| Gene symbol |
Time course plot |
Accession |
Description |
Treatment |
15min |
30min |
1hr |
2hr |
4hr |
8hr |
20hr |
AK050363
| 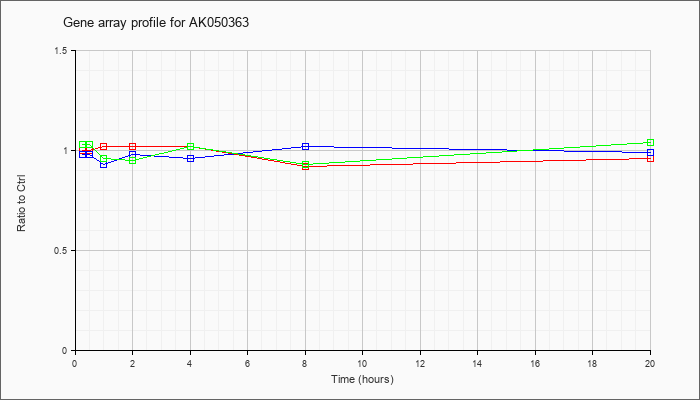 |
AK050363 |
adult male liver tumor cDNA, RIKEN full-length enriched library, clone:C730040N22 product:unclassifiable, full insert sequence. [AK050363] |
KLA | 1.00 |
.98 |
1.02 |
1.02 |
1.02 |
.92 |
.96 |
| ATP | .98 |
1.03 |
.93 |
.98 |
.96 |
1.02 |
.99 |
| KLA/ATP | 1.03 |
.96 |
.96 |
.95 |
1.02 |
.93 |
1.04 |
|
| Abl1 |  |
AK083248 |
adult male hippocampus cDNA, RIKEN full-length enriched library, clone:C630031E05 product:v-abl Abelson murine leukemia oncogene 1, full insert sequence. [AK083248] |
KLA | 1.07 |
1.07 |
.92 |
1.01 |
.98 |
1.07 |
.98 |
| ATP | .98 |
.99 |
.94 |
.90 |
.99 |
1.07 |
1.04 |
| KLA/ATP | 1.04 |
1.01 |
1.03 |
.98 |
.97 |
.95 |
.99 |
|
| Abl1 |  |
NM_009594 |
c-abl oncogene 1, receptor tyrosine kinase (Abl1), transcript variant 2, mRNA [NM_009594] |
KLA | .63 |
.62 |
.65 |
.71 |
.96 |
1.17 |
1.05 |
| ATP | 1.01 |
1.16 |
1.03 |
1.14 |
1.12 |
.98 |
.98 |
| KLA/ATP | .66 |
.64 |
.67 |
.73 |
.84 |
.90 |
1.05 |
|
| Akt1 | 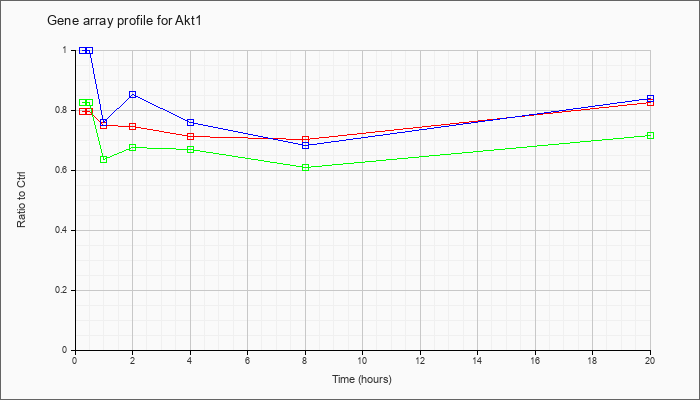 |
NM_009652 |
thymoma viral proto-oncogene 1 (Akt1), mRNA [NM_009652] |
KLA | .80 |
.80 |
.75 |
.75 |
.71 |
.70 |
.82 |
| ATP | 1.00 |
1.02 |
.76 |
.85 |
.76 |
.68 |
.84 |
| KLA/ATP | .82 |
.80 |
.64 |
.68 |
.67 |
.61 |
.72 |
|
| Akt2 | 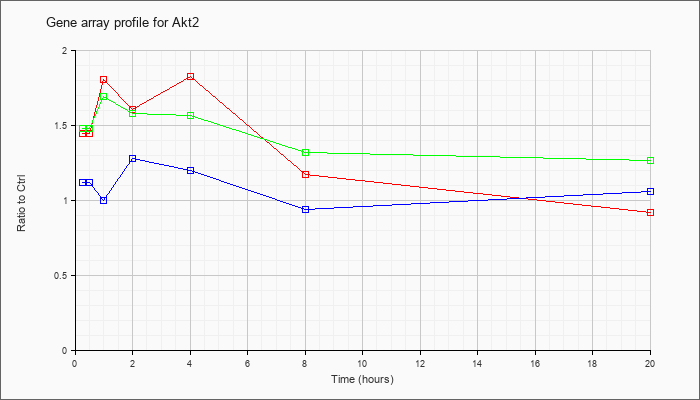 |
NM_007434 |
thymoma viral proto-oncogene 2 (Akt2), transcript variant 2, mRNA [NM_007434] |
KLA | 1.45 |
1.49 |
1.81 |
1.61 |
1.83 |
1.17 |
.92 |
| ATP | 1.12 |
1.12 |
1.00 |
1.28 |
1.20 |
.94 |
1.06 |
| KLA/ATP | 1.48 |
1.70 |
1.69 |
1.58 |
1.57 |
1.32 |
1.27 |
|
| Akt3 | 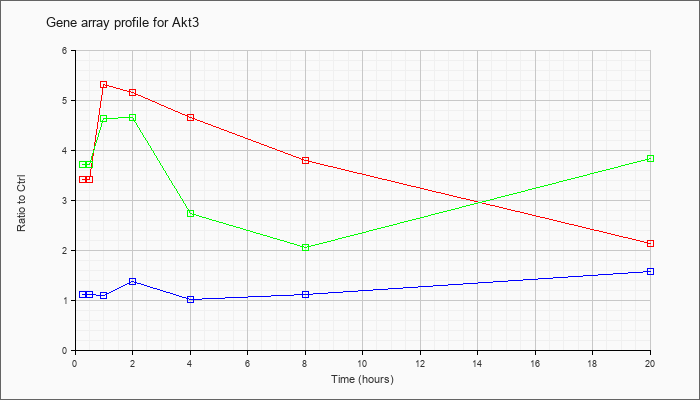 |
NM_011785 |
thymoma viral proto-oncogene 3 (Akt3), mRNA [NM_011785] |
KLA | 3.42 |
3.45 |
5.31 |
5.16 |
