| Gene symbol |
Time course plot |
Accession |
Description |
Treatment |
15min |
30min |
1hr |
2hr |
4hr |
8hr |
20hr |
1700055N
04Rik | 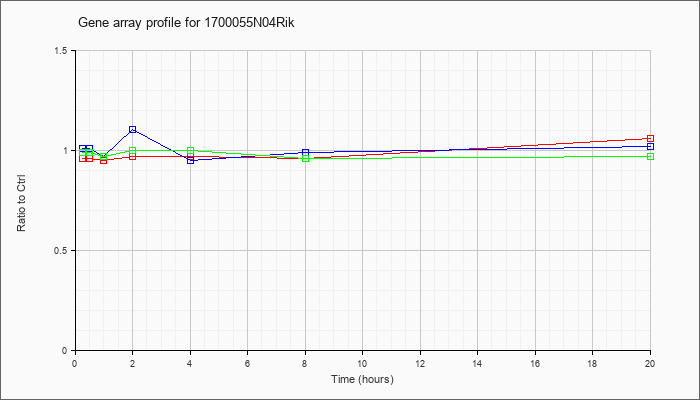 |
AK081788 |
16 days embryo head cDNA, RIKEN full-length enriched library, clone:C130077J09 product:weakly similar to ALDEHYDE DEHYDROGENASE 7 (EC 1.2.1.5) [Homo sapiens], full insert sequence [AK081788] |
KLA | .96 |
.98 |
.95 |
.97 |
.97 |
.96 |
1.06 |
| ATP | 1.01 |
1.00 |
.97 |
1.10 |
.95 |
.99 |
1.02 |
| KLA/ATP | .99 |
1.00 |
.97 |
1.00 |
1.00 |
.96 |
.97 |
|
AI790298
| 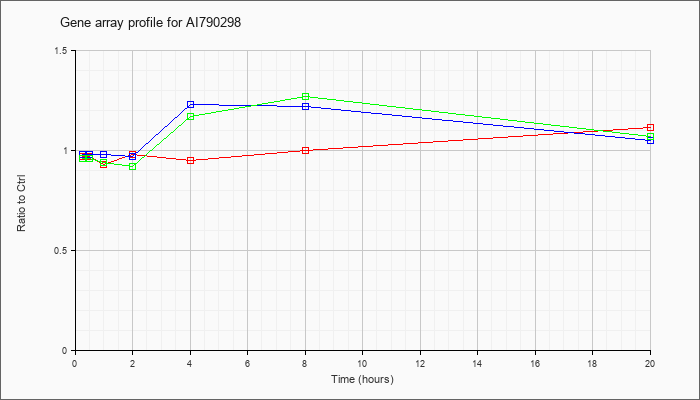 |
AK129347 |
mRNA for mKIAA1394 protein [AK129347] |
KLA | .97 |
.95 |
.93 |
.98 |
.95 |
1.00 |
1.11 |
| ATP | .98 |
1.03 |
.98 |
.97 |
1.23 |
1.22 |
1.05 |
| KLA/ATP | .96 |
.99 |
.94 |
.92 |
1.17 |
1.27 |
1.07 |
|
| Abat | 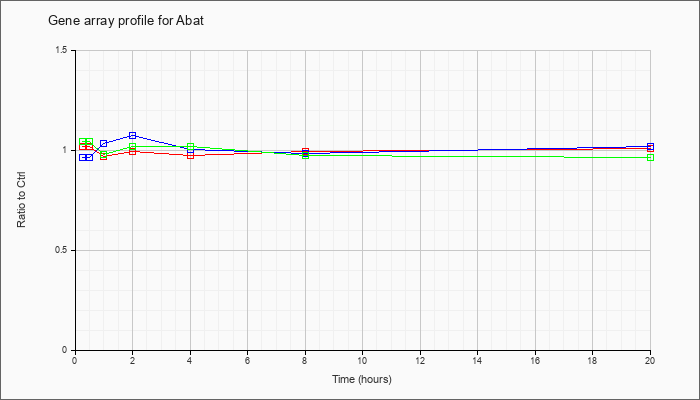 |
AK083062 |
12 days embryo spinal cord cDNA, RIKEN full-length enriched library, clone:C530041H11 product:hypothetical protein, full insert sequence. [AK083062] |
KLA | 1.01 |
1.09 |
.97 |
.99 |
1.03 |
1.02 |
1.06 |
| ATP | .96 |
.99 |
1.00 |
1.11 |
1.04 |
.98 |
1.06 |
| KLA/ATP | 1.06 |
1.03 |
.96 |
1.06 |
1.10 |
1.06 |
.96 |
|
| Abat |  |
NM_172961 |
4-aminobutyrate aminotransferase (Abat), nuclear gene encoding mitochondrial protein, mRNA [NM_172961] |
KLA | 1.03 |
.96 |
.97 |
1.00 |
.92 |
.97 |
.96 |
| ATP | .97 |
.93 |
1.07 |
1.04 |
.97 |
.99 |
.98 |
