| Gene symbol |
Time course plot |
Accession |
Description |
Treatment |
15min |
30min |
1hr |
2hr |
4hr |
8hr |
20hr |
1500003O
03Rik | 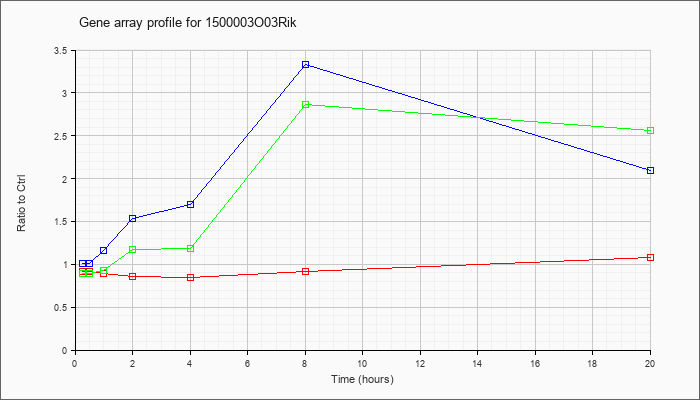 |
NM_019769 |
RIKEN cDNA 1500003O03 gene (1500003O03Rik), mRNA [NM_019769] |
KLA | .92 |
.89 |
.89 |
.86 |
.85 |
.92 |
1.08 |
| ATP | 1.01 |
1.16 |
1.16 |
1.53 |
1.70 |
3.33 |
2.09 |
| KLA/ATP | .89 |
.97 |
.93 |
1.17 |
1.19 |
2.86 |
2.57 |
|
2010110P
09Rik | 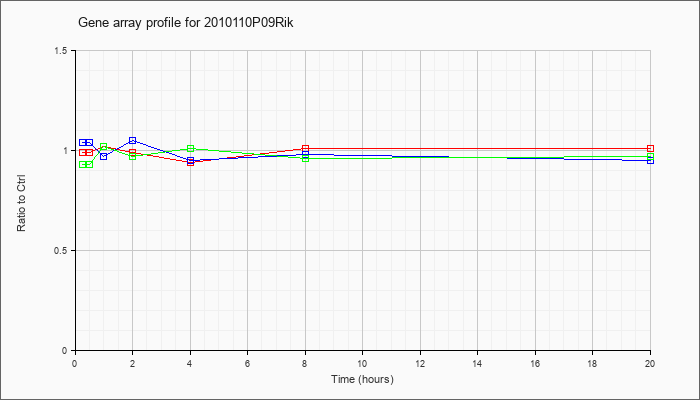 |
NM_027363 |
RIKEN cDNA 2010110P09 gene (2010110P09Rik), mRNA [NM_027363] |
KLA | .99 |
.98 |
1.02 |
.99 |
.94 |
1.01 |
1.01 |
| ATP | 1.04 |
1.05 |
.97 |
1.05 |
.95 |
.98 |
.95 |
| KLA/ATP | .93 |
1.00 |
1.02 |
.97 |
1.01 |
.96 |
.97 |
|
AK036998
|  |
AK036998 |
adult female vagina cDNA, RIKEN full-length enriched library, clone:9930034M05 product:unclassifiable, full insert sequence. [AK036998] |
KLA | 1.43 |
1.09 |
.89 |
.79 |
.96 |
.97 |
.71 |
| ATP | 1.04 |
1.04 |
.71 |
.64 |
.90 |
.82 |
.82 |
| KLA/ATP | 1.34 |
1.23 |
1.01 |
.89 |
1.10 |
.82 |
.75 |
|
| Acp5 |  |
NM_007388 |
acid phosphatase 5, tartrate resistant (Acp5), transcript variant 3, mRNA [NM_007388] |
KLA | 3.62 |
3.55 |
3.27 |
3.26 |
3.31 |
3.93 |
5.79 |
| ATP | .96 |
1.00 |
1.29 |
1.96 |
2.30 |
3.05 |
3.81 |
| KLA/ATP | 3.61 |
3.85 |
4.05 |
5.38 |
4.56 |
5.15 |
6.77 |
|
| Akt1 | 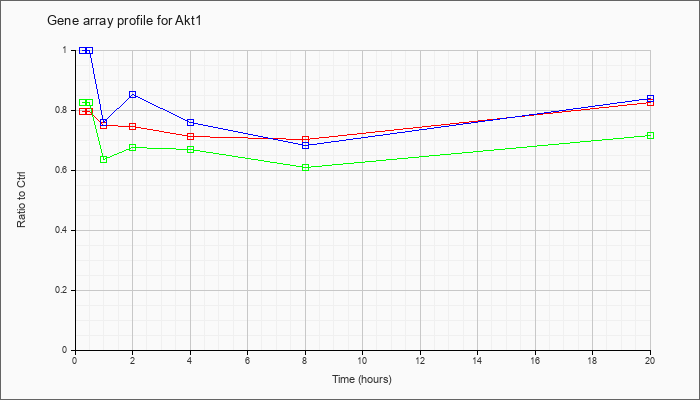 |
NM_009652 |
thymoma viral proto-oncogene 1 (Akt1), mRNA [NM_009652] |
KLA | .80 |
.80 |
.75 |
.75 |
.71 |
.70 |
.82 |
| ATP | 1.00 |
1.02 |
.76 |
.85 |
.76 |
.68 |
.84 |
| KLA/ATP | .82 |
.80 |
.64 |
.68 |
.67 |
.61 |
.72 |
|
| Akt2 | 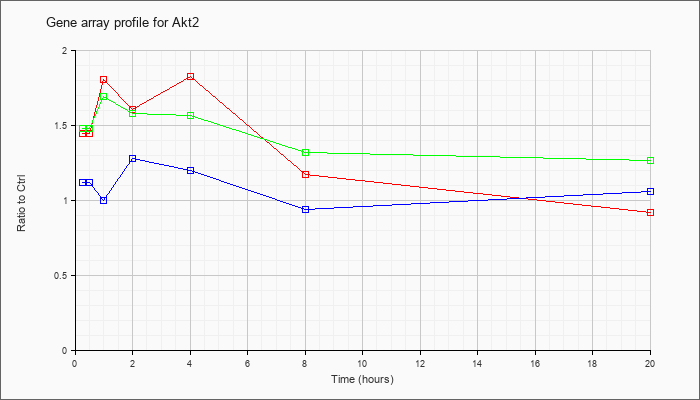 |
NM_007434 |
thymoma viral proto-oncogene 2 (Akt2), transcript variant 2, mRNA [NM_007434] |
KLA | 1.45 |
1.49 |
1.81 |
1.61 |
1.83 |
1.17 |
.92 |
| ATP | 1.12 |
1.12 |
1.00 |
1.28 |
1.20 |
.94 |
1.06 |
| KLA/ATP | 1.48 |
1.70 |
1.69 |
1.58 |
1.57 |
1.32 |
1.27 |
|
| Akt3 | 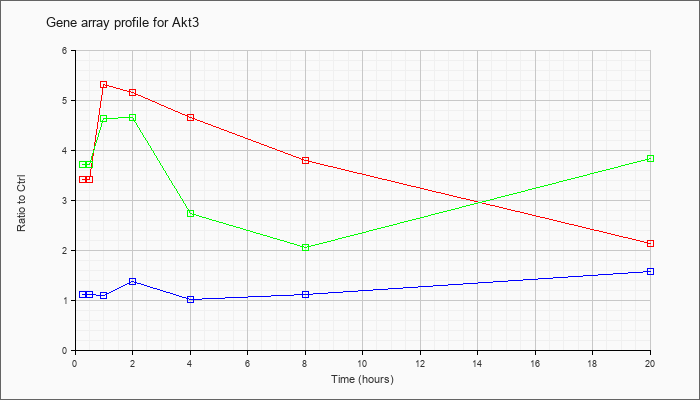 |
NM_011785 |
thymoma viral proto-oncogene 3 (Akt3), mRNA [NM_011785] |
KLA | 3.42 |
3.45 |
5.31 |
5.16 |
4.65 |
3.80 |
2.12 |
| ATP | 1.12 |
1.22 |
1.10 |
1.37 |
1.02 |
1.11 |
1.56 |
| KLA/ATP | 3.72 |
4.17 |
4.62 |
4.64 |
2.74 |
2.05 |
3.82 |
|
BB229853
|  |
BB229853 |
gb|BB229853 RIKEN full-length enriched, 3 days neonate thymus Mus musculus cDNA clone A630023O21 3. [BB229853] |
KLA | 6.32 |
5.94 |
6.13 |
6.41 |
7.39 |
12.33 |
6.08 |
| ATP | .39 |
.24 |
.33 |
.38 |
1.18 |
5.75 |
2.85 |
| KLA/ATP | 2.27 |
2.02 |
3.25 |
4.04 |
7.11 |
10.22 |
13.93 |
|
| Blnk | 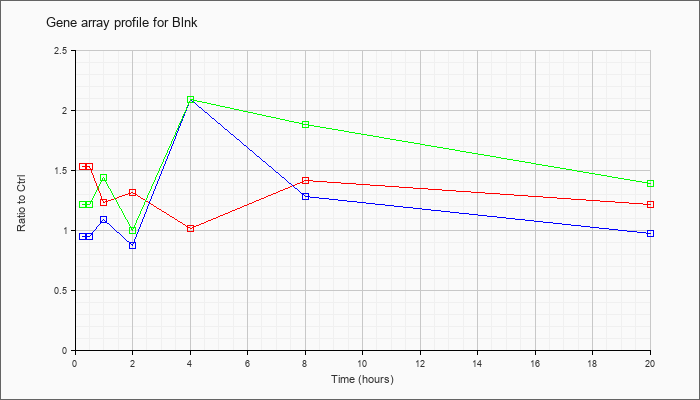 |
NM_008528 |
B-cell linker (Blnk), mRNA [NM_008528] |
KLA | 1.53 |
1.39 |
1.23 |
1.31 |
1.01 |
1.41 |
1.21 |
| ATP | .95 |
.81 |
1.09 |
.87 |
2.09 |
1.28 |
.97 |
| KLA/ATP | 1.21 |
1.23 |
1.44 |
1.00 |
2.09 |
1.88 |
1.39 |
|
| Btk | 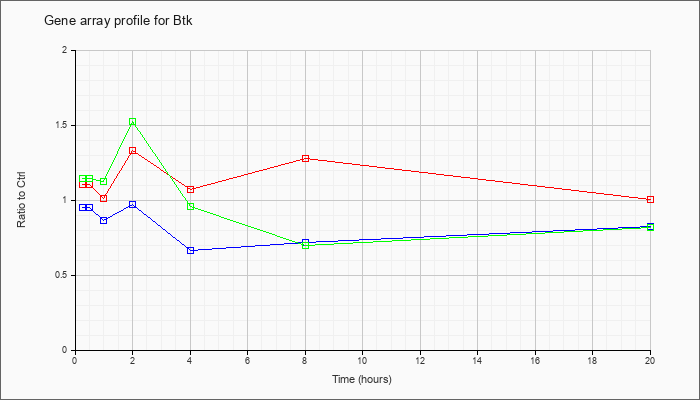 |
NM_013482 |
Bruton agammaglobulinemia tyrosine kinase (Btk), mRNA [NM_013482] |
KLA | 1.10 |
1.24 |
1.01 |
1.33 |
1.07 |
1.27 |
1.01 |
| ATP | .95 |
1.05 |
.86 |
.97 |
.66 |
.72 |
.82 |
| KLA/ATP | 1.14 |
1.35 |
1.12 |
1.52 |
.96 |
.70 |
.82 |
|
| Calcr | 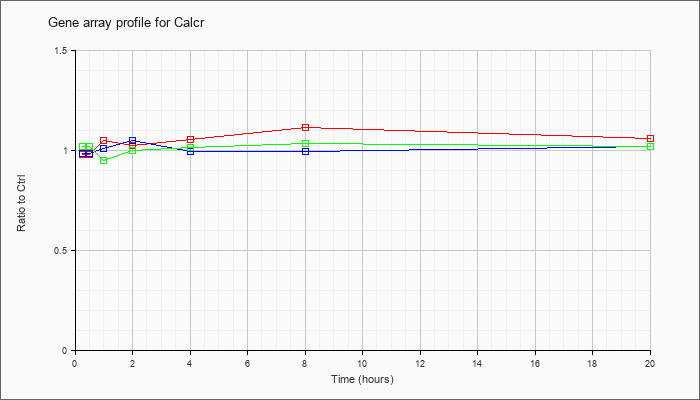 |
NM_007588 |
calcitonin receptor (Calcr), transcript variant a, mRNA [NM_007588] |
KLA | .98 |
1.01 |
1.05 |
1.03 |
1.06 |
1.12 |
1.06 |
| ATP | .99 |
.97 |
1.01 |
1.05 |
1.00 |
1.00 |
1.02 |
| KLA/ATP | 1.02 |
1.02 |
.95 |
1.00 |
1.02 |
1.04 |
1.02 |
|
| Camk4 | 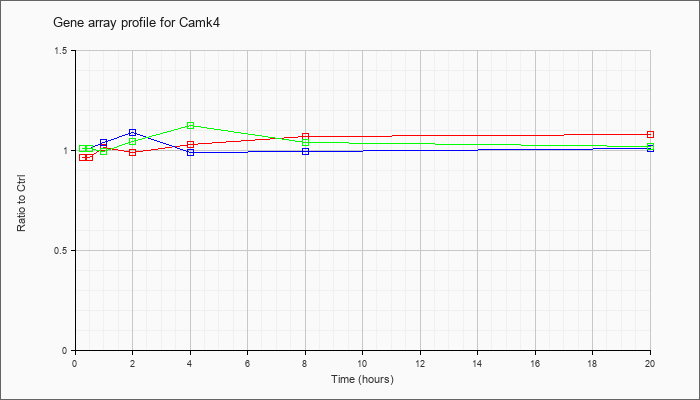 |
AK043457 |
7 days neonate cerebellum cDNA, RIKEN full-length enriched library, clone:A730097A20 product:CALCIUM/CALMODULIN-DEPENDENT PROTEIN KINASE TYPE IV CATALYTIC CHAIN (EC 2.7.1.123) (CAM KINASE-GR) (CAMK IV) [CONTAINS: CALSPERMIN] homolog [Mus |
KLA | .98 |
1.04 |
1.02 |
.93 |
1.03 |
1.07 |
1.07 |
| ATP | 1.01 |
.97 |
.97 |
1.10 |
1.00 |
.99 |
1.05 |
| KLA/ATP | 1.00 |
.96 |
1.02 |
.98 |
1.13 |
1.07 |
1.00 |
|
| Camk4 |  |
NM_009793 |
calcium/calmodulin-dependent protein kinase IV (Camk4), mRNA [NM_009793] |
KLA | .96 |
1.02 |
1.01 |
1.02 |
1.03 |
1.07 |
1.08 |
| ATP | 1.01 |
1.03 |
1.07 |
1.08 |
.99 |
1.00 |
.99 |
| KLA/ATP | 1.01 |
.95 |
.98 |
1.08 |
1.12 |
1.02 |
1.03 |
|
| Chuk | 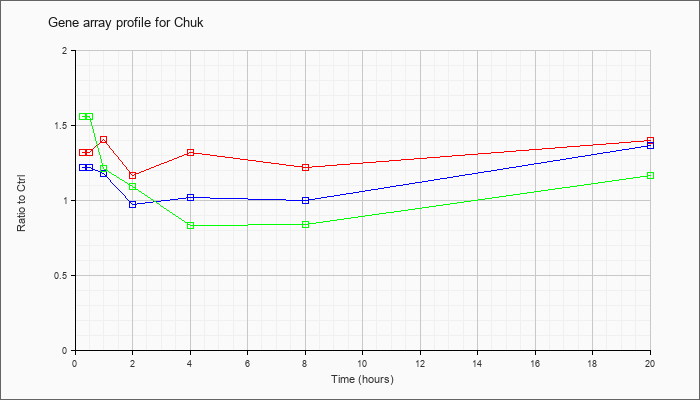 |
NM_007700 |
conserved helix-loop-helix ubiquitous kinase (Chuk), mRNA [NM_007700] |
KLA | 1.32 |
1.42 |
1.41 |
1.17 |
1.32 |
1.22 |
1.40 |
| ATP | 1.22 |
1.30 |
1.18 |
.97 |
1.02 |
1.00 |
1.37 |
| KLA/ATP | 1.56 |
1.61 |
1.21 |
1.09 |
.83 |
.84 |
1.17 |
|
| Creb1 | 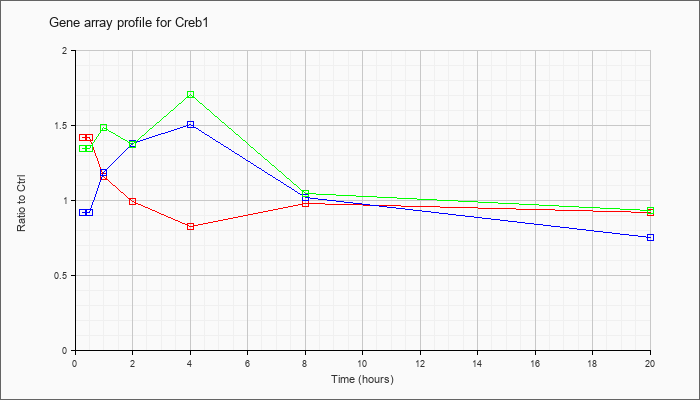 |
NM_009952 |
cAMP responsive element binding protein 1 (Creb1), transcript variant B, mRNA [NM_009952] |
KLA | 1.42 |
1.33 |
1.16 |
.99 |
.83 |
.97 |
.92 |
| ATP | .92 |
.91 |
1.18 |
1.38 |
1.51 |
1.02 |
.75 |
| KLA/ATP | 1.34 |
1.31 |
1.49 |
1.37 |
1.71 |
1.05 |
.93 |
|
| Csf1 | 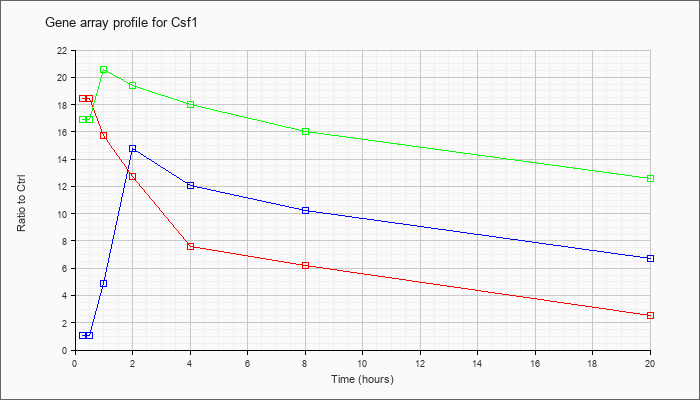 |
NM_007778 |
colony stimulating factor 1 (macrophage) (Csf1), transcript variant 1, mRNA [NM_007778] |
KLA | 18.44 |
16.94 |
15.74 |
12.71 |
7.63 |
6.20 |
2.51 |
| ATP | 1.04 |
1.22 |
4.90 |
14.80 |
12.10 |
10.26 |
6.73 |
| KLA/ATP | 16.93 |
18.17 |
20.56 |
19.41 |
18.03 |
15.99 |
12.59 |
|
| Csf1r | 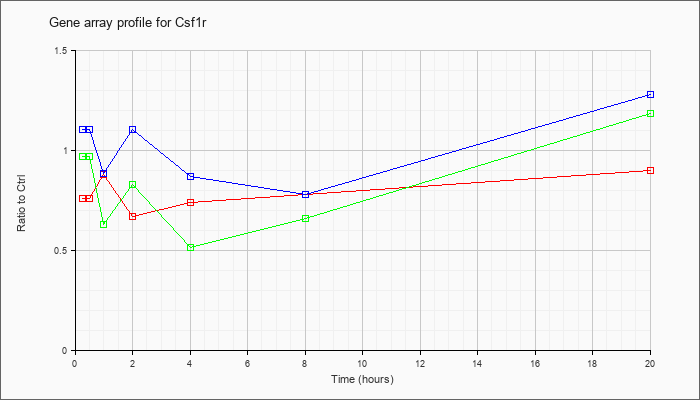 |
NM_001037859 |
colony stimulating factor 1 receptor (Csf1r), mRNA [NM_001037859] |
KLA | .76 |
.81 |
.88 |
.67 |
.74 |
.78 |
.90 |
| ATP | 1.10 |
1.22 |
.88 |
1.10 |
.87 |
.78 |
1.28 |
| KLA/ATP | .97 |
.87 |
.63 |
.83 |
.51 |
.66 |
1.18 |
|
| Ctsk | 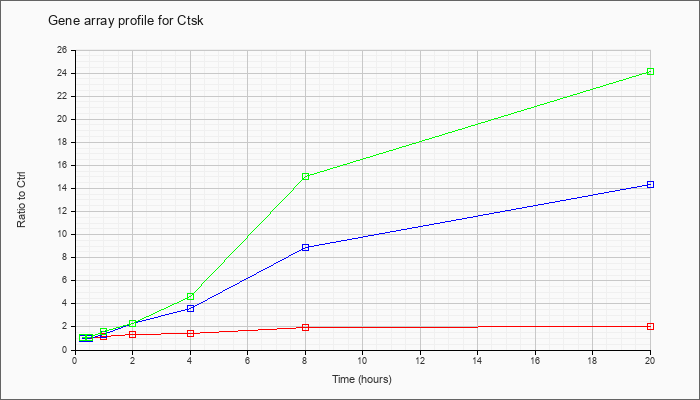 |
NM_007802 |
cathepsin K (Ctsk), mRNA [NM_007802] |
KLA | 1.02 |
1.14 |
1.16 |
1.37 |
1.40 |
1.95 |
2.05 |
| ATP | 1.04 |
1.06 |
1.34 |
2.32 |
3.59 |
8.92 |
14.36 |
| KLA/ATP | 1.08 |
1.23 |
1.62 |
2.29 |
4.63 |
15.04 |
24.14 |
|
| Cyba | 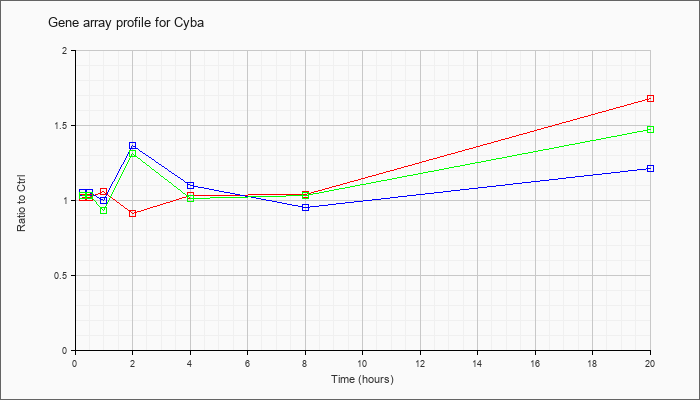 |
NM_007806 |
cytochrome b-245, alpha polypeptide (Cyba), mRNA [NM_007806] |
KLA | 1.02 |
1.05 |
1.06 |
.91 |
1.03 |
1.04 |
1.68 |
| ATP | 1.05 |
1.11 |
1.00 |
1.36 |
1.10 |
.95 |
1.21 |
| KLA/ATP | 1.03 |
1.11 |
.93 |
1.31 |
1.01 |
1.03 |
1.47 |
|
| Cybb | 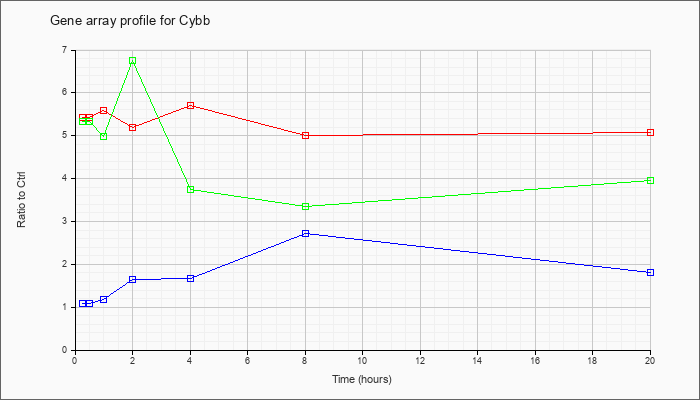 |
NM_007807 |
cytochrome b-245, beta polypeptide (Cybb), mRNA [NM_007807] |
KLA | 5.43 |
5.43 |
5.59 |
5.19 |
5.71 |
5.00 |
5.08 |