4.65 |
3.80 |
2.12 |
| ATP | 1.12 |
1.22 |
1.10 |
1.37 |
1.02 |
1.11 |
1.56 |
| KLA/ATP | 3.72 |
4.17 |
4.62 |
4.64 |
2.74 |
2.05 |
3.82 |
|
| Arhgdia | 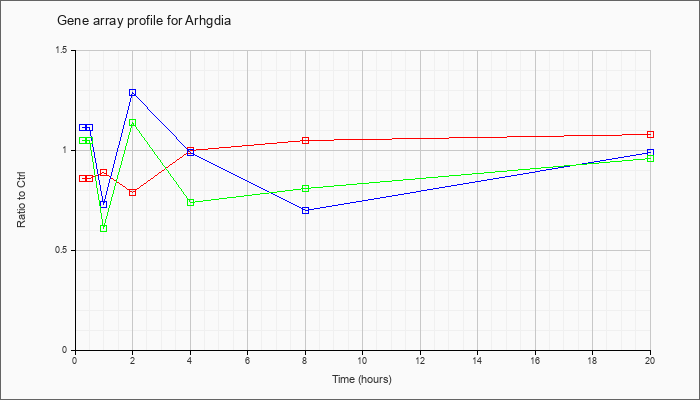 |
NM_133796 |
Rho GDP dissociation inhibitor (GDI) alpha (Arhgdia), mRNA [NM_133796] |
KLA | .86 |
.92 |
.89 |
.79 |
1.00 |
1.05 |
1.08 |
| ATP | 1.11 |
1.31 |
.73 |
1.29 |
.99 |
.70 |
.99 |
| KLA/ATP | 1.05 |
1.11 |
.61 |
1.14 |
.74 |
.81 |
.96 |
|
| Arhgdib | 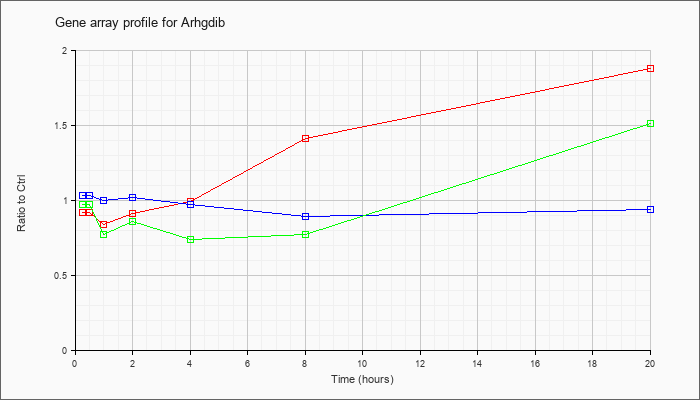 |
NM_007486 |
Rho, GDP dissociation inhibitor (GDI) beta (Arhgdib), mRNA [NM_007486] |
KLA | .92 |
.96 |
.84 |
.91 |
.99 |
1.41 |
1.88 |
| ATP | 1.03 |
1.09 |
1.00 |
1.02 |
.97 |
.89 |
.94 |
| KLA/ATP | .97 |
.89 |
.77 |
.86 |
.74 |
.77 |
1.51 |
|
| Arhgdig | 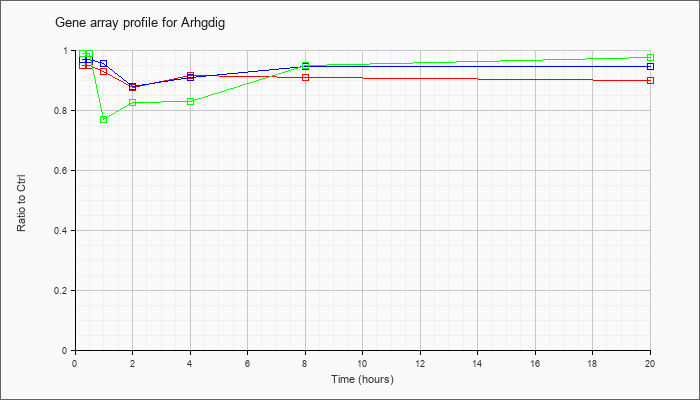 |
NM_008113 |
Rho GDP dissociation inhibitor (GDI) gamma (Arhgdig), mRNA [NM_008113] |
KLA | .95 |
.92 |
.93 |
.88 |
.92 |
.91 |
.90 |
| ATP | .97 |
1.04 |
.96 |
.88 |
.91 |
.95 |
.95 |
| KLA/ATP | .99 |
.96 |
.77 |
.83 |
.83 |
.95 |
.98 |
|
| Atf4 | 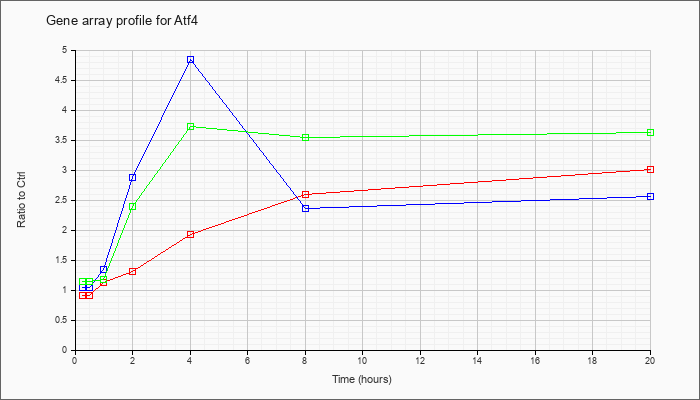 |
NM_009716 |
activating transcription factor 4 (Atf4), mRNA [NM_009716] |
KLA | .91 |
1.06 |
1.13 |
1.31 |
1.92 |
2.59 |
3.01 |
| ATP | 1.05 |
1.14 |
1.34 |
2.88 |
4.84 |
2.36 |
2.56 |
| KLA/ATP | 1.14 |
1.08 |
1.17 |
2.40 |
3.72 |
3.54 |
3.62 |
|
BB217526
| 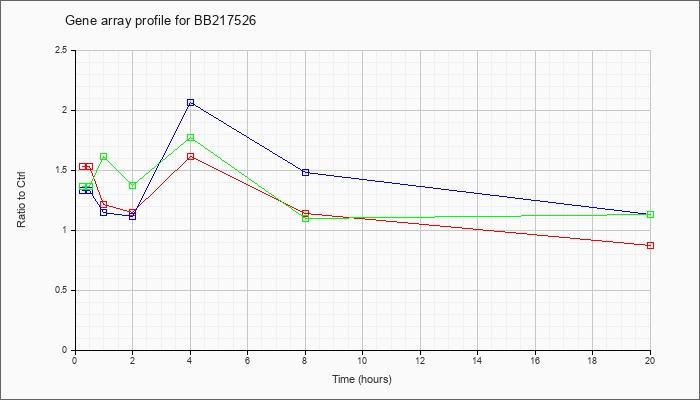 |
AK083592 |
9 days embryo whole body cDNA, RIKEN full-length enriched library, clone:D030051F04 product:unclassifiable, full insert sequence [AK083592] |
KLA | 1.53 |
1.19 |
1.21 |
1.15 |
1.61 |
1.14 |
.87 |
| ATP | 1.33 |
1.11 |
1.15 |
1.11 |
2.06 |
1.48 |
1.13 |
| KLA/ATP | 1.36 |
1.18 |
1.61 |
1.37 |
1.77 |
1.10 |
1.13 |
|
BG242006
| 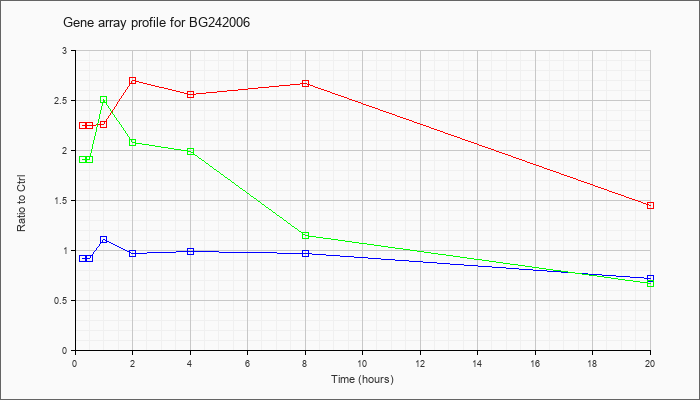 |
BG242006 |
gb|602354740F1 NCI_CGAP_Mam1 Mus musculus cDNA clone IMAGE:4483064 5. [BG242006] |
KLA | 2.25 |