| KLA/ATP | 1.03 |
1.01 |
1.00 |
.98 |
.94 |
.89 |
.97 |
|
| Acadm | 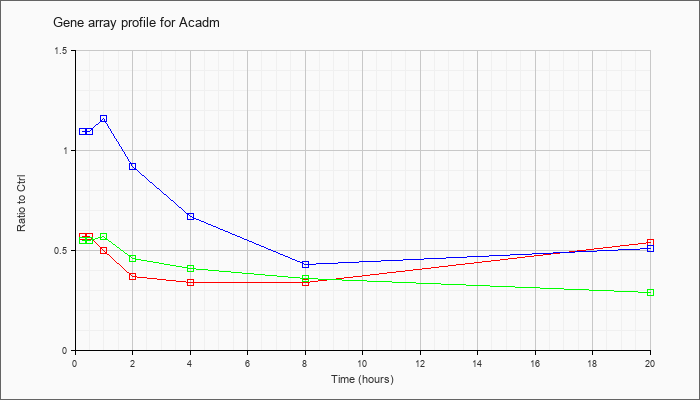 |
NM_007382 |
acyl-Coenzyme A dehydrogenase, medium chain (Acadm), nuclear gene encoding mitochondrial protein, mRNA [NM_007382] |
KLA | .57 |
.50 |
.50 |
.37 |
.34 |
.34 |
.54 |
| ATP | 1.09 |
1.08 |
1.16 |
.92 |
.67 |
.43 |
.51 |
| KLA/ATP | .55 |
.56 |
.57 |
.46 |
.41 |
.36 |
.29 |
|
| Aldh1a3 | 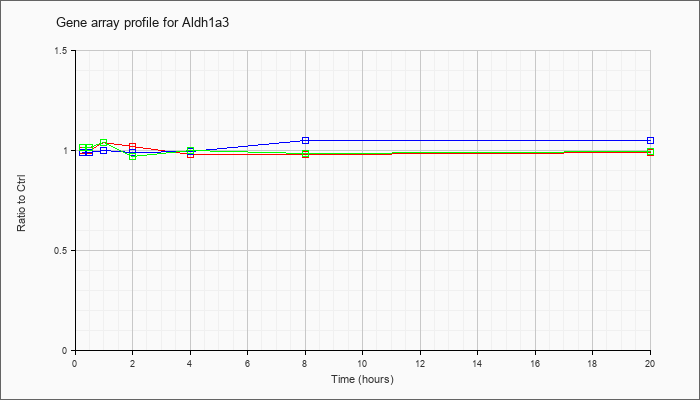 |
AK086764 |
15 days embryo head cDNA, RIKEN full-length enriched library, clone:D930050B08 product:aldehyde dehydrogenase family 1, subfamily A3, full insert sequence. [AK086764] |
KLA | .97 |
1.05 |
1.05 |
1.01 |
1.00 |
.98 |
.97 |
| ATP | 1.00 |
1.03 |
.99 |
.92 |
1.01 |
1.09 |
1.02 |
| KLA/ATP | 1.01 |
.99 |
1.06 |
1.00 |
.97 |
.98 |
1.00 |
|
| Aldh1a3 |  |
NM_053080 |
aldehyde dehydrogenase family 1, subfamily A3 (Aldh1a3), mRNA [NM_053080] |
KLA | 1.03 |
.96 |
1.03 |
1.03 |
.96 |
.98 |
1.01 |
| ATP | .98 |
1.04 |
1.01 |
1.06 |
.98 |
1.01 |
1.08 |
| KLA/ATP | 1.02 |
1.00 |
1.02 |
.94 |
1.03 |
.99 |
.99 |
|
| Aldh1b1 | 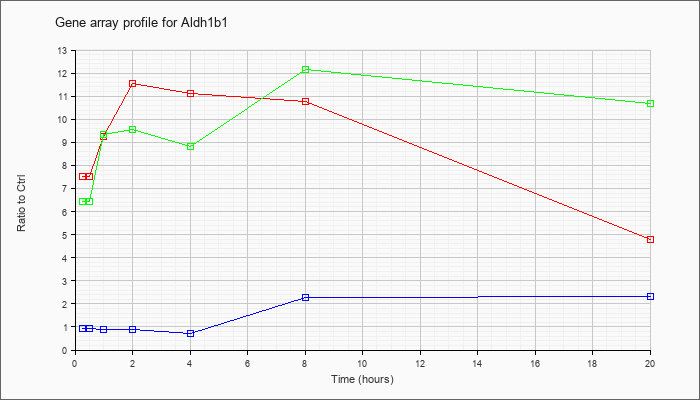 |
NM_028270 |
aldehyde dehydrogenase 1 family, member B1 (Aldh1b1), nuclear gene encoding mitochondrial protein, mRNA [NM_028270] |
KLA | 7.51 |
7.83 |
9.27 |
11.57 |
11.10 |
10.78 |
4.79 |
| ATP | .93 |
.91 |
.90 |
.89 |
.73 |
2.27 |
2.30 |
| KLA/ATP | 6.45 |
7.40 |
9.33 |
9.54 |
8.82 |
12.15 |
10.67 |
|