| ATP | 1.08 |
1.18 |
1.19 |
1.64 |
1.68 |
2.73 |
1.82 |
| KLA/ATP | 5.34 |
5.62 |
4.98 |
6.75 |
3.74 |
3.36 |
3.96 |
|
| Cyld | 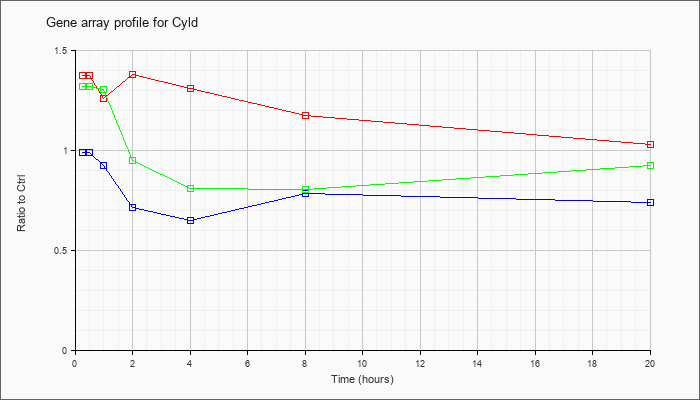 |
AK147204 |
15 days pregnant adult female placenta cDNA, RIKEN full-length enriched library, clone:M421001K23 product:cylindromatosis (turban tumor syndrome), full insert sequence [AK147204] |
KLA | .85 |
.83 |
.73 |
.65 |
.62 |
.61 |
.48 |
| ATP | .88 |
.79 |
.69 |
.47 |
.44 |
.43 |
.21 |
| KLA/ATP | .75 |
.66 |
.63 |
.42 |
.57 |
.41 |
.21 |
|
| Cyld |  |
NM_173369 |
cylindromatosis (turban tumor syndrome) (Cyld), mRNA [NM_173369] |
KLA | 1.64 |
1.65 |
1.53 |
1.74 |
1.66 |
1.46 |
1.30 |
| ATP | 1.05 |
1.14 |
1.04 |
.84 |
.75 |
.96 |
1.00 |
| KLA/ATP | 1.60 |
1.77 |
1.64 |
1.22 |
.93 |
1.00 |
1.28 |
|
EG232801
| 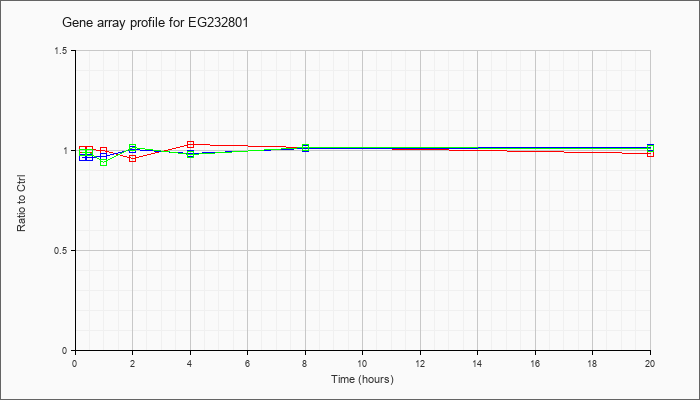 |
NM_001081239 |
predicted gene, EG232801 (EG232801), mRNA [NM_001081239] |
KLA | 1.01 |
1.01 |
1.00 |
.96 |
1.03 |
1.02 |
.99 |
| ATP | .97 |
1.00 |
.97 |
1.01 |
.99 |
1.01 |
1.02 |
| KLA/ATP | .99 |
1.01 |
.94 |
1.02 |
.98 |
1.02 |
1.01 |
|
| Fcgr1 | 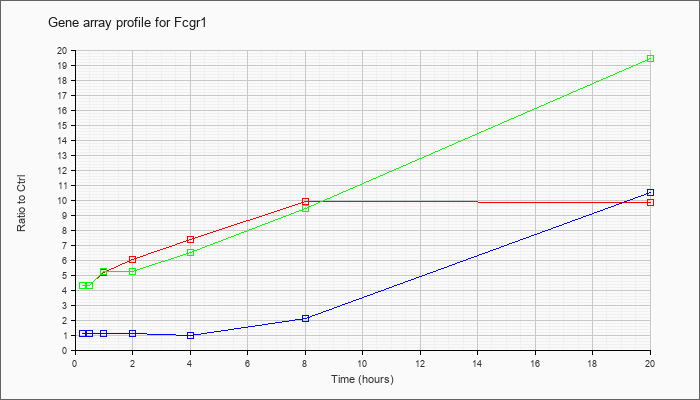 |
AF143181 |
strain AB/H (Biozzi) high affinity immunoglobulin gamma Fc receptor I (Fcgr1) mRNA, Fcgr1-d allele, complete cds. [AF143181] |
KLA | 4.27 |
4.21 |
5.59 |
6.14 |
8.13 |
9.76 |
10.98 |
| ATP | 1.23 |
1.12 |
1.08 |
1.36 |
1.09 |
2.09 |
11.81 |
| KLA/ATP | 4.56 |
4.93 |
4.95 |
5.88 |
6.66 |
9.64 |
23.09 |
|
| Fcgr1 |  |
NM_010186 |
Fc receptor, IgG, high affinity I (Fcgr1), mRNA [NM_010186] |
KLA | 4.31 |
4.37 |
4.70 |
5.97 |
6.59 |
10.06 |
8.67 |
| ATP | 1.01 |
.88 |
1.16 |
.89 |
.90 |
2.15 |
9.23 |
| KLA/ATP | 4.01 |
4.10 |
5.52 |
4.54 |
6.31 |
9.24 |
15.83 |
|
| Fcgr2b | 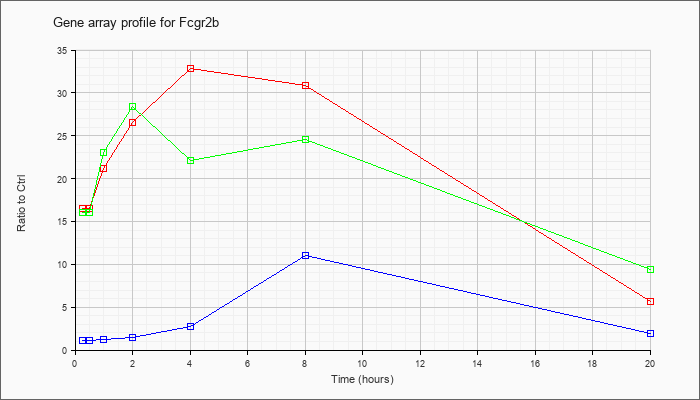 |
NM_001077189 |
Fc receptor, IgG, low affinity IIb (Fcgr2b), transcript variant 1, mRNA [NM_001077189] |
KLA | 16.93 |
17.57 |
21.75 |
27.43 |
34.13 |
31.88 |
5.70 |
| ATP | 1.11 |
1.09 |
1.17 |
1.46 |
2.81 |
11.26 |
1.92 |
| KLA/ATP | 16.51 |
17.91 |
23.67 |
29.22 |
22.67 |
25.33 |
9.54 |
|
| Fcgr2b |  |
NM_010187 |
Fc receptor, IgG, low affinity IIb (Fcgr2b), transcript variant 2, mRNA [NM_010187] |
KLA | 11.41 |
11.67 |
14.06 |
15.47 |
17.28 |
18.63 |
4.69 |
| ATP | 1.05 |
1.06 |
1.18 |
1.57 |
2.27 |
8.97 |
2.10 |
| KLA/ATP | 10.98 |
12.01 |
15.03 |
18.38 |
15.15 |
16.00 |
8.33 |
|
| Fcgr3 | 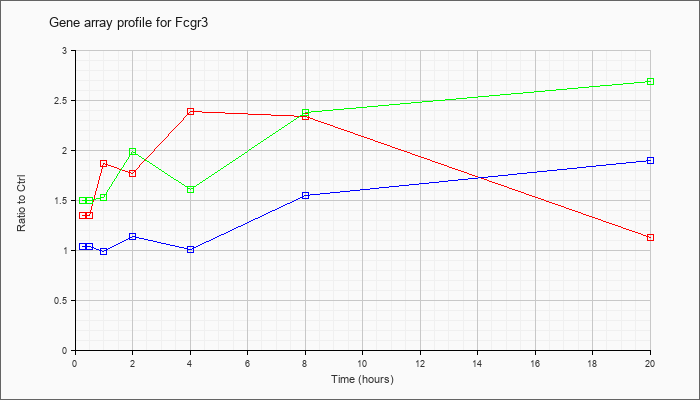 |
NM_010188 |
Fc receptor, IgG, low affinity III (Fcgr3), mRNA [NM_010188] |
KLA | 1.35 |
1.47 |
1.87 |
1.77 |
2.39 |
2.34 |
1.13 |
| ATP | 1.04 |
1.10 |
.99 |
1.14 |
1.01 |
1.55 |
1.90 |
| KLA/ATP | 1.50 |
1.53 |
1.53 |
1.99 |
1.61 |
2.38 |
2.69 |
|
| Fcgr4 | 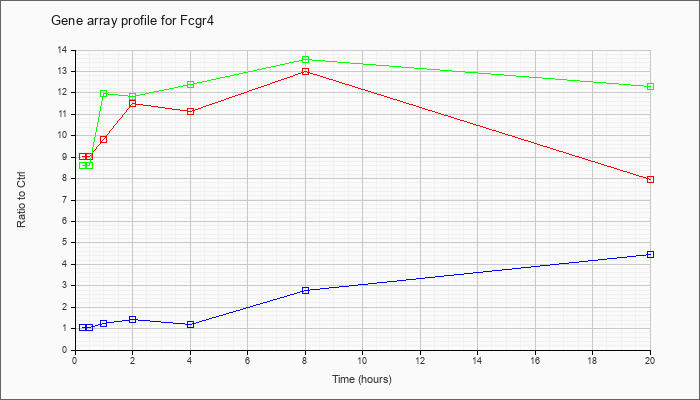 |
NM_144559 |
Fc receptor, IgG, low affinity IV (Fcgr4), mRNA [NM_144559] |
KLA | 9.03 |
8.68 |
9.82 |
11.51 |
11.12 |
13.01 |
7.95 |
| ATP | 1.04 |
.95 |
1.22 |
1.43 |
1.17 |
2.80 |
4.48 |
| KLA/ATP | 8.63 |
8.84 |
11.99 |
11.84 |
12.40 |
13.56 |
12.31 |
|
| Fhl2 | 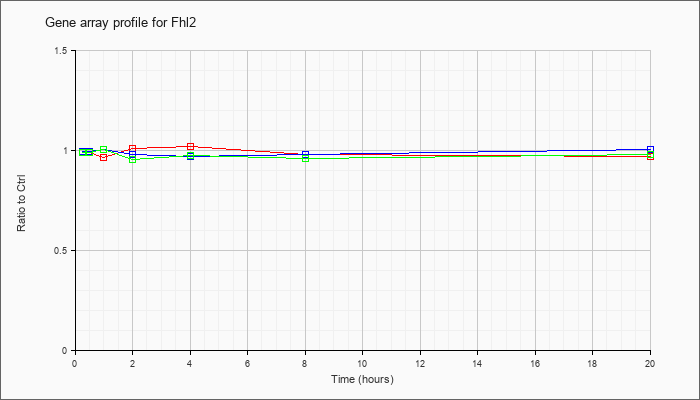 |
AK081770 |
16 days embryo head cDNA, RIKEN full-length enriched library, clone:C130075G22 product:unclassifiable, full insert sequence [AK081770] |
KLA | 1.00 |
1.00 |
.97 |
1.00 |
.96 |
.97 |
1.01 |
| ATP | 1.01 |
.95 |
.98 |
1.02 |
.98 |
1.06 |
1.06 |
| KLA/ATP | 1.00 |
.97 |
.99 |
.99 |
.99 |
1.00 |
.99 |
|
| Fhl2 |  |
NM_010212 |
four and a half LIM domains 2 (Fhl2), mRNA [NM_010212] |
KLA | .97 |
1.02 |
.96 |
1.02 |
1.13 |
1.01 |
.89 |
| ATP | .96 |
1.00 |
1.05 |
.91 |
.94 |
.82 |
.89 |
| KLA/ATP | .97 |
1.04 |
1.03 |
.89 |
.94 |
.87 |
.95 |
|
| Fos | 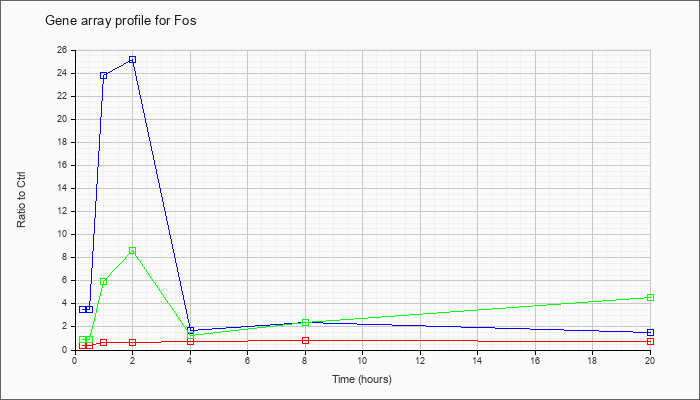 |
NM_010234 |
FBJ osteosarcoma oncogene (Fos), mRNA [NM_010234] |
KLA | .42 |
.33 |
.64 |
.61 |
.71 |
.85 |
.71 |
| ATP | 3.48 |
5.46 |
23.78 |
25.22 |
1.71 |
2.36 |
1.50 |
| KLA/ATP | .90 |
1.74 |
5.97 |
8.62 |
1.28 |
2.42 |
4.52 |
|
| Fosb | 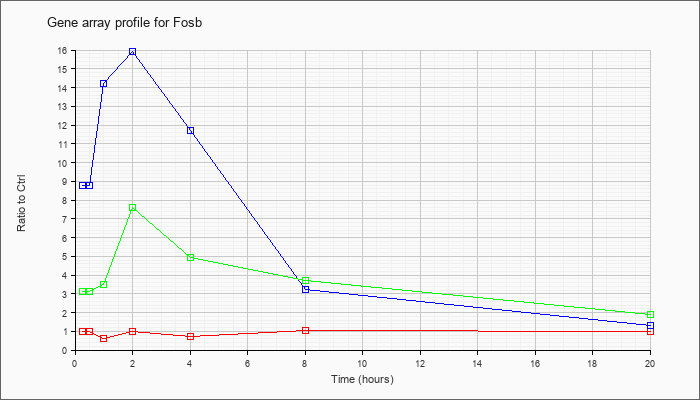 |
NM_008036 |
FBJ osteosarcoma oncogene B (Fosb), mRNA [NM_008036] |
KLA | .99 |
.88 |
.61 |
1.01 |
.74 |
1.03 |
1.00 |
| ATP | 8.75 |
11.87 |
14.21 |
15.92 |
11.71 |
3.22 |
1.29 |
| KLA/ATP | 3.13 |
5.07 |
3.50 |
7.61 |
4.95 |
3.70 |
1.87 |
|
| Fosl1 | 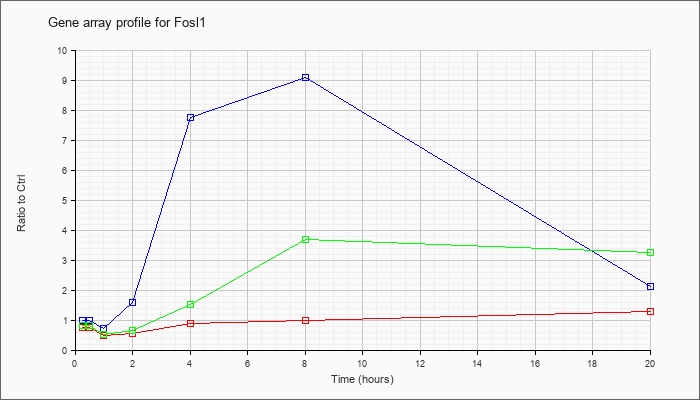 |
NM_010235 |
fos-like antigen 1 (Fosl1), mRNA [NM_010235] |
KLA | .75 |
.72 |
.50 |
.55 |
.87 |
.97 |
1.29 |
| ATP | 1.00 |
.99 |
.72 |
1.57 |
7.76 |
9.10 |
2.11 |
| KLA/ATP | .83 |
.75 |
.53 |
.65 |
1.53 |
3.70 |
3.26 |
|
| Fosl2 | 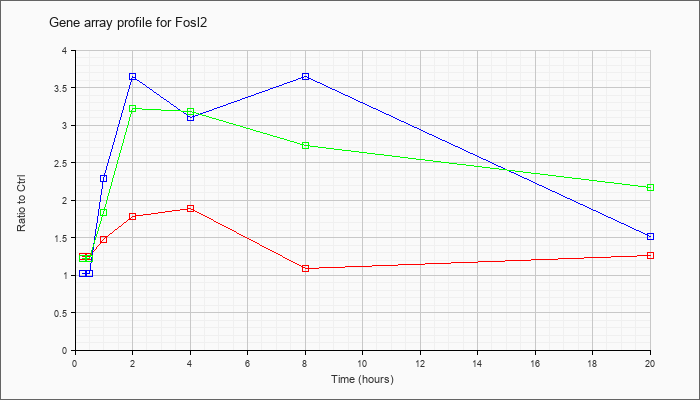 |
NM_008037 |
fos-like antigen 2 (Fosl2), mRNA [NM_008037] |
KLA | 1.25 |
1.36 |
1.47 |
1.78 |
1.89 |
1.09 |
1.26 |
| ATP | 1.02 |
1.33 |
2.29 |
3.65 |
3.11 |
3.65 |
1.52 |
| KLA/ATP | 1.22 |
1.46 |
1.84 |
3.23 |
3.18 |
2.73 |
2.17 |
|
| Fyn | 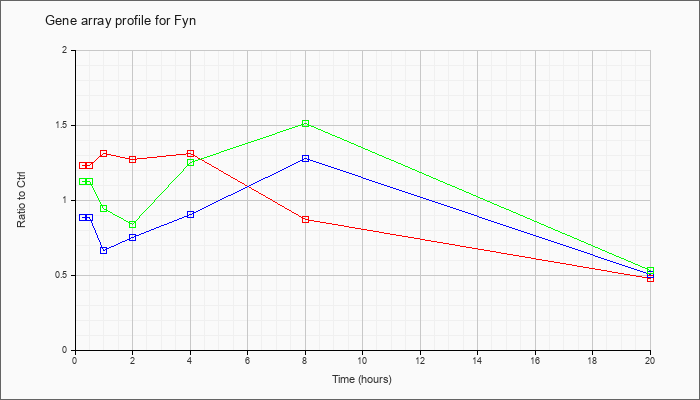 |
U70324 |
Fyn(T) (L-fyn(T)) mRNA, complete cds. [U70324] |
KLA | 1.23 |
1.27 |
1.31 |
1.27 |
1.31 |
.87 |
.48 |
| ATP | .88 |
.71 |
.66 |
.75 |
.90 |
1.28 |
.51 |
| KLA/ATP | 1.12 |
1.01 |
.94 |
.84 |
1.25 |
1.51 |
.53 |
|
| Gab2 |  |
NM_010248 |
growth factor receptor bound protein 2-associated protein 2 (Gab2), mRNA [NM_010248] |
KLA | 1.01 |
1.04 |
1.04 |
1.02 |
1.12 |
1.16 |
1.14 |
| ATP | 1.02 |
1.06 |
1.00 |
1.13 |
1.16 |
1.19 |
1.23 |
| KLA/ATP | 1.06 |
1.05 |
1.03 |
1.02 |
1.03 |
1.04 |
1.29 |
|
| Grb2 | 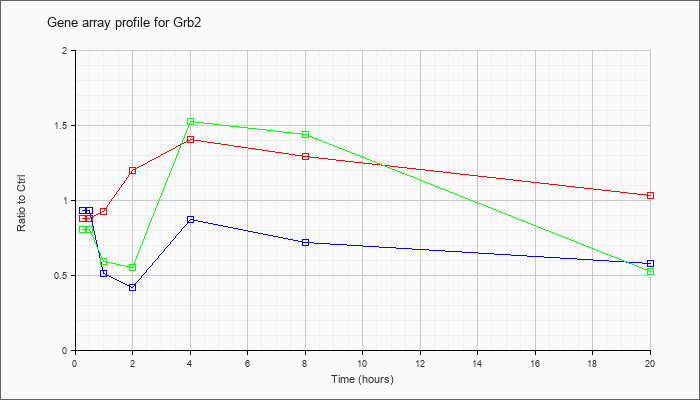 |
NM_008163 |
growth factor receptor bound protein 2 (Grb2), mRNA [NM_008163] |
KLA | .92 |
.88 |
.90 |
1.27 |
1.43 |
1.22 |
1.00 |
| ATP | 1.02 |
.83 |
.51 |
.42 |
.92 |
.70 |
.57 |
| KLA/ATP | .84 |
.77 |
.61 |
.56 |
1.56 |
1.49 |
.52 |
|
| Grb2 |  |
U07617 |
Grb2 adaptor protein (grb2) mRNA, complete cds. [U07617] |
KLA | .83 |
.80 |
.95 |
1.13 |
1.38 |
1.36 |
1.06 |
| ATP | .84 |
.74 |
.51 |
.41 |
.82 |
.74 |
.59 |
| KLA/ATP | .77 |
.70 |
.57 |
.54 |
1.49 |
1.39 |
.53 |
|
| Ifnar1 | 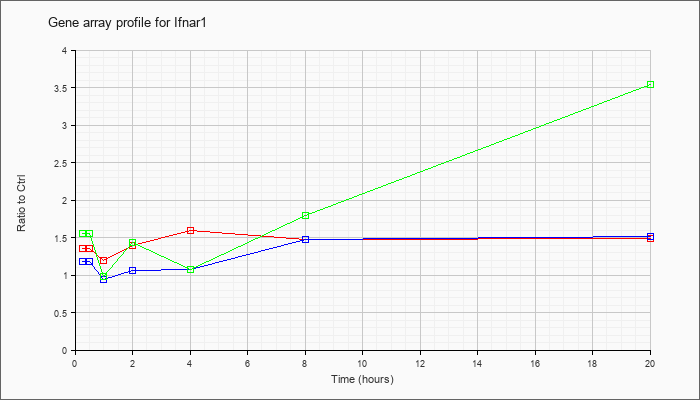 |
NM_010508 |
interferon (alpha and beta) receptor 1 (Ifnar1), mRNA [NM_010508] |
KLA | 1.35 |
1.29 |
1.19 |
1.39 |
1.60 |
1.47 |
1.49 |
| ATP | 1.18 |
1.25 |
.94 |
1.06 |
1.07 |
1.48 |
1.52 |
| KLA/ATP | 1.56 |
1.57 |
.98 |
1.43 |
1.08 |
1.79 |
3.54 |
|
| Ifnar2 | 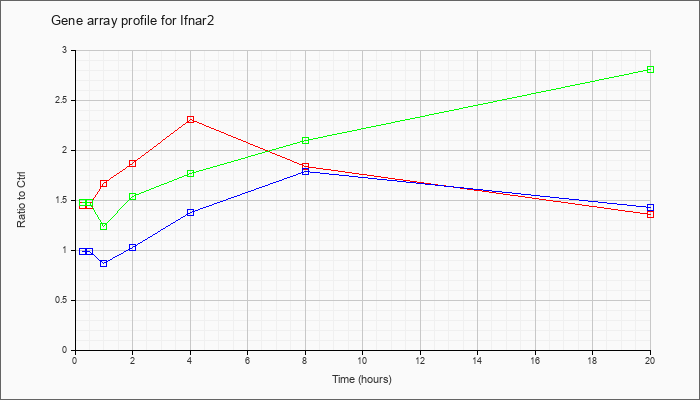 |
NM_010509 |
