| Gene symbol |
Time course plot |
Accession |
Description |
Treatment |
15min |
30min |
1hr |
2hr |
4hr |
8hr |
20hr |
| Agxt | 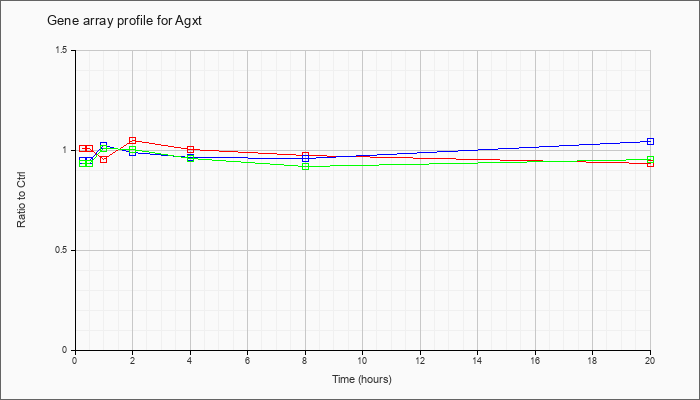 |
NM_016702 |
alanine-glyoxylate aminotransferase (Agxt), mRNA [NM_016702] |
KLA | 1.01 |
.95 |
.96 |
1.05 |
1.01 |
.98 |
.94 |
| ATP | .95 |
1.03 |
1.03 |
.99 |
.97 |
.96 |
1.05 |
| KLA/ATP | .94 |
.99 |
1.01 |
1.01 |
.96 |
.92 |
.96 |
|
| Agxt2 | 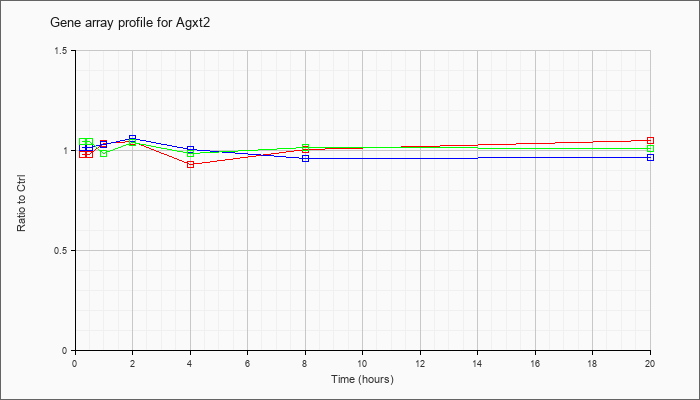 |
NM_001031851 |
alanine-glyoxylate aminotransferase 2 (Agxt2), nuclear gene encoding mitochondrial protein, mRNA [NM_001031851] |
KLA | .98 |
.99 |
1.04 |
1.05 |
.93 |
1.01 |
1.05 |
| ATP | 1.02 |
1.03 |
1.03 |
1.06 |
1.01 |
.96 |
.97 |
| KLA/ATP | 1.05 |
.98 |
.99 |
1.04 |
.99 |
1.02 |
1.01 |
|
| Alas1 | 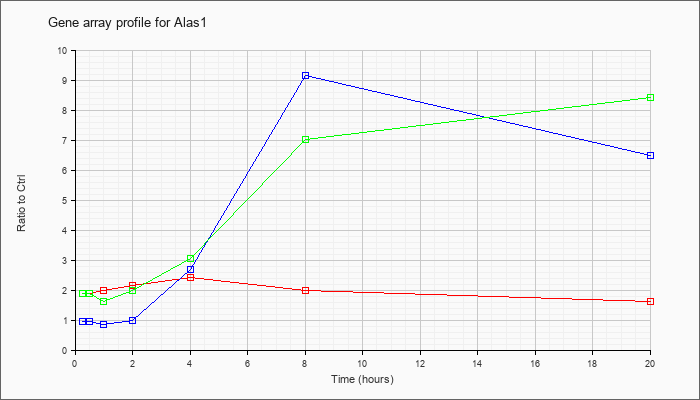 |
NM_020559 |
aminolevulinic acid synthase 1 (Alas1), mRNA [NM_020559] |
KLA | 1.90 |
2.05 |
1.99 |
2.15 |
2.41 |
2.00 |
1.62 |
| ATP | .96 |
1.03 |
.87 |
.98 |
2.70 |
9.15 |
6.49 |
| KLA/ATP | 1.88 |
2.04 |
1.63 |
1.99 |
3.06 |
7.03 |
8.41 |
|
| Alas2 |  |
NM_009653 |
aminolevulinic acid synthase 2, erythroid (Alas2), nuclear gene encoding mitochondrial protein, transcript variant 1, mRNA [NM_009653] |
KLA | .94 |
1.05 |
1.03 |
.98 |
.93 |
.99 |
1.00 |
| ATP | 1.00 |
1.03 |
1.02 |
1.05 |
.95 |
.96 |
1.04 |
| KLA/ATP | 1.07 |
1.08 |
1.03 |
.94 |
.97 |
.98 |
.98 |
|
| Aldh7a1 | 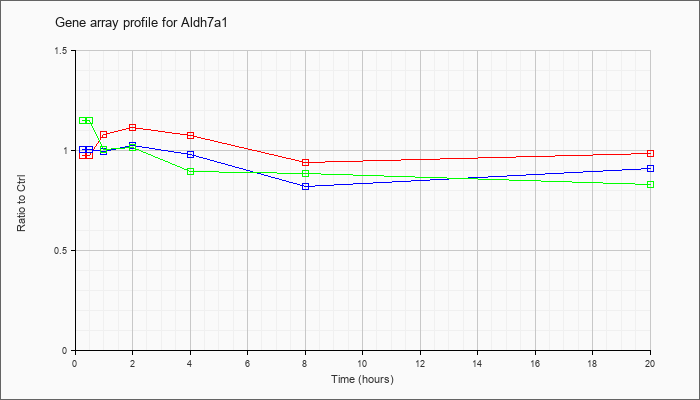 |
NM_138600 |
aldehyde dehydrogenase family 7, member A1 (Aldh7a1), mRNA [NM_138600] |
KLA | .98 |
1.03 |
1.08 |
1.12 |
1.08 |
.94 |
.99 |
| ATP | 1.01 |
1.03 |