| Aldh2 | 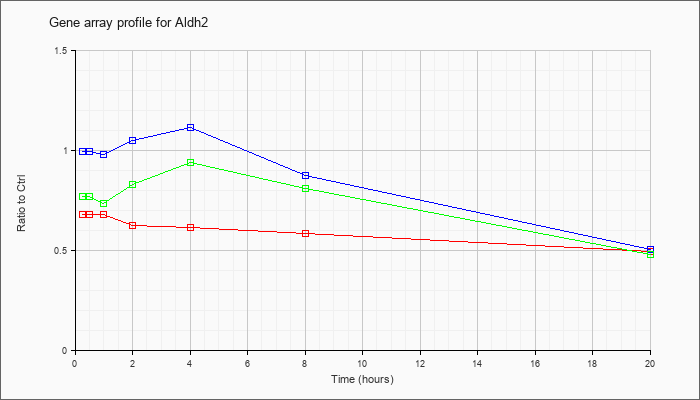 |
AK163452 |
adult male corpora quadrigemina cDNA, RIKEN full-length enriched library, clone:B230105J09 product:aldehyde dehydrogenase 2, mitochondrial, full insert sequence [AK163452] |
KLA | .62 |
.65 |
.59 |
.60 |
.63 |
.64 |
.49 |
| ATP | 1.01 |
.91 |
1.06 |
.93 |
1.18 |
.93 |
.41 |
| KLA/ATP | .70 |
.68 |
.70 |
.84 |
1.16 |
.96 |
.45 |
|
| Aldh2 |  |
NM_009656 |
aldehyde dehydrogenase 2, mitochondrial (Aldh2), nuclear gene encoding mitochondrial protein, mRNA [NM_009656] |
KLA | .74 |
.81 |
.77 |
.65 |
.60 |
.53 |
.50 |
| ATP | .98 |
1.10 |
.90 |
1.17 |
1.04 |
.82 |
.60 |
| KLA/ATP | .84 |
.86 |
.77 |
.82 |
.72 |
.66 |
.51 |
|
| Aldh3a1 | 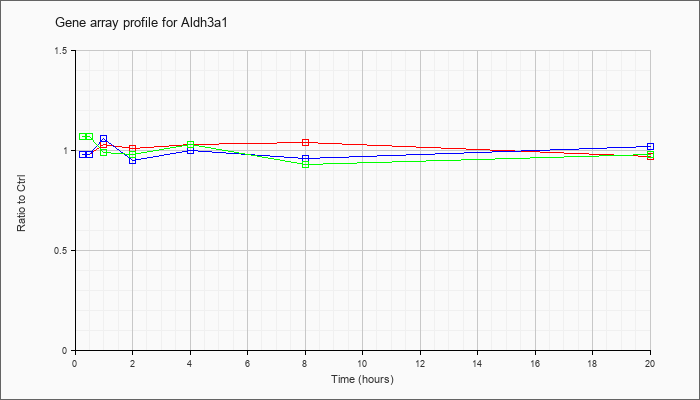 |
NM_007436 |
aldehyde dehydrogenase family 3, subfamily A1 (Aldh3a1), transcript variant 1, mRNA [NM_007436] |
KLA | .98 |
.98 |
1.03 |
1.01 |
1.03 |
1.04 |
.97 |
| ATP | .98 |
1.01 |
1.06 |
.95 |
1.00 |
.96 |
1.02 |
| KLA/ATP | 1.07 |
1.08 |
.99 |
.98 |
1.03 |
.93 |
.98 |
|
| Aldh3a2 | 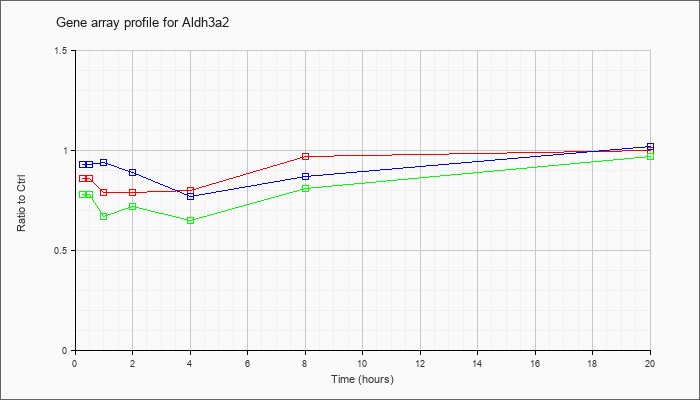 |
NM_007437 |
aldehyde dehydrogenase family 3, subfamily A2 (Aldh3a2), mRNA [NM_007437] |
KLA | .86 |
.83 |
.79 |
.79 |
.80 |
.97 |
1.00 |
| ATP | .93 |
.98 |
.94 |
.89 |
.77 |
.87 |
1.02 |
| KLA/ATP | .78 |
.81 |
.67 |
.72 |
.65 |
.81 |
.97 |
|
| Aldh3b1 | 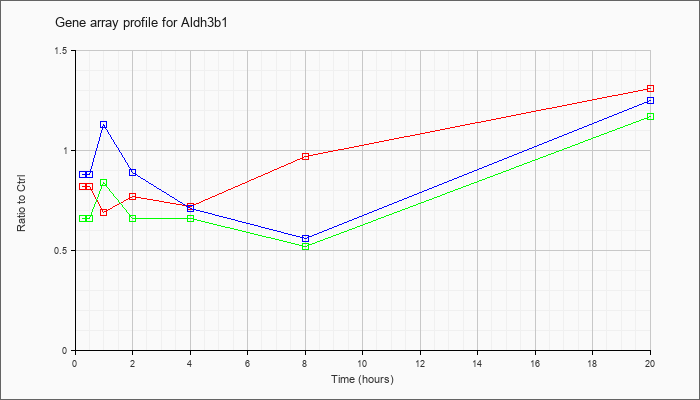 |
NM_026316 |
aldehyde dehydrogenase 3 family, member B1 (Aldh3b1), mRNA [NM_026316] |