interferon (alpha and beta) receptor 2 (Ifnar2), transcript variant 1, mRNA [NM_010509] |
KLA | 1.45 |
1.43 |
1.67 |
1.87 |
2.31 |
1.84 |
1.36 |
| ATP | .99 |
.92 |
.87 |
1.03 |
1.38 |
1.79 |
1.43 |
| KLA/ATP | 1.48 |
1.48 |
1.24 |
1.54 |
1.77 |
2.10 |
2.81 |
|
| Ifnb1 |  |
NM_010510 |
interferon beta 1, fibroblast (Ifnb1), mRNA [NM_010510] |
KLA | 49.82 |
46.57 |
34.05 |
16.05 |
5.94 |
7.31 |
8.49 |
| ATP | .99 |
1.02 |
1.89 |
4.78 |
6.31 |
4.44 |
1.79 |
| KLA/ATP | 49.22 |
61.83 |
134.08 |
144.05 |
110.96 |
31.78 |
11.24 |
|
| Ifng | 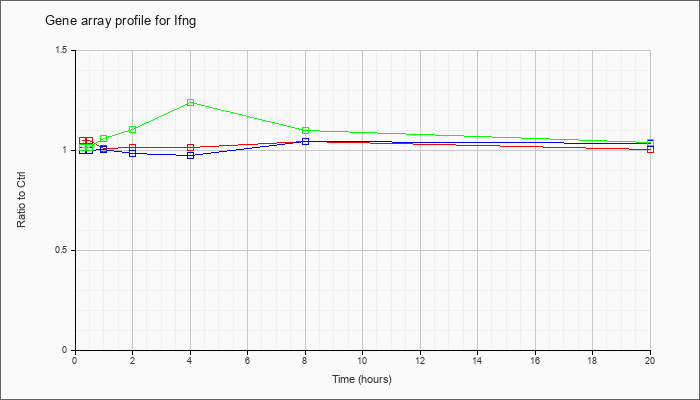 |
NM_008337 |
interferon gamma (Ifng), mRNA [NM_008337] |
KLA | 1.05 |
1.02 |
1.01 |
1.01 |
1.01 |
1.04 |
1.00 |
| ATP | 1.00 |
1.01 |
1.01 |
.98 |
.97 |
1.04 |
1.03 |
| KLA/ATP | 1.01 |
1.01 |
1.06 |
1.10 |
1.24 |
1.10 |
1.04 |
|
| Ifngr1 | 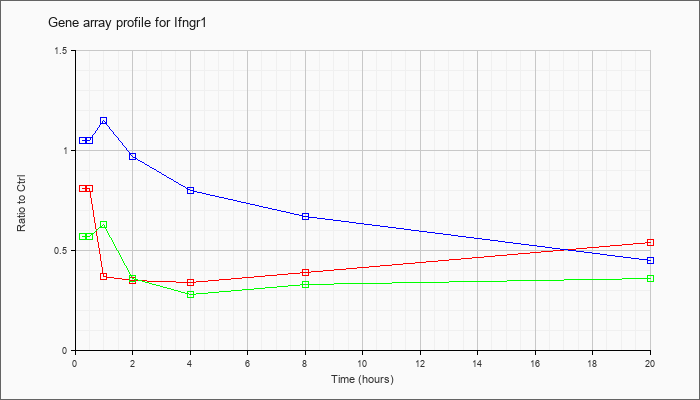 |
NM_010511 |
interferon gamma receptor 1 (Ifngr1), mRNA [NM_010511] |
KLA | .81 |
.60 |
.37 |
.35 |
.34 |
.39 |
.54 |
| ATP | 1.05 |
.96 |
1.15 |
.97 |
.80 |
.67 |
.45 |
| KLA/ATP | .57 |
.57 |
.63 |
.36 |
.28 |
.33 |
.36 |
|
| Ifngr2 | 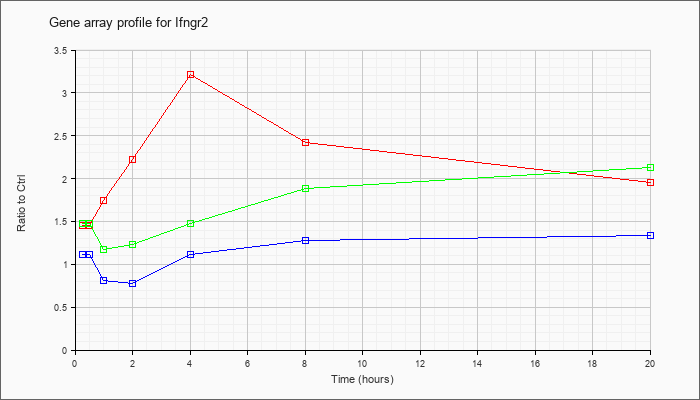 |
NM_008338 |
interferon gamma receptor 2 (Ifngr2), mRNA [NM_008338] |
KLA | 1.45 |
1.49 |
1.74 |
2.22 |
3.22 |
2.42 |
1.96 |
| ATP | 1.11 |
1.03 |
.81 |
.78 |
1.12 |
1.28 |
1.34 |
| KLA/ATP | 1.48 |
1.41 |
1.17 |
1.23 |
1.48 |
1.89 |
2.13 |
|
| Ikbkb | 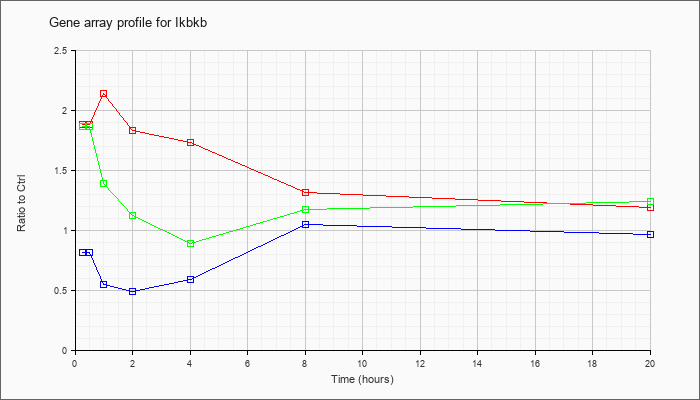 |
NM_010546 |
inhibitor of kappaB kinase beta (Ikbkb), mRNA [NM_010546] |
KLA | 1.88 |
2.10 |
2.14 |
1.83 |
1.73 |
1.31 |
1.19 |
| ATP | .81 |
.74 |
.55 |
.49 |
.59 |
1.05 |
.96 |
| KLA/ATP | 1.86 |
1.73 |
1.39 |
1.12 |
.89 |
1.17 |
1.24 |
|
| Ikbkg | 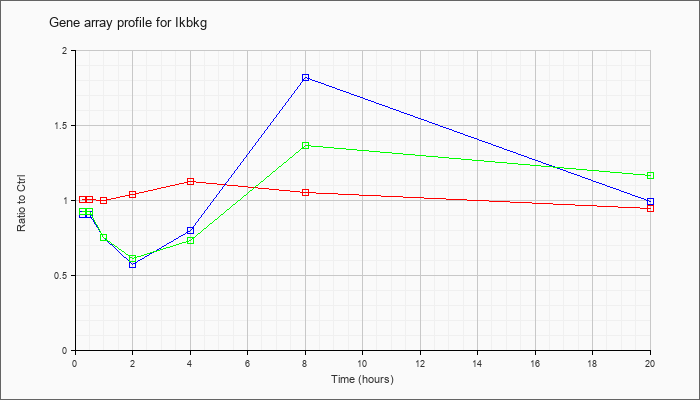 |
AK042138 |
3 days neonate thymus cDNA, RIKEN full-length enriched library, clone:A630062P04 product:unclassifiable, full insert sequence. [AK042138] |
KLA | .92 |
.87 |
.78 |
.78 |
.87 |
.88 |
.62 |
| ATP | .84 |
.71 |
.80 |
.50 |
1.15 |
2.33 |
.61 |
| KLA/ATP | .72 |
.69 |
.74 |
.48 |
1.06 |
1.97 |
.68 |
|
| Ikbkg |  |
NM_010547 |
inhibitor of kappaB kinase gamma (Ikbkg), transcript variant 1, mRNA [NM_010547] |
KLA | 1.51 |
1.63 |
1.74 |
1.83 |
1.83 |
1.73 |
1.72 |
| ATP | .80 |
.79 |
.82 |
.75 |
.66 |
2.58 |
2.37 |
| KLA/ATP | 1.43 |
1.42 |
1.17 |
1.04 |
.71 |
1.54 |
3.17 |
|
| Ikbkg |  |
NM_178590 |
inhibitor of kappaB kinase gamma (Ikbkg), transcript variant 2, mRNA [NM_178590] |
KLA | .84 |
.93 |
.86 |
.91 |
1.03 |
.89 |
.88 |
| ATP | 1.03 |
.98 |
.68 |
.56 |
.51 |
.93 |
.68 |
| KLA/ATP | .89 |
.81 |
.55 |
.52 |
.41 |
.67 |
.65 |
|
| Il1a | 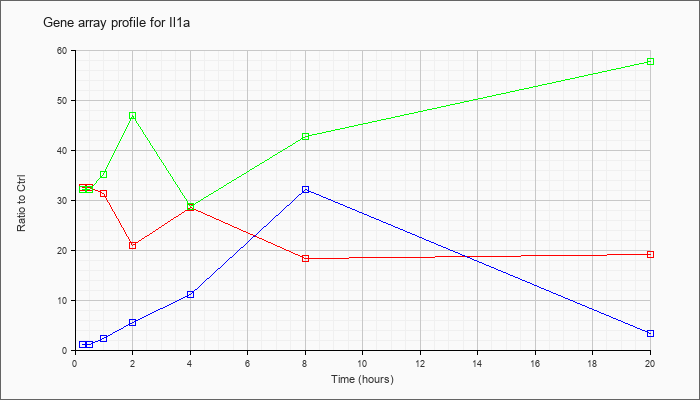 |
NM_010554 |
interleukin 1 alpha (Il1a), mRNA [NM_010554] |
KLA | 32.45 |
35.22 |
31.30 |
20.95 |
28.57 |
18.36 |
19.02 |
| ATP | 1.04 |
1.02 |
2.21 |
5.46 |
11.06 |
32.07 |
3.40 |
| KLA/ATP | 32.02 |
41.20 |
35.12 |
46.91 |
28.76 |
42.72 |
57.80 |
|
| Il1b | 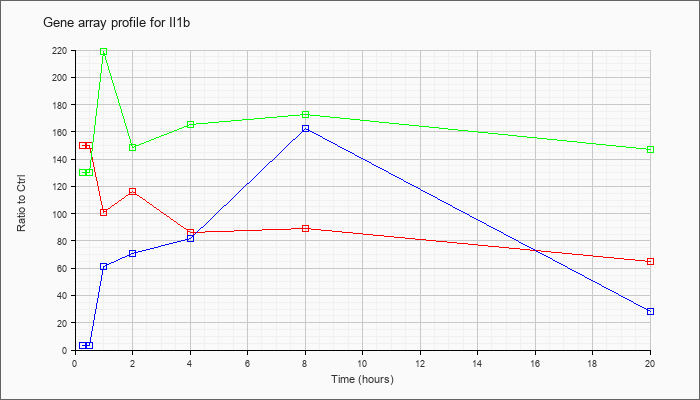 |
NM_008361 |
interleukin 1 beta (Il1b), mRNA [NM_008361] |
KLA | 149.78 |
133.59 |
100.91 |
116.58 |
86.48 |
89.37 |
64.72 |
| ATP | 3.49 |
13.96 |
61.44 |
70.91 |
82.12 |
162.79 |
28.18 |
| KLA/ATP | 130.32 |
139.00 |
219.12 |
148.17 |
165.44 |
172.57 |
146.98 |
|
| Il1r1 |  |
NM_008362 |
interleukin 1 receptor, type I (Il1r1), transcript variant 1, mRNA [NM_008362] |
KLA | 1.20 |
1.08 |
1.22 |
1.52 |
2.19 |
2.32 |
1.61 |
| ATP | 1.10 |
.96 |
1.18 |
1.56 |
8.14 |
9.56 |
7.54 |
| KLA/ATP | 1.22 |
1.13 |
1.37 |
1.85 |
4.54 |
6.85 |
8.96 |
|
| Irf9 | 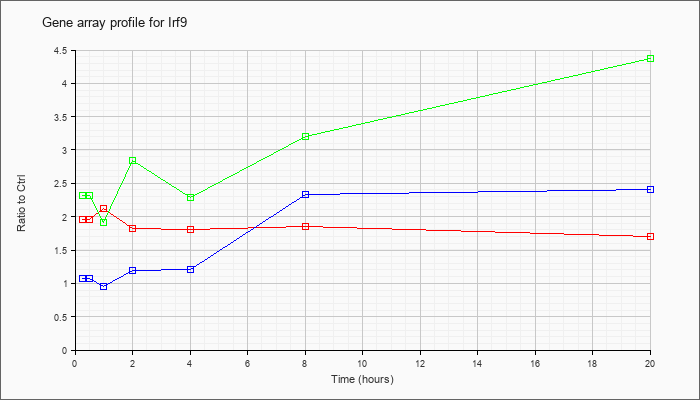 |
NM_008394 |
interferon regulatory factor 9 (Irf9), mRNA [NM_008394] |
KLA | 1.96 |
1.99 |
2.13 |
1.83 |
1.81 |
1.85 |
1.71 |
| ATP | 1.08 |
1.06 |
.96 |
1.19 |
1.21 |
2.33 |
2.42 |
| KLA/ATP | 2.31 |
2.30 |
1.92 |
2.85 |
2.29 |
3.20 |
4.37 |
|
| Itgb3 | 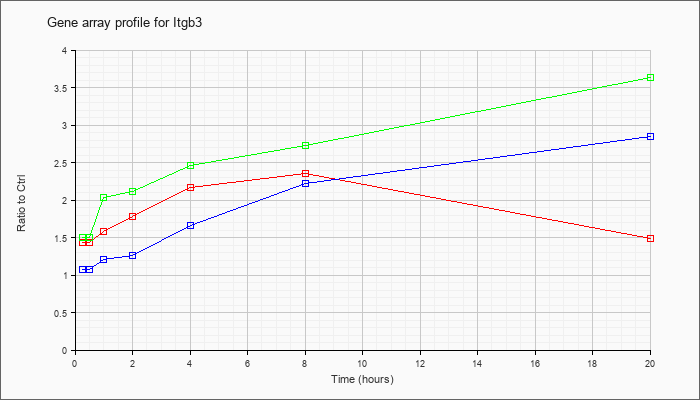 |
AF026509 |
integrin beta3 subunit mRNA, complete cds. [AF026509] |
KLA | 2.38 |
2.64 |
2.65 |
3.15 |
4.02 |
4.46 |
2.83 |
| ATP | 1.21 |
1.17 |
1.25 |
1.62 |
2.33 |
3.09 |
4.78 |
| KLA/ATP | 2.48 |
2.63 |
3.39 |
3.72 |
4.30 |
4.71 |
6.79 |
|
| Itgb3 |  |
NM_016780 |
integrin beta 3 (Itgb3), mRNA [NM_016780] |
KLA | .50 |
.51 |
.52 |
.41 |
.31 |
.25 |
.15 |
| ATP | .94 |
.88 |
1.17 |
.89 |
.99 |
1.36 |
.91 |
| KLA/ATP | .51 |
.54 |
.67 |
.52 |
.61 |
.74 |
.48 |
|
| Jak1 | 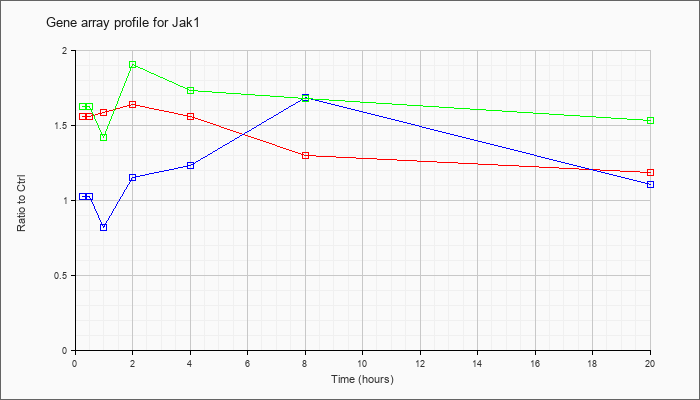 |
AK029232 |
0 day neonate head cDNA, RIKEN full-length enriched library, clone:4831433J01 product:Janus kinase 1, full insert sequence. [AK029232] |
KLA | 1.25 |
1.13 |
1.19 |
1.31 |
1.24 |
1.17 |
1.14 |
| ATP | 1.01 |
1.11 |
.96 |
1.38 |
1.06 |
1.26 |
1.05 |
| KLA/ATP | 1.19 |
1.29 |
1.27 |
1.59 |
1.18 |
1.38 |
1.24 |
|
| Jak1 |  |
NM_146145 |
Janus kinase 1 (Jak1), mRNA [NM_146145] |
KLA | 1.71 |
1.75 |
1.79 |
1.81 |
1.72 |
1.37 |
1.21 |
| ATP | 1.03 |
1.01 |
.75 |
1.04 |
1.32 |
1.90 |
1.14 |
| KLA/ATP | 1.84 |
1.73 |
1.49 |
2.07 |
2.01 |
1.83 |
1.68 |
|
| Jun | 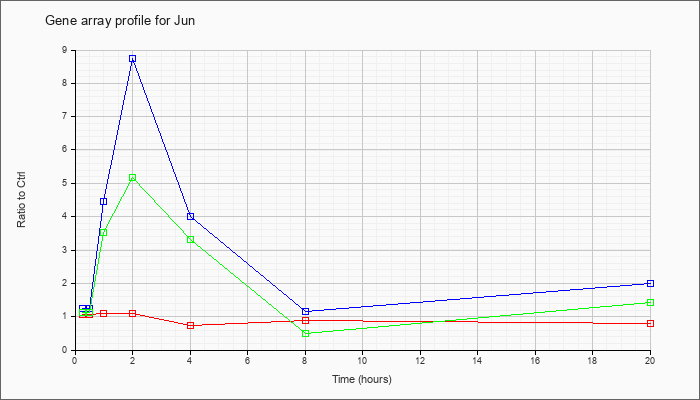 |
NM_010591 |
Jun oncogene (Jun), mRNA [NM_010591] |
KLA | 1.08 |
.96 |
1.09 |
1.09 |
.75 |
.88 |
.81 |
| ATP | 1.26 |
1.48 |
4.45 |
8.73 |
4.02 |
1.16 |
1.99 |
| KLA/ATP | 1.12 |
1.38 |
3.52 |
5.17 |
3.32 |
.49 |
1.42 |
|
| Junb | 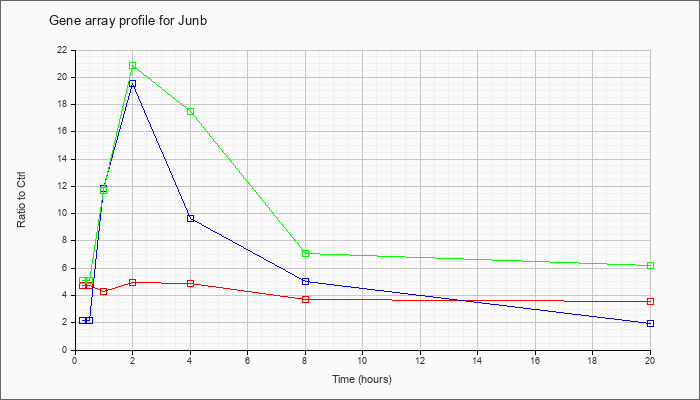 |
NM_008416 |
Jun-B oncogene (Junb), mRNA [NM_008416] |
KLA | 4.71 |
4.37 |
4.30 |
4.98 |
4.85 |
3.70 |
3.53 |
| ATP | 2.20 |
3.40 |
11.86 |
19.56 |
9.63 |
5.04 |
1.94 |
| KLA/ATP | 5.09 |
6.66 |
11.69 |
20.90 |
17.53 |
7.06 |
6.21 |
|
| Jund | 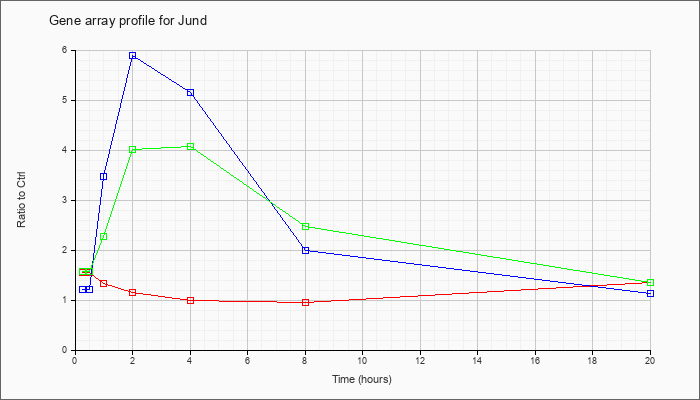 |
NM_010592 |
Jun proto-oncogene related gene d (Jund), mRNA [NM_010592] |
KLA | 1.56 |
1.42 |
1.33 |
1.15 |
1.00 |
.95 |
1.36 |
| ATP | 1.21 |
1.51 |
3.47 |
5.90 |
5.16 |
1.99 |
1.13 |
| KLA/ATP | 1.58 |
1.80 |
2.28 |
4.01 |
4.07 |
2.47 |
1.35 |
|
| Lck | 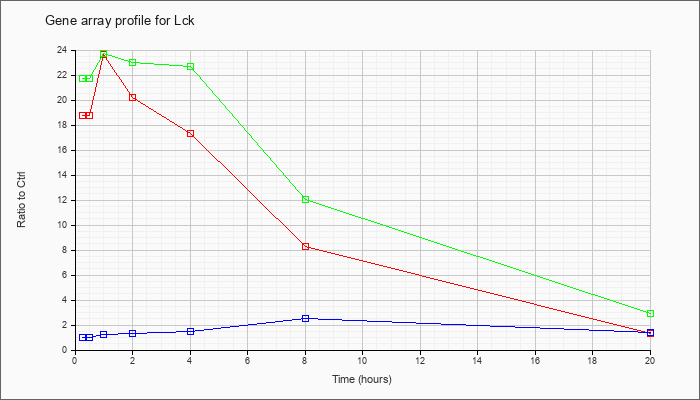 |
NM_010693 |
lymphocyte protein tyrosine kinase (Lck), mRNA [NM_010693] |
KLA | 18.74 |
17.97 |
23.64 |
20.20 |
17.34 |
8.29 |
1.29 |
| ATP | 1.00 |
1.04 |
1.28 |
1.30 |
1.48 |
2.55 |
1.44 |
| KLA/ATP | 21.76 |
24.22 |
23.73 |
23.04 |
22.66 |
12.06 |
2.90 |
|
| Lcp2 | 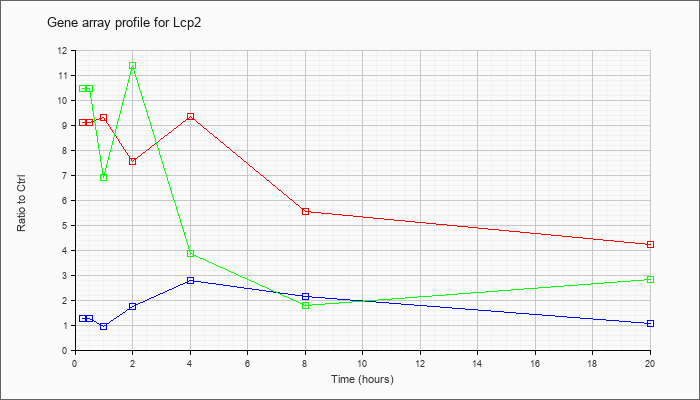 |
NM_010696 |
lymphocyte cytosolic protein 2 (Lcp2), mRNA [NM_010696] |