1.00 |
1.03 |
.98 |
.82 |
.91 |
| KLA/ATP | 1.15 |
1.12 |
1.01 |
1.02 |
.90 |
.89 |
.83 |
|
| Aoc2 | 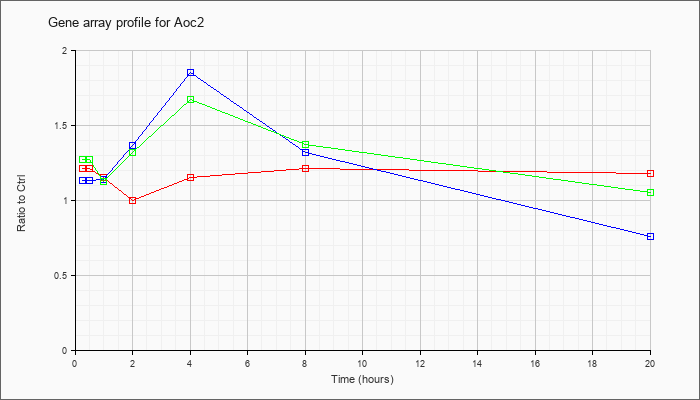 |
NM_178932 |
amine oxidase, copper containing 2 (retina-specific) (Aoc2), mRNA [NM_178932] |
KLA | 1.21 |
1.19 |
1.15 |
1.00 |
1.15 |
1.21 |
1.18 |
| ATP | 1.13 |
1.13 |
1.14 |
1.36 |
1.85 |
1.32 |
.76 |
| KLA/ATP | 1.27 |
1.34 |
1.12 |
1.32 |
1.67 |
1.37 |
1.05 |
|
| Aoc3 | 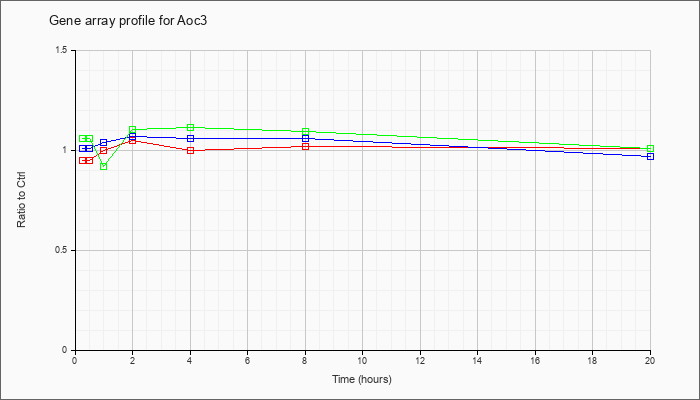 |
NM_009675 |
amine oxidase, copper containing 3 (Aoc3), mRNA [NM_009675] |
KLA | .95 |
1.06 |
1.00 |
1.05 |
1.00 |
1.02 |
1.01 |
| ATP | 1.01 |
.90 |
1.04 |
1.07 |
1.06 |
1.06 |
.97 |
| KLA/ATP | 1.06 |
.98 |
.92 |
1.10 |
1.11 |
1.09 |
1.01 |
|
| Bhmt | 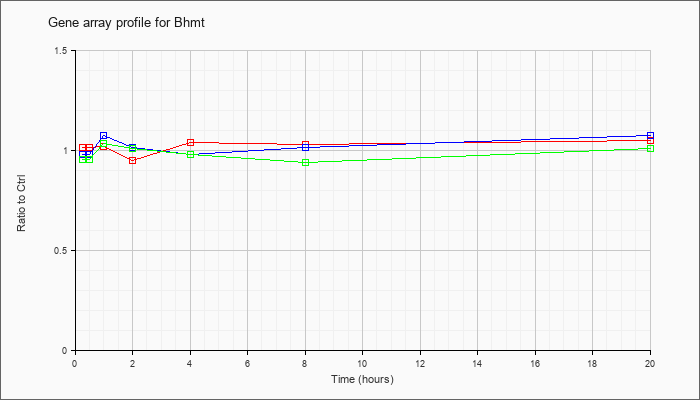 |
NM_016668 |
betaine-homocysteine methyltransferase (Bhmt), mRNA [NM_016668] |
KLA | 1.02 |
1.04 |
1.02 |
.95 |
1.04 |
1.03 |
1.05 |
| ATP | .98 |
.96 |
1.08 |
1.02 |
.98 |
1.02 |
1.08 |
| KLA/ATP | .96 |
1.02 |
1.04 |
1.01 |
.98 |
.94 |
1.01 |
|
| Bpgm | 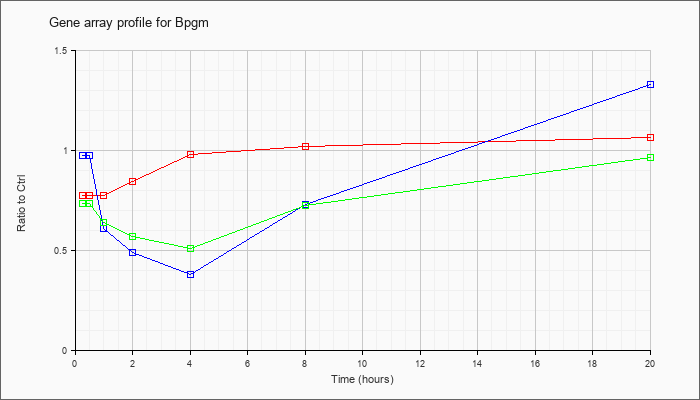 |
NM_007563 |
2,3-bisphosphoglycerate mutase (Bpgm), mRNA [NM_007563] |
KLA | .78 |
.85 |
.78 |
.85 |
.98 |
1.02 |
1.07 |
| ATP | .98 |
.81 |
.61 |
.49 |
.38 |
.73 |
1.33 |
| KLA/ATP | .74 |
.74 |
.64 |
.57 |
.51 |
.73 |
.97 |
|
| Cbs | 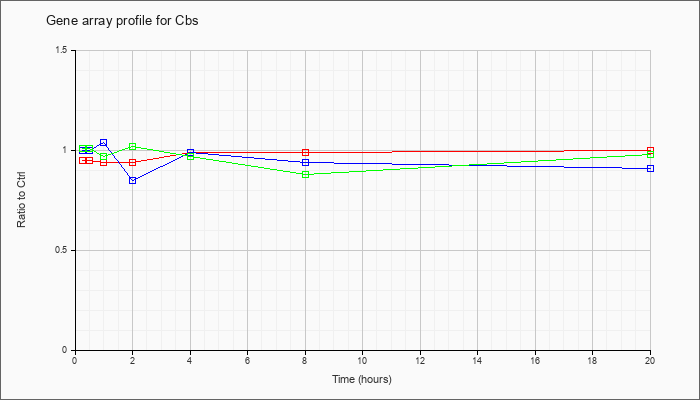 |
NM_144855 |
cystathionine beta-synthase (Cbs), transcript variant 1, mRNA [NM_144855] |
KLA | .95 |
1.02 |
.94 |
.94 |
.99 |
.99 |
1.00 |
| ATP | 1.00 |
1.14 |
1.04 |
.85 |
.99 |
.94 |
.91 |
| KLA/ATP | 1.01 |
1.08 |
.97 |
1.02 |
.97 |
.88 |
.98 |
|