2.23 |
2.26 |
2.70 |
2.56 |
2.67 |
1.45 |
| ATP | .92 |
.87 |
1.11 |
.97 |
.99 |
.97 |
.72 |
| KLA/ATP | 1.91 |
2.04 |
2.51 |
2.08 |
1.99 |
1.15 |
.67 |
|
| Bad |  |
AK029400 |
0 day neonate head cDNA, RIKEN full-length enriched library, clone:4833429M11 product:Bcl-associated death promoter, full insert sequence. [AK029400] |
KLA | 1.15 |
1.22 |
1.26 |
1.31 |
1.40 |
1.67 |
1.59 |
| ATP | .95 |
1.06 |
1.12 |
1.06 |
.78 |
.89 |
1.27 |
| KLA/ATP | 1.25 |
1.39 |
1.33 |
1.29 |
1.11 |
.91 |
1.38 |
|
| Bax | 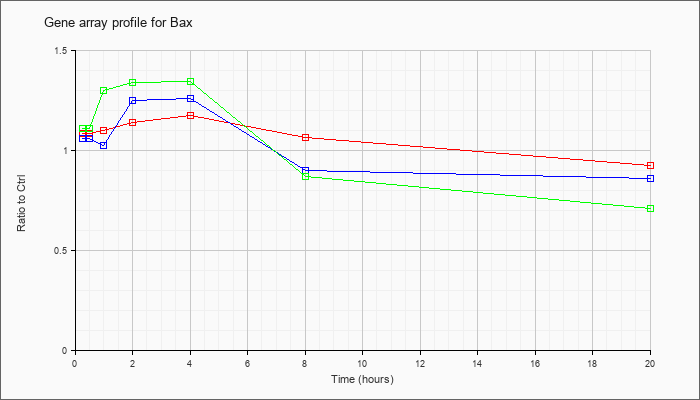 |
NM_007527 |
Bcl2-associated X protein (Bax), mRNA [NM_007527] |
KLA | 1.08 |
1.14 |
1.10 |
1.14 |
1.17 |
1.06 |
.92 |
| ATP | 1.06 |
1.05 |
1.02 |
1.25 |
1.26 |
.90 |
.86 |
| KLA/ATP | 1.11 |
1.19 |
1.30 |
1.34 |
1.34 |
.87 |
.71 |
|
| Bcl2 | 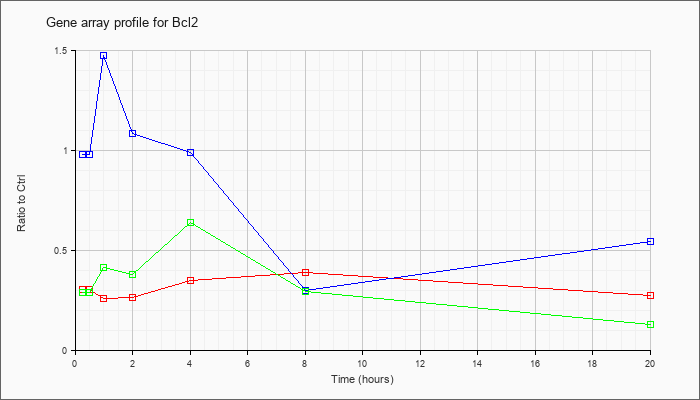 |
NM_009741 |
B-cell leukemia/lymphoma 2 (Bcl2), transcript variant 1, mRNA [NM_009741] |
KLA | .30 |
.26 |
.25 |
.25 |
.34 |
.38 |
.26 |
| ATP | .97 |
.89 |
1.50 |
1.04 |
.99 |
.29 |
.52 |
| KLA/ATP | .28 |
.26 |
.40 |
.33 |
.62 |
.29 |
.12 |
|
| Bcl2 |  |
NM_177410 |
B-cell leukemia/lymphoma 2 (Bcl2), transcript variant 2, mRNA [NM_177410] |
KLA | .33 |
.33 |
.37 |
.42 |
.49 |
.46 |
.39 |
| ATP | 1.07 |
1.12 |
1.22 |
1.56 |
1.00 |
.38 |
.76 |
| KLA/ATP | .35 |
.34 |
.54 |
.89 |
.76 |
.36 |
.23 |
|
| Bdnf | 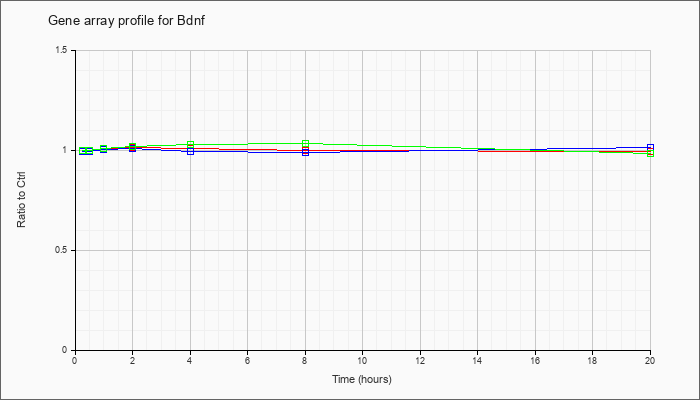 |
NM_007540 |
brain derived neurotrophic factor (Bdnf), transcript variant 1, mRNA [NM_007540] |
KLA | 1.00 |
1.01 |
1.01 |
1.01 |
1.01 |
1.00 |
.99 |
| ATP | 1.00 |
1.00 |
1.00 |
1.01 |
.99 |
.99 |
1.01 |
| KLA/ATP | 1.00 |
1.01 |
1.01 |
1.02 |
1.03 |
1.03 |
.98 |
|
| Braf |  |
NM_139294 |
Braf transforming gene (Braf), mRNA [NM_139294] |
KLA | 1.52 |
1.57 |
1.55 |
1.33 |
1.83 |
1.52 |
1.60 |
| ATP | 1.08 |
1.17 |
1.02 |
1.37 |
.95 |
1.15 |
.95 |
| KLA/ATP | 1.64 |
1.69 |