KLA | 9.10 |
9.37 |
9.29 |
7.53 |
9.33 |
5.54 |
4.23 |
| ATP | 1.25 |
1.38 |
.95 |
1.73 |
2.80 |
2.13 |
1.06 |
| KLA/ATP | 10.47 |
10.89 |
6.89 |
11.38 |
3.88 |
1.80 |
2.82 |
|
| Lilrb3 | 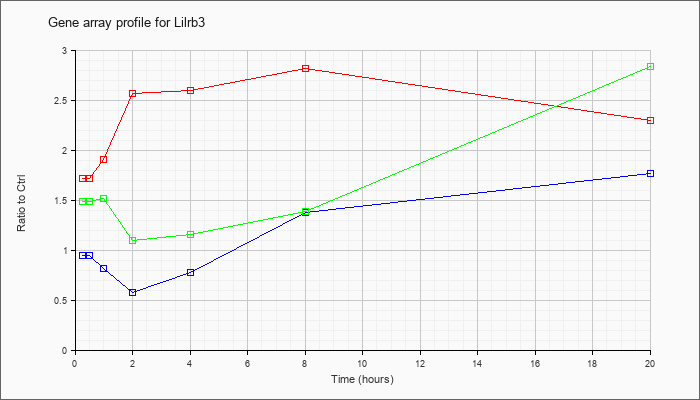 |
NM_011095 |
leukocyte immunoglobulin-like receptor, subfamily B (with TM and ITIM domains), member 3 (Lilrb3), mRNA [NM_011095] |
KLA | 1.72 |
1.76 |
1.91 |
2.57 |
2.60 |
2.82 |
2.30 |
| ATP | .95 |
.85 |
.82 |
.58 |
.78 |
1.38 |
1.77 |
| KLA/ATP | 1.49 |
1.42 |
1.52 |
1.10 |
1.16 |
1.39 |
2.84 |
|
| Map2k1 | 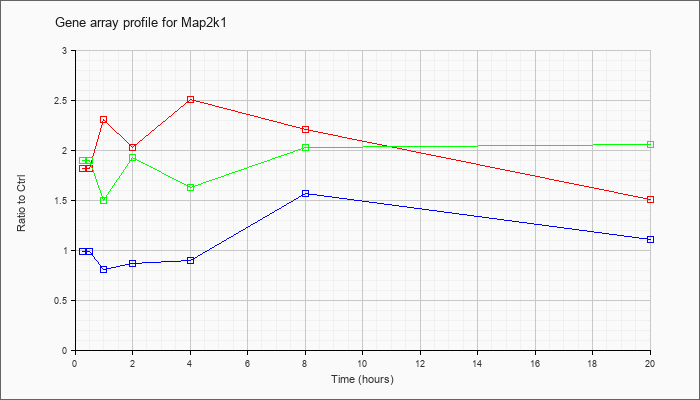 |
NM_008927 |
mitogen-activated protein kinase kinase 1 (Map2k1), mRNA [NM_008927] |
KLA | 1.82 |
2.00 |
2.31 |
2.03 |
2.51 |
2.20 |
1.51 |
| ATP | .99 |
1.14 |
.81 |
.87 |
.90 |
1.57 |
1.11 |
| KLA/ATP | 1.90 |
2.03 |
1.50 |
1.93 |
1.63 |
2.03 |
2.06 |
|
| Map2k6 | 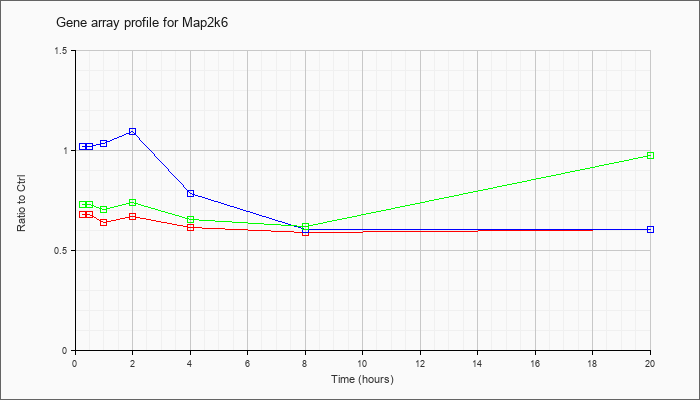 |
NM_011943 |
mitogen-activated protein kinase kinase 6 (Map2k6), mRNA [NM_011943] |
KLA | .68 |
.67 |
.64 |
.67 |
.62 |
.59 |
.61 |
| ATP | 1.02 |
1.12 |
1.04 |
1.10 |
.79 |
.61 |
.61 |
| KLA/ATP | .73 |
.69 |
.71 |
.74 |
.66 |
.62 |
.98 |
|
| Map2k7 | 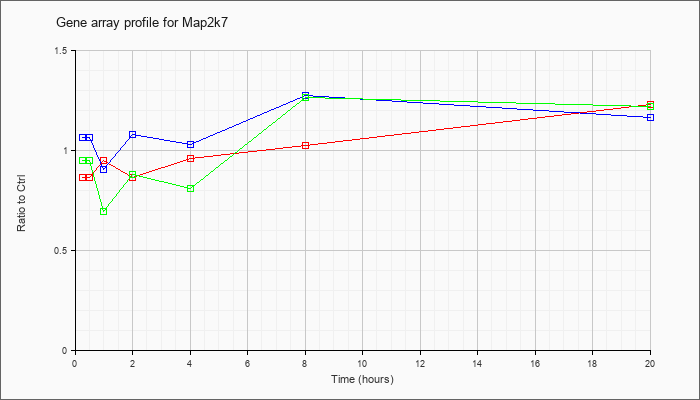 |
BC070467 |
mitogen activated protein kinase kinase 7, mRNA (cDNA clone MGC:96654 IMAGE:30615302), complete cds [BC070467] |
KLA | .77 |
.79 |
.88 |
.77 |
.97 |
.96 |
1.27 |
| ATP | 1.11 |
1.26 |
.93 |
1.19 |
1.13 |
1.45 |
1.16 |
| KLA/ATP | .89 |
.88 |
.59 |
.89 |
.86 |
1.32 |
1.28 |
|
| Map2k7 |  |
NM_001042557 |
mitogen-activated protein kinase kinase 7 (Map2k7), transcript variant 1, mRNA [NM_001042557] |
KLA | .96 |
.95 |
1.02 |
.96 |
.95 |
1.09 |
1.19 |
| ATP | 1.02 |
1.07 |
.88 |
.97 |
.93 |
1.10 |
1.17 |
| KLA/ATP | 1.01 |
.99 |
.80 |
.87 |
.76 |
1.21 |
1.16 |
|
| Map3k14 | 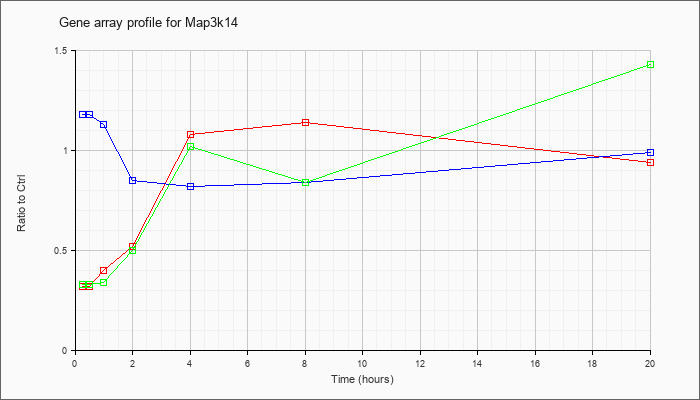 |
NM_016896 |
mitogen-activated protein kinase kinase kinase 14 (Map3k14), mRNA [NM_016896] |
KLA | .32 |
.30 |
.40 |
.52 |
1.08 |
1.14 |
.94 |
| ATP | 1.18 |
1.24 |
1.13 |
.85 |
.82 |
.84 |
.99 |
| KLA/ATP | .33 |
.30 |
.34 |
.50 |
1.02 |
.84 |
1.43 |
|
| Map3k7 | 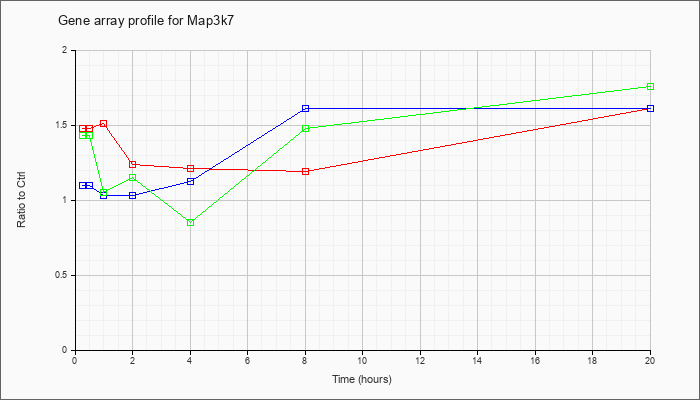 |
NM_172688 |
mitogen-activated protein kinase kinase kinase 7 (Map3k7), mRNA [NM_172688] |
KLA | 1.48 |
1.49 |
1.51 |
1.24 |
1.21 |
1.19 |
1.61 |
| ATP | 1.10 |
1.16 |
1.03 |
1.03 |
1.12 |
1.61 |
1.61 |
| KLA/ATP | 1.43 |
1.56 |
1.05 |
1.15 |
.85 |
1.48 |
1.76 |
|
Map3k7ip
1 | 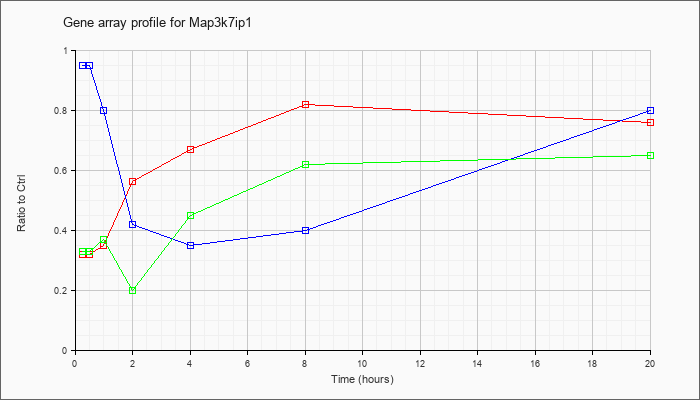 |
NM_025609 |
mitogen-activated protein kinase kinase kinase 7 interacting protein 1 (Map3k7ip1), mRNA [NM_025609] |
KLA | .32 |
.30 |
.35 |
.56 |
.67 |
.82 |
.76 |
| ATP | .95 |
.81 |
.80 |
.42 |
.35 |
.40 |
.80 |
| KLA/ATP | .33 |
.30 |
.37 |
.20 |
.45 |
.62 |
.65 |
|
Map3k7ip
2 | 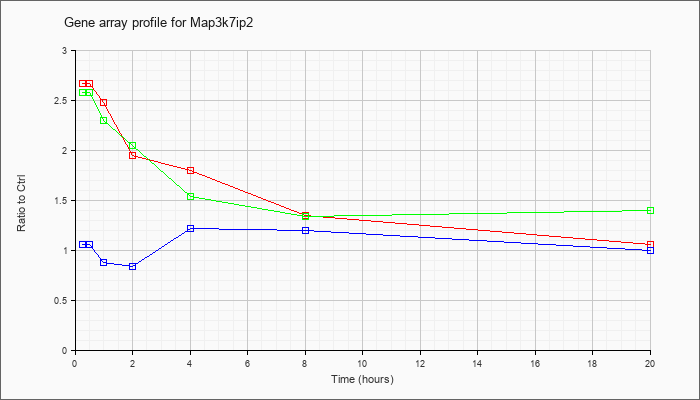 |
AK086230 |
15 days embryo head cDNA, RIKEN full-length enriched library, clone:D930015C19 product:inferred: Mus musculus, Similar to TAK1-binding protein 2; KIAA0733 protein, clone MGC:8010 IMAGE:3586132, mRNA, complete cds, full insert sequence. [A |
KLA | 1.14 |
1.02 |
1.07 |
1.03 |
1.13 |
1.02 |
.96 |
| ATP | 1.00 |
.88 |
.92 |
.90 |
1.06 |
.96 |
.94 |
| KLA/ATP | 1.05 |
1.00 |
.90 |
.98 |
1.02 |
.97 |
.97 |
|
Map3k7ip
2 |  |
NM_138667 |
mitogen-activated protein kinase kinase kinase 7 interacting protein 2 (Map3k7ip2), mRNA [NM_138667] |
KLA | 3.43 |
3.22 |
3.19 |
2.41 |
2.13 |
1.51 |
1.10 |
| ATP | 1.08 |
1.08 |
.86 |
.81 |
1.30 |
1.32 |
1.02 |
| KLA/ATP | 3.34 |
3.66 |
3.00 |
2.58 |
1.80 |
1.52 |
1.61 |
|
| Mapk1 | 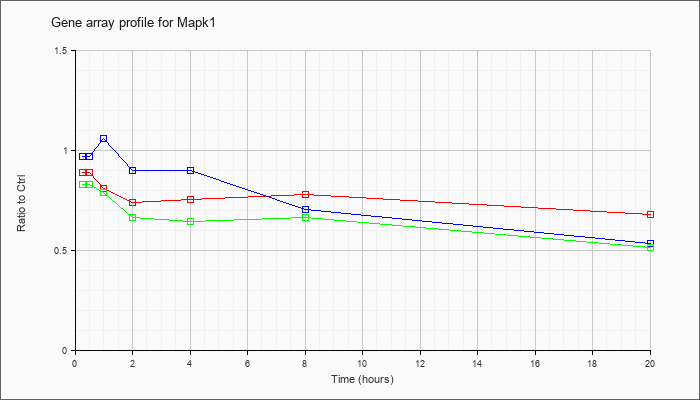 |
NM_001038663 |
mitogen-activated protein kinase 1 (Mapk1), transcript variant 2, mRNA [NM_001038663] |
KLA | .97 |
1.03 |
.93 |
.86 |
.79 |
1.01 |
1.04 |
| ATP | .96 |
1.18 |
1.01 |
1.13 |
.84 |
1.04 |
1.16 |
| KLA/ATP | .99 |
.99 |
.84 |
.87 |
.63 |
.96 |
1.26 |
|
| Mapk1 |  |
NM_011949 |
mitogen-activated protein kinase 1 (Mapk1), transcript variant 1, mRNA [NM_011949] |
KLA | .88 |
.80 |
.80 |
.73 |
.75 |
.76 |
.65 |
| ATP | .97 |
.96 |
1.06 |
.88 |
.90 |
.68 |
.48 |
| KLA/ATP | .82 |
.81 |
.78 |
.65 |
.64 |
.64 |
.45 |
|
| Mapk10 | 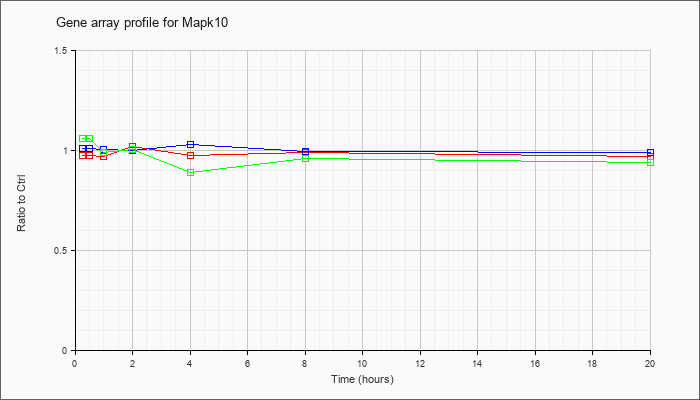 |
AK076990 |
adult male testis cDNA, RIKEN full-length enriched library, clone:4930597C01 product:mitogen activated protein kinase 10, full insert sequence. [AK076990] |
KLA | 1.03 |
1.05 |
.98 |
1.04 |
.94 |
.95 |
1.02 |
| ATP | 1.00 |
.99 |
1.06 |
.97 |
1.01 |
.96 |
.99 |
| KLA/ATP | 1.07 |
1.01 |
.95 |
1.05 |
.82 |
.95 |
.87 |
|
| Mapk10 |  |
NM_009158 |
mitogen-activated protein kinase 10 (Mapk10), transcript variant 1, mRNA [NM_009158] |
KLA | .95 |
1.00 |
.97 |
1.01 |
.99 |
1.01 |
.95 |
| ATP | 1.01 |
1.02 |
.98 |
1.01 |
1.04 |
1.01 |
.99 |
| KLA/ATP | 1.05 |
.99 |
1.01 |
.98 |
.92 |
.97 |
.98 |
|
| Mapk11 | 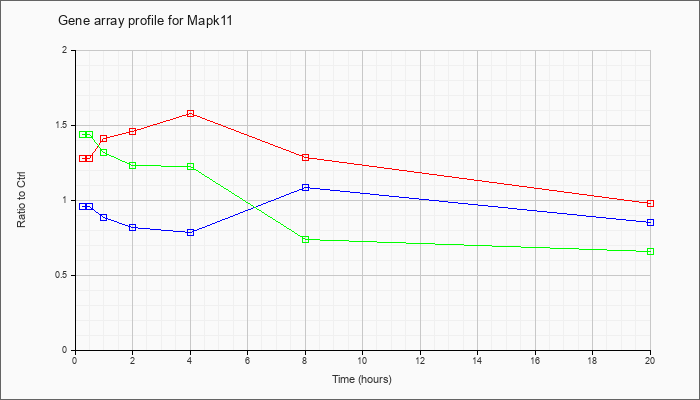 |
NM_011161 |
mitogen-activated protein kinase 11 (Mapk11), mRNA [NM_011161] |
KLA | 1.28 |
1.30 |
1.41 |
1.46 |
1.58 |
1.29 |
.98 |
| ATP | .96 |
.95 |
.89 |
.82 |
.79 |
1.09 |
.85 |
| KLA/ATP | 1.44 |
1.42 |
1.32 |
1.23 |
1.23 |
.74 |
.66 |
|
| Mapk12 | 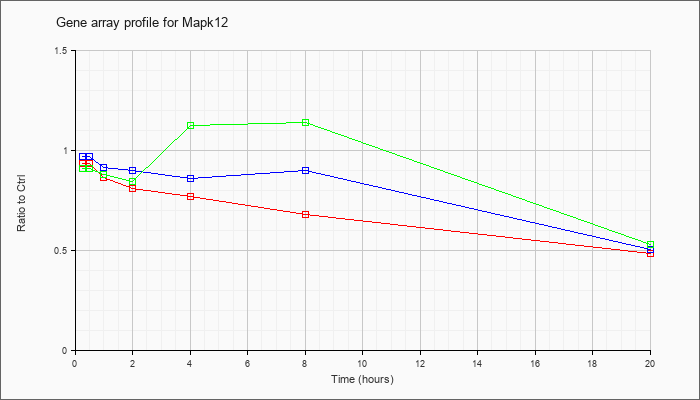 |
NM_013871 |
mitogen-activated protein kinase 12 (Mapk12), mRNA [NM_013871] |
KLA | .94 |
.86 |
.87 |
.81 |
.77 |
.68 |
.49 |
| ATP | .97 |
.83 |
.92 |
.90 |
.86 |
.90 |
.51 |
| KLA/ATP | .91 |
.89 |
.88 |
.85 |
1.12 |
1.14 |
.53 |
|
| Mapk13 | 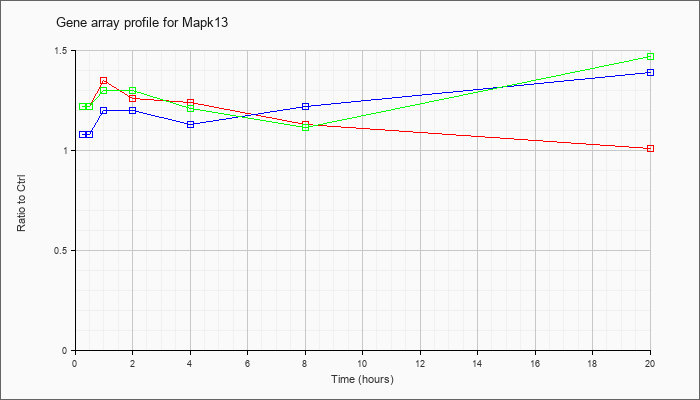 |
NM_011950 |
mitogen-activated protein kinase 13 (Mapk13), mRNA [NM_011950] |
KLA | 1.22 |
1.21 |
1.35 |
1.26 |
1.24 |
1.13 |
1.01 |
| ATP | 1.08 |
1.00 |
1.20 |
1.20 |
1.13 |
1.22 |
1.39 |
| KLA/ATP | 1.22 |
1.27 |
1.30 |
1.30 |
1.21 |
1.11 |
1.47 |
|
| Mapk14 | 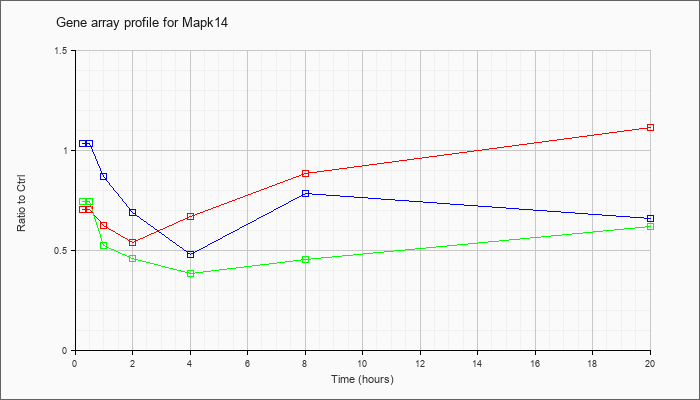 |
NM_011951 |
mitogen-activated protein kinase 14 (Mapk14), mRNA [NM_011951] |
KLA | .70 |
.70 |
.62 |
.54 |
.67 |
.88 |
1.11 |
| ATP | 1.03 |
.98 |
.87 |
.69 |
.48 |
.78 |
.66 |
| KLA/ATP | .74 |
.68 |
.52 |
.46 |
.38 |
.45 |
.62 |
|
| Mapk3 | 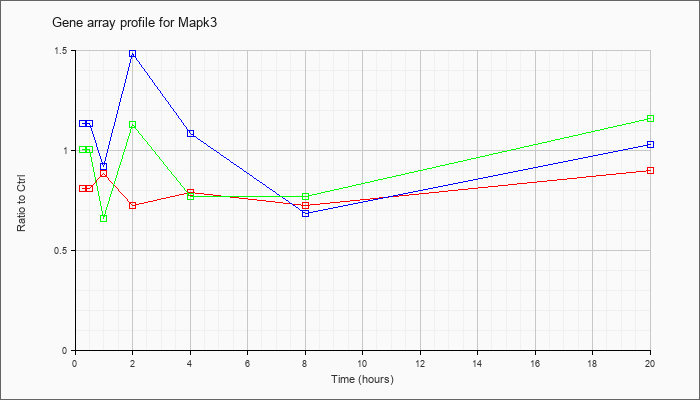 |
NM_011952 |
mitogen-activated protein kinase 3 (Mapk3), mRNA [NM_011952] |
KLA | .81 |
.85 |
.88 |
.72 |
.79 |
.73 |
.90 |
| ATP | 1.13 |
1.30 |
.92 |
1.48 |
1.08 |
.68 |
1.03 |
| KLA/ATP | 1.01 |
1.01 |