| Chdh | 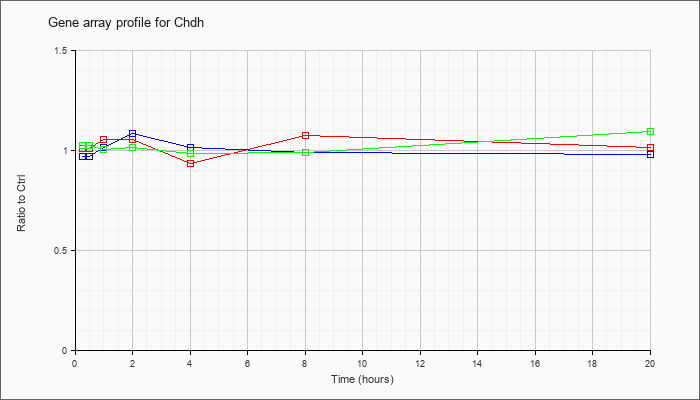 |
AK052715 |
0 day neonate kidney cDNA, RIKEN full-length enriched library, clone:D630034H06 product:CHOLINE DEHYDROGENASE (FRAGMENT) homolog [Homo sapiens], full insert sequence [AK052715] |
KLA | 1.05 |
1.02 |
1.04 |
1.10 |
1.06 |
1.17 |
.88 |
| ATP | .96 |
1.01 |
.99 |
1.00 |
1.01 |
.98 |
.97 |
| KLA/ATP | .99 |
1.01 |
1.00 |
1.04 |
.99 |
.99 |
1.06 |
|
| Chdh |  |
NM_175343 |
choline dehydrogenase (Chdh), mRNA [NM_175343] |
KLA | .97 |
.99 |
1.07 |
1.01 |
.81 |
.98 |
1.15 |
| ATP | .98 |
1.06 |
1.04 |
1.17 |
1.02 |
1.00 |
.99 |
| KLA/ATP | 1.06 |
1.01 |
1.01 |
.99 |
.98 |
.99 |
1.12 |
|
| Cth | 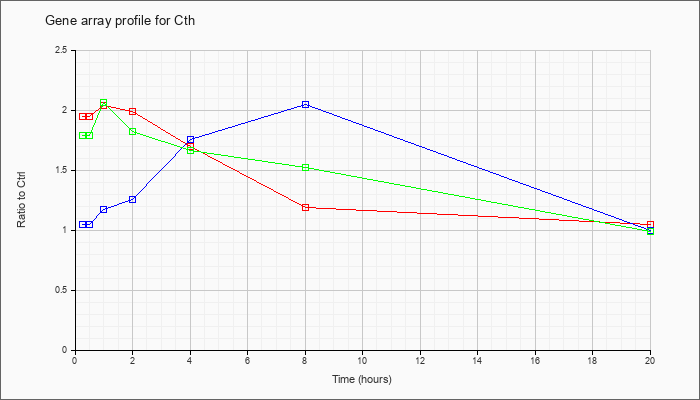 |
NM_145953 |
cystathionase (cystathionine gamma-lyase) (Cth), mRNA [NM_145953] |
KLA | 1.95 |
1.88 |
2.04 |
1.99 |
1.70 |
1.19 |
1.05 |
| ATP | 1.05 |
1.03 |
1.17 |
1.26 |
1.76 |
2.05 |
1.00 |
| KLA/ATP | 1.79 |
2.03 |
2.06 |
1.83 |
1.66 |
1.53 |
.99 |
|
| Dao1 | 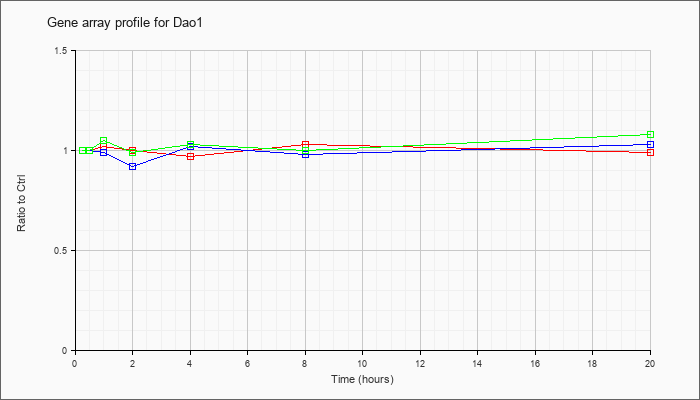 |
NM_010018 |
D-amino acid oxidase 1 (Dao1), mRNA [NM_010018] |
KLA | 1.00 |
.95 |
1.02 |
1.00 |
.97 |
1.03 |
.99 |
| ATP | 1.00 |
.93 |
.99 |
.92 |
1.02 |
.98 |
1.03 |
| KLA/ATP | 1.00 |
1.01 |
1.05 |
.99 |
1.03 |
1.00 |
1.08 |
|
| Dld | 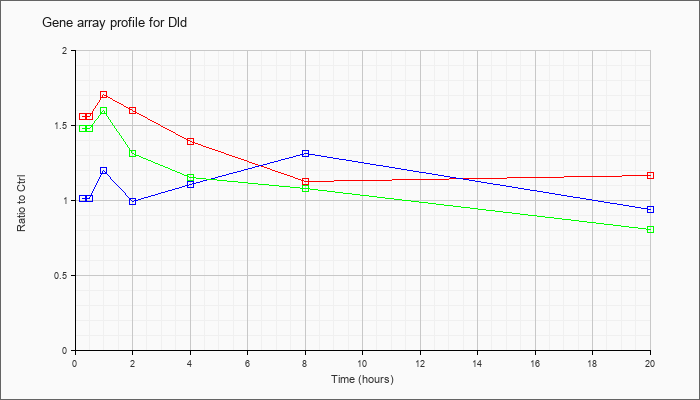 |
NM_007861 |
dihydrolipoamide dehydrogenase (Dld), mRNA [NM_007861] |
KLA | 1.60 |
1.66 |
1.84 |
1.66 |
1.47 |
1.16 |
1.17 |
| ATP | 1.05 |
1.08 |
1.21 |
1.06 |
1.15 |
1.24 |
.93 |
| KLA/ATP | 1.52 |
1.63 |
1.65 |
1.42 |
1.17 |
1.05 |
.79 |
|
| Dld |  |
U73445 |
dihydrolipoamide dehydrogenase (Dld) mRNA, complete cds. [U73445] |