KLA | .82 |
.77 |
.69 |
.77 |
.72 |
.97 |
1.31 |
| ATP | .88 |
.85 |
1.13 |
.89 |
.71 |
.56 |
1.25 |
| KLA/ATP | .66 |
.72 |
.84 |
.66 |
.66 |
.52 |
1.17 |
|
| Aldh3b2 | 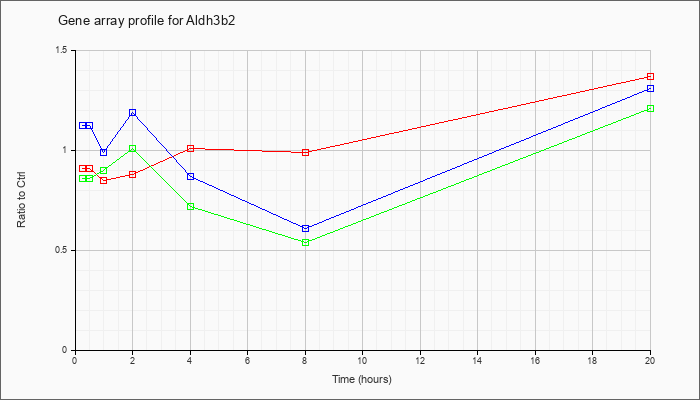 |
XM_129134 |
gb|PREDICTED: Mus musculus aldehyde dehydrogenase 3 family, member B2 (Aldh3b2), mRNA [XM_129134] |
KLA | .91 |
.93 |
.85 |
.88 |
1.01 |
.99 |
1.37 |
| ATP | 1.12 |
1.11 |
.99 |
1.19 |
.87 |
.61 |
1.31 |
| KLA/ATP | .86 |
1.02 |
.90 |
1.01 |
.72 |
.54 |
1.21 |
|
| Aldh6a1 | 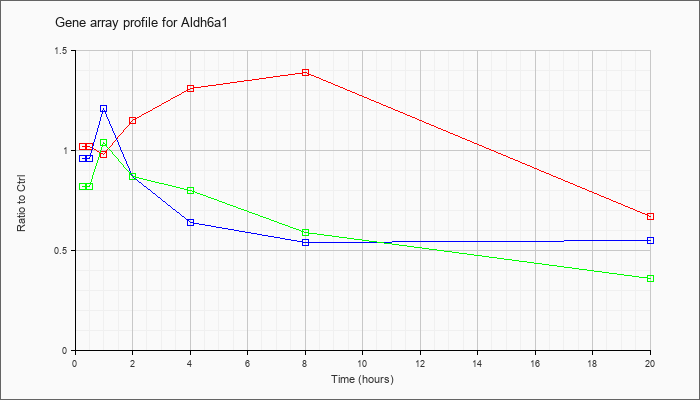 |
NM_134042 |
aldehyde dehydrogenase family 6, subfamily A1 (Aldh6a1), nuclear gene encoding mitochondrial protein, mRNA [NM_134042] |
KLA | 1.02 |
.97 |
.98 |
1.15 |
1.31 |
1.39 |
.67 |
| ATP | .96 |
.92 |
1.21 |
.87 |
.64 |
.54 |
.55 |
| KLA/ATP | .82 |
.97 |
1.04 |
.87 |
.80 |
.59 |
.36 |
|
| Aldh7a1 | 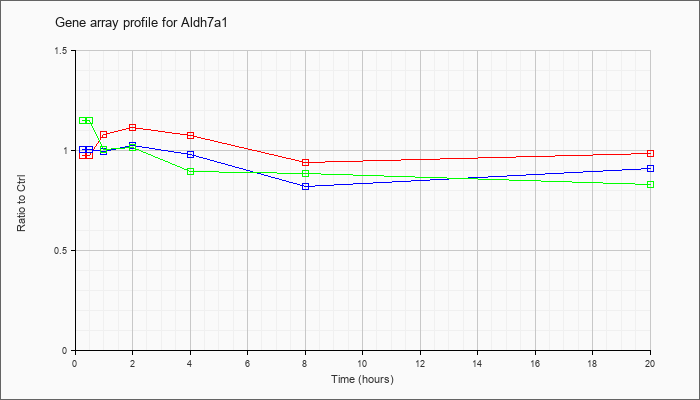 |
NM_138600 |
aldehyde dehydrogenase family 7, member A1 (Aldh7a1), mRNA [NM_138600] |
KLA | .98 |
1.03 |
1.08 |
1.12 |
1.08 |
.94 |
.99 |
| ATP | 1.01 |
1.03 |
1.00 |
1.03 |
.98 |
.82 |
.91 |
| KLA/ATP | 1.15 |
1.12 |
1.01 |
1.02 |
.90 |
.89 |
.83 |
|
| Aldh9a1 | 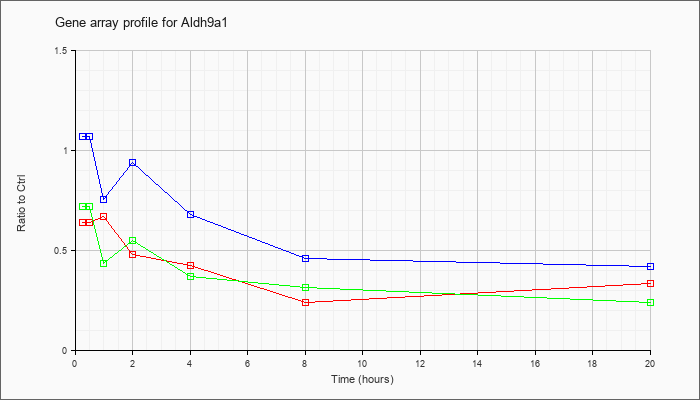 |
NM_019993 |
aldehyde dehydrogenase 9, subfamily A1 (Aldh9a1), mRNA [NM_019993] |
KLA | .64 |
.64 |
.67 |
.48 |
.42 |
.24 |
.33 |
| ATP | 1.07 |
1.02 |
.75 |
.94 |
.68 |
.46 |
.42 |
| KLA/ATP | .72 |
.65 |
.43 |
