.02 |
1.04 |
.90 |
1.03 |
.99 |
.99 |
| KLA/ATP | 1.06 |
1.03 |
.97 |
1.03 |
1.03 |
.98 |
1.04 |
|
H2-M10.6
| 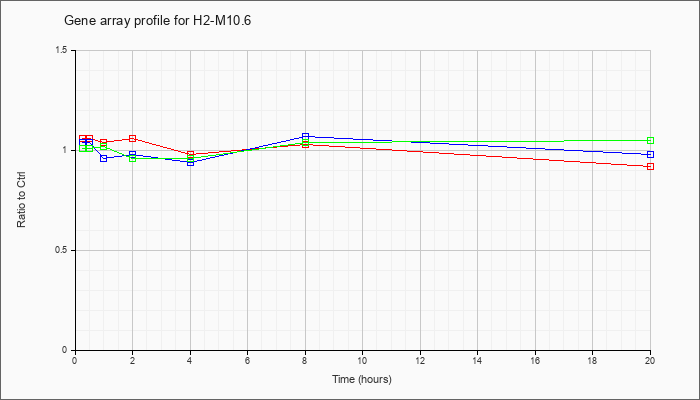 |
NM_201611 |
histocompatibility 2, M region locus 10.6 (H2-M10.6), mRNA [NM_201611] |
KLA | 1.06 |
.99 |
1.04 |
1.06 |
.98 |
1.03 |
.92 |
| ATP | 1.04 |
1.09 |
.96 |
.98 |
.94 |
1.07 |
.98 |
| KLA/ATP | 1.01 |
1.06 |
1.02 |
.96 |
.96 |
1.04 |
1.05 |
|
| H2-M11 | 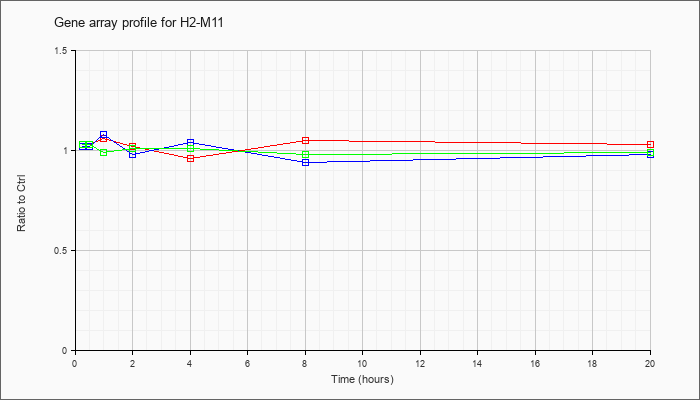 |
NM_177635 |
histocompatibility 2, M region locus 11 (H2-M11), mRNA [NM_177635] |
KLA | 1.03 |
1.07 |
1.06 |
1.02 |
.96 |
1.05 |
1.03 |
| ATP | 1.02 |
.96 |
1.08 |
.98 |
1.04 |
.94 |
.98 |
| KLA/ATP | 1.03 |
1.03 |
.99 |
1.01 |
1.01 |
.98 |
.99 |
|
| H2-M3 | 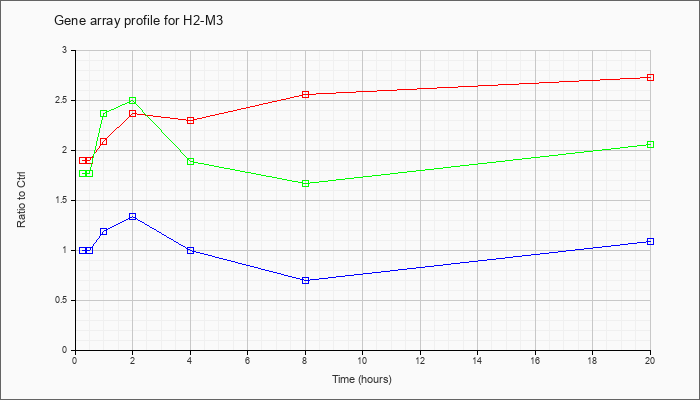 |
NM_013819 |
histocompatibility 2, M region locus 3 (H2-M3), mRNA [NM_013819] |
KLA | 1.90 |
2.04 |
2.09 |
2.37 |
2.30 |
2.56 |
2.73 |
| ATP | 1.00 |
1.00 |
1.19 |
1.34 |
1.00 |
.70 |
1.09 |
| KLA/ATP | 1.77 |
1.94 |
2.37 |
2.49 |
1.89 |
1.67 |
2.06 |
|
| H2-M9 | 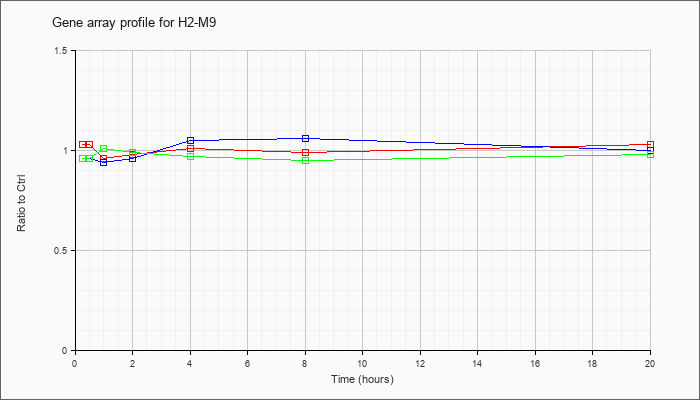 |
NM_008205 |
histocompatibility 2, M region locus 9 (H2-M9), mRNA [NM_008205] |
KLA | 1.03 |
.99 |
.96 |
.98 |
1.01 |
.99 |
1.03 |
| ATP | .96 |
1.06 |
.94 |
.96 |