KLA | 1.48 |
1.43 |
1.43 |
1.47 |
1.25 |
1.05 |
1.16 |
| ATP | .94 |
.93 |
1.18 |
.86 |
1.02 |
1.46 |
.97 |
| KLA/ATP | 1.39 |
1.37 |
1.50 |
1.10 |
1.11 |
1.14 |
.85 |
|
| Dmgdh |  |
NM_028772 |
dimethylglycine dehydrogenase precursor (Dmgdh), nuclear gene encoding mitochondrial protein, mRNA [NM_028772] |
KLA | .97 |
1.02 |
.98 |
.99 |
1.02 |
.92 |
.99 |
| ATP | 1.04 |
.94 |
.91 |
1.03 |
1.04 |
.94 |
1.01 |
| KLA/ATP | .95 |
.97 |
.99 |
.99 |
.98 |
1.06 |
1.02 |
|
| Gamt | 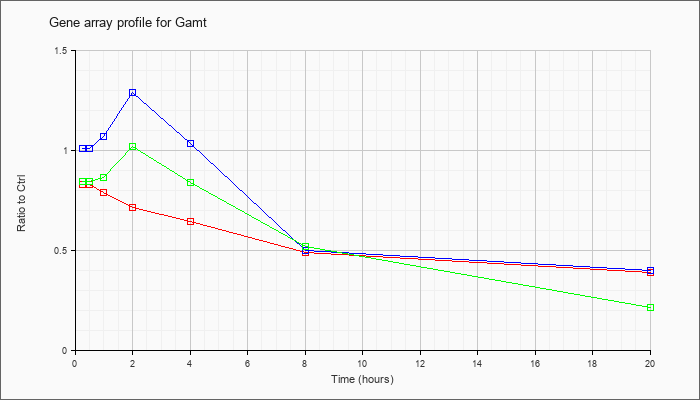 |
NM_010255 |
guanidinoacetate methyltransferase (Gamt), mRNA [NM_010255] |
KLA | .83 |
.81 |
.79 |
.71 |
.64 |
.49 |
.39 |
| ATP | 1.01 |
1.05 |
1.07 |
1.29 |
1.03 |
.50 |
.40 |
| KLA/ATP | .84 |
.88 |
.86 |
1.02 |
.84 |
.52 |
.21 |
|
| Gatm |  |
NM_025961 |
glycine amidinotransferase (L-arginine:glycine amidinotransferase) (Gatm), nuclear gene encoding mitochondrial protein, mRNA [NM_025961] |
KLA | 1.12 |
1.10 |
1.08 |
1.22 |
1.29 |
1.64 |
.65 |
| ATP | 1.02 |
1.14 |
1.14 |
1.48 |
.82 |
.21 |
.15 |
| KLA/ATP | 1.15 |
1.11 |
1.08 |
1.36 |
.67 |
.26 |
.17 |
|
| Gcat | 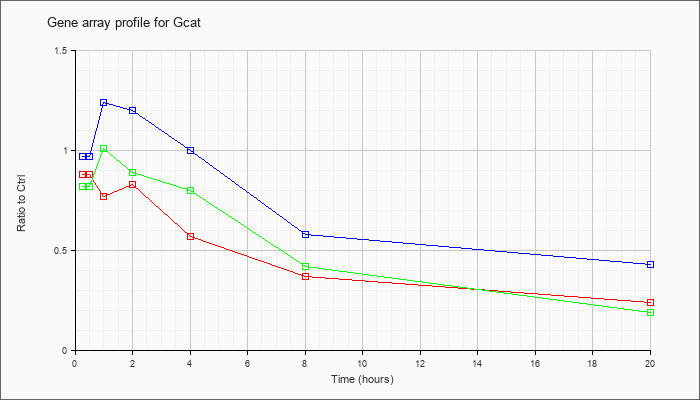 |
NM_013847 |
glycine C-acetyltransferase (2-amino-3-ketobutyrate-coenzyme A ligase) (Gcat), nuclear gene encoding mitochondrial protein, mRNA [NM_013847] |
KLA | .88 |
.88 |
.77 |
.83 |
.57 |
.37 |
.24 |
| ATP | .97 |
.91 |
1.24 |
1.20 |
1.00 |
.58 |
.43 |
| KLA/ATP | .82 |
.73 |
1.01 |
.89 |
.80 |
.42 |
.19 |
|
| Gldc | 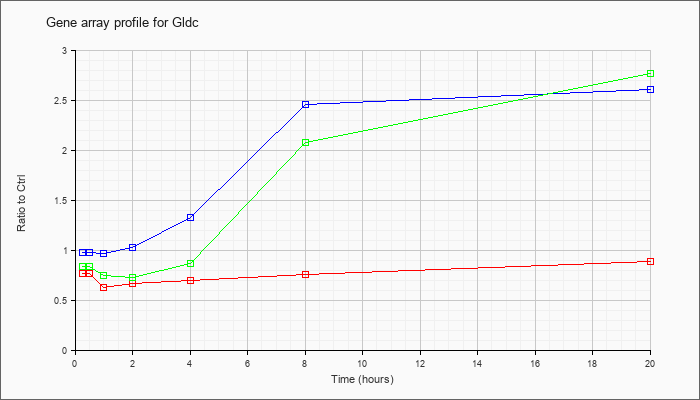 |
NM_138595 |
glycine decarboxylase (Gldc), mRNA [NM_138595] |
KLA | .77 |
.75 |