.55 |
.37 |
.31 |
.24 |
|
| Aoc2 | 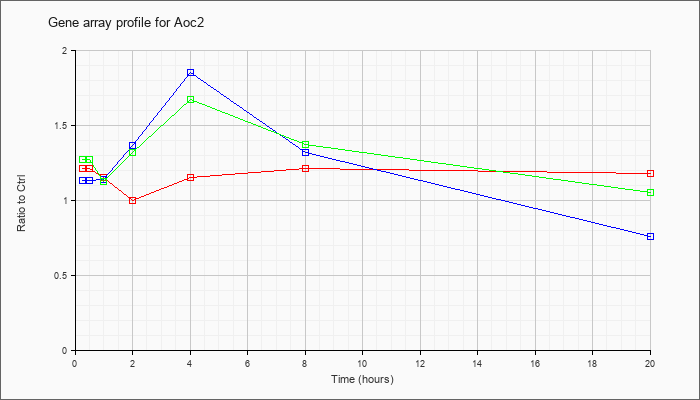 |
NM_178932 |
amine oxidase, copper containing 2 (retina-specific) (Aoc2), mRNA [NM_178932] |
KLA | 1.21 |
1.19 |
1.15 |
1.00 |
1.15 |
1.21 |
1.18 |
| ATP | 1.13 |
1.13 |
1.14 |
1.36 |
1.85 |
1.32 |
.76 |
| KLA/ATP | 1.27 |
1.34 |
1.12 |
1.32 |
1.67 |
1.37 |
1.05 |
|
| Aoc3 | 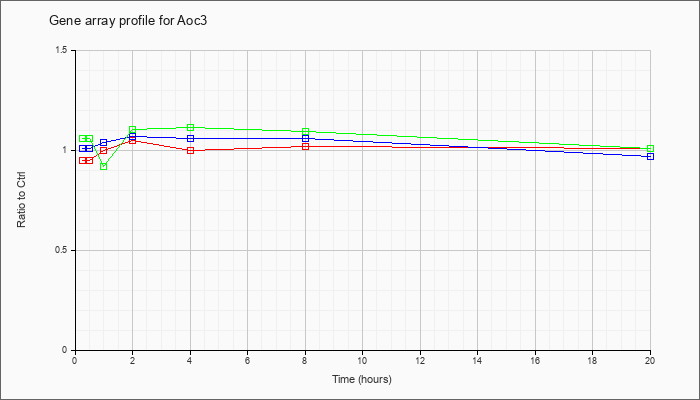 |
NM_009675 |
amine oxidase, copper containing 3 (Aoc3), mRNA [NM_009675] |
KLA | .95 |
1.06 |
1.00 |
1.05 |
1.00 |
1.02 |
1.01 |
| ATP | 1.01 |
.90 |
1.04 |
1.07 |
1.06 |
1.06 |
.97 |
| KLA/ATP | 1.06 |
.98 |
.92 |
1.10 |
1.11 |
1.09 |
1.01 |
|
| Cndp1 | 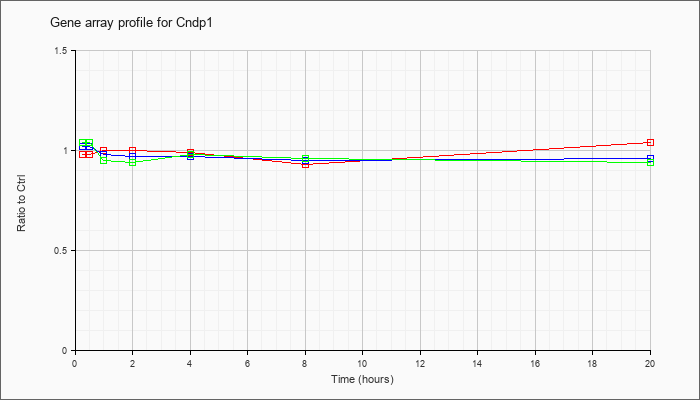 |
NM_177450 |
carnosine dipeptidase 1 (metallopeptidase M20 family) (Cndp1), mRNA [NM_177450] |
KLA | .98 |
1.00 |
1.00 |
1.00 |
.99 |
.93 |
1.04 |
| ATP | 1.02 |
.96 |
.98 |
.97 |
.97 |
.95 |
.96 |
| KLA/ATP | 1.04 |
1.04 |
.95 |
.94 |
.98 |
.96 |
.94 |
|
| Cndp2 |  |
NM_023149 |
CNDP dipeptidase 2 (metallopeptidase M20 family) (Cndp2), mRNA [NM_023149] |
KLA | .88 |
.96 |
1.08 |
.98 |
1.35 |
1.73 |
1.59 |
| ATP | 1.09 |
1.27 |
.77 |
1.02 |
1.04 |
1.30 |
1.67 |
| KLA/ATP | 1.17 |
1.04 |
.63 |
1.09 |
.95 |
1.78 |
2.70 |
|
| Dpyd | 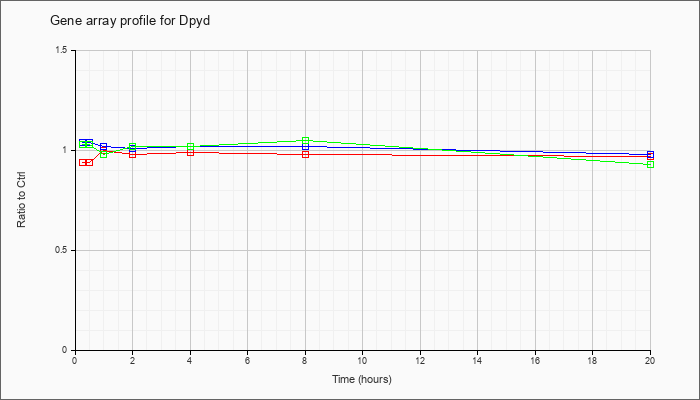 |
NM_170778 |
dihydropyrimidine dehydrogenase (Dpyd), mRNA [NM_170778] |
KLA | .94 |
.95 |
1.00 |
.98 |
.99 |
.98 |
.97 |
| ATP | 1.04 |
.95 |
1.02 |
1.01 |
1.02 |