1.05 |
1.06 |
1.00 |
| KLA/ATP | .96 |
1.01 |
1.01 |
.99 |
.97 |
.95 |
.98 |
|
| H2-Oa | 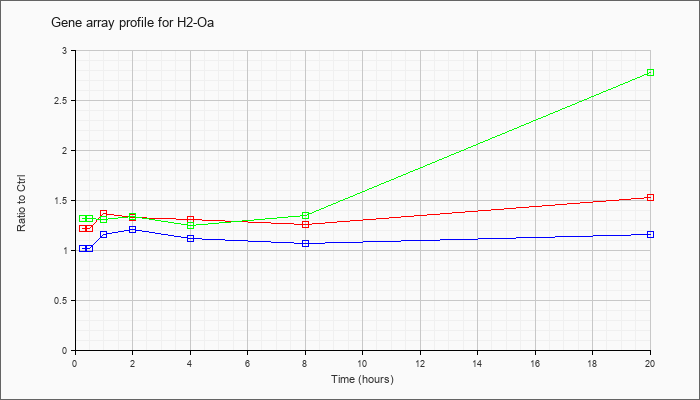 |
NM_008206 |
histocompatibility 2, O region alpha locus (H2-Oa), mRNA [NM_008206] |
KLA | 1.22 |
1.32 |
1.37 |
1.33 |
1.31 |
1.26 |
1.53 |
| ATP | 1.02 |
1.05 |
1.16 |
1.21 |
1.12 |
1.07 |
1.16 |
| KLA/ATP | 1.32 |
1.34 |
1.31 |
1.34 |
1.25 |
1.35 |
2.78 |
|
| H2-Ob | 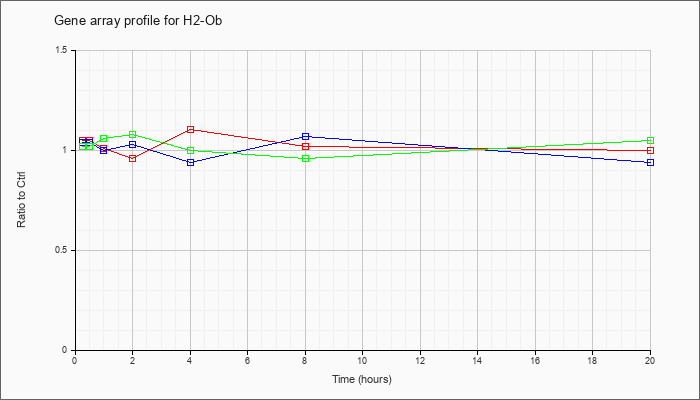 |
NM_010389 |
histocompatibility 2, O region beta locus (H2-Ob), mRNA [NM_010389] |
KLA | 1.05 |
1.07 |
1.01 |
.96 |
1.10 |
1.02 |
1.00 |
| ATP | 1.04 |
.99 |
1.00 |
1.03 |
.94 |
1.07 |
.94 |
| KLA/ATP | 1.02 |
.97 |
1.06 |
1.08 |
1.00 |
.96 |
1.05 |
|
| H2-Q1 | 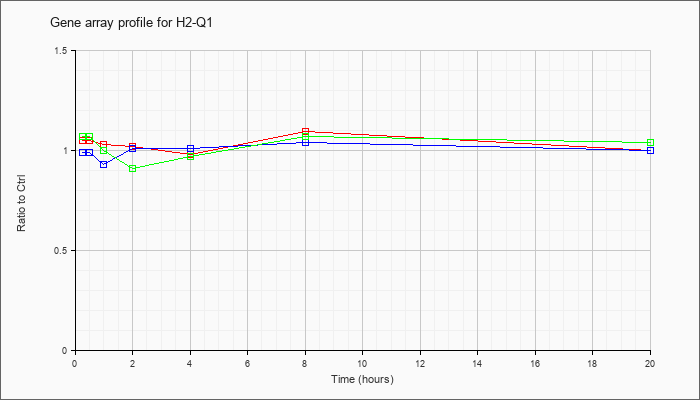 |
NM_010390 |
histocompatibility 2, Q region locus 1 (H2-Q1), mRNA [NM_010390] |
KLA | 1.05 |
1.08 |
1.03 |
1.02 |
.98 |
1.09 |
1.00 |
| ATP | .99 |
.92 |
.93 |
1.01 |
1.01 |
1.04 |
1.00 |
| KLA/ATP | 1.07 |
1.00 |
1.00 |
.91 |
.97 |
1.07 |
1.04 |
|
| H2-Q10 | 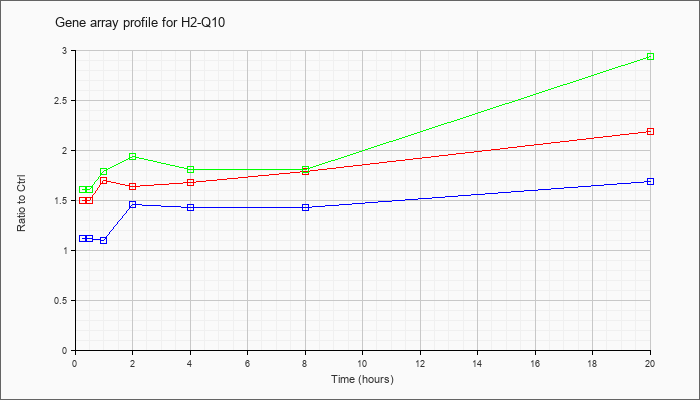 |
NM_010391 |
histocompatibility 2, Q region locus 10 (H2-Q10), mRNA [NM_010391] |
KLA | 1.50 |
1.50 |
1.70 |
1.64 |
1.67 |
1.79 |
2.18 |
| ATP | 1.12 |
1.11 |
1.09 |
1.46 |
1.42 |