.63 |
.67 |
.70 |
.76 |
.89 |
| ATP | .98 |
1.02 |
.97 |
1.03 |
1.33 |
2.46 |
2.61 |
| KLA/ATP | .84 |
.69 |
.75 |
.73 |
.87 |
2.08 |
2.77 |
|
| Glyctk | 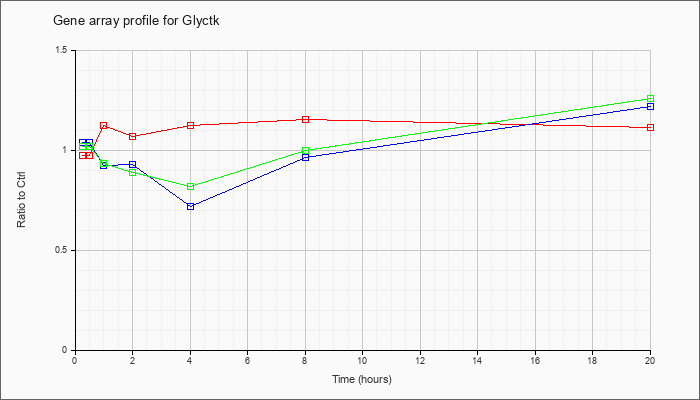 |
NM_174846 |
glycerate kinase (Glyctk), transcript variant 1, mRNA [NM_174846] |
KLA | .98 |
1.05 |
1.12 |
1.07 |
1.13 |
1.16 |
1.11 |
| ATP | 1.04 |
1.07 |
.93 |
.93 |
.72 |
.97 |
1.22 |
| KLA/ATP | 1.02 |
1.02 |
.94 |
.89 |
.82 |
1.00 |
1.26 |
|
| Gnmt | 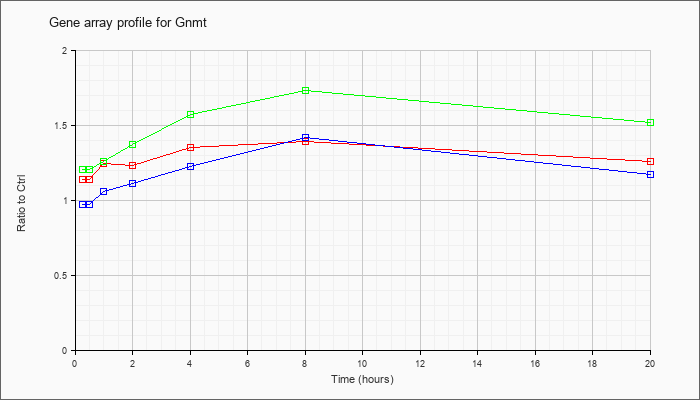 |
NM_010321 |
glycine N-methyltransferase (Gnmt), mRNA [NM_010321] |
KLA | 1.14 |
1.20 |
1.25 |
1.23 |
1.35 |
1.39 |
1.26 |
| ATP | .97 |
.99 |
1.06 |
1.11 |
1.23 |
1.42 |
1.17 |
| KLA/ATP | 1.21 |
1.20 |
1.26 |
1.37 |
1.57 |
1.73 |
1.52 |
|
| Grhpr | 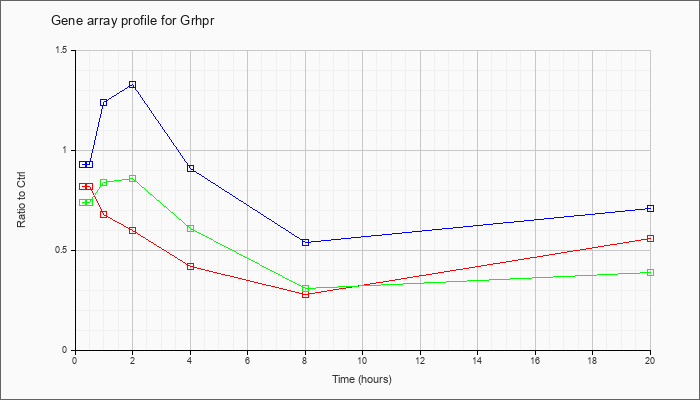 |
NM_080289 |
glyoxylate reductase/hydroxypyruvate reductase (Grhpr), mRNA [NM_080289] |
KLA | .82 |
.79 |
.68 |
.60 |
.42 |
.28 |
.56 |
| ATP | .93 |
.99 |
1.24 |
1.33 |
.91 |
.54 |
.71 |
| KLA/ATP | .74 |
.78 |
.84 |
.86 |
.61 |
.31 |
.39 |
|
| Maoa | 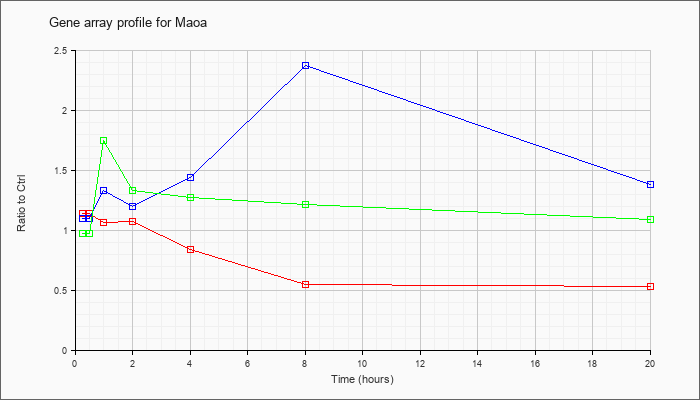 |
NM_173740 |
monoamine oxidase A (Maoa), nuclear gene encoding mitochondrial protein, mRNA [NM_173740] |
KLA | 1.14 |
1.11 |
1.06 |
1.07 |
.84 |
.55 |
.53 |
| ATP | 1.10 |
1.07 |
1.33 |
1.20 |
1.44 |
2.37 |
1.38 |
| KLA/ATP | .97 |
1.11 |
1.75 |
1.33 |
1.27 |
1.21 |
1.09 |
|
| Maob | 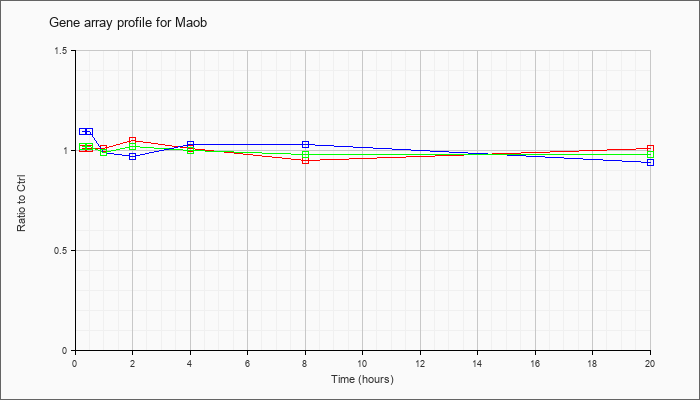 |
NM_172778 |
monoamine oxidase B (Maob), nuclear gene encoding mitochondrial protein, mRNA [NM_172778] |
KLA | 1.01 |
.97 |
1.01 |
1.05 |
1.01 |
.95 |
1.01 |
| ATP | 1.09 |
.98 |
.99 |