1.02 |
.98 |
| KLA/ATP | 1.03 |
.98 |
.98 |
1.02 |
1.02 |
1.05 |
.93 |
|
| Echs1 | 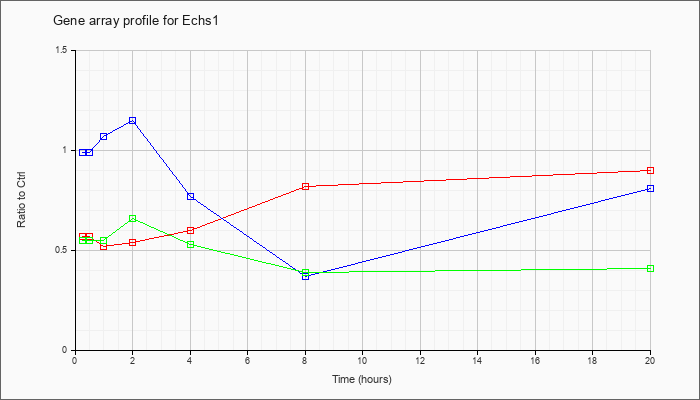 |
NM_053119 |
enoyl Coenzyme A hydratase, short chain, 1, mitochondrial (Echs1), nuclear gene encoding mitochondrial protein, mRNA [NM_053119] |
KLA | .57 |
.55 |
.52 |
.54 |
.60 |
.82 |
.90 |
| ATP | .99 |
.97 |
1.07 |
1.15 |
.77 |
.37 |
.81 |
| KLA/ATP | .55 |
.56 |
.55 |
.66 |
.53 |
.39 |
.41 |
|
| Ehhadh | 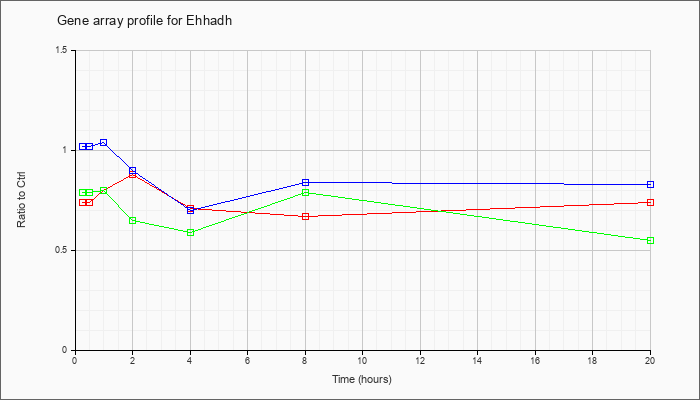 |
NM_023737 |
enoyl-Coenzyme A, hydratase/3-hydroxyacyl Coenzyme A dehydrogenase (Ehhadh), mRNA [NM_023737] |
KLA | .74 |
.77 |
.80 |
.88 |
.71 |
.67 |
.74 |
| ATP | 1.02 |
1.06 |
1.04 |
.90 |
.70 |
.84 |
.83 |
| KLA/ATP | .79 |
.81 |
.80 |
.65 |
.59 |
.79 |
.55 |
|
| Gad1 | 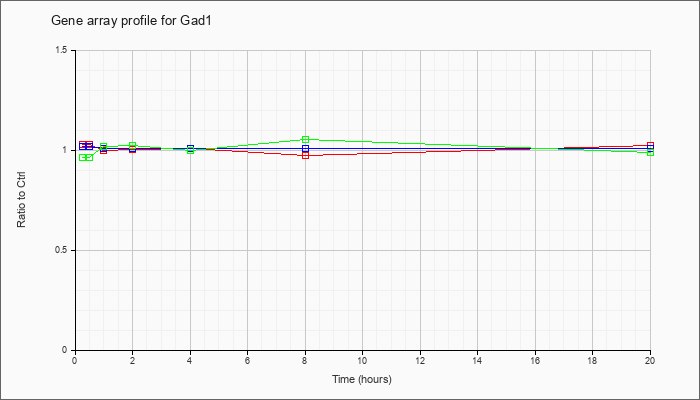 |
AK054554 |
10 days neonate olfactory brain cDNA, RIKEN full-length enriched library, clone:E530004P22 product:glutamic acid decarboxylase 1, full insert sequence. [AK054554] |
KLA | 1.03 |
1.00 |
1.00 |
1.00 |
1.00 |
.98 |
1.10 |
| ATP | 1.03 |
.91 |
1.01 |
1.05 |
.99 |
.97 |
1.08 |
| KLA/ATP | .97 |
.99 |
.99 |
1.10 |
1.02 |
1.09 |
.96 |
|
| Gad1 |  |
NM_008077 |
glutamic acid decarboxylase 1 (Gad1), mRNA [NM_008077] |
KLA | 1.03 |
.99 |
1.00 |
1.01 |
1.01 |
.97 |
.99 |
| ATP | 1.02 |
1.02 |
1.01 |
.99 |
1.02 |
1.03 |
.98 |
| KLA/ATP | .96 |
1.03 |
1.04 |
.99 |
.99 |
1.04 |
1.01 |
|
| Gad2 | 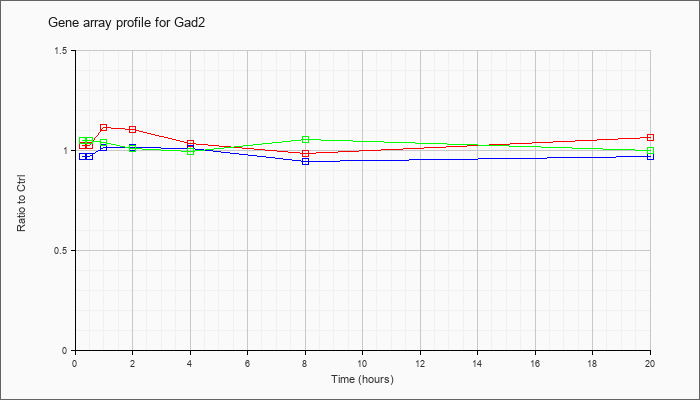 |