1.43 |
1.69 |
| KLA/ATP | 1.61 |
1.72 |
1.79 |
1.94 |
1.81 |
1.81 |
2.93 |
|
| H2-Q2 | 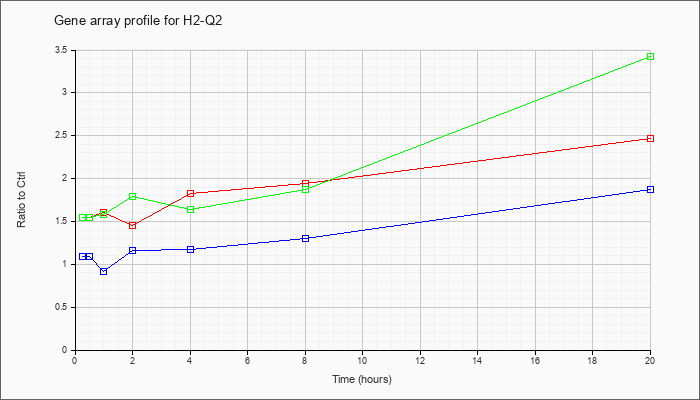 |
NM_010392 |
histocompatibility 2, Q region locus 2 (H2-Q2), mRNA [NM_010392] |
KLA | 1.55 |
1.52 |
1.61 |
1.45 |
1.83 |
1.94 |
2.47 |
| ATP | 1.09 |
1.13 |
.92 |
1.16 |
1.17 |
1.30 |
1.88 |
| KLA/ATP | 1.55 |
1.70 |
1.58 |
1.80 |
1.64 |
1.87 |
3.43 |
|
| H2-Q7 | 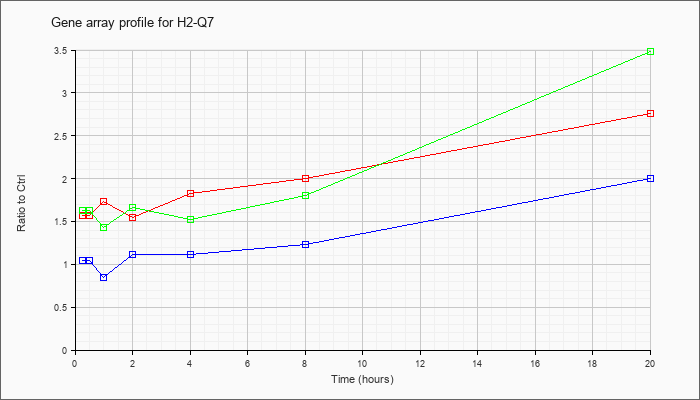 |
NM_010394 |
histocompatibility 2, Q region locus 7 (H2-Q7), mRNA [NM_010394] |
KLA | 1.57 |
1.58 |
1.74 |
1.55 |
1.82 |
2.00 |
2.76 |
| ATP | 1.05 |
1.09 |
.84 |
1.12 |
1.12 |
1.23 |
2.00 |
| KLA/ATP | 1.63 |
1.75 |
1.43 |
1.67 |
1.53 |
1.80 |
3.48 |
|
| H2-Q8 | 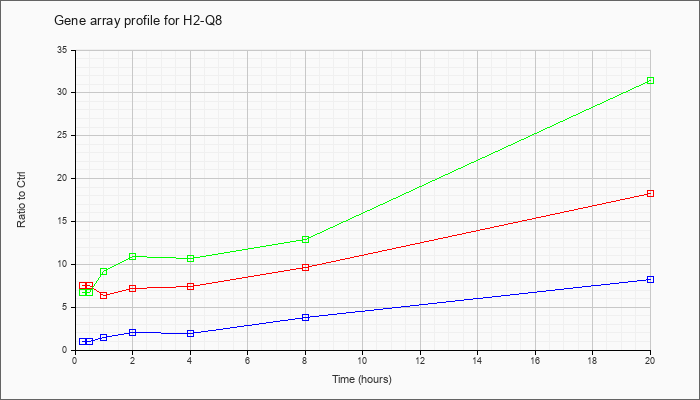 |
NM_023124 |
histocompatibility 2, Q region locus 8 (H2-Q8), mRNA [NM_023124] |
KLA | 7.48 |
6.46 |
6.41 |
7.15 |
7.46 |
9.60 |
18.27 |
| ATP | 1.01 |
.98 |
1.48 |
2.05 |
1.93 |
3.83 |
8.26 |
| KLA/ATP | 6.72 |
7.10 |
9.11 |
10.95 |
10.71 |
12.88 |
31.43 |
|
| H2-T22 | 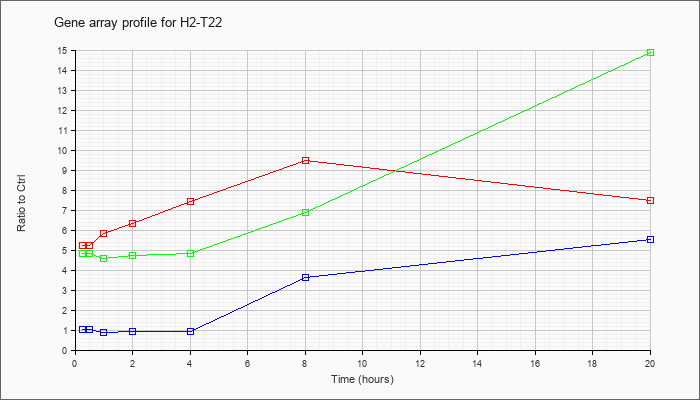 |
NM_010397 |
histocompatibility 2, T region locus 22 (H2-T22), mRNA [NM_010397] |
KLA | 5.21 |
5.40 |
5.81 |
6.31 |
7.42 |
9.49 |
7.50 |
| ATP | 1.01 |
.95 |
.88 |
.92 |
.93 |