.97 |
1.03 |
1.03 |
.94 |
| KLA/ATP | 1.02 |
1.05 |
.99 |
1.02 |
1.00 |
.98 |
.98 |
|
Mm.30384
7 | 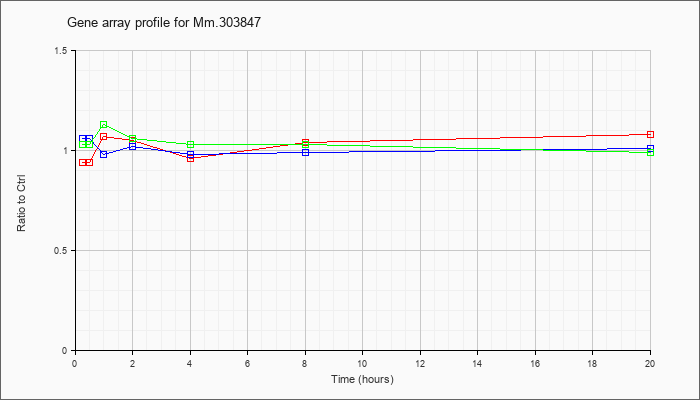 |
73661177 |
Unknown |
KLA | .94 |
1.00 |
1.07 |
1.05 |
.96 |
1.04 |
1.08 |
| ATP | 1.06 |
1.01 |
.98 |
1.02 |
.98 |
.99 |
1.01 |
| KLA/ATP | 1.03 |
.94 |
1.13 |
1.06 |
1.03 |
1.03 |
.99 |
|
Mm.42309
9 | 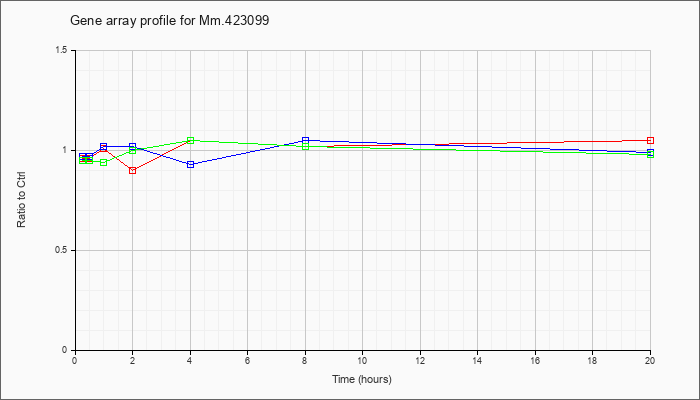 |
74325340 |
Unknown |
KLA | .96 |
1.03 |
1.01 |
.90 |
1.05 |
1.02 |
1.05 |
| ATP | .97 |
.94 |
1.02 |
1.02 |
.93 |
1.05 |
.99 |
| KLA/ATP | .95 |
.97 |
.94 |
1.00 |
1.05 |
1.02 |
.98 |
|
| Mm.7329 | 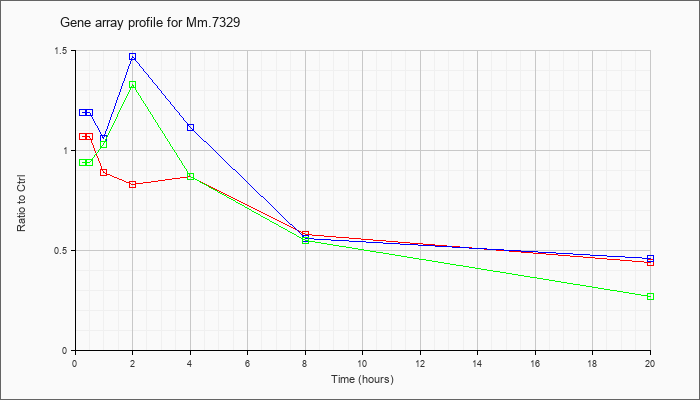 |
158508431 |
Unknown |
KLA | 1.07 |
1.00 |
.89 |
.83 |
.87 |
.58 |
.44 |
| ATP | 1.19 |
1.31 |
1.06 |
1.47 |
1.11 |
.56 |
.46 |
| KLA/ATP | .94 |
1.10 |
1.03 |
1.33 |
.87 |
.55 |
.27 |
|
| Pgam1 | 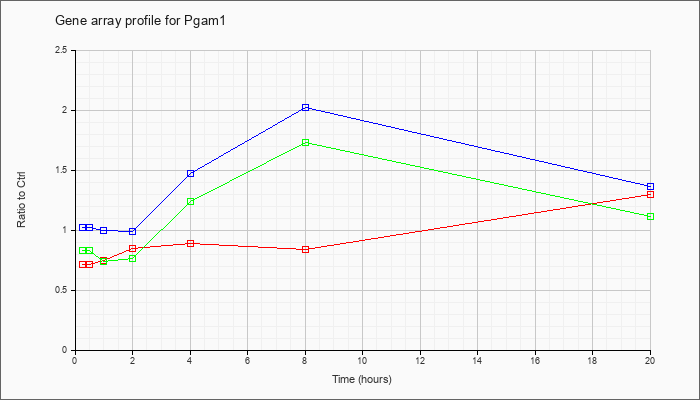 |
NM_023418 |
phosphoglycerate mutase 1 (Pgam1), mRNA [NM_023418] |
KLA | .71 |
.69 |
.75 |
.85 |
.89 |
.84 |
1.30 |
| ATP | 1.02 |
1.10 |
1.00 |
.99 |
1.47 |
2.02 |
1.36 |
| KLA/ATP | .83 |
.83 |
.74 |
.76 |
1.24 |
1.73 |
1.11 |
|
| Pgam2 | 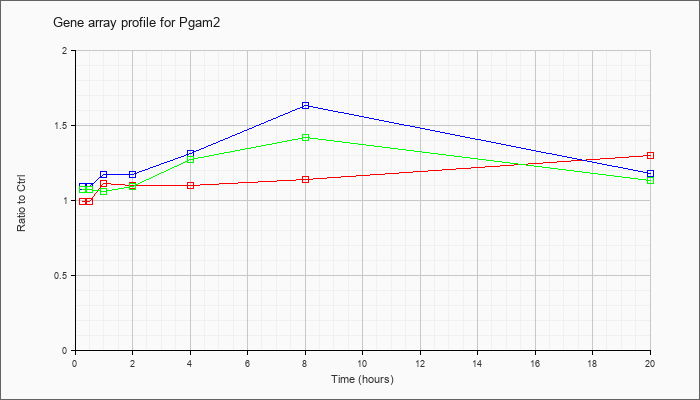 |
NM_018870 |
phosphoglycerate mutase 2 (Pgam2), mRNA [NM_018870] |
KLA | .99 |
1.05 |
1.11 |
1.10 |
1.10 |
1.14 |
1.30 |
| ATP | 1.09 |
1.08 |
1.17 |
1.17 |
1.31 |
1.63 |
1.18 |
| KLA/ATP | 1.07 |
1.08 |
1.06 |
1.09 |
1.27 |
1.42 |
1.13 |
|
| Phgdh | 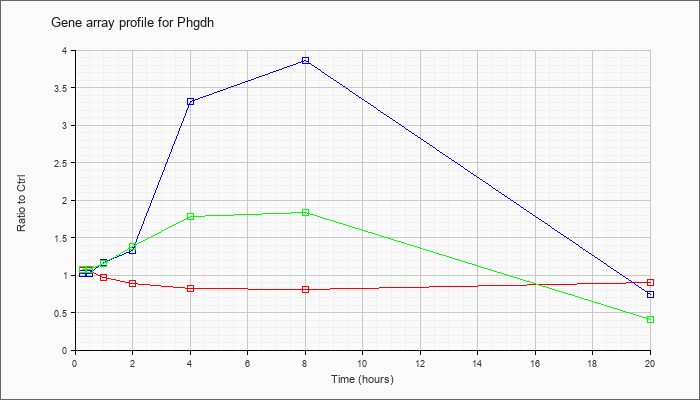 |
NM_016966 |
3-phosphoglycerate dehydrogenase (Phgdh), mRNA [NM_016966] |
KLA | 1.06 |
1.11 |
.97 |
.89 |
.82 |
.80 |
.91 |
| ATP | 1.02 |
1.03 |
1.17 |