.66 |
1.13 |
.77 |
.77 |
1.16 |
|
| Mapk8 |  |
AK162915 |
adult male spinal cord cDNA, RIKEN full-length enriched library, clone:A330031O19 product:mitogen activated protein kinase 8, full insert sequence [AK162915] |
KLA | .94 |
.86 |
.80 |
.83 |
.77 |
.85 |
.83 |
| ATP | 1.08 |
1.12 |
.94 |
.80 |
.83 |
.71 |
.75 |
| KLA/ATP | .83 |
.81 |
.80 |
.74 |
.74 |
.75 |
.67 |
|
| Mapk8 |  |
AK163829 |
16 days embryo head cDNA, RIKEN full-length enriched library, clone:C130070A06 product:mitogen activated protein kinase 8, full insert sequence [AK163829] |
KLA | .65 |
.61 |
.66 |
.70 |
.90 |
1.02 |
1.03 |
| ATP | .99 |
.94 |
1.68 |
1.44 |
1.75 |
1.84 |
1.10 |
| KLA/ATP | .63 |
.67 |
1.11 |
1.00 |
2.01 |
1.68 |
1.15 |
|
| Mapk8 |  |
NM_016700 |
mitogen-activated protein kinase 8 (Mapk8), mRNA [NM_016700] |
KLA | .96 |
1.00 |
.94 |
.96 |
.94 |
.99 |
1.04 |
| ATP | 1.01 |
1.05 |
1.01 |
.99 |
1.03 |
.98 |
1.02 |
| KLA/ATP | .96 |
.97 |
.96 |
.97 |
.93 |
.94 |
.96 |
|
| Mapk9 | 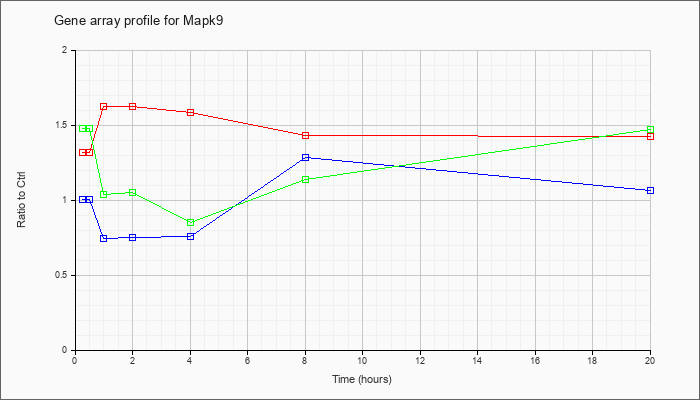 |
NM_016961 |
mitogen-activated protein kinase 9 (Mapk9), transcript variant 2, mRNA [NM_016961] |
KLA | 1.28 |
1.37 |
1.59 |
1.60 |
1.61 |
1.43 |
1.38 |
| ATP | 1.03 |
.99 |
.74 |
.71 |
.79 |
1.20 |
1.01 |
| KLA/ATP | 1.53 |
1.30 |
1.07 |
.99 |
.83 |
1.10 |
1.40 |
|
| Mapk9 |  |
NM_207692 |
mitogen-activated protein kinase 9 (Mapk9), transcript variant 1, mRNA [NM_207692] |
KLA | 1.39 |
1.53 |
1.69 |
1.69 |
1.55 |
1.44 |
1.51 |
| ATP | .95 |
.92 |
.75 |
.85 |
.70 |
1.46 |
1.17 |
| KLA/ATP | 1.37 |
1.35 |
.99 |
1.18 |
.91 |
1.22 |
1.62 |
|
| Mitf | 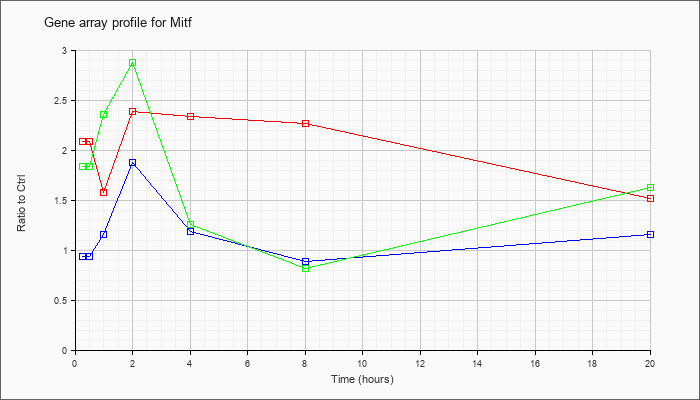 |
NM_008601 |
microphthalmia-associated transcription factor (Mitf), transcript variant 2, mRNA [NM_008601] |
KLA | 2.09 |
2.20 |
1.58 |
2.38 |
2.33 |
2.27 |
1.52 |
| ATP | .93 |
.99 |
1.15 |
1.87 |
1.18 |
.89 |
1.16 |
| KLA/ATP | 1.83 |
2.28 |
2.35 |
2.87 |
1.26 |
.81 |
1.63 |
|
Mm.15898
1 | 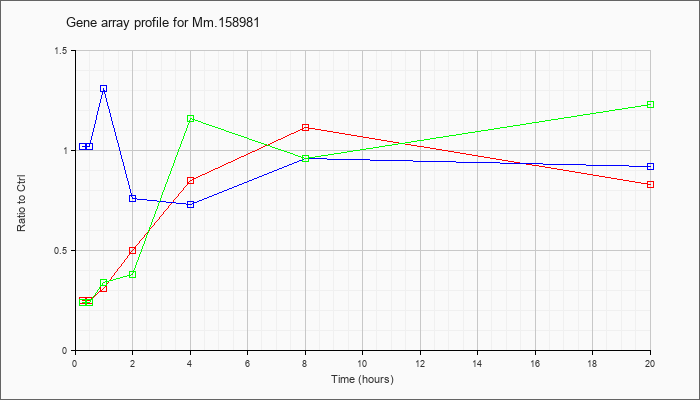 |
142388182 |
Unknown |
KLA | .25 |
.22 |
.31 |
.50 |
.85 |
1.11 |
.83 |
| ATP | 1.02 |
.94 |
1.31 |
.76 |
.73 |
.96 |
.92 |
| KLA/ATP | .24 |
.22 |
.34 |
.38 |
1.16 |
.96 |
1.23 |
|
| Mm.1741 | 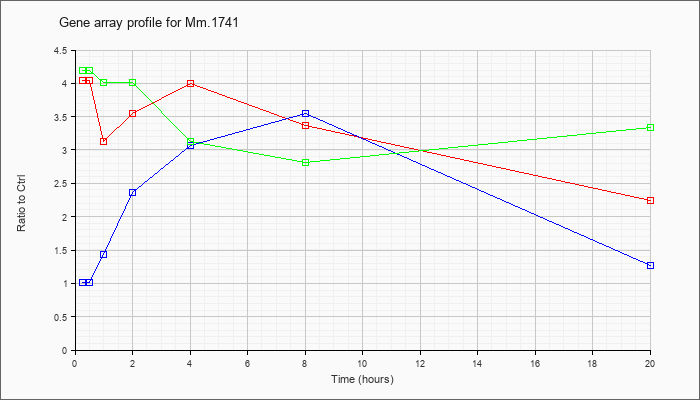 |
31982052 |
Unknown |
KLA | 4.05 |
3.90 |
3.12 |
3.55 |
4.00 |
3.37 |
2.25 |
| ATP | 1.01 |
1.01 |
1.44 |
2.36 |
3.07 |
3.54 |
1.27 |
| KLA/ATP | 4.19 |
3.74 |
4.02 |
4.02 |
3.13 |
2.82 |
3.33 |
|
Mm.21312
8 | 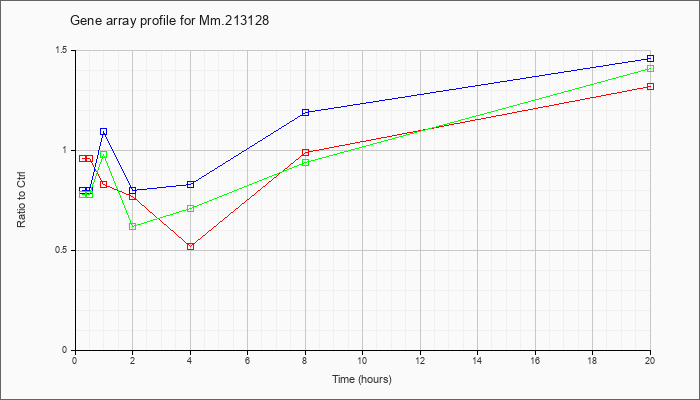 |
141801344 |
Unknown |
KLA | .96 |
.95 |
.83 |
.77 |
.52 |
.99 |
1.32 |
| ATP | .80 |
.82 |
1.09 |
.80 |
.83 |
1.19 |
1.46 |
| KLA/ATP | .78 |
.92 |
.98 |
.62 |
.71 |
.94 |
1.41 |
|
Mm.24833
5 |  |
110350004 |
Unknown |
KLA | .83 |
.79 |
.63 |
.93 |
.75 |
1.10 |
1.03 |
| ATP | 10.10 |
12.76 |
17.13 |
18.61 |
13.01 |
3.37 |
1.36 |
| KLA/ATP | 3.05 |
5.96 |
4.94 |
8.73 |
5.80 |
4.08 |
1.91 |
|
Mm.24936
4 | 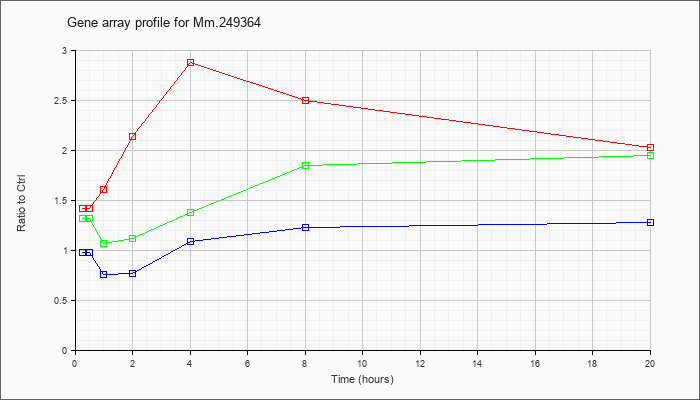 |
133892283 |
Unknown |
KLA | 1.42 |
1.41 |
1.61 |
2.14 |
2.88 |
2.49 |
2.03 |
| ATP | .98 |
.98 |
.76 |
.77 |
1.09 |
1.23 |
1.28 |
| KLA/ATP | 1.32 |
1.32 |
1.07 |
1.12 |
1.38 |
1.85 |
1.95 |
|
Mm.25381
9 | 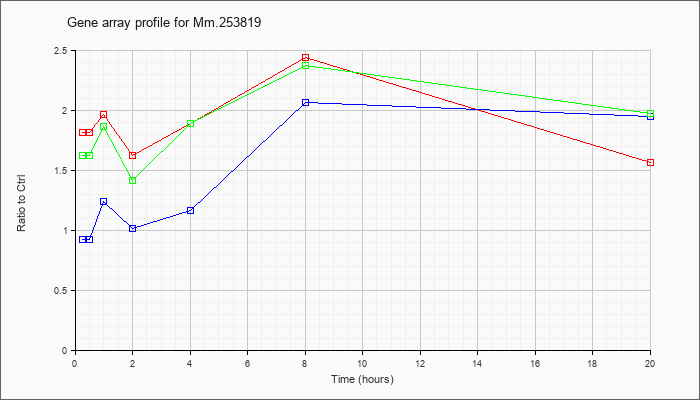 |
118130676 |
Unknown |
KLA | 1.81 |
1.88 |
1.96 |
1.62 |
1.89 |
2.44 |
1.56 |
| ATP | .92 |
.91 |
1.24 |
1.01 |
1.16 |
2.06 |
1.95 |
| KLA/ATP | 1.62 |
1.86 |
1.86 |
1.41 |
1.89 |
2.37 |
1.97 |
|
Mm.25806
1 |  |
156766060 |
Unknown |
KLA | 1.08 |
.96 |
1.05 |
.97 |
1.01 |
1.05 |
.88 |
| ATP | .94 |
1.00 |
.99 |
.97 |
.99 |
1.02 |
1.02 |
| KLA/ATP | 1.00 |
1.01 |
.95 |
1.00 |
.95 |
.98 |
1.01 |
|
Mm.27970
| 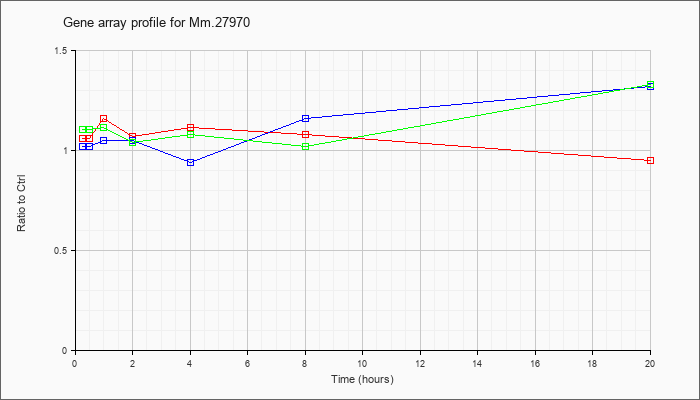 |
13994135 |
Unknown |
KLA | 1.06 |
1.11 |
1.16 |
1.07 |
1.11 |
1.08 |
.95 |
| ATP | 1.02 |
1.09 |
1.05 |
1.05 |
.94 |
1.16 |
1.32 |
| KLA/ATP | 1.10 |
1.19 |
1.11 |
1.04 |
1.08 |
1.02 |
1.33 |
|
Mm.32653
1 |  |
6679332 |
Unknown |
KLA | 1.26 |
1.23 |
1.50 |
1.57 |
1.97 |
2.07 |
1.91 |
| ATP | .99 |
1.06 |
.80 |
.92 |
.91 |
1.19 |
2.08 |
| KLA/ATP | 1.28 |
1.26 |
1.00 |
1.26 |
.89 |
1.17 |
2.90 |
|
Mm.33138
9 | 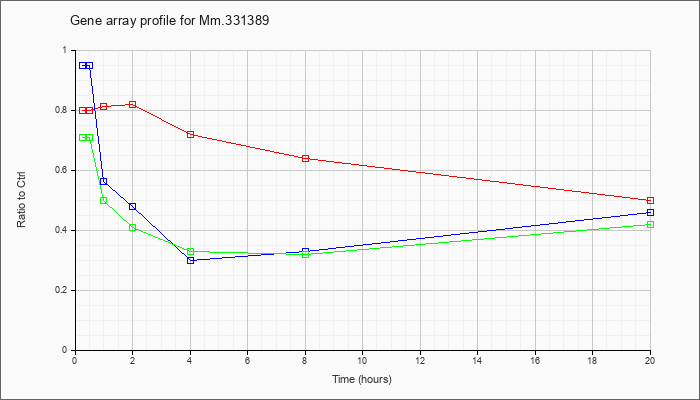 |
158508469 |
Unknown |
KLA | .80 |
.79 |
.81 |
.82 |
.72 |
.64 |
.50 |
| ATP | .95 |
.85 |
.56 |
.48 |
.30 |
.33 |
.46 |
| KLA/ATP | .71 |
.64 |
.50 |
.41 |
.33 |
.32 |
.42 |
|
Mm.42529
6 | 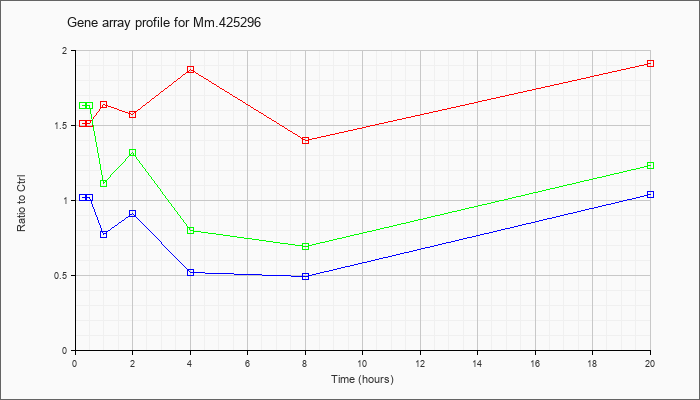 |
170172552 |
Unknown |
KLA | 1.51 |
1.59 |
1.64 |
1.57 |
1.87 |
1.40 |
1.91 |
| ATP | 1.02 |
1.04 |
.77 |
.91 |
.52 |
.49 |
1.04 |
| KLA/ATP | 1.63 |
1.53 |
1.11 |
1.32 |
.80 |
.69 |
1.23 |
|
Mm.45876
9 |  |
6755063 |
Unknown |
KLA | .99 |
.91 |
.94 |
.94 |
.97 |
1.03 |
1.32 |
| ATP | .95 |
.99 |
1.10 |
1.09 |
1.22 |
1.16 |
1.94 |
| KLA/ATP | .91 |
.90 |
.86 |
.93 |
.89 |
.68 |
1.80 |
|
Mm.46002
4 | 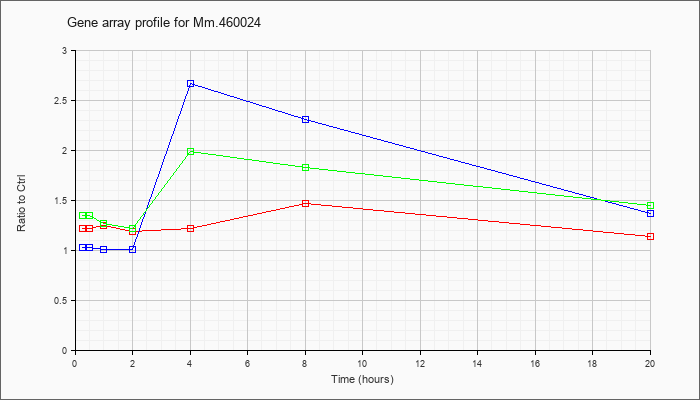 |
38348245 |
Unknown |
KLA | 1.22 |
1.25 |
1.25 |
1.19 |
1.22 |
1.47 |
1.14 |
| ATP | 1.03 |
.94 |
1.01 |
1.01 |
2.67 |
2.31 |
1.37 |
| KLA/ATP | 1.35 |
1.19 |
1.27 |
1.22 |
1.99 |
1.83 |
1.45 |
|
| Ncf1 |  |
NM_010876 |
neutrophil cytosolic factor 1 (Ncf1), mRNA [NM_010876] |
KLA | 1.62 |
1.71 |
1.81 |
1.70 |
1.99 |
1.64 |
2.07 |
| ATP | 1.02 |
1.06 |
.78 |
1.02 |
.64 |
.56 |
1.08 |
| KLA/ATP | 1.87 |
1.81 |
1.22 |
1.73 |
1.00 |
.80 |
1.49 |
|
| Ncf2 | 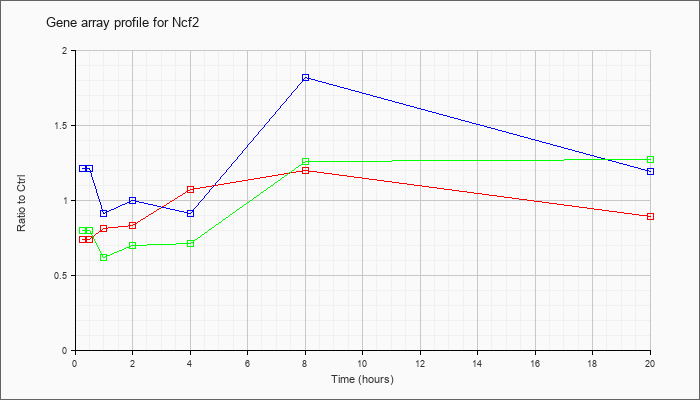 |
NM_010877 |
neutrophil cytosolic factor 2 (Ncf2), mRNA [NM_010877] |
KLA | .74 |
.69 |
.81 |
.83 |
1.07 |
1.20 |
.89 |
| ATP | 1.21 |
1.13 |
.91 |
1.00 |
.91 |
1.82 |
1.19 |
| KLA/ATP | .80 |
.74 |
.62 |
.70 |
.71 |
1.26 |
1.27 |
|
| Ncf4 | 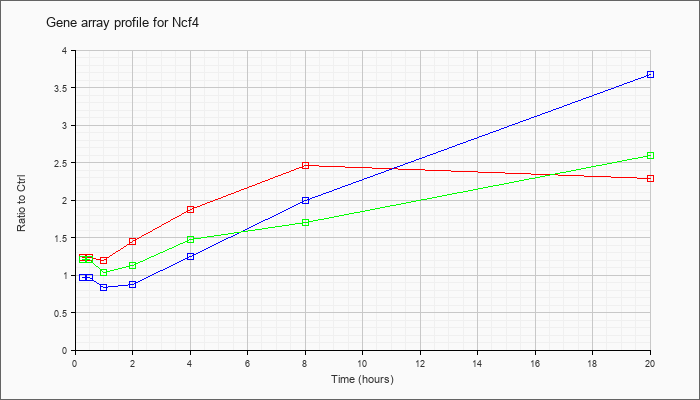 |
NM_008677 |
neutrophil cytosolic factor 4 (Ncf4), mRNA [NM_008677] |
KLA | 1.23 |
1.30 |
1.19 |
1.45 |
1.88 |
2.46 |
2.29 |
| ATP | .97 |
.83 |
.83 |
.87 |
1.25 |
1.99 |
3.67 |
| KLA/ATP | 1.21 |
1.11 |
1.04 |
1.13 |
1.48 |
1.70 |
2.60 |
|
| Nfatc1 |  |
NM_016791 |
nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 1 (Nfatc1), transcript variant 1, mRNA [NM_016791] |
KLA | .71 |
.66 |
.68 |
.62 |
.63 |
.67 |
.81 |
| ATP | 1.12 |
1.29 |
1.15 |
1.21 |
.83 |
.63 |
1.25 |
| KLA/ATP | .78 |
.79 |
.65 |
.74 |
.59 |
.45 |
.92 |
|
| Nfatc1 |  |
NM_198429 |
nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 1 (Nfatc1), transcript variant 2, mRNA [NM_198429] |