3.65 |
5.53 |
| KLA/ATP | 4.83 |
5.12 |
4.58 |
4.74 |
4.81 |
6.89 |
14.89 |
|
| H2-T23 | 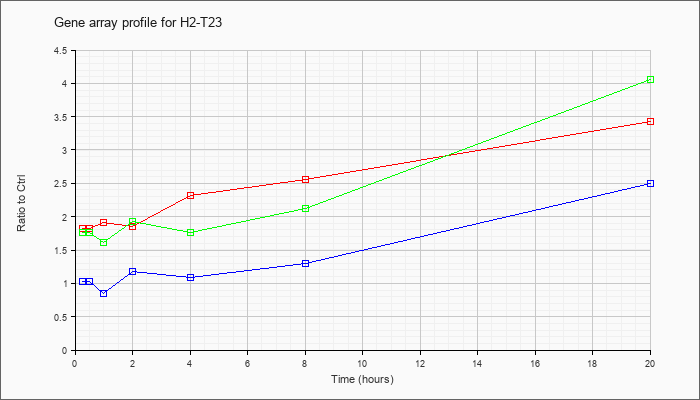 |
NM_010398 |
histocompatibility 2, T region locus 23 (H2-T23), mRNA [NM_010398] |
KLA | 1.83 |
1.80 |
1.92 |
1.86 |
2.32 |
2.56 |
3.43 |
| ATP | 1.03 |
1.08 |
.85 |
1.18 |
1.09 |
1.30 |
2.49 |
| KLA/ATP | 1.76 |
1.92 |
1.62 |
1.93 |
1.76 |
2.13 |
4.06 |
|
| H2-T24 | 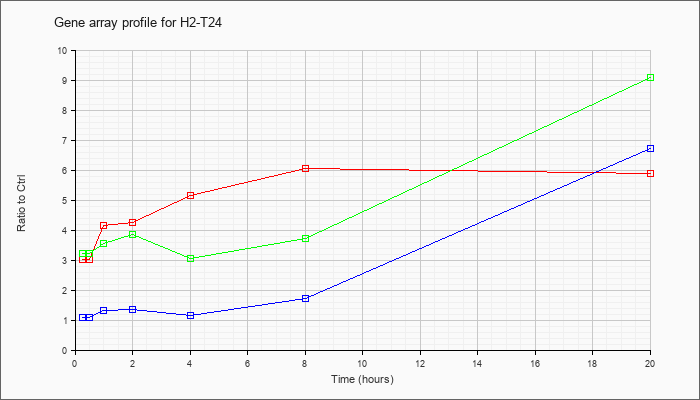 |
NM_008207 |
histocompatibility 2, T region locus 24 (H2-T24), mRNA [NM_008207] |
KLA | 3.02 |
3.26 |
4.14 |
4.26 |
5.15 |
6.06 |
5.89 |
| ATP | 1.08 |
1.09 |
1.32 |
1.35 |
1.17 |
1.71 |
6.71 |
| KLA/ATP | 3.23 |
3.31 |
3.54 |
3.85 |
3.07 |
3.73 |
9.10 |
|
| H2-T3 | 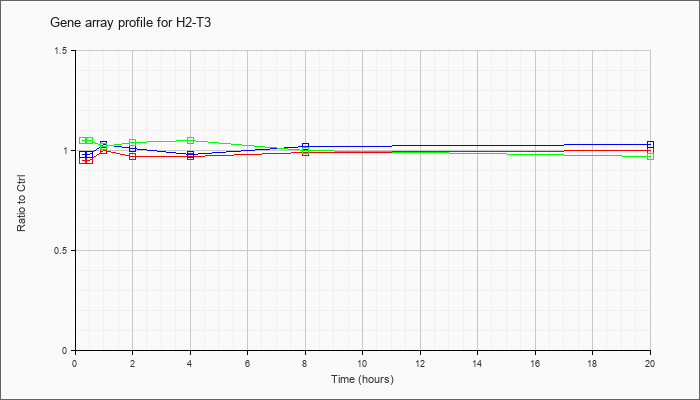 |
NM_008208 |
histocompatibility 2, T region locus 3 (H2-T3), mRNA [NM_008208] |
KLA | .95 |
1.04 |
1.00 |
.97 |
.97 |
.99 |
1.00 |
| ATP | .98 |
1.08 |
1.03 |
1.01 |
.98 |
1.02 |
1.03 |
| KLA/ATP | 1.05 |
.96 |
1.02 |
1.04 |
1.05 |
1.00 |
.97 |
|
| Ifng | 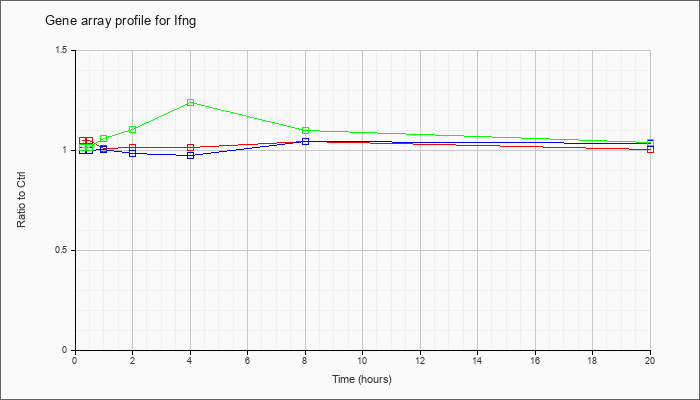 |
NM_008337 |
interferon gamma (Ifng), mRNA [NM_008337] |
KLA | 1.05 |
1.02 |
1.01 |
1.01 |
1.01 |
1.04 |
1.00 |
| ATP | 1.00 |
1.01 |
1.01 |
.98 |
.97 |
1.04 |
1.03 |
| KLA/ATP | 1.01 |