1.33 |
3.31 |
3.86 |
.74 |
| KLA/ATP | 1.07 |
1.06 |
1.15 |
1.38 |
1.78 |
1.84 |
.40 |
|
| Pipox | 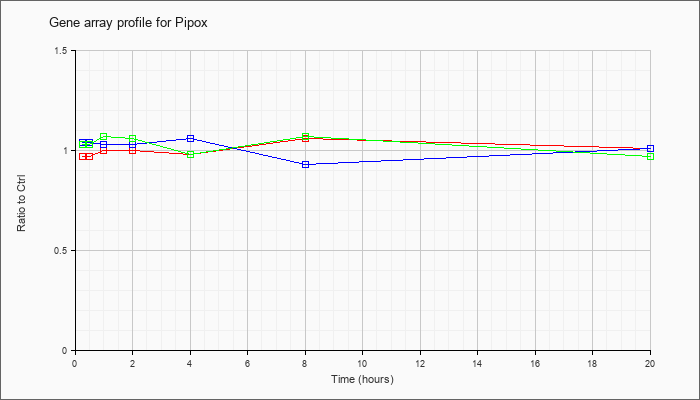 |
NM_008952 |
pipecolic acid oxidase (Pipox), mRNA [NM_008952] |
KLA | .97 |
.94 |
1.00 |
1.00 |
.98 |
1.06 |
1.01 |
| ATP | 1.04 |
1.01 |
1.03 |
1.03 |
1.06 |
.93 |
1.01 |
| KLA/ATP | 1.03 |
1.12 |
1.07 |
1.06 |
.98 |
1.07 |
.97 |
|
| Psat1 |  |
NM_177420 |
phosphoserine aminotransferase 1 (Psat1), mRNA [NM_177420] |
KLA | 1.02 |
1.21 |
1.24 |
1.27 |
1.64 |
1.59 |
.38 |
| ATP | 1.02 |
1.01 |
1.04 |
1.17 |
1.27 |
1.09 |
.43 |
| KLA/ATP | 1.16 |
1.11 |
1.04 |
1.37 |
1.04 |
.70 |
.24 |
|
| Psph |  |
NM_133900 |
phosphoserine phosphatase (Psph), mRNA [NM_133900] |
KLA | .94 |
.88 |
.76 |
.88 |
.65 |
.87 |
.75 |
| ATP | .77 |
.61 |
.53 |
.35 |
1.23 |
1.85 |
.73 |
| KLA/ATP | .71 |
.64 |
.54 |
.31 |
.46 |
.78 |
.61 |
|
| Sardh | 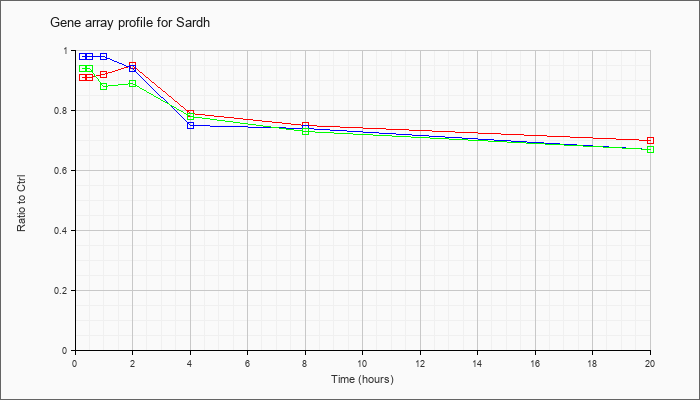 |
NM_138665 |
sarcosine dehydrogenase (Sardh), mRNA [NM_138665] |
KLA | .91 |
.92 |
.92 |
.95 |
.79 |
.75 |
.70 |
| ATP | .98 |
.90 |
.98 |
.94 |
.75 |
.74 |
.67 |
| KLA/ATP | .94 |
.92 |
.88 |
.89 |
.78 |
.73 |
.67 |
|
| Sds | 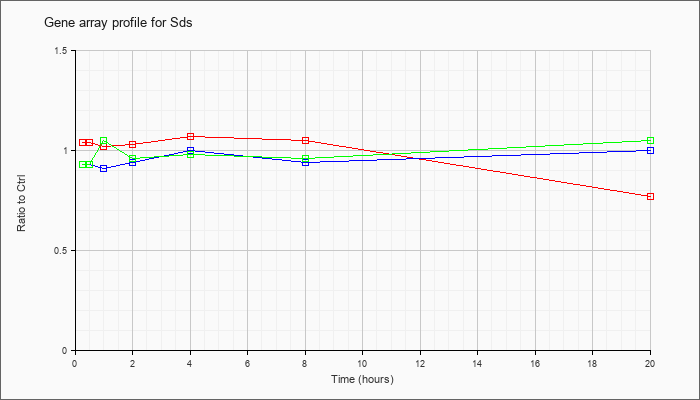 |
NM_145565 |
serine dehydratase (Sds), mRNA [NM_145565] |
KLA | 1.04 |
1.04 |
1.02 |
1.03 |
1.07 |
1.05 |
.77 |
| ATP | .93 |
.97 |
.91 |
.94 |
1.00 |
.94 |
1.00 |
| KLA/ATP | .93 |
.95 |
1.05 |
.96 |
.98 |
.96 |
1.05 |
|
| Shmt1 | 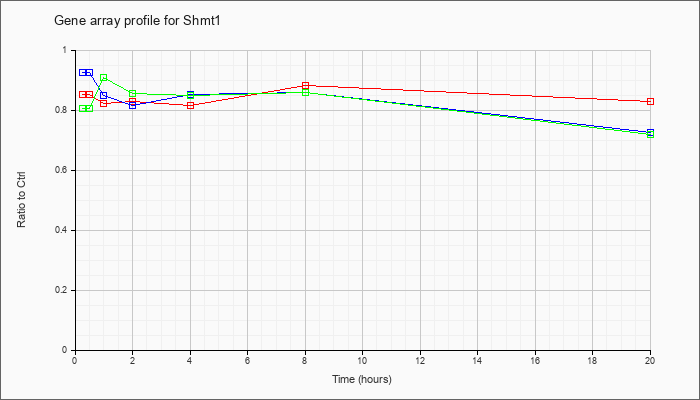 |
NM_009171 |
serine hydroxymethyltransferase 1 (soluble) (Shmt1), mRNA [NM_009171] |
KLA | .85 |
.89 |
.82 |
.83 |
.82 |
.88 |
.83 |