NM_008078 |
glutamic acid decarboxylase 2 (Gad2), mRNA [NM_008078] |
KLA | 1.03 |
1.07 |
1.11 |
1.11 |
1.04 |
.99 |
1.07 |
| ATP | .97 |
.99 |
1.02 |
1.02 |
1.01 |
.95 |
.97 |
| KLA/ATP | 1.05 |
1.08 |
1.04 |
1.01 |
1.00 |
1.06 |
1.00 |
|
| Hadha | 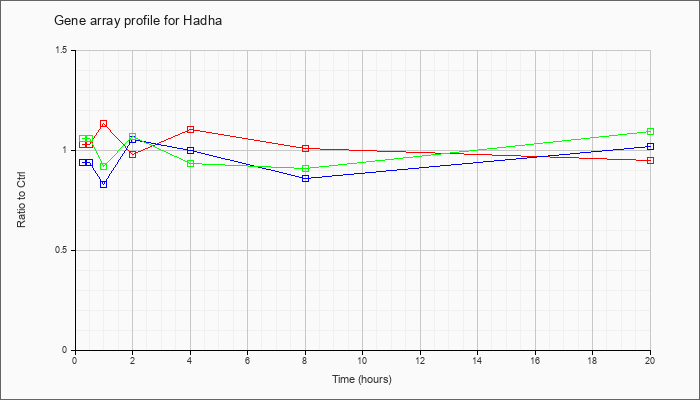 |
AK050856 |
9 days embryo whole body cDNA, RIKEN full-length enriched library, clone:D030026J11 product:hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), alpha subunit, full insert |
KLA | .96 |
.92 |
.91 |
.80 |
.97 |
.92 |
.90 |
| ATP | .72 |
.64 |
.63 |
.71 |
.91 |
.93 |
.93 |
| KLA/ATP | .79 |
.72 |
.68 |
.73 |
.87 |
.90 |
.88 |
|
| Hadha |  |
NM_178878 |
hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), alpha subunit (Hadha), nuclear gene encoding mitochondrial protein, mRNA [NM_178878] |
KLA | 1.10 |
1.17 |
1.36 |
1.16 |
1.23 |
1.10 |
1.00 |
| ATP | 1.16 |
1.22 |
1.03 |
1.40 |
1.09 |
.79 |
1.11 |
| KLA/ATP | 1.33 |
1.35 |
1.16 |
1.41 |
1.00 |
.92 |
1.31 |
|
| Hibch | 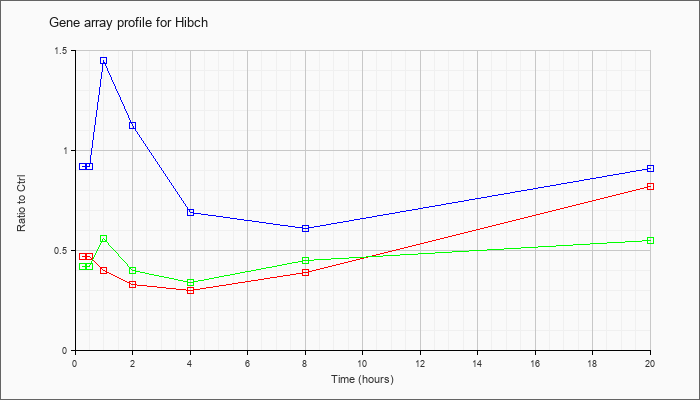 |
NM_146108 |
3-hydroxyisobutyryl-Coenzyme A hydrolase (Hibch), mRNA [NM_146108] |
KLA | .47 |
.44 |
.40 |
.33 |
.30 |
.39 |
.82 |
| ATP | .92 |
.92 |
1.45 |
1.12 |
.69 |
.61 |
.91 |
| KLA/ATP | .42 |
.41 |
.56 |
.40 |
.34 |
.45 |
.55 |
|
| Mlycd | 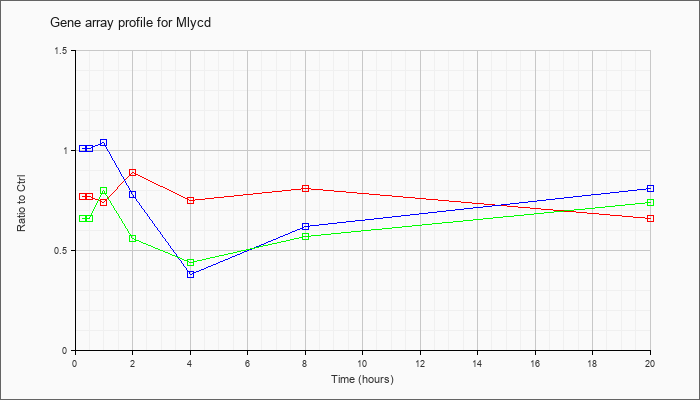 |
NM_019966 |
malonyl-CoA decarboxylase (Mlycd), nuclear gene encoding mitochondrial protein, mRNA [NM_019966] |