1.01 |
1.06 |
1.10 |
1.24 |
1.10 |
1.04 |
|
| Il1a | 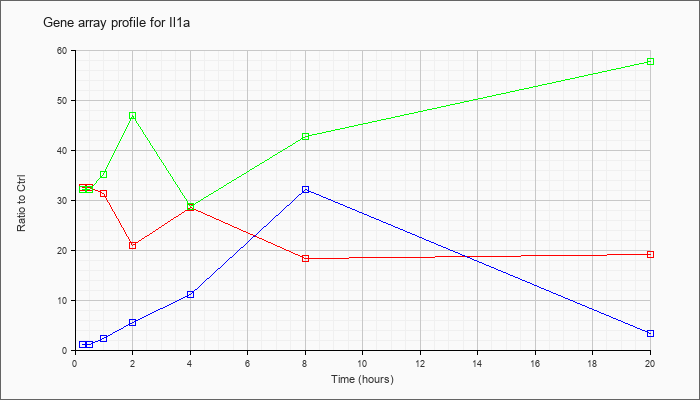 |
NM_010554 |
interleukin 1 alpha (Il1a), mRNA [NM_010554] |
KLA | 32.45 |
35.22 |
31.30 |
20.95 |
28.57 |
18.36 |
19.02 |
| ATP | 1.04 |
1.02 |
2.21 |
5.46 |
11.06 |
32.07 |
3.40 |
| KLA/ATP | 32.02 |
41.20 |
35.12 |
46.91 |
28.76 |
42.72 |
57.80 |
|
| Il1b | 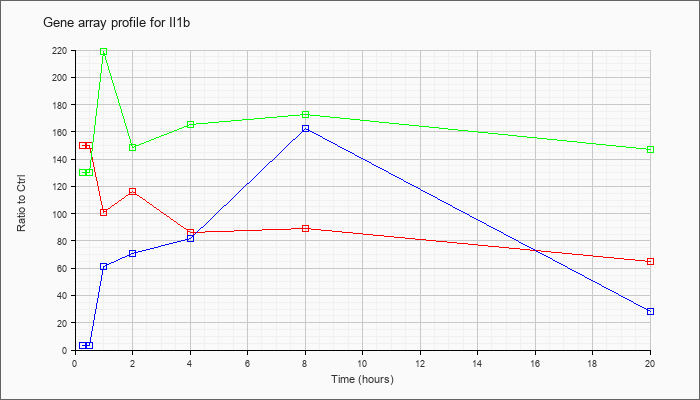 |
NM_008361 |
interleukin 1 beta (Il1b), mRNA [NM_008361] |
KLA | 149.78 |
133.59 |
100.91 |
116.58 |
86.48 |
89.37 |
64.72 |
| ATP | 3.49 |
13.96 |
61.44 |
70.91 |
82.12 |
162.79 |
28.18 |
| KLA/ATP | 130.32 |
139.00 |
219.12 |
148.17 |
165.44 |
172.57 |
146.98 |
|
| Il2 | 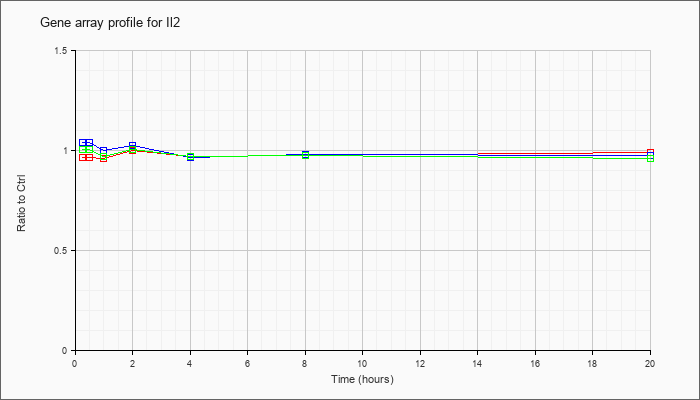 |
NM_008366 |
interleukin 2 (Il2), mRNA [NM_008366] |
KLA | .96 |
.97 |
.96 |
1.00 |
.97 |
.97 |
.99 |
| ATP | 1.04 |
1.02 |
1.00 |
1.03 |
.97 |
.98 |
.97 |
| KLA/ATP | 1.01 |
.98 |
.97 |
1.00 |
.97 |
.97 |
.96 |
|
| Il6 | 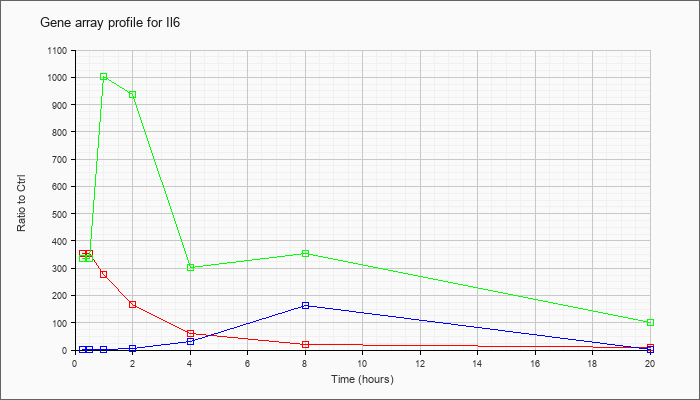 |
NM_031168 |
interleukin 6 (Il6), mRNA [NM_031168] |
KLA | 352.99 |
378.70 |
276.81 |
168.47 |
59.99 |
18.66 |
8.27 |
| ATP | 1.04 |
1.08 |
2.28 |
3.69 |
30.40 |
161.93 |
2.11 |
| KLA/ATP | 334.28 |
528.66 |
###.## |
935.86 |
302.45 |
355.36 |
99.65 |