1.39 |
1.51 |
1.04 |
1.31 |
1.26 |
|
C330002I
19Rik |  |
AK083260 |
adult male hippocampus cDNA, RIKEN full-length enriched library, clone:C630031M24 product:KIDINS220 homolog [Rattus norvegicus], full insert sequence. [AK083260] |
KLA | 1.04 |
1.08 |
1.18 |
1.10 |
1.09 |
.89 |
.66 |
| ATP | .88 |
.67 |
.71 |
.62 |
.67 |
.71 |
.75 |
| KLA/ATP | .96 |
.72 |
.75 |
.68 |
.83 |
.74 |
.57 |
|
C330002I
19Rik |  |
NM_001081378 |
RIKEN cDNA C330002I19 gene (C330002I19Rik), mRNA [NM_001081378] |
KLA | .88 |
.98 |
.93 |
1.00 |
.95 |
.83 |
.79 |
| ATP | 1.09 |
1.11 |
1.16 |
.72 |
.89 |
.87 |
.67 |
| KLA/ATP | 1.04 |
1.08 |
1.24 |
.82 |
1.06 |
.69 |
.76 |
|
| Calm1 | 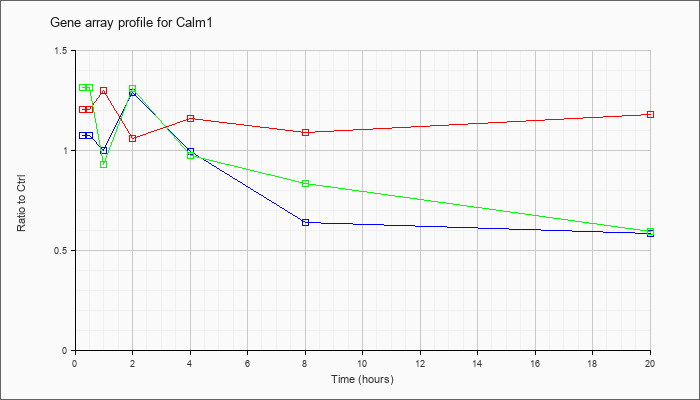 |
NM_009790 |
calmodulin 1 (Calm1), mRNA [NM_009790] |
KLA | 1.20 |
1.26 |
1.30 |
1.06 |
1.16 |
1.09 |
1.18 |
| ATP | 1.07 |
1.20 |
1.00 |
1.29 |
.99 |
.64 |
.58 |
| KLA/ATP | 1.31 |
1.31 |
.93 |
1.31 |
.97 |
.83 |
.59 |
|
| Calm2 | 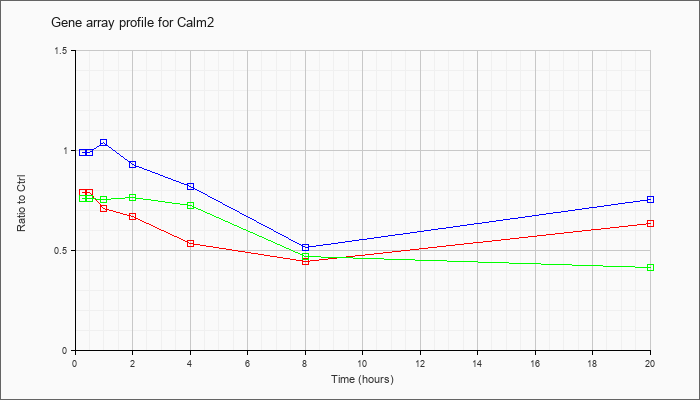 |
NM_007589 |
calmodulin 2 (Calm2), mRNA [NM_007589] |
KLA | .79 |
.81 |
.71 |
.67 |
.54 |
.45 |
.64 |
| ATP | .99 |
1.03 |
1.04 |
.93 |
.82 |
.52 |
.76 |
| KLA/ATP | .76 |
.78 |
.76 |
.77 |
.73 |
.47 |
.42 |
|
| Calm3 | 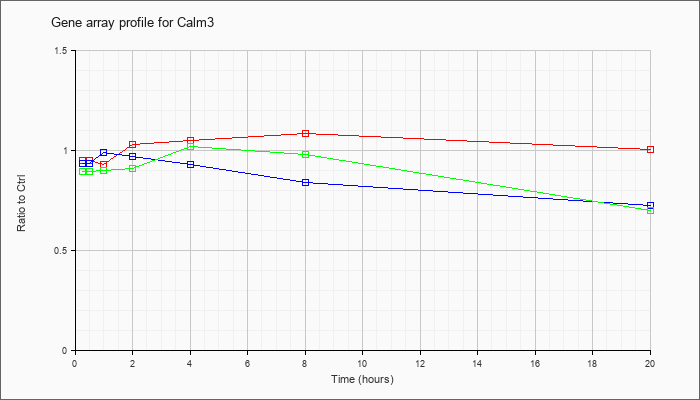 |
NM_007590 |
calmodulin 3 (Calm3), mRNA [NM_007590] |
KLA | .95 |
.96 |
.93 |
1.03 |
1.05 |
1.08 |
1.00 |
| ATP | .93 |
.96 |
.99 |
.97 |
.93 |
.84 |
.72 |
| KLA/ATP | .89 |
.93 |
.90 |
.91 |
1.02 |
.98 |
.70 |
|
| Calm4 | 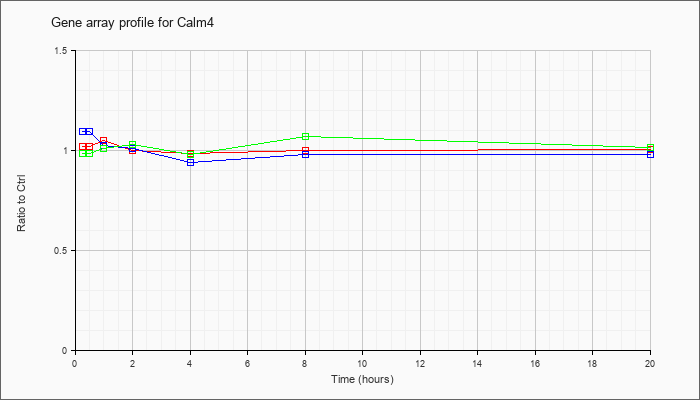 |
NM_020036 |
calmodulin 4 (Calm4), mRNA [NM_020036] |
KLA | 1.02 |
.96 |
1.05 |
1.00 |
.99 |
1.00 |
1.01 |
| ATP | 1.10 |
1.05 |
1.03 |
1.01 |
.94 |
.98 |
.98 |
| KLA/ATP | .99 |
1.12 |
1.01 |
1.03 |
.98 |
1.07 |
1.02 |
|
| Calm5 | 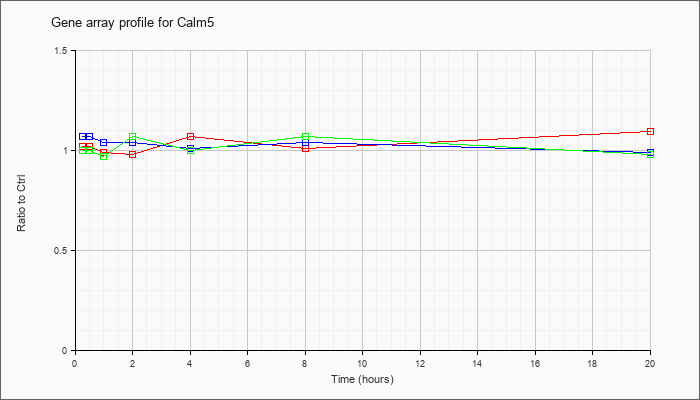 |
NM_001008706 |
calmodulin 5 (Calm5), mRNA [NM_001008706] |
KLA | 1.02 |
1.11 |
.99 |
.98 |
1.07 |