KLA | .49 |
.41 |
.50 |
.43 |
.50 |
.78 |
.69 |
| ATP | 1.05 |
.81 |
.84 |
.54 |
.60 |
.38 |
.69 |
| KLA/ATP | .46 |
.37 |
.45 |
.32 |
.62 |
.39 |
.61 |
|
| Nfkb1 | 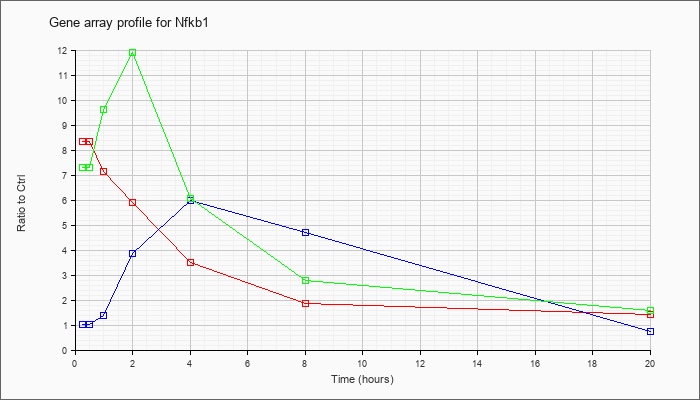 |
BC050841 |
nuclear factor of kappa light chain gene enhancer in B-cells 1, p105, mRNA (cDNA clone IMAGE:6392793), complete cds. [BC050841] |
KLA | 5.81 |
4.96 |
4.77 |
3.21 |
3.37 |
2.00 |
1.71 |
| ATP | 1.39 |
1.52 |
3.25 |
12.63 |
13.64 |
4.80 |
1.08 |
| KLA/ATP | 7.21 |
8.69 |
19.53 |
23.93 |
11.70 |
2.85 |
2.01 |
|
| Nfkb1 |  |
M57999 |
gb|Mouse transcription factor NF-kappa-B DNA binding subunit mRNA, complete cds. [M57999] |
KLA | 8.55 |
7.87 |
7.35 |
6.13 |
3.51 |
1.85 |
1.38 |
| ATP | .98 |
.96 |
1.21 |
3.06 |
5.29 |
4.67 |
.69 |
| KLA/ATP | 7.32 |
7.20 |
8.71 |
10.82 |
5.55 |
2.77 |
1.54 |
|
| Nfkb2 |  |
NM_019408 |
nuclear factor of kappa light polypeptide gene enhancer in B-cells 2, p49/p100 (Nfkb2), mRNA [NM_019408] |
KLA | 7.31 |
7.92 |
7.95 |
6.42 |
6.41 |
4.42 |
2.81 |
| ATP | .99 |
.98 |
.72 |
1.48 |
2.80 |
5.31 |
1.44 |
| KLA/ATP | 8.25 |
7.93 |
6.51 |
7.68 |
5.08 |
4.38 |
4.86 |
|
| Nfkbia | 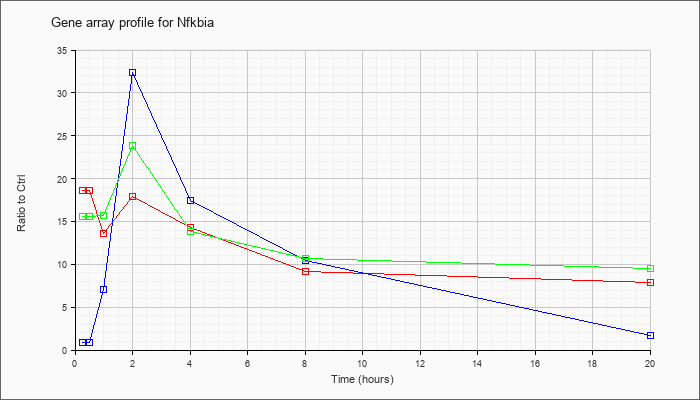 |
NM_010907 |
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha (Nfkbia), mRNA [NM_010907] |
KLA | 18.65 |
18.46 |
13.63 |
17.88 |
14.25 |
9.17 |
7.92 |
| ATP | .91 |
1.09 |
7.11 |
32.39 |
17.44 |
10.49 |
1.72 |
| KLA/ATP | 15.57 |
17.66 |
15.73 |
23.91 |
13.83 |
10.73 |
9.48 |
|
| Nox1 | 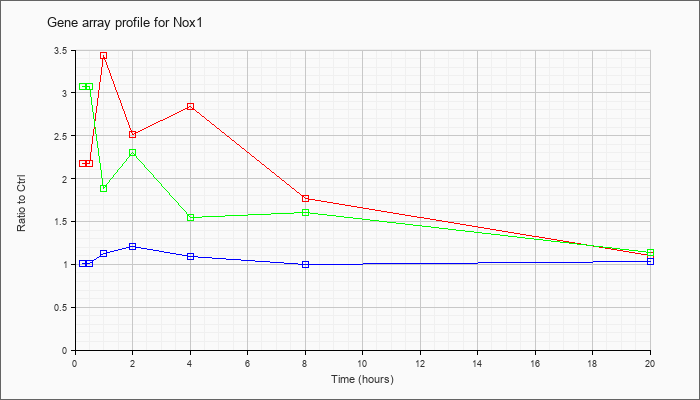 |
NM_172203 |
NADPH oxidase 1 (Nox1), mRNA [NM_172203] |
KLA | 2.18 |
2.93 |
3.44 |
2.51 |
2.84 |
1.77 |
1.10 |
| ATP | 1.01 |
.99 |
1.13 |
1.21 |
1.09 |
1.00 |
1.03 |
| KLA/ATP | 3.07 |
3.07 |
1.88 |
2.30 |
1.55 |
1.60 |
1.14 |
|
| Nox3 | 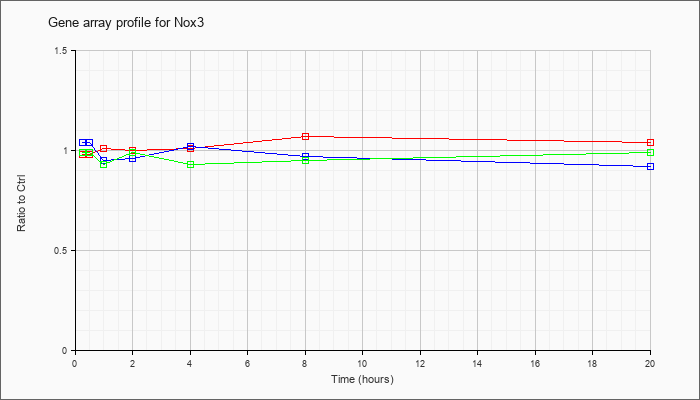 |
NM_198958 |
NADPH oxidase 3 (Nox3), mRNA [NM_198958] |
KLA | .98 |
.99 |
1.01 |
1.00 |
1.01 |
1.07 |
1.04 |
| ATP | 1.04 |
.95 |
.95 |
.96 |
1.02 |
.97 |
.92 |
| KLA/ATP | .99 |
1.03 |
.93 |
.99 |
.93 |
.95 |
.99 |
|
| Oscar | 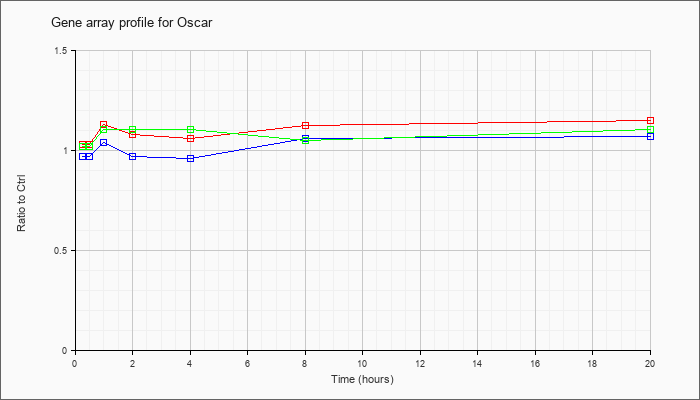 |
NM_175632 |
osteoclast associated receptor (Oscar), mRNA [NM_175632] |
KLA | 1.03 |
1.03 |
1.13 |
1.08 |
1.06 |
1.12 |
1.15 |
| ATP | .97 |
.98 |
1.04 |
.97 |
.96 |
1.06 |
1.07 |
| KLA/ATP | 1.02 |
1.08 |
1.10 |
1.10 |
1.10 |
1.05 |
1.10 |
|
| Pik3ca | 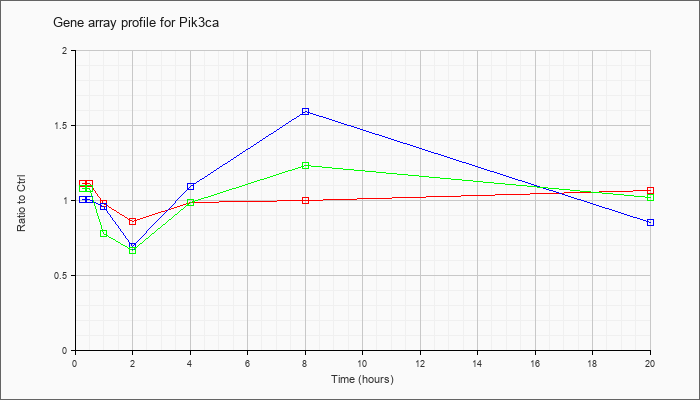 |
ENSMUST00000108243 |
ens|Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha isoform (EC 2.7.1.153) (PI3-kinase p110 subunit alpha) (PtdIns-3- kinase p110) (PI3K). [Source:Uniprot/SWISSPROT;Acc:P42337] [ENSMUST00000108243] |
KLA | 1.07 |
1.01 |
.91 |
.90 |
.98 |
.96 |
.86 |
| ATP | 1.03 |
.87 |
1.12 |
.63 |
1.22 |
1.86 |
.60 |
| KLA/ATP | 1.05 |
.82 |
.91 |
.61 |
1.17 |
1.26 |
.63 |
|
| Pik3ca |  |
NM_008839 |
phosphatidylinositol 3-kinase, catalytic, alpha polypeptide (Pik3ca), mRNA [NM_008839] |
KLA | 1.15 |
1.11 |
1.05 |
.82 |
.99 |
1.03 |
1.27 |
| ATP | .98 |
1.00 |
.79 |
.75 |
.96 |
1.32 |
1.10 |
| KLA/ATP | 1.10 |
.98 |
.64 |
.72 |
.80 |
1.20 |
1.41 |
|
| Pik3cb | 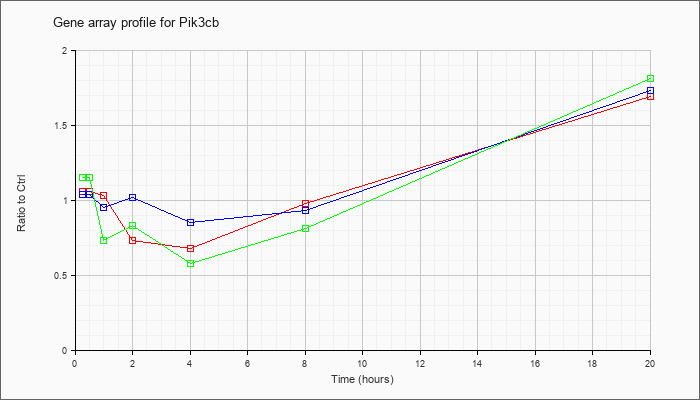 |
NM_029094 |
phosphatidylinositol 3-kinase, catalytic, beta polypeptide (Pik3cb), mRNA [NM_029094] |
KLA | 1.06 |
1.06 |
1.03 |
.73 |
.68 |
.98 |
1.69 |
| ATP | 1.04 |
1.21 |
.95 |
1.02 |
.85 |
.93 |
1.73 |
| KLA/ATP | 1.15 |
1.27 |
.73 |
.83 |
.58 |
.81 |
1.81 |
|
| Pik3cd | 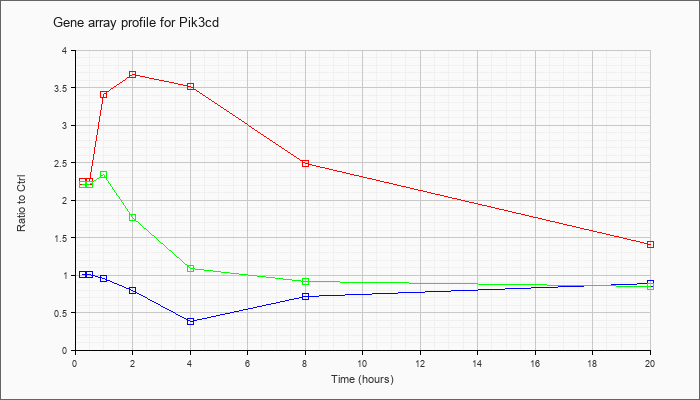 |
NM_008840 |
phosphatidylinositol 3-kinase catalytic delta polypeptide (Pik3cd), transcript variant 1, mRNA [NM_008840] |
KLA | 2.28 |
2.42 |
3.10 |
4.17 |
3.35 |
2.31 |
1.34 |
| ATP | 1.02 |
.91 |
.99 |
.63 |
.30 |
.67 |
.80 |
| KLA/ATP | 2.14 |
2.23 |
2.70 |
1.51 |
1.17 |
.85 |
.72 |
|
| Pik3cd |  |
U86587 |
phosphatidylinositol 3-kinase catalytic subunit p110 delta mRNA, complete cds. [U86587] |
KLA | 2.21 |
2.24 |
3.71 |
3.17 |
3.67 |
2.66 |
1.48 |
| ATP | 1.00 |
1.03 |
.93 |
.96 |
.46 |
.75 |
.97 |
| KLA/ATP | 2.27 |
2.32 |
1.97 |
2.02 |
1.01 |
.99 |
.98 |
|
| Pik3cg | 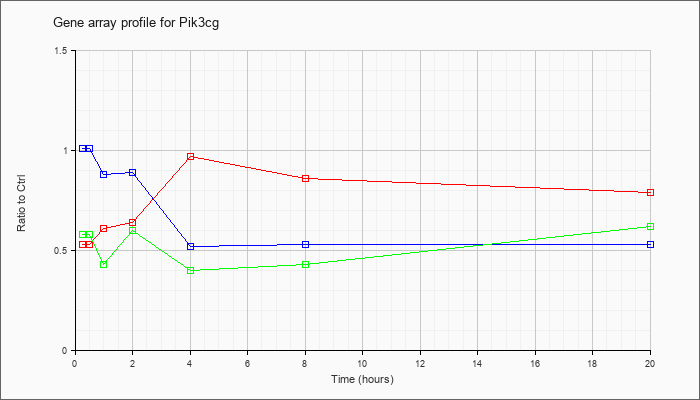 |
NM_020272 |
phosphoinositide-3-kinase, catalytic, gamma polypeptide (Pik3cg), mRNA [NM_020272] |
KLA | .53 |
.53 |
.61 |
.64 |
.97 |
.86 |
.79 |
| ATP | 1.01 |
1.13 |
.88 |
.89 |
.52 |
.53 |
.53 |
| KLA/ATP | .58 |
.53 |
.43 |
.60 |
.40 |
.43 |
.62 |
|
| Pik3r1 | 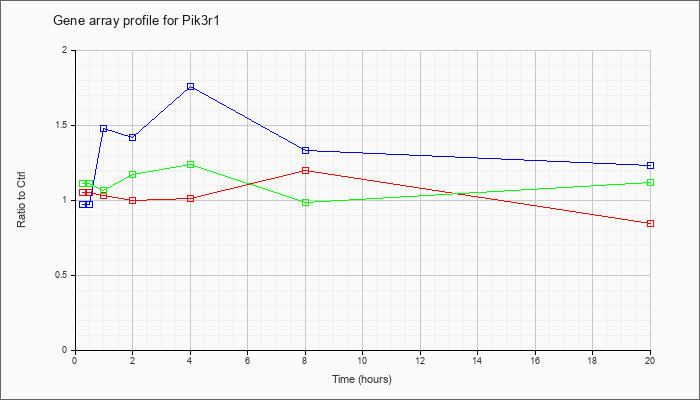 |
NM_001077495 |
phosphatidylinositol 3-kinase, regulatory subunit, polypeptide 1 (p85 alpha) (Pik3r1), transcript variant 2, mRNA [NM_001077495] |
KLA | 1.05 |
1.07 |
1.03 |
1.00 |
1.01 |
1.20 |
.85 |
| ATP | .97 |
1.02 |
1.48 |
1.42 |
1.76 |
1.33 |
1.23 |
| KLA/ATP | 1.11 |
1.12 |
1.06 |
1.17 |
1.24 |
.99 |
1.12 |
|
| Pik3r2 | 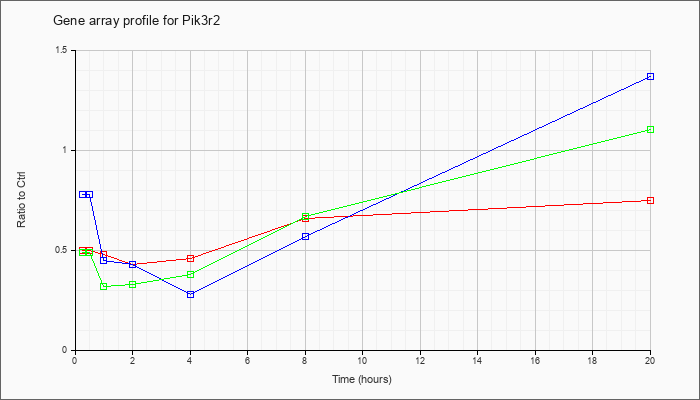 |
NM_008841 |
phosphatidylinositol 3-kinase, regulatory subunit, polypeptide 2 (p85 beta) (Pik3r2), mRNA [NM_008841] |
KLA | .50 |
.50 |
.48 |
.43 |
.46 |
.66 |
.75 |
| ATP | .78 |
.68 |
.45 |
.43 |
.28 |
.57 |
1.37 |
| KLA/ATP | .49 |
.38 |
.32 |
.33 |
.38 |
.67 |
1.10 |
|
| Pik3r3 | 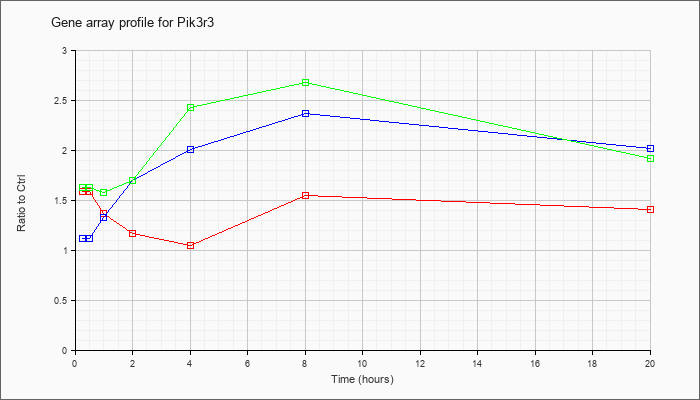 |
NM_181585 |
phosphatidylinositol 3 kinase, regulatory subunit, polypeptide 3 (p55) (Pik3r3), mRNA [NM_181585] |
KLA | 1.59 |
1.49 |
1.37 |
1.17 |
1.05 |
1.55 |
1.41 |
| ATP | 1.12 |
1.00 |
1.33 |
1.70 |
2.01 |
2.37 |
2.02 |
| KLA/ATP | 1.63 |
1.54 |
1.58 |
1.70 |
2.43 |
2.68 |
1.92 |
|
| Pik3r5 | 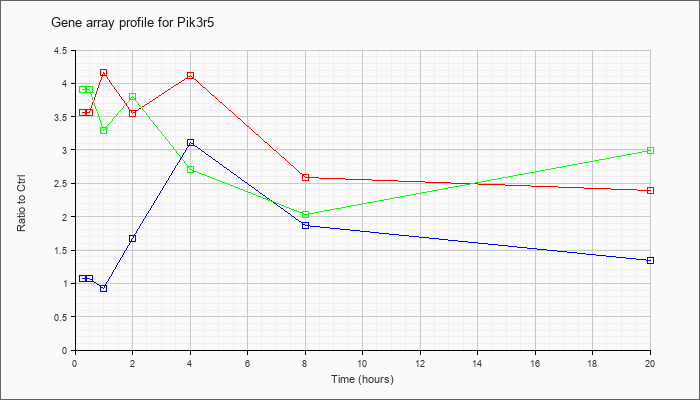 |
NM_177320 |
phosphoinositide-3-kinase, regulatory subunit 5, p101 (Pik3r5), mRNA [NM_177320] |
KLA | 3.57 |
3.58 |
4.16 |
3.54 |
4.11 |
2.59 |
2.39 |
| ATP | 1.08 |
1.08 |
.93 |
1.67 |
3.11 |
1.88 |
1.34 |
| KLA/ATP | 3.91 |
3.70 |
3.30 |
3.80 |
2.70 |
2.04 |
3.00 |
|
| Pira2 | 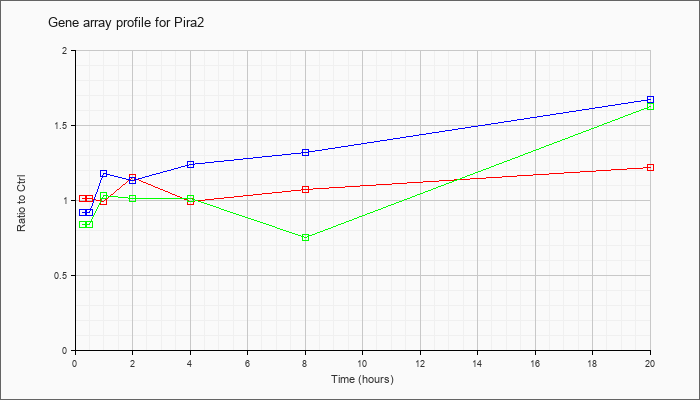 |
NM_011089 |
paired-Ig-like receptor A2 (Pira2), mRNA [NM_011089] |
KLA | 1.01 |
.97 |
.99 |
1.15 |
.99 |
1.07 |
1.22 |
| ATP | .92 |
.83 |
1.18 |
1.13 |
1.24 |
1.32 |
1.67 |
| KLA/ATP | .84 |
.85 |
1.03 |
1.01 |
1.01 |
.75 |
1.62 |
|
| Pira3 | 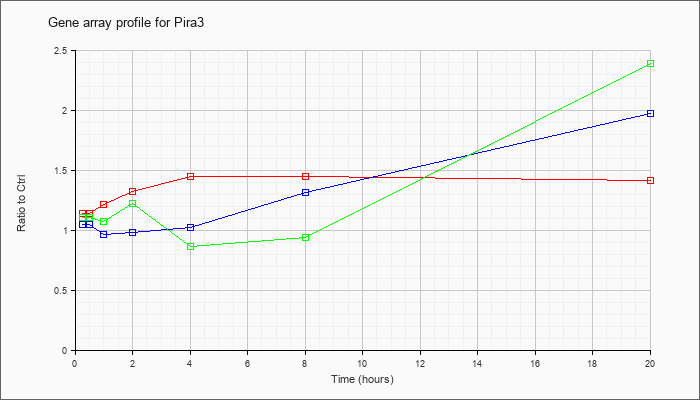 |
NM_011090 |
paired-Ig-like receptor A3 (Pira3), mRNA [NM_011090] |
KLA | 1.14 |
1.10 |
1.21 |
1.32 |
1.45 |
1.45 |
1.42 |
| ATP | 1.05 |
1.11 |
.96 |
.98 |
1.02 |
1.31 |
1.98 |
| KLA/ATP | 1.11 |
1.13 |