|
| Klra3 | 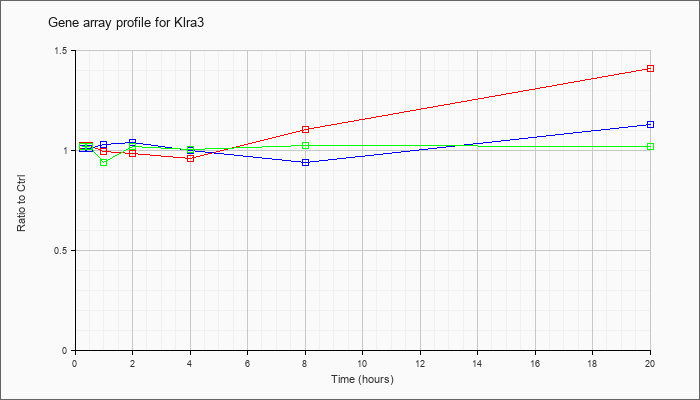 |
NM_010648 |
killer cell lectin-like receptor, subfamily A, member 3 (Klra3), mRNA [NM_010648] |
KLA | 1.02 |
1.00 |
.99 |
.98 |
.96 |
1.10 |
1.41 |
| ATP | 1.01 |
1.01 |
1.03 |
1.04 |
1.00 |
.94 |
1.13 |
| KLA/ATP | 1.02 |
1.00 |
.94 |
1.02 |
1.00 |
1.02 |
1.02 |
|
| Klra7 | 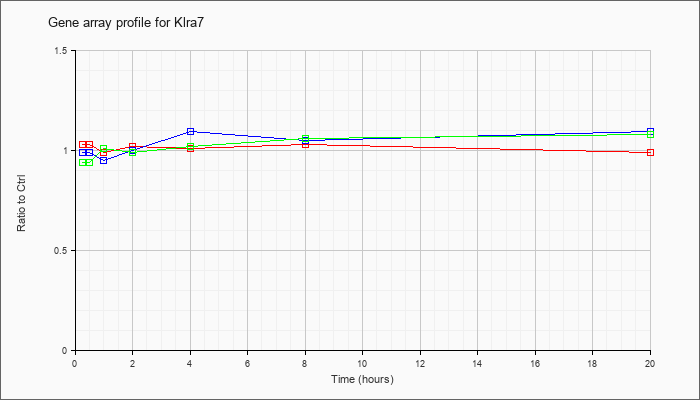 |
NM_014194 |
killer cell lectin-like receptor, subfamily A, member 7 (Klra7), transcript variant 2, mRNA [NM_014194] |
KLA | 1.03 |
.96 |
.99 |
1.02 |
1.01 |
1.03 |
.99 |
| ATP | .99 |
.95 |
.95 |
1.00 |
1.09 |
1.05 |
1.09 |
| KLA/ATP | .94 |
.99 |
1.01 |
.99 |
1.02 |
1.06 |
1.08 |
|
| Klra9 | 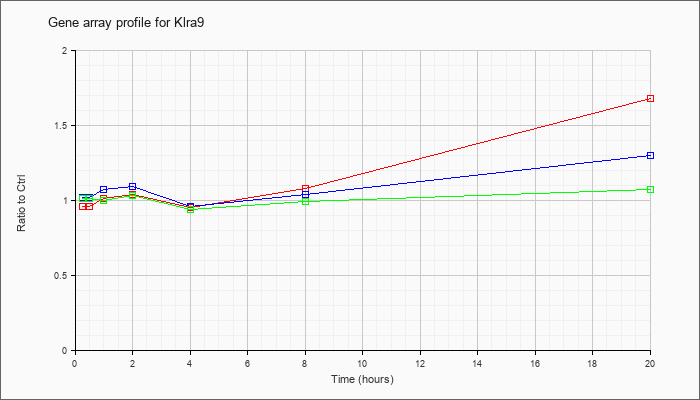 |
NM_010651 |
killer cell lectin-like receptor subfamily A, member 9 (Klra9), mRNA [NM_010651] |
KLA | .96 |
.90 |
1.01 |
1.04 |
.95 |
1.08 |
1.68 |
| ATP | 1.02 |
1.05 |
1.07 |
1.09 |
.96 |
1.04 |
1.30 |
| KLA/ATP | 1.01 |
.98 |
1.00 |
1.03 |
.94 |
.99 |
1.07 |
|
| Klrc1 | 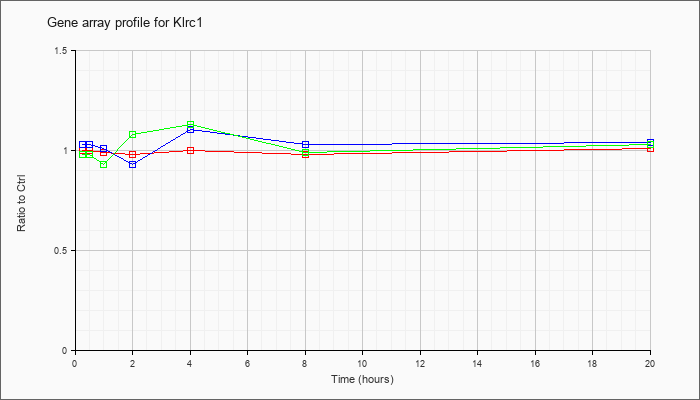 |
NM_010652 |
killer cell lectin-like receptor subfamily C, member 1 (Klrc1), mRNA [NM_010652] |
KLA | 1.00 |
.96 |
.99 |
.98 |
1.00 |
.98 |
1.01 |
| ATP | 1.03 |
.97 |
1.01 |
.93 |
1.10 |
1.03 |
1.04 |