1.01 |
1.09 |
| ATP | 1.07 |
1.05 |
1.04 |
1.04 |
1.01 |
1.04 |
.99 |
| KLA/ATP | 1.00 |
1.00 |
.97 |
1.07 |
1.00 |
1.07 |
.98 |
|
| Calml3 |  |
NM_027416 |
calmodulin-like 3 (Calml3), mRNA [NM_027416] |
KLA | 1.01 |
.95 |
1.05 |
.91 |
1.01 |
.98 |
.89 |
| ATP | .91 |
1.05 |
1.04 |
1.01 |
.93 |
1.07 |
.99 |
| KLA/ATP | 1.01 |
.96 |
.97 |
.96 |
.97 |
.92 |
.99 |
|
| Camk2a | 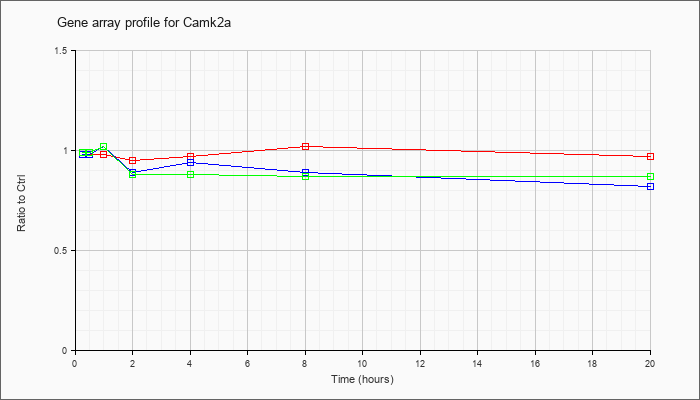 |
NM_177407 |
calcium/calmodulin-dependent protein kinase II alpha (Camk2a), transcript variant 2, mRNA [NM_177407] |
KLA | .98 |
.97 |
.98 |
.95 |
.97 |
1.02 |
.97 |
| ATP | .98 |
1.11 |
1.02 |
.89 |
.94 |
.89 |
.82 |
| KLA/ATP | .99 |
.95 |
1.02 |
.88 |
.88 |
.87 |
.87 |
|
| Camk2b | 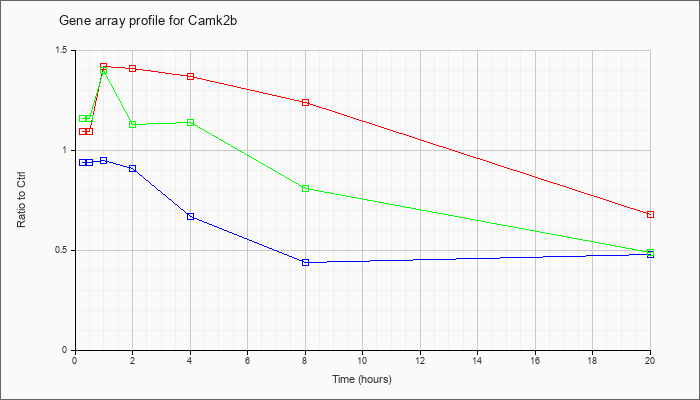 |
NM_007595 |
calcium/calmodulin-dependent protein kinase II, beta (Camk2b), mRNA [NM_007595] |
KLA | 1.09 |
1.28 |
1.42 |
1.41 |
1.37 |
1.24 |
.68 |
| ATP | .94 |
.87 |
.95 |
.91 |
.67 |
.44 |
.48 |
| KLA/ATP | 1.16 |
1.24 |
1.40 |
1.13 |
1.14 |
.81 |
.49 |
|
| Camk2d | 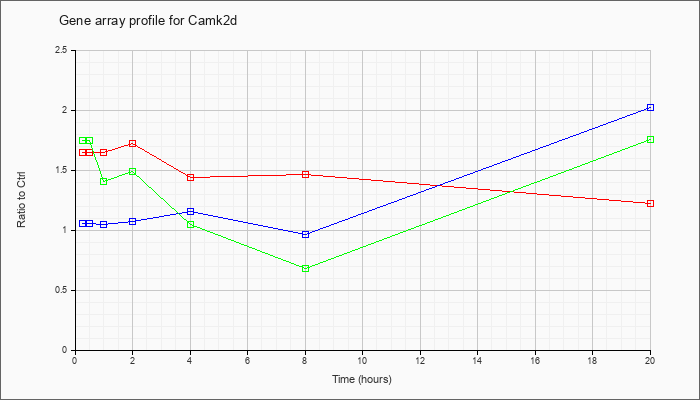 |
AK032524 |
adult male olfactory brain cDNA, RIKEN full-length enriched library, clone:6430579P05 product:calcium/calmodulin-dependent protein kinase II, delta, full insert sequence. [AK032524] |
KLA | 1.31 |
1.45 |
1.42 |
1.21 |
1.34 |
1.18 |
1.16 |
| ATP | 1.04 |
1.29 |
1.39 |
1.24 |
.94 |
.91 |
1.22 |
| KLA/ATP | 1.46 |
1.64 |
1.42 |
1.24 |
1.00 |
.93 |
1.10 |
|
| Camk2d |  |
NM_001025439 |
calcium/calmodulin-dependent protein kinase II, delta (Camk2d), transcript variant 1, mRNA [NM_001025439] |
KLA | 1.76 |
1.75 |
1.72 |
1.89 |
1.47 |
1.55 |
1.24 |
| ATP | 1.06 |
1.06 |
.93 |
1.01 |
1.22 |
.98 |
2.28 |
| KLA/ATP | 1.84 |
1.78 |
1.40 |
1.57 |
1.07 |
.60 |
1.97 |
|
| Camk2g | 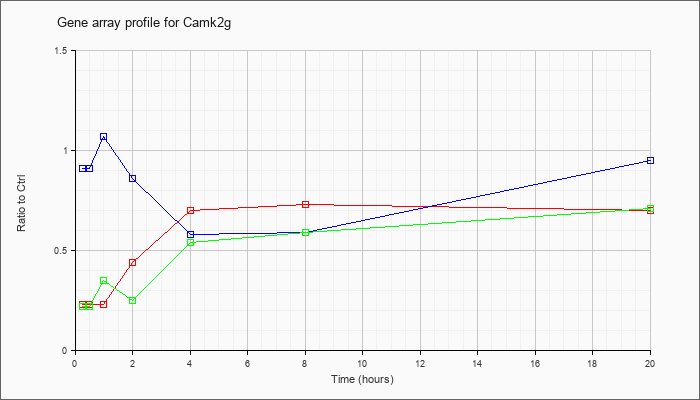 |
NM_178597 |
calcium/calmodulin-dependent protein kinase II gamma (Camk2g), transcript variant 1, mRNA [NM_178597] |