| ATP | .93 |
.81 |
.85 |
.82 |
.85 |
.86 |
.72 |
| KLA/ATP | .81 |
.83 |
.91 |
.86 |
.85 |
.86 |
.72 |
|
| Shmt2 | 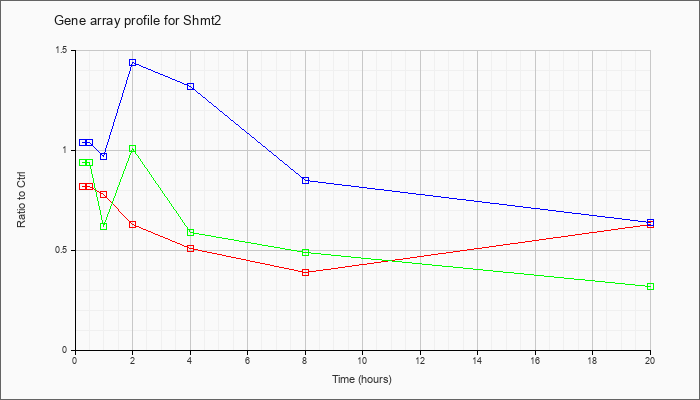 |
NM_028230 |
serine hydroxymethyltransferase 2 (mitochondrial) (Shmt2), nuclear gene encoding mitochondrial protein, mRNA [NM_028230] |
KLA | .82 |
.90 |
.78 |
.63 |
.51 |
.39 |
.63 |
| ATP | 1.04 |
1.21 |
.97 |
1.44 |
1.32 |
.85 |
.64 |
| KLA/ATP | .94 |
.92 |
.62 |
1.01 |
.59 |
.49 |
.32 |
|
| Srr | 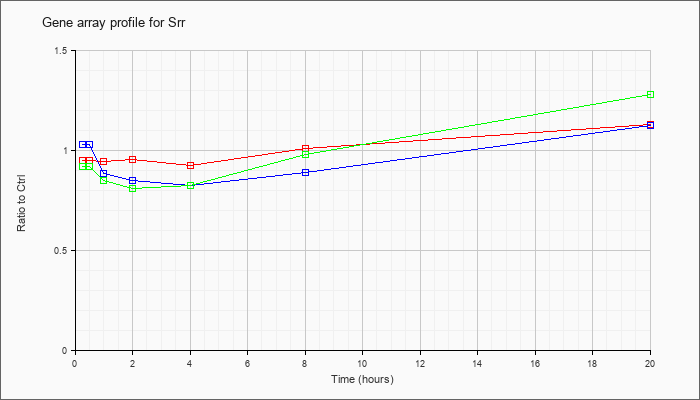 |
AK002636 |
adult male kidney cDNA, RIKEN full-length enriched library, clone:0610015N12 product:serine racemase, full insert sequence. [AK002636] |
KLA | .86 |
.84 |
.89 |
.89 |
.84 |
.91 |
.85 |
| ATP | .98 |
.88 |
.88 |
.73 |
.78 |
.84 |
.76 |
| KLA/ATP | .83 |
.80 |
.81 |
.70 |
.90 |
.78 |
.84 |
|
| Srr |  |
NM_013761 |
serine racemase (Srr), mRNA [NM_013761] |
KLA | .98 |
.93 |
.96 |
.97 |
.95 |
1.04 |
1.22 |
| ATP | 1.04 |
.93 |
.88 |
.89 |
.84 |
.90 |
1.24 |
| KLA/ATP | .95 |
.92 |
.86 |
.85 |
.80 |
1.04 |
1.42 |
|
| Tdh | 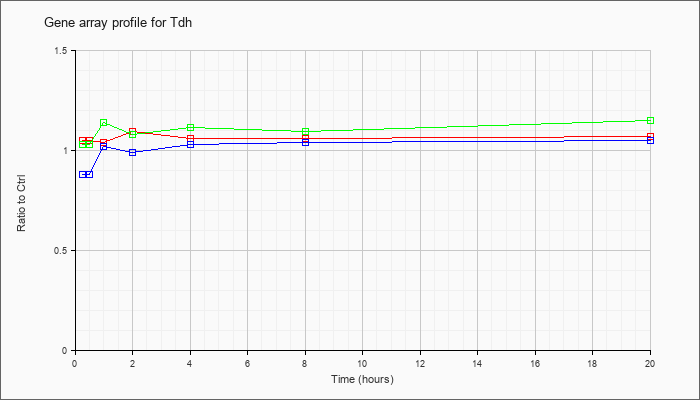 |
NM_021480 |
L-threonine dehydrogenase (Tdh), mRNA [NM_021480] |
KLA | 1.05 |
1.01 |
1.04 |
1.09 |
1.06 |
1.06 |
1.07 |
| ATP | .88 |
.99 |
1.02 |
.99 |
1.03 |
1.04 |
1.05 |
| KLA/ATP | 1.03 |
.91 |
1.14 |
1.08 |
1.11 |
1.09 |
1.15 |
|
| Tha1 | 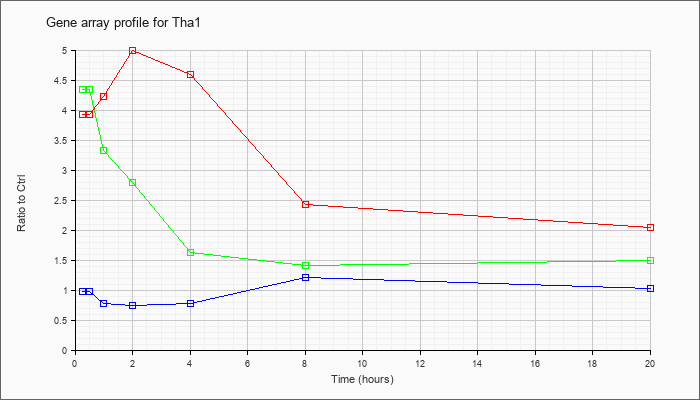 |
NM_027919 |
threonine aldolase 1 (Tha1), mRNA [NM_027919] |
KLA | 3.92 |
4.62 |
4.23 |
4.99 |
4.59 |
2.42 |
2.04 |
| ATP | .98 |
.92 |
.78 |
.74 |
.78 |
1.21 |
1.03 |
| KLA/ATP | 4.35 |
4.64 |
3.33 |
2.80 |
1.63 |
1.41 |
1.50 |
|