KLA | .77 |
.70 |
.74 |
.89 |
.75 |
.81 |
.66 |
| ATP | 1.01 |
.91 |
1.04 |
.78 |
.38 |
.62 |
.81 |
| KLA/ATP | .66 |
.66 |
.80 |
.56 |
.44 |
.57 |
.74 |
|
Mm.26684
3 | 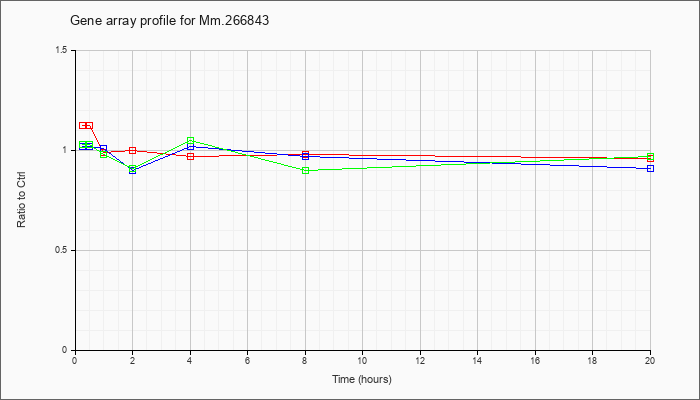 |
94405693 |
Unknown |
KLA | 1.12 |
.99 |
.99 |
1.00 |
.97 |
.98 |
.96 |
| ATP | 1.02 |
1.08 |
1.01 |
.90 |
1.02 |
.97 |
.91 |
| KLA/ATP | 1.03 |
.96 |
.98 |
.91 |
1.05 |
.90 |
.97 |
|
Mm.34137
| 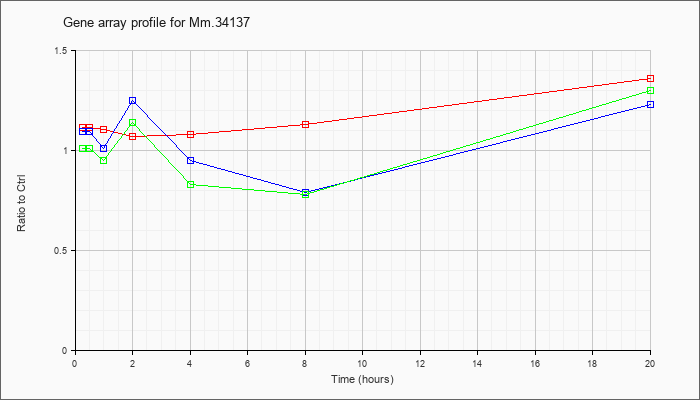 |
94405716 |
Unknown |
KLA | 1.11 |
1.12 |
1.10 |
1.07 |
1.08 |
1.13 |
1.36 |
| ATP | 1.09 |
1.14 |
1.01 |
1.25 |
.95 |
.79 |
1.23 |
| KLA/ATP | 1.01 |
1.24 |
.95 |
1.14 |
.83 |
.78 |
1.30 |
|
| Sms | 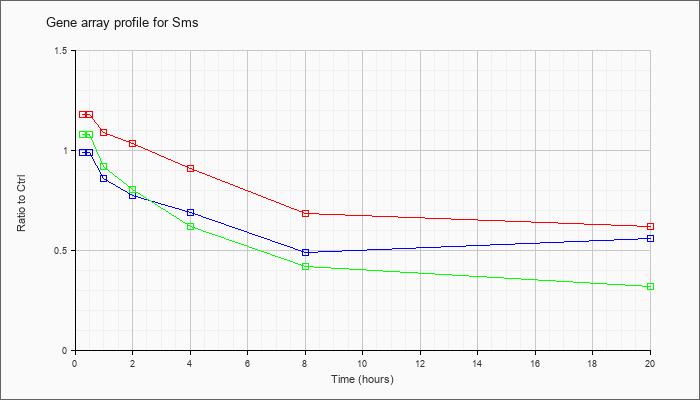 |
NM_009214 |
spermine synthase (Sms), mRNA [NM_009214] |
KLA | 1.18 |
1.13 |
1.09 |
1.03 |
.91 |
.68 |
.62 |
| ATP | .99 |
1.03 |
.86 |
.77 |
.69 |
.49 |
.56 |
| KLA/ATP | 1.08 |
1.13 |
.92 |
.80 |
.62 |
.42 |
.32 |
|
| Srm |  |
NM_009272 |
spermidine synthase (Srm), mRNA [NM_009272] |
KLA | .61 |
.68 |
.55 |
.41 |
.31 |
.32 |
.57 |
| ATP | .95 |
1.06 |
.92 |
1.19 |
.94 |
.50 |
.61 |
| KLA/ATP | .70 |
.65 |
.55 |
.59 |
.44 |
.26 |
.17 |
|
| Upb1 | 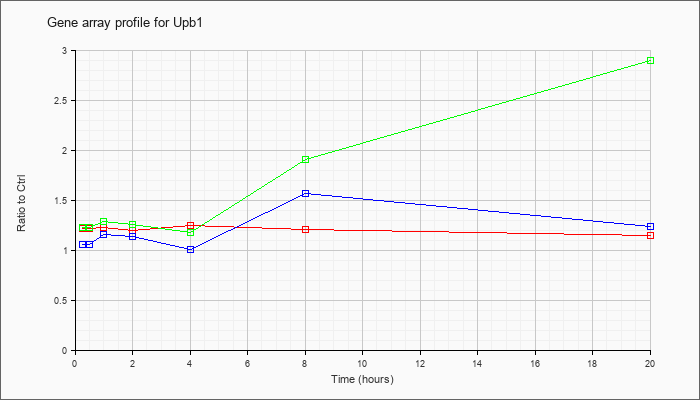 |
NM_133995 |
ureidopropionase, beta (Upb1), mRNA [NM_133995] |
KLA | 1.22 |
1.30 |
1.23 |
1.20 |
1.25 |
1.21 |
1.15 |
| ATP | 1.06 |
1.13 |
1.16 |
1.14 |
1.01 |
1.57 |
1.24 |
| KLA/ATP | 1.23 |
1.24 |
1.29 |
1.26 |
1.18 |
1.91 |
2.90 |
|