KLA | .23 |
.24 |
.23 |
.44 |
.70 |
.73 |
.70 |
| ATP | .91 |
.85 |
1.07 |
.86 |
.58 |
.59 |
.95 |
| KLA/ATP | .22 |
.20 |
.35 |
.25 |
.54 |
.59 |
.71 |
|
| Camk4 | 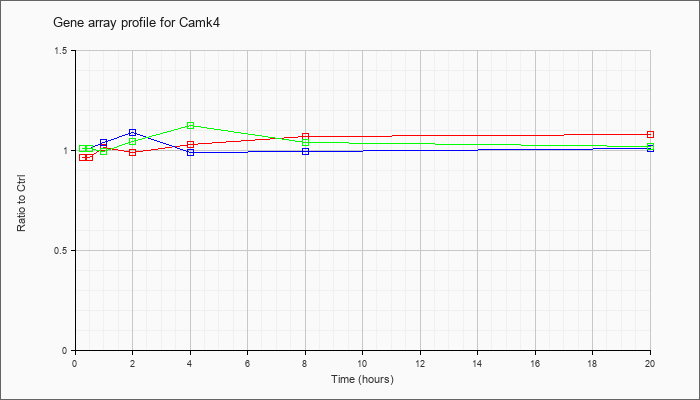 |
AK043457 |
7 days neonate cerebellum cDNA, RIKEN full-length enriched library, clone:A730097A20 product:CALCIUM/CALMODULIN-DEPENDENT PROTEIN KINASE TYPE IV CATALYTIC CHAIN (EC 2.7.1.123) (CAM KINASE-GR) (CAMK IV) [CONTAINS: CALSPERMIN] homolog [Mus |
KLA | .98 |
1.04 |
1.02 |
.93 |
1.03 |
1.07 |
1.07 |
| ATP | 1.01 |
.97 |
.97 |
1.10 |
1.00 |
.99 |
1.05 |
| KLA/ATP | 1.00 |
.96 |
1.02 |
.98 |
1.13 |
1.07 |
1.00 |
|
| Camk4 |  |
NM_009793 |
calcium/calmodulin-dependent protein kinase IV (Camk4), mRNA [NM_009793] |
KLA | .96 |
1.02 |
1.01 |
1.02 |
1.03 |
1.07 |
1.08 |
| ATP | 1.01 |
1.03 |
1.07 |
1.08 |
.99 |
1.00 |
.99 |
| KLA/ATP | 1.01 |
.95 |
.98 |
1.08 |
1.12 |
1.02 |
1.03 |
|
| Cdc42 | 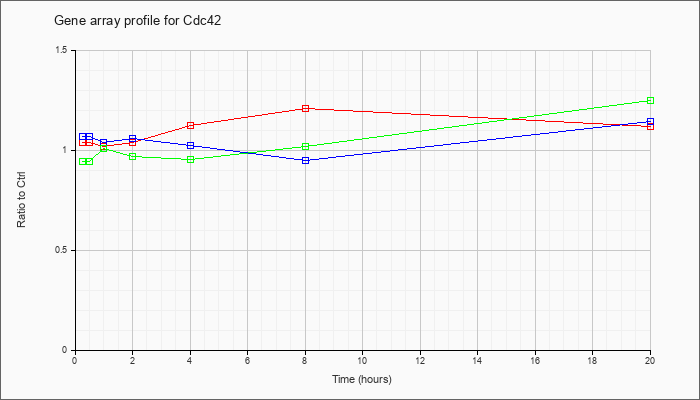 |
NM_009861 |
cell division cycle 42 homolog (S. cerevisiae) (Cdc42), mRNA [NM_009861] |
KLA | 1.04 |
1.01 |
1.02 |
1.04 |
1.12 |
1.21 |
1.12 |
| ATP | 1.07 |
1.10 |
1.04 |
1.06 |
1.02 |
.95 |
1.14 |
| KLA/ATP | .94 |
1.04 |
1.01 |
.97 |
.95 |
1.02 |
1.25 |
|
| Crk | 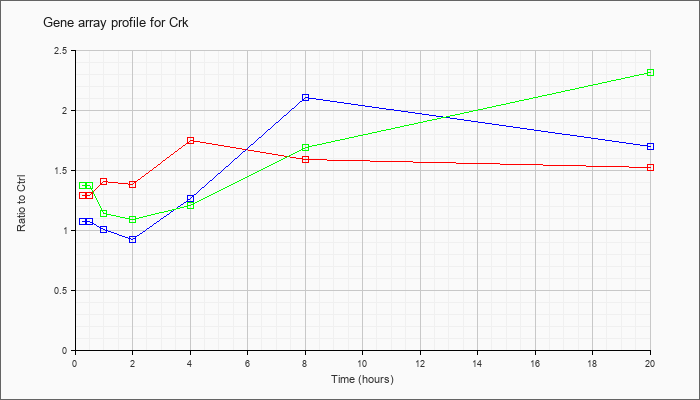 |
NM_133656 |
v-crk sarcoma virus CT10 oncogene homolog (avian) (Crk), mRNA [NM_133656] |
KLA | 1.29 |
1.32 |
1.41 |
1.38 |
1.75 |
1.59 |
1.52 |
| ATP | 1.08 |
1.17 |
1.01 |
.93 |
1.27 |
2.11 |
1.70 |
| KLA/ATP | 1.37 |
1.49 |
1.14 |
1.09 |
1.21 |
1.69 |
2.32 |
|
| Crkl | 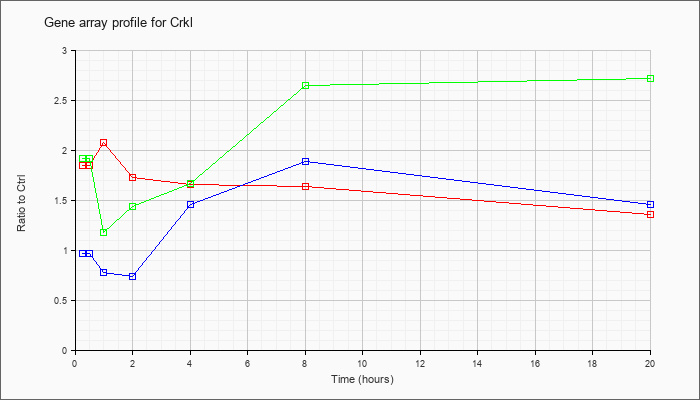 |
NM_007764 |
v-crk sarcoma virus CT10 oncogene homolog (avian)-like (Crkl), mRNA [NM_007764] |
KLA | 1.85 |
1.97 |
2.08 |
1.73 |
1.66 |
1.64 |
1.36 |
| ATP | .97 |
.94 |
.78 |
.74 |
1.46 |
1.89 |
1.46 |
| KLA/ATP | 1.92 |
1.88 |
1.18 |
1.44 |
1.67 |
2.65 |
2.72 |
|