1.08 |
1.22 |
.87 |
.94 |
2.39 |
|
| Pira6 | 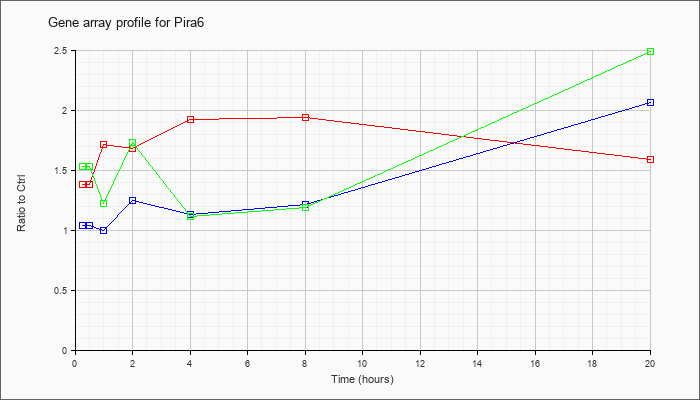 |
NM_011093 |
paired-Ig-like receptor A6 (Pira6), transcript variant 2, mRNA [NM_011093] |
KLA | 1.38 |
1.43 |
1.71 |
1.68 |
1.92 |
1.94 |
1.59 |
| ATP | 1.04 |
1.09 |
1.00 |
1.25 |
1.13 |
1.21 |
2.06 |
| KLA/ATP | 1.53 |
1.45 |
1.22 |
1.73 |
1.11 |
1.19 |
2.49 |
|
| Plcg2 | 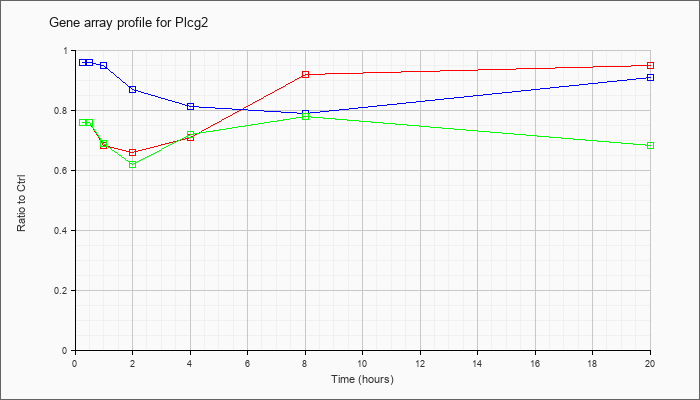 |
NM_172285 |
phospholipase C, gamma 2 (Plcg2), mRNA [NM_172285] |
KLA | .76 |
.74 |
.68 |
.66 |
.71 |
.92 |
.95 |
| ATP | .96 |
.90 |
.95 |
.87 |
.81 |
.79 |
.91 |
| KLA/ATP | .76 |
.71 |
.69 |
.62 |
.72 |
.78 |
.68 |
|
| Pparg | 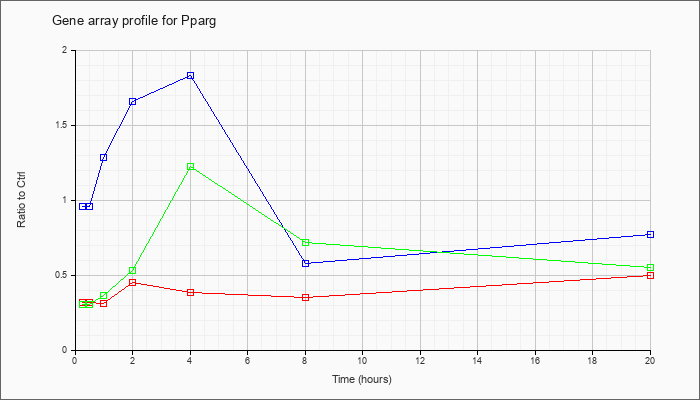 |
NM_011146 |
peroxisome proliferator activated receptor gamma (Pparg), mRNA [NM_011146] |
KLA | .32 |
.28 |
.31 |
.45 |
.38 |
.35 |
.49 |
| ATP | .96 |
1.01 |
1.28 |
1.65 |
1.83 |
.57 |
.77 |
| KLA/ATP | .30 |
.29 |
.36 |
.53 |
1.23 |
.72 |
.55 |
|
| Ppp3ca | 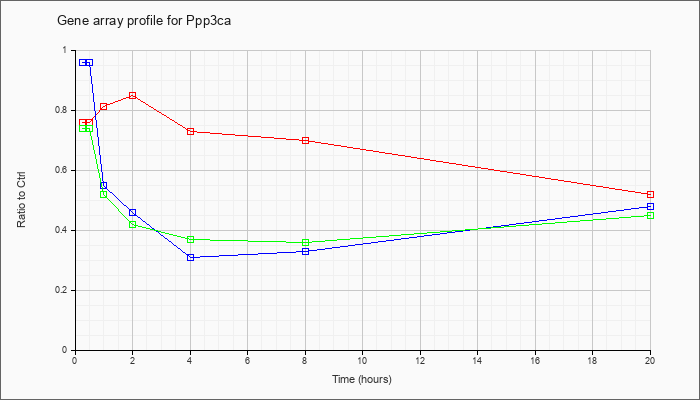 |
NM_008913 |
protein phosphatase 3, catalytic subunit, alpha isoform (Ppp3ca), mRNA [NM_008913] |
KLA | .76 |
.85 |
.81 |
.85 |
.73 |
.70 |
.52 |
| ATP | .96 |
.78 |
.55 |
.46 |
.31 |
.33 |
.48 |
| KLA/ATP | .74 |
.64 |
.52 |
.42 |
.37 |
.36 |
.45 |
|
| Ppp3cb | 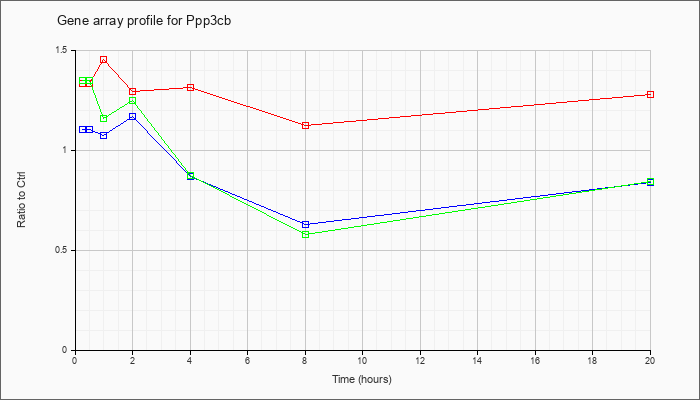 |
AK135203 |
12 days embryo male wolffian duct includes surrounding region cDNA, RIKEN full-length enriched library, clone:6720421K10 product:protein phosphatase 3, catalytic subunit, beta isoform, full insert sequence. [AK135203] |
KLA | 1.74 |
1.57 |
1.91 |
1.59 |
1.47 |
1.26 |
1.40 |
| ATP | 1.17 |
1.03 |
1.30 |
1.00 |
.81 |
.62 |
1.37 |
| KLA/ATP | 1.68 |
1.70 |
1.91 |
1.37 |
1.09 |
.47 |
1.13 |
|
| Ppp3cb |  |
AK171904 |
activated spleen cDNA, RIKEN full-length enriched library, clone:F830018L06 product:Protein phosphatase 3 catalytic subunit beta3 (EC 3.1.3.16) (Serine [AK171904] |
KLA | 1.29 |
1.29 |
1.21 |
1.15 |
1.22 |
1.13 |
1.28 |
| ATP | .90 |
.86 |
.80 |
.98 |
.94 |
.73 |
.92 |
| KLA/ATP | 1.06 |
1.03 |
.77 |
1.04 |
.92 |
.68 |
.90 |
|
| Ppp3cb |  |
NM_008914 |
protein phosphatase 3, catalytic subunit, beta isoform (Ppp3cb), mRNA [NM_008914] |
KLA | 1.21 |
1.29 |
1.38 |
1.24 |
1.29 |
1.08 |
1.23 |
| ATP | 1.15 |
1.31 |
1.09 |
1.29 |
.86 |
.60 |
.64 |
| KLA/ATP | 1.33 |
1.40 |
1.04 |
1.28 |
.79 |
.58 |
.73 |
|
| Ppp3cc | 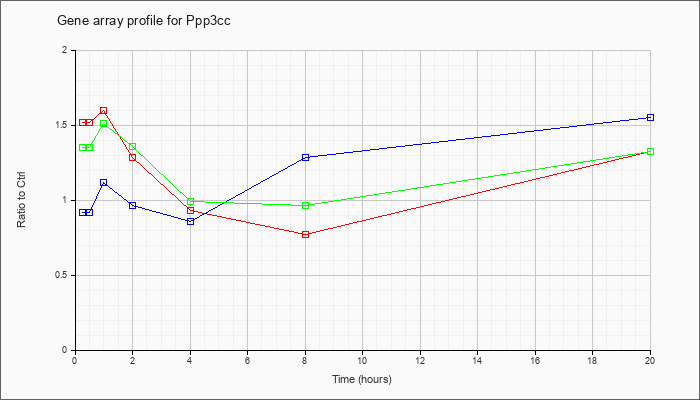 |
AK037563 |
16 days neonate thymus cDNA, RIKEN full-length enriched library, clone:A130026K22 product:unclassifiable, full insert sequence [AK037563] |
KLA | 1.27 |
1.18 |
1.21 |
1.15 |
1.01 |
.92 |
1.36 |
| ATP | .96 |
1.09 |
1.10 |
1.03 |
.85 |
1.06 |
1.47 |
| KLA/ATP | 1.17 |
1.24 |
1.07 |
1.13 |
.91 |
.88 |
1.36 |
|
| Ppp3cc |  |
NM_008915 |
protein phosphatase 3, catalytic subunit, gamma isoform (Ppp3cc), mRNA [NM_008915] |
KLA | 1.64 |
1.81 |
1.79 |
1.35 |
.90 |
.70 |
1.31 |
| ATP | .90 |
.90 |
1.13 |
.93 |
.86 |
1.40 |
1.60 |
| KLA/ATP | 1.45 |
1.62 |
1.74 |
1.47 |
1.04 |
1.01 |
1.31 |
|
| Ppp3r1 | 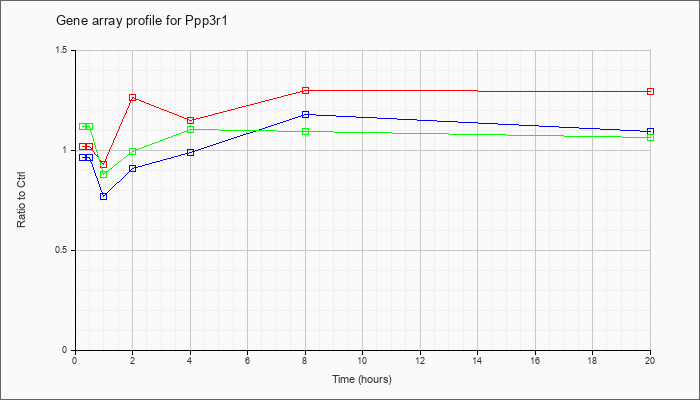 |
NM_024459 |
protein phosphatase 3, regulatory subunit B, alpha isoform (calcineurin B, type I) (Ppp3r1), mRNA [NM_024459] |
KLA | 1.02 |
1.17 |
.93 |
1.27 |
1.15 |
1.30 |
1.30 |
| ATP | .97 |
.95 |
.77 |
.91 |
.99 |
1.18 |
1.10 |
| KLA/ATP | 1.12 |
1.04 |
.88 |
1.00 |
1.10 |
1.10 |
1.07 |
|
| Ppp3r2 | 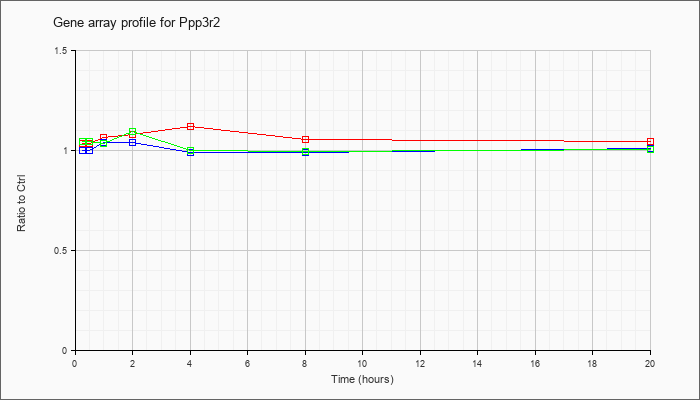 |
NM_001004025 |
protein phosphatase 3, regulatory subunit B, alpha isoform (calcineurin B, type II) (Ppp3r2), mRNA [NM_001004025] |
KLA | 1.03 |
1.05 |
1.06 |
1.08 |
1.12 |
1.05 |
1.04 |
| ATP | 1.00 |
1.02 |
1.04 |
1.04 |
.99 |
.99 |
1.01 |
| KLA/ATP | 1.04 |
1.12 |
1.03 |
1.09 |
1.00 |
.99 |
1.00 |
|
| Rac1 | 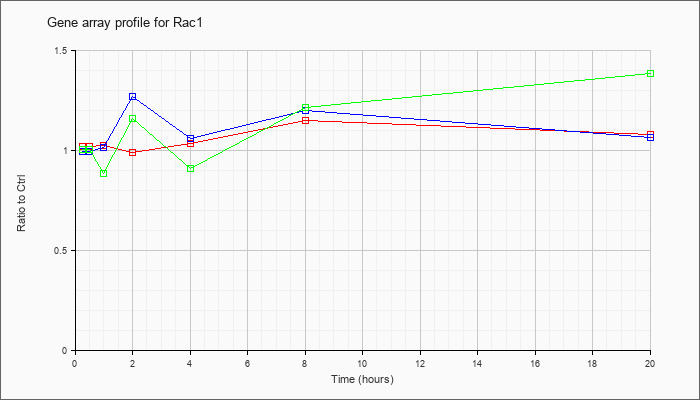 |
NM_009007 |
RAS-related C3 botulinum substrate 1 (Rac1), mRNA [NM_009007] |
KLA | 1.02 |
1.01 |
1.02 |
.99 |
1.03 |
1.15 |
1.08 |
| ATP | .99 |
1.15 |
1.01 |
1.27 |
1.06 |
1.20 |
1.06 |
| KLA/ATP | 1.00 |
1.04 |
.88 |
1.16 |
.91 |
1.21 |
1.38 |
|
| Rela | 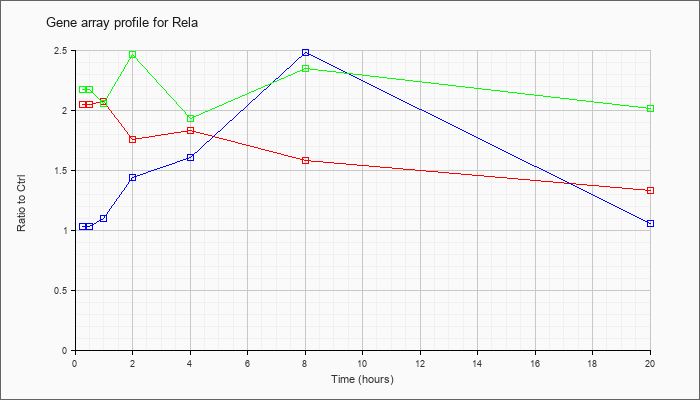 |
NM_009045 |
v-rel reticuloendotheliosis viral oncogene homolog A (avian) (Rela), mRNA [NM_009045] |
KLA | 2.05 |
2.05 |
2.07 |
1.75 |
1.83 |
1.58 |
1.33 |
| ATP | 1.03 |
1.08 |
1.10 |
1.44 |
1.61 |
2.48 |
1.05 |
| KLA/ATP | 2.17 |
2.27 |
2.05 |
2.47 |
1.93 |
2.35 |
2.02 |
|
| Relb |  |
NM_009046 |
avian reticuloendotheliosis viral (v-rel) oncogene related B (Relb), mRNA [NM_009046] |
KLA | 4.39 |
4.46 |
3.54 |
3.47 |
4.36 |
3.18 |
2.72 |
| ATP | 1.01 |
1.07 |
1.23 |
2.66 |
3.23 |
3.50 |
1.24 |
| KLA/ATP | 4.43 |
4.52 |
3.56 |
4.10 |
2.85 |
2.92 |
3.30 |
|
| Sfpi1 | 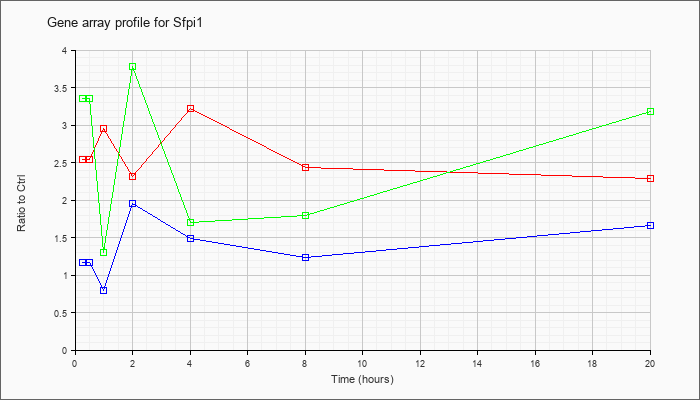 |
NM_011355 |
SFFV proviral integration 1 (Sfpi1), mRNA [NM_011355] |
KLA | 2.54 |
2.77 |
2.96 |
2.31 |
3.22 |
2.44 |
2.29 |
| ATP | 1.17 |
1.55 |
.79 |
1.96 |
1.49 |
1.23 |
1.66 |
| KLA/ATP | 3.35 |
3.40 |
1.30 |
3.78 |
1.70 |
1.80 |
3.18 |
|
| Sirpa | 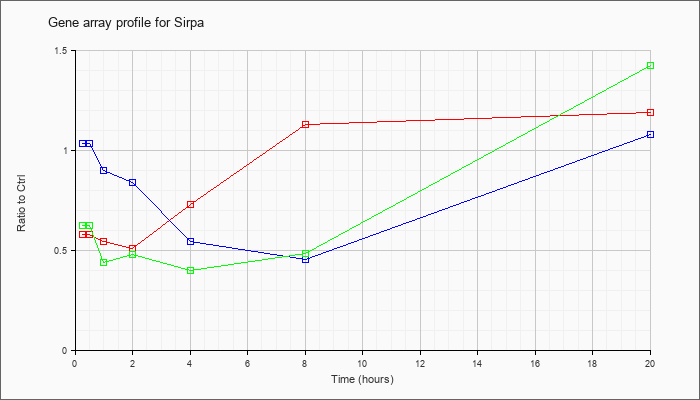 |
NM_007547 |
signal-regulatory protein alpha (Sirpa), mRNA [NM_007547] |
KLA | .58 |
.56 |
.55 |
.51 |
.73 |
1.13 |
1.19 |
| ATP | 1.03 |
1.10 |
.90 |
.84 |
.54 |
.45 |
1.08 |
| KLA/ATP | .62 |
.62 |
.44 |
.48 |
.40 |
.49 |
1.42 |
|
| Sirpb1 | 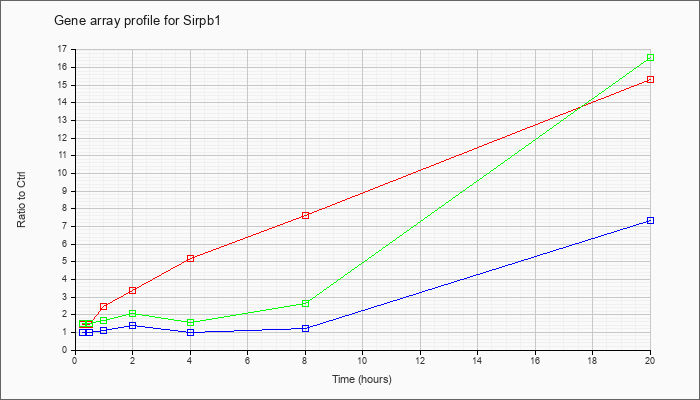 |
NM_001002898 |
signal-regulatory protein beta 1 (Sirpb1), transcript variant 3, mRNA [NM_001002898] |
KLA | 1.42 |
1.48 |
2.48 |
3.40 |
5.19 |
7.61 |
15.34 |
| ATP | .97 |
.99 |
1.13 |
1.41 |
1.02 |
1.23 |
7.36 |
| KLA/ATP | 1.53 |
1.50 |
1.68 |
2.07 |
1.58 |
2.62 |
16.58 |
|
| Socs1 | 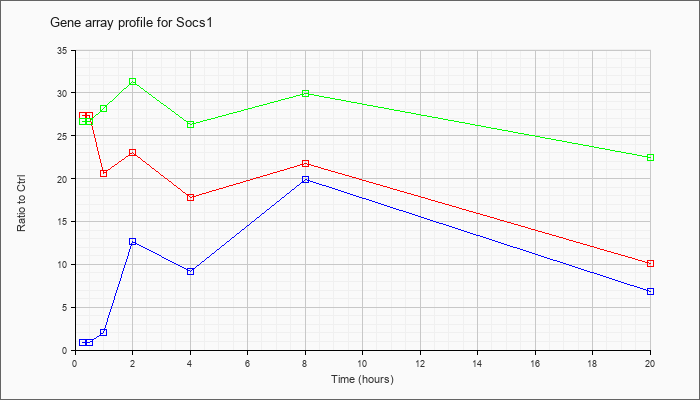 |
NM_009896 |
suppressor of cytokine signaling 1 (Socs1), mRNA [NM_009896] |
KLA | 27.36 |
22.16 |
20.64 |
23.08 |
17.74 |
21.71 |
10.12 |
| ATP | .91 |
.72 |
2.03 |
12.69 |
9.16 |
19.87 |
6.80 |
| KLA/ATP | 26.70 |
24.56 |
28.15 |
31.28 |
26.32 |
29.95 |
22.51 |
|
| Socs3 | 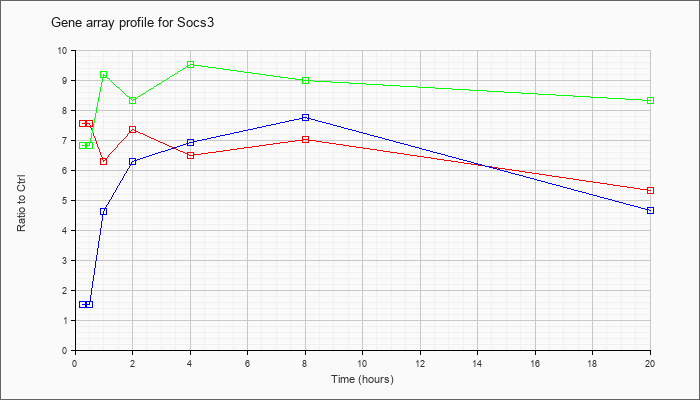 |
NM_007707 |
suppressor of cytokine signaling 3 (Socs3), mRNA [NM_007707] |
KLA | 7.56 |
6.68 |
6.29 |
7.36 |
6.47 |
7.03 |
5.33 |
| ATP | 1.51 |
1.75 |
4.63 |
6.27 |
6.92 |
7.74 |
4.64 |
| KLA/ATP | 6.82 |
6.77 |
9.19 |
8.32 |
9.51 |
8.98 |
8.33 |
|
| Sqstm1 |  |
NM_011018 |
sequestosome 1 (Sqstm1), mRNA [NM_011018] |