| KLA/ATP | .98 |
.99 |
.93 |
1.08 |
1.13 |
.99 |
1.03 |
|
| Klrd1 | 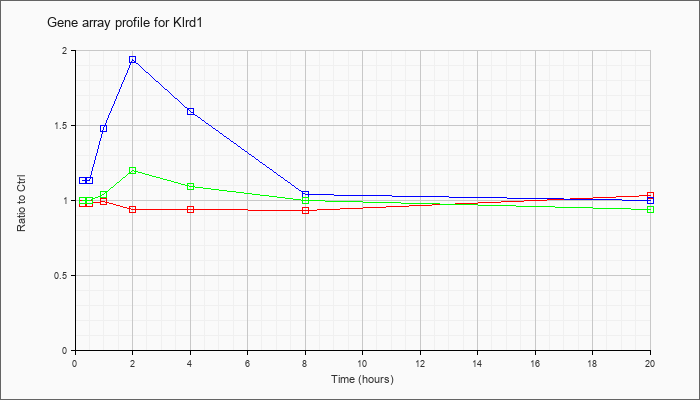 |
NM_010654 |
killer cell lectin-like receptor, subfamily D, member 1 (Klrd1), mRNA [NM_010654] |
KLA | .98 |
.99 |
.99 |
.94 |
.94 |
.93 |
1.03 |
| ATP | 1.13 |
1.28 |
1.48 |
1.94 |
1.59 |
1.04 |
1.00 |
| KLA/ATP | 1.00 |
.95 |
1.04 |
1.20 |
1.09 |
1.00 |
.94 |
|
LOC67670
8 | 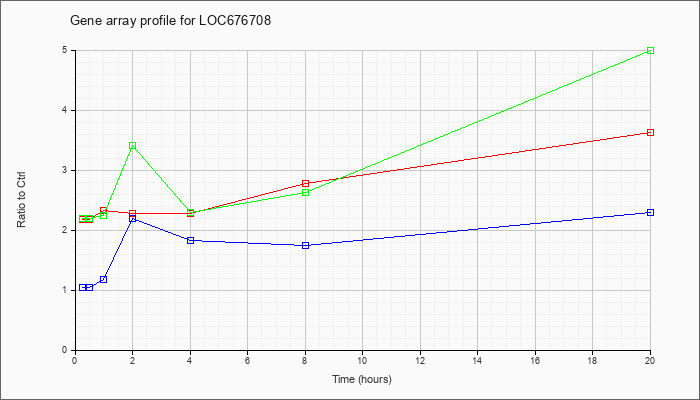 |
XM_992322 |
ref|PREDICTED: Mus musculus similar to class I (Qa) Q2k antigen (LOC676708), mRNA [XM_992322] |
KLA | 2.18 |
2.22 |
2.32 |
2.27 |
2.28 |
2.77 |
3.62 |
| ATP | 1.04 |
1.17 |
1.17 |
2.19 |
1.83 |
1.75 |
2.30 |
| KLA/ATP | 2.20 |
2.44 |
2.25 |
3.41 |
2.30 |
2.63 |
5.00 |
|
Mm.19506
1 | 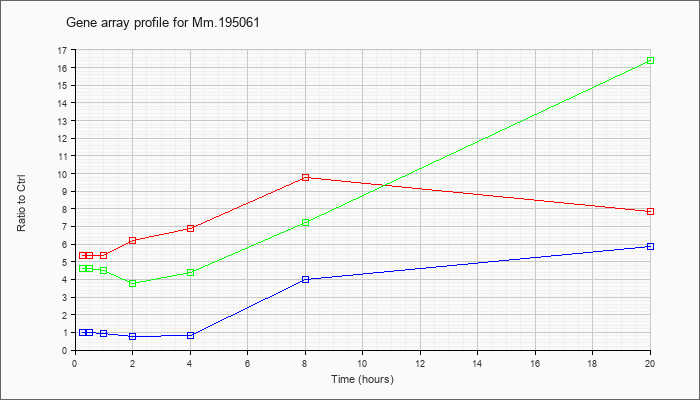 |
76253707 |
Unknown |
KLA | 5.35 |
4.83 |
5.36 |
6.20 |
6.87 |
9.80 |
7.83 |
| ATP | .97 |
.90 |
.96 |
.77 |
.80 |
3.98 |
5.87 |
| KLA/ATP | 4.63 |
4.51 |
4.51 |
3.75 |
4.39 |
7.21 |
16.38 |
|
Mm.25500
3 | 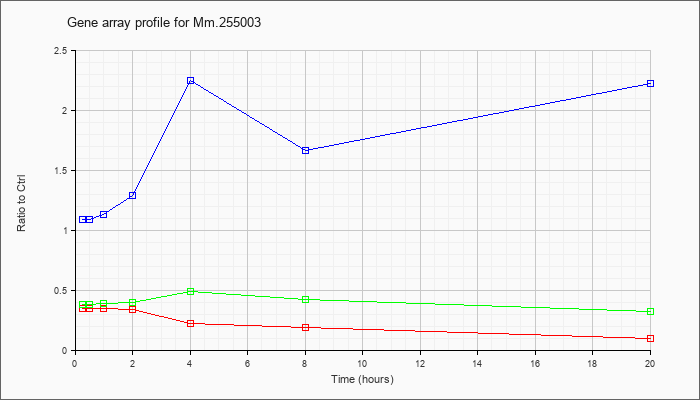 |
161484596 |
Unknown |
KLA | .35 |
.34 |
.35 |
.34 |
.22 |
.19 |
.10 |
| ATP | 1.09 |
1.27 |
1.13 |
1.29 |
2.25 |
1.66 |
2.22 |
| KLA/ATP | .38 |
.36 |
.39 |
.40 |
.49 |
.42 |
.32 |
|
Mm.43967