| Fasl | 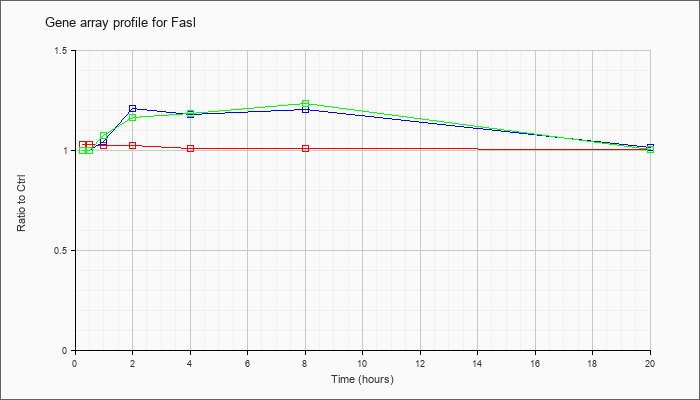 |
NM_010177 |
Fas ligand (TNF superfamily, member 6) (Fasl), mRNA [NM_010177] |
KLA | 1.03 |
1.01 |
1.03 |
1.02 |
1.01 |
1.01 |
1.00 |
| ATP | 1.00 |
.99 |
1.04 |
1.21 |
1.18 |
1.20 |
1.01 |
| KLA/ATP | 1.00 |
1.02 |
1.07 |
1.16 |
1.18 |
1.23 |
1.00 |
|
| Foxo3a | 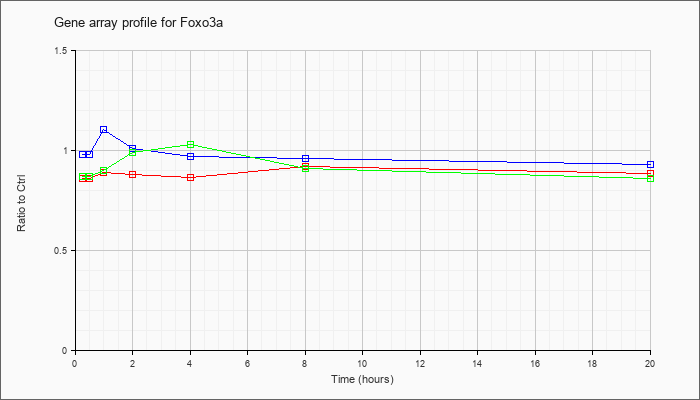 |
AK079567 |
adult male hypothalamus cDNA, RIKEN full-length enriched library, clone:A230087I01 product:unclassifiable, full insert sequence. [AK079567] |
KLA | .82 |
.81 |
.91 |
.83 |
.86 |
.86 |
.81 |
| ATP | .98 |
1.00 |
1.13 |
1.03 |
.97 |
.90 |
.82 |
| KLA/ATP | .86 |
.80 |
.90 |
.99 |
1.07 |
.84 |
.73 |
|
| Foxo3a |  |
NM_019740 |
forkhead box O3a (Foxo3a), mRNA [NM_019740] |
KLA | .94 |
.92 |
.86 |
.97 |
.87 |
1.04 |
1.04 |
| ATP | .99 |
1.09 |
1.06 |
.97 |
.96 |
1.07 |
1.15 |
| KLA/ATP | .90 |
.79 |
.90 |
.99 |
.95 |
1.04 |
1.11 |
|
| Frs2 | 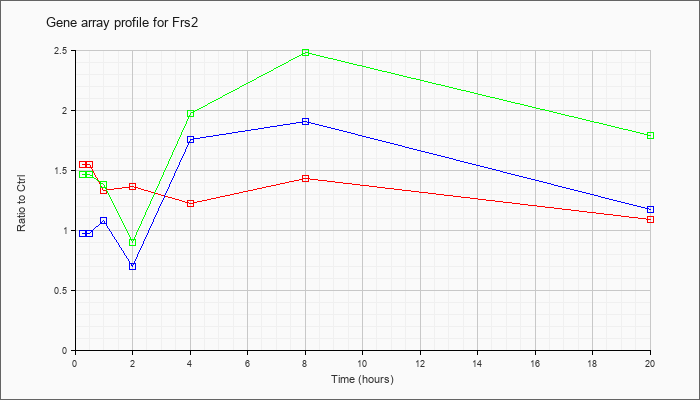 |
NM_177798 |
fibroblast growth factor receptor substrate 2 (Frs2), mRNA [NM_177798] |
KLA | 1.54 |
1.42 |
1.33 |
1.36 |
1.22 |
1.43 |
1.09 |
| ATP | .97 |
.98 |
1.08 |
.70 |
1.76 |
1.90 |
1.17 |
| KLA/ATP | 1.46 |
1.35 |
1.38 |
.89 |
1.97 |
2.48 |
1.79 |
|
| Gab1 | 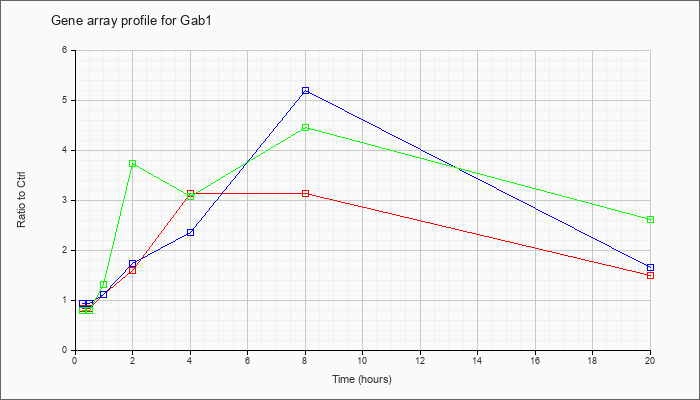 |
NM_021356 |
growth factor receptor bound protein 2-associated protein 1 (Gab1), mRNA [NM_021356] |
KLA | .83 |
.95 |
1.12 |
1.59 |
3.13 |
3.14 |
1.50 |
| ATP | .94 |
.96 |
1.12 |
1.74 |
2.35 |
5.20 |
1.66 |
| KLA/ATP | .79 |
.78 |
1.32 |
3.74 |
3.08 |
4.44 |
2.62 |
|
| Grb2 | 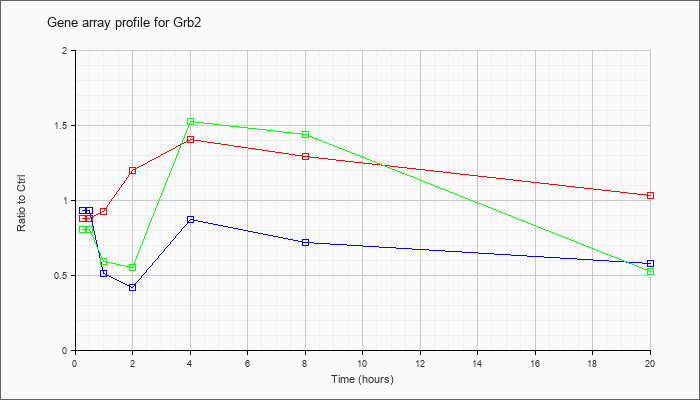 |
NM_008163 |
growth factor receptor bound protein 2 (Grb2), mRNA [NM_008163] |
KLA | .92 |
.88 |
.90 |
1.27 |
1.43 |
1.22 |
1.00 |
| ATP | 1.02 |
.83 |
.51 |
.42 |
.92 |
.70 |
.57 |
| KLA/ATP | .84 |
.77 |
.61 |
.56 |
1.56 |
1.49 |
.52 |
|
| Grb2 |  |
U07617 |
Grb2 adaptor protein (grb2) mRNA, complete cds. [U07617] |