KLA | 3.50 |
3.07 |
2.12 |
1.55 |
.75 |
1.00 |
2.24 |
| ATP | 1.03 |
.98 |
1.63 |
2.91 |
6.11 |
9.76 |
1.45 |
| KLA/ATP | 3.46 |
3.23 |
3.50 |
2.93 |
3.38 |
4.26 |
2.74 |
|
| Stat1 |  |
NM_009283 |
signal transducer and activator of transcription 1 (Stat1), mRNA [NM_009283] |
KLA | 2.90 |
2.72 |
3.00 |
3.64 |
4.58 |
6.51 |
5.58 |
| ATP | 1.01 |
.92 |
.89 |
.72 |
.51 |
2.22 |
3.07 |
| KLA/ATP | 2.73 |
2.65 |
2.34 |
1.95 |
1.99 |
4.70 |
8.66 |
|
| Stat2 | 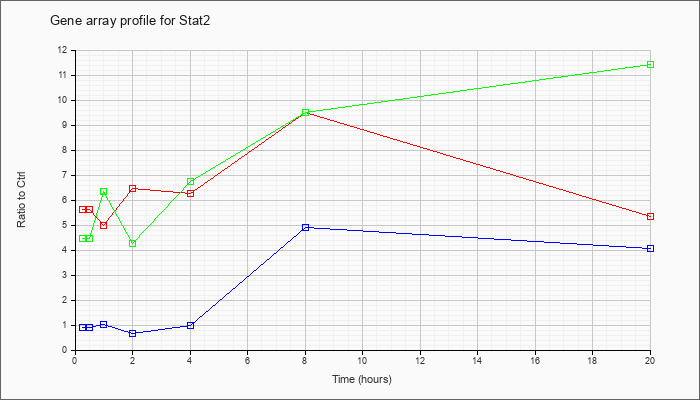 |
NM_019963 |
signal transducer and activator of transcription 2 (Stat2), mRNA [NM_019963] |
KLA | 5.64 |
4.80 |
4.99 |
6.48 |
6.28 |
9.50 |
5.36 |
| ATP | .90 |
.80 |
1.03 |
.66 |
.97 |
4.90 |
4.07 |
| KLA/ATP | 4.47 |
4.88 |
6.34 |
4.27 |
6.76 |
9.49 |
11.43 |
|
| Syk | 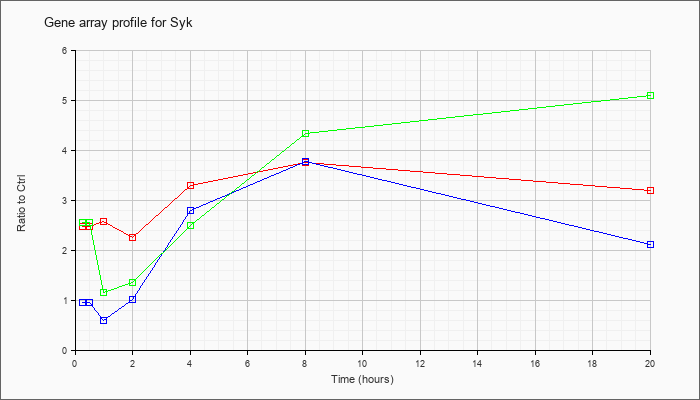 |
NM_011518 |
spleen tyrosine kinase (Syk), mRNA [NM_011518] |
KLA | 2.48 |
2.59 |
2.57 |
2.26 |
3.29 |
3.75 |
3.19 |
| ATP | .96 |
.85 |
.59 |
1.01 |
2.80 |
3.77 |
2.11 |
| KLA/ATP | 2.56 |
2.09 |
1.16 |
1.35 |
2.50 |
4.33 |
5.10 |
|
| Tec | 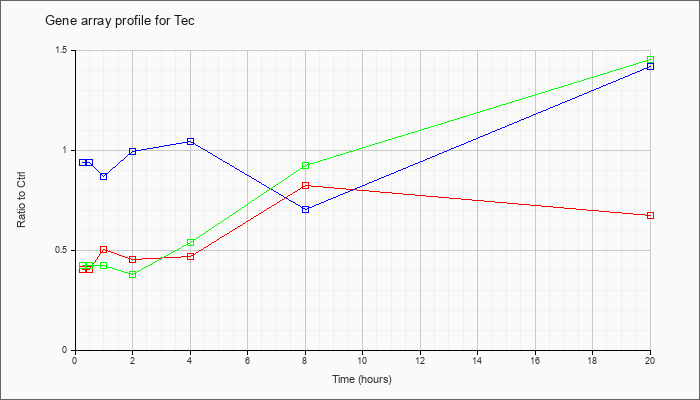 |
NM_001113460 |
cytoplasmic tyrosine kinase, Dscr28C related (Drosophila) (Tec), transcript variant 1, mRNA [NM_001113460] |
KLA | .41 |
.41 |
.51 |
.46 |
.47 |
.83 |
.68 |
| ATP | .94 |
.91 |
.87 |
1.00 |
1.05 |
.71 |
1.42 |
| KLA/ATP | .43 |
.42 |
.43 |
.38 |
.54 |
.93 |
1.46 |
|
| Tgfb1 | 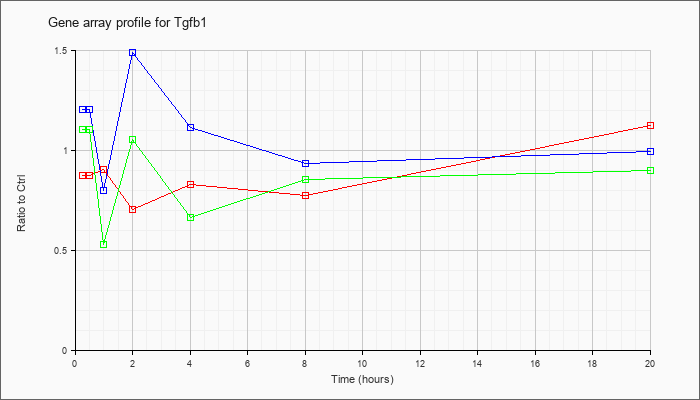 |
NM_011577 |
transforming growth factor, beta 1 (Tgfb1), mRNA [NM_011577] |
KLA | .87 |
.89 |
.90 |
.70 |
.83 |
.77 |
1.12 |
| ATP | 1.20 |
1.41 |
.80 |
1.49 |
1.11 |
.93 |
.99 |
| KLA/ATP | 1.10 |
1.20 |
.53 |
1.05 |
.66 |
.85 |
.90 |
|
| Tgfb2 | 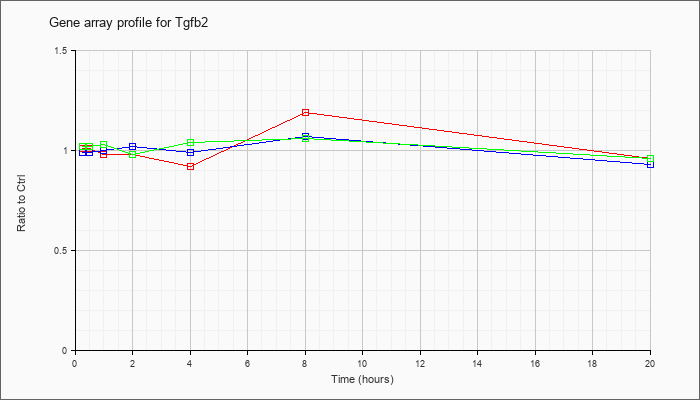 |
NM_009367 |
transforming growth factor, beta 2 (Tgfb2), mRNA [NM_009367] |
KLA | 1.01 |
.91 |
.98 |
.98 |
.92 |
1.19 |
.96 |
| ATP | .99 |
1.03 |
1.00 |
1.02 |
.99 |
1.07 |
.93 |
| KLA/ATP | 1.02 |
.99 |
1.03 |
.98 |
1.04 |
1.06 |
.96 |
|
| Tgfbr1 | 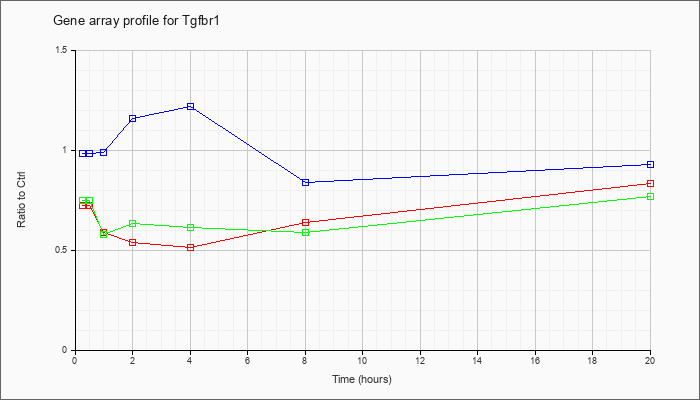 |
NM_009370 |
transforming growth factor, beta receptor I (Tgfbr1), mRNA [NM_009370] |
KLA | .73 |
.74 |
.59 |
.54 |
.52 |
.64 |
.84 |
| ATP | .99 |
1.12 |
.99 |
1.16 |
1.22 |
.84 |
.93 |
| KLA/ATP | .75 |
.78 |
.58 |
.64 |
.62 |
.59 |
.77 |
|
| Tgfbr2 | 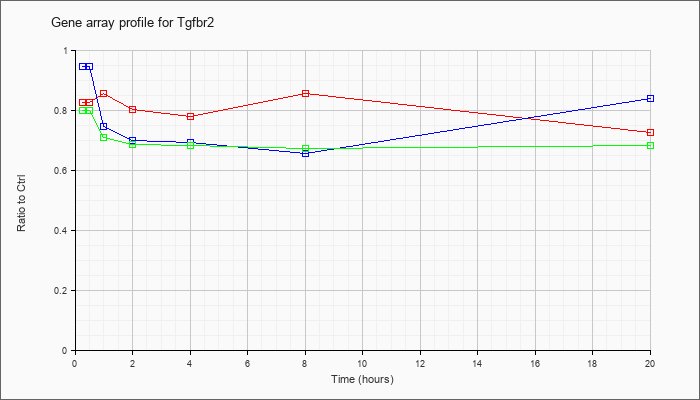 |
AK090393 |
10 days pregnant adult female ovary and uterus cDNA, RIKEN full-length enriched library, clone:G630098D03 product:transforming growth factor, beta receptor II, full insert sequence. [AK090393] |
KLA | .98 |
.89 |
1.00 |
.96 |
.94 |
1.00 |
.92 |
| ATP | .99 |
.99 |
.90 |
.94 |
.93 |
.96 |
1.02 |
| KLA/ATP | .92 |
.96 |
.98 |
.98 |
.98 |
.95 |
1.01 |
|
| Tgfbr2 |  |
NM_009371 |
transforming growth factor, beta receptor II (Tgfbr2), transcript variant 1, mRNA [NM_009371] |
KLA | .55 |
.50 |
.58 |
.50 |
.52 |
.55 |
.44 |
| ATP | .98 |
.75 |
.50 |
.32 |
.41 |
.29 |
.50 |
| KLA/ATP | .59 |
.45 |
.32 |
.23 |
.31 |
.27 |
.25 |
|
| Tgfbr2 |  |
NM_029575 |
transforming growth factor, beta receptor II (Tgfbr2), transcript variant 2, mRNA [NM_029575] |
KLA | .95 |
.93 |
.99 |
.95 |
.88 |
1.02 |
.82 |
| ATP | .87 |
.93 |
.84 |
.84 |
.74 |
.72 |
1.00 |
| KLA/ATP | .89 |
.85 |
.83 |
.85 |
.76 |
.80 |
.79 |
|
| Tnf | 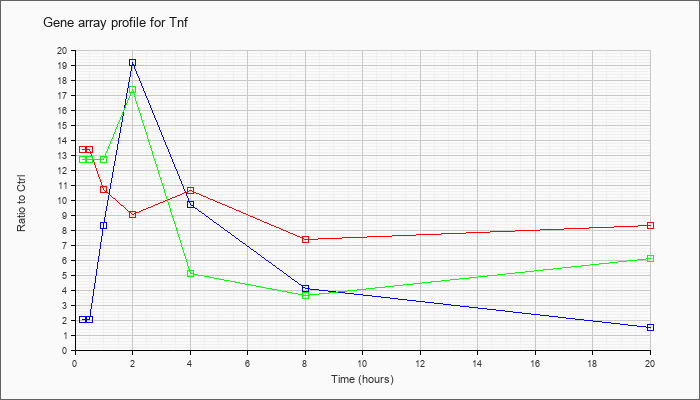 |
NM_013693 |
tumor necrosis factor (Tnf), mRNA [NM_013693] |
KLA | 13.35 |
12.76 |
10.71 |
9.03 |
10.64 |
7.39 |
8.32 |
| ATP | 2.01 |
2.48 |
8.27 |
19.20 |
9.72 |
4.09 |
1.49 |
| KLA/ATP | 12.72 |
15.76 |
12.73 |
17.34 |
5.10 |
3.61 |
6.07 |
|
Tnfrsf11
a | 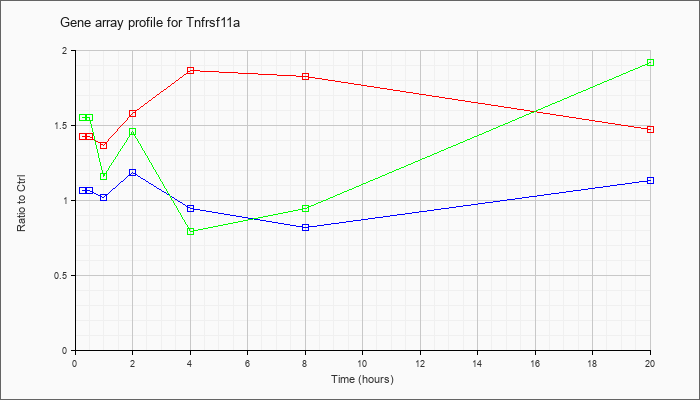 |
NM_009399 |
tumor necrosis factor receptor superfamily, member 11a (Tnfrsf11a), mRNA [NM_009399] |
KLA | 1.43 |
1.33 |
1.36 |
1.58 |
1.87 |
1.83 |
1.47 |
| ATP | 1.07 |
1.17 |
1.02 |
1.19 |
.95 |
.82 |
1.13 |
| KLA/ATP | 1.55 |
1.46 |
1.16 |
1.46 |
.79 |
.95 |
1.92 |
|
Tnfrsf11
b |  |
NM_008764 |
tumor necrosis factor receptor superfamily, member 11b (osteoprotegerin) (Tnfrsf11b), mRNA [NM_008764] |
KLA | 1.00 |
.98 |
.98 |
.98 |
1.02 |
.94 |
.90 |
| ATP | .99 |
.98 |
1.06 |
.97 |
.94 |
.99 |
.98 |
| KLA/ATP | .97 |
1.04 |
1.09 |
.94 |
.95 |
.92 |
1.01 |
|
Tnfrsf1a
| 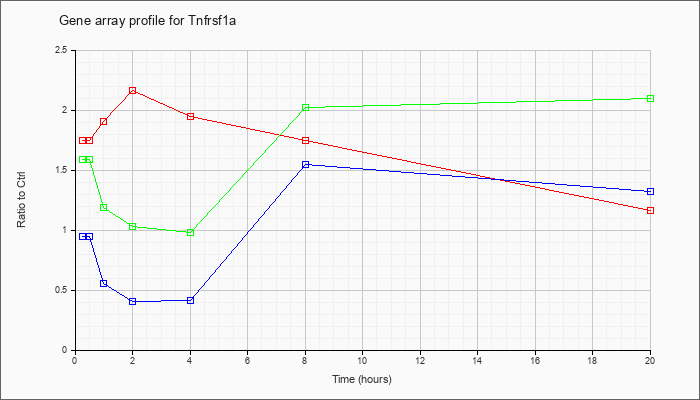 |
NM_011609 |
tumor necrosis factor receptor superfamily, member 1a (Tnfrsf1a), mRNA [NM_011609] |
KLA | 1.74 |
1.71 |
1.90 |
2.16 |
1.95 |
1.74 |
1.16 |
| ATP | .95 |
.83 |
.55 |
.41 |
.41 |
1.55 |
1.32 |
| KLA/ATP | 1.58 |
1.49 |
1.19 |
1.03 |
.98 |
2.02 |
2.10 |
|
| Tnfsf11 | 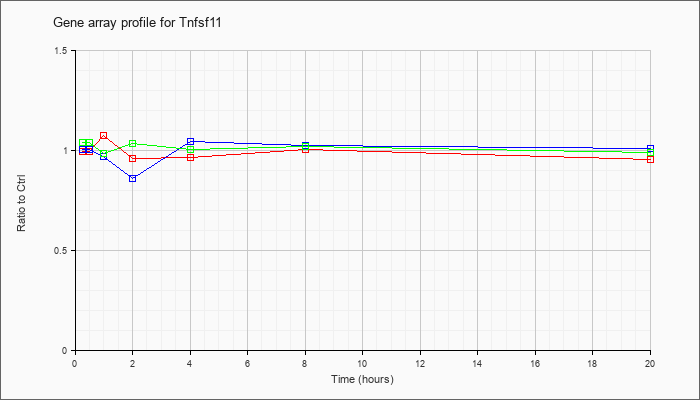 |
NM_011613 |
tumor necrosis factor (ligand) superfamily, member 11 (Tnfsf11), mRNA [NM_011613] |
KLA | 1.00 |
1.00 |
1.08 |
.96 |
.97 |
1.01 |
.96 |
| ATP | 1.01 |
1.05 |
.97 |
.86 |
1.05 |
1.03 |
1.01 |
| KLA/ATP | 1.04 |
1.05 |
.99 |
1.04 |
1.01 |
1.02 |
.99 |
|
| Traf2 |  |
NM_009422 |
Tnf receptor-associated factor 2 (Traf2), mRNA [NM_009422] |
KLA | 3.38 |
3.62 |
3.49 |
3.36 |
3.49 |
3.39 |
2.68 |
| ATP | .97 |
1.04 |
1.00 |
1.10 |
1.54 |
2.40 |
1.55 |
| KLA/ATP | 3.38 |
3.16 |
2.93 |
2.74 |
2.67 |
3.11 |
4.70 |
|
| Traf6 | 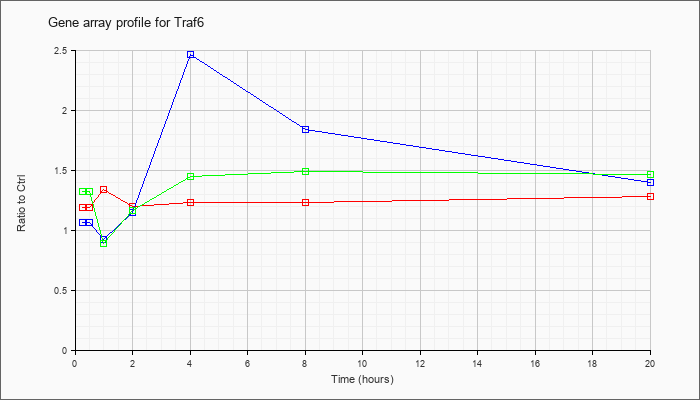 |
NM_009424 |
Tnf receptor-associated factor 6 (Traf6), mRNA [NM_009424] |
KLA | 1.19 |
1.28 |
1.34 |
1.20 |
1.23 |
1.23 |
1.28 |
| ATP | 1.06 |
1.12 |
.92 |
1.15 |
2.46 |
1.84 |
1.40 |
| KLA/ATP | 1.32 |
1.37 |
.89 |
1.16 |
1.45 |
1.49 |
1.46 |
|
| Trem2 | 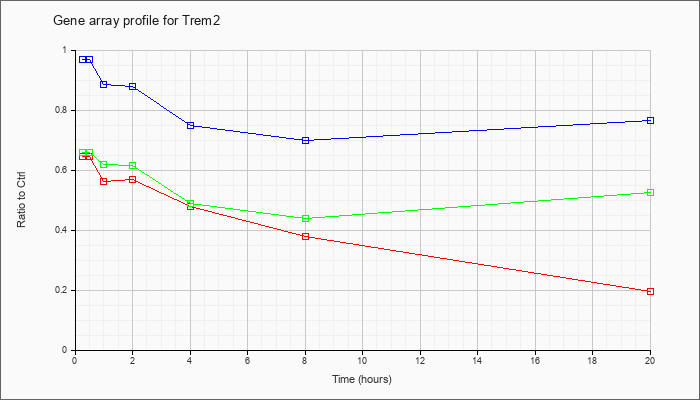 |
NM_031254 |
triggering receptor expressed on myeloid cells 2 (Trem2), mRNA [NM_031254] |
KLA | .65 |
.62 |
.56 |
.57 |
.48 |
.38 |
.20 |
| ATP | .97 |
.98 |
.89 |
.88 |
.75 |
.70 |
.77 |
| KLA/ATP | .66 |
.64 |
.62 |
.62 |
.49 |
.44 |
.53 |
|
| Tyk2 | 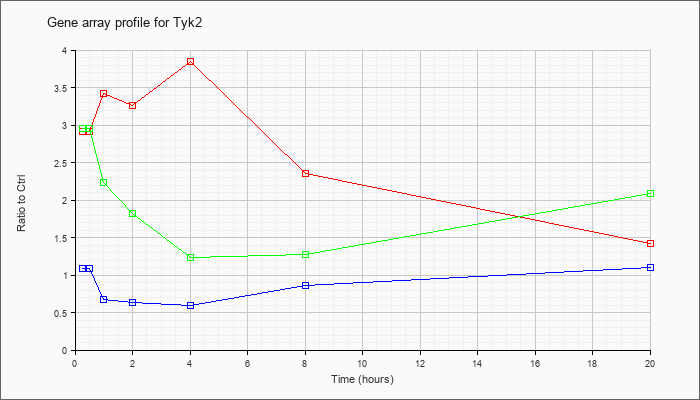 |
NM_018793 |
tyrosine kinase 2 (Tyk2), mRNA [NM_018793] |
KLA | 2.91 |
2.95 |
3.42 |
3.26 |
3.85 |
2.35 |
1.42 |
| ATP | 1.09 |
1.01 |
.68 |
.64 |
.59 |
.86 |
1.10 |
| KLA/ATP | 2.95 |
3.04 |
2.23 |
1.82 |
1.24 |
1.27 |
2.09 |
|
| Tyrobp |  |
NM_011662 |
TYRO protein tyrosine kinase binding protein (Tyrobp), mRNA [NM_011662] |
KLA | .80 |
.78 |
.84 |
.69 |
.72 |
.69 |
.92 |
| ATP | 1.10 |
1.12 |
.90 |
1.11 |
.83 |
.71 |
.90 |
| KLA/ATP | .92 |
.90 |
.64 |
.87 |
.65 |
.61 |
.92 |
|