5 | 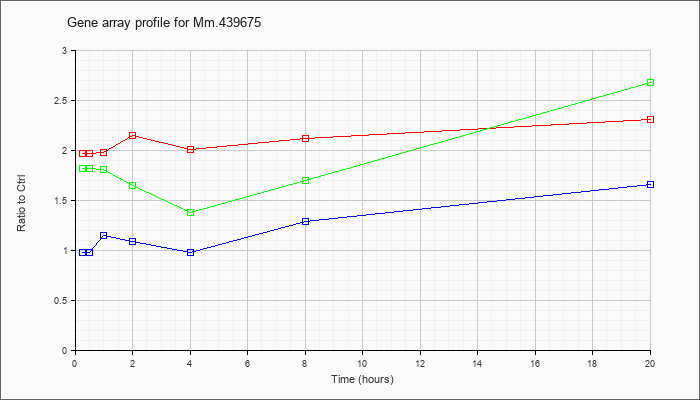 |
133778954 |
Unknown |
KLA | 1.76 |
1.74 |
1.70 |
1.72 |
1.75 |
2.14 |
3.19 |
| ATP | 1.00 |
1.01 |
1.18 |
1.28 |
1.25 |
1.48 |
1.84 |
| KLA/ATP | 1.67 |
1.64 |
1.70 |
1.79 |
1.63 |
2.04 |
3.36 |
|
Mm.43967
5 |  |
149275074 |
Unknown |
KLA | 2.17 |
2.23 |
2.26 |
2.58 |
2.26 |
2.10 |
1.43 |
| ATP | .96 |
.91 |
1.12 |
.90 |
.70 |
1.10 |
1.47 |
| KLA/ATP | 1.96 |
2.10 |
1.92 |
1.51 |
1.12 |
1.35 |
1.99 |
|
Mm.44165
1 |  |
160333403 |
Unknown |
KLA | 5.45 |
5.27 |
5.02 |
6.34 |
6.63 |
10.05 |
8.10 |
| ATP | .91 |
.84 |
.96 |
.75 |
.78 |
3.91 |
6.03 |
| KLA/ATP | 4.29 |
4.38 |
4.50 |
3.55 |
4.22 |
7.22 |
17.63 |
|
| Prf1 | 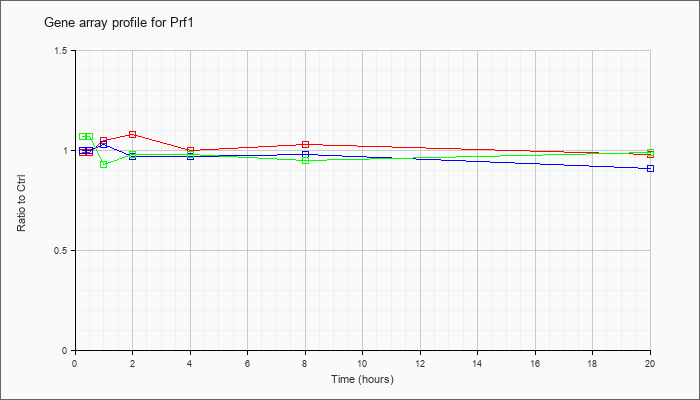 |
NM_011073 |
perforin 1 (pore forming protein) (Prf1), mRNA [NM_011073] |
KLA | .99 |
1.02 |
1.05 |
1.08 |
1.00 |
1.03 |
.98 |
| ATP | 1.00 |
.97 |
1.03 |
.97 |
.97 |
.98 |
.91 |
| KLA/ATP | 1.07 |
.99 |
.93 |
.98 |
.98 |
.95 |
.99 |
|
| Tnf | 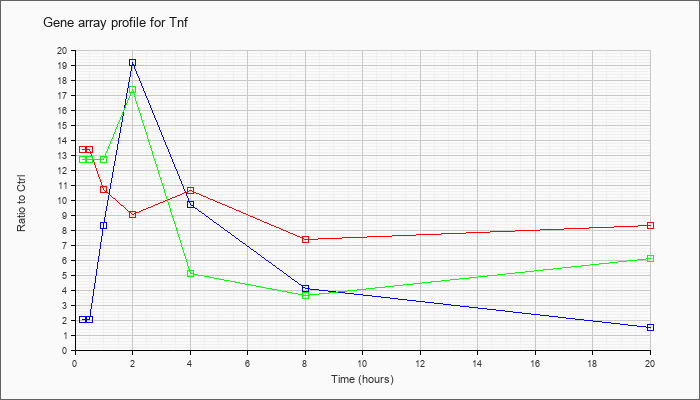 |
NM_013693 |
tumor necrosis factor (Tnf), mRNA [NM_013693] |
KLA | 13.35 |
12.76 |
10.71 |
9.03 |
10.64 |
7.39 |
8.32 |
| ATP | 2.01 |
2.48 |
8.27 |
19.20 |
9.72 |
4.09 |
1.49 |
| KLA/ATP | 12.72 |
15.76 |
12.73 |
17.34 |
5.10 |
3.61 |
6.07 |
|