KLA | .83 |
.80 |
.95 |
1.13 |
1.38 |
1.36 |
1.06 |
| ATP | .84 |
.74 |
.51 |
.41 |
.82 |
.74 |
.59 |
| KLA/ATP | .77 |
.70 |
.57 |
.54 |
1.49 |
1.39 |
.53 |
|
| Gsk3b | 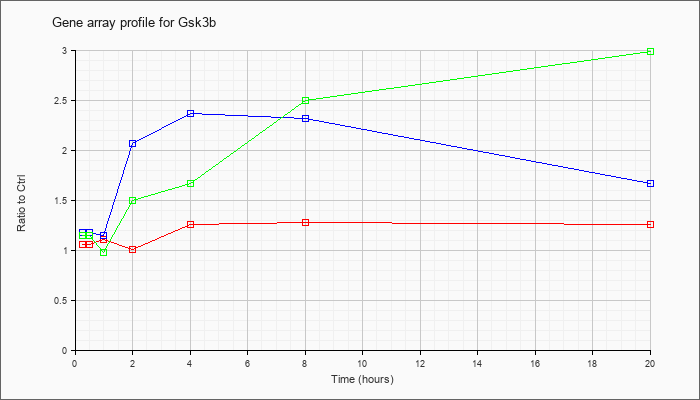 |
NM_019827 |
glycogen synthase kinase 3 beta (Gsk3b), mRNA [NM_019827] |
KLA | 1.06 |
1.03 |
1.11 |
1.01 |
1.26 |
1.28 |
1.26 |
| ATP | 1.18 |
1.27 |
1.15 |
2.07 |
2.37 |
2.32 |
1.67 |
| KLA/ATP | 1.15 |
1.21 |
.98 |
1.50 |
1.67 |
2.50 |
2.99 |
|
| Hras1 | 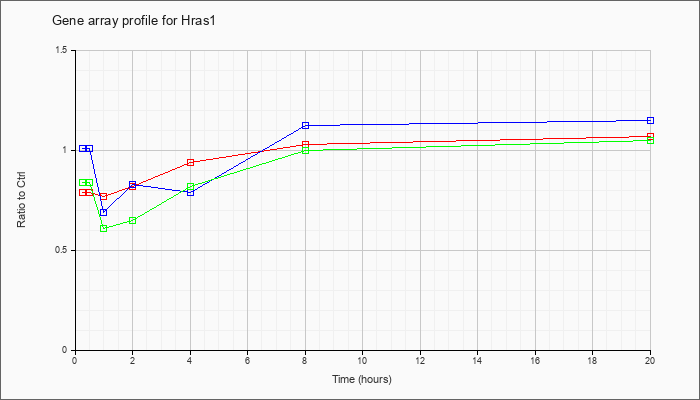 |
NM_008284 |
Harvey rat sarcoma virus oncogene 1 (Hras1), mRNA [NM_008284] |
KLA | .79 |
.82 |
.77 |
.82 |
.94 |
1.03 |
1.07 |
| ATP | 1.01 |
.95 |
.69 |
.83 |
.79 |
1.12 |
1.15 |
| KLA/ATP | .84 |
.77 |
.61 |
.65 |
.82 |
1.00 |
1.05 |
|
| Ikbkb | 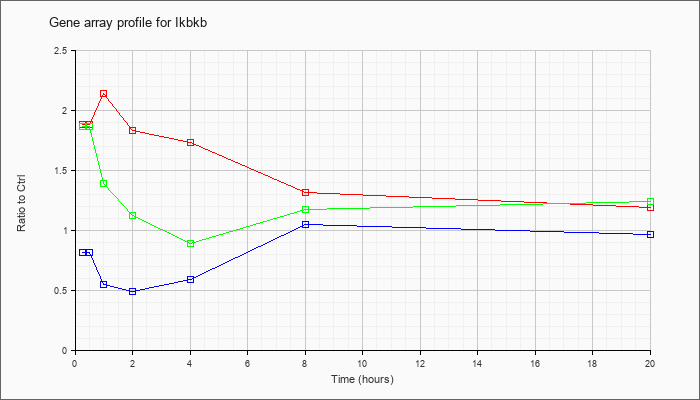 |
NM_010546 |
inhibitor of kappaB kinase beta (Ikbkb), mRNA [NM_010546] |
KLA | 1.88 |
2.10 |
2.14 |
1.83 |
1.73 |
1.31 |
1.19 |
| ATP | .81 |
.74 |
.55 |
.49 |
.59 |
1.05 |
.96 |
| KLA/ATP | 1.86 |
1.73 |
1.39 |
1.12 |
.89 |
1.17 |
1.24 |
|
| Irak1 | 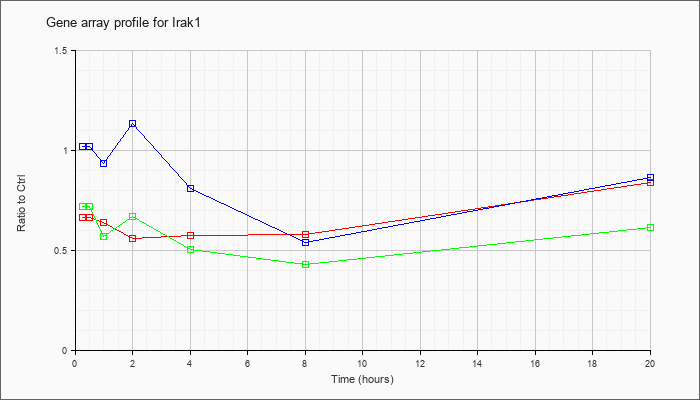 |
NM_008363 |
interleukin-1 receptor-associated kinase 1 (Irak1), mRNA [NM_008363] |
KLA | .67 |
.67 |
.64 |
.56 |
.58 |
.58 |
.84 |
| ATP | 1.02 |
1.10 |
.94 |
1.14 |
.81 |
.54 |
.87 |
| KLA/ATP | .72 |
.72 |
.57 |
.67 |
.51 |
.43 |
.62 |
|
| Irak2 | 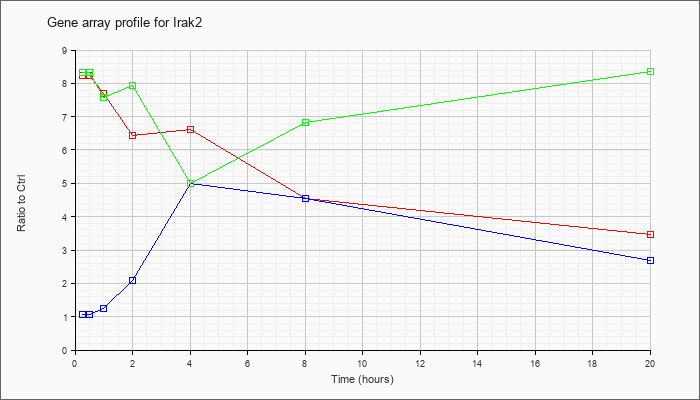 |
NM_172161 |
interleukin-1 receptor-associated kinase 2 (Irak2), transcript variant 1, mRNA [NM_172161] |
KLA | 8.24 |
8.89 |
7.70 |
6.43 |
6.61 |
4.53 |
3.48 |
| ATP | 1.06 |
1.22 |
1.26 |
2.10 |
4.99 |
4.55 |
2.69 |
| KLA/ATP | 8.32 |
9.13 |
7.57 |
7.95 |
5.01 |
6.84 |
8.36 |
|
| Irak3 |  |
NM_028679 |
interleukin-1 receptor-associated kinase 3 (Irak3), mRNA [NM_028679] |
KLA | 2.04 |
2.22 |
2.80 |
3.81 |
6.02 |
4.81 |
6.76 |
| ATP | 1.13 |
1.14 |
1.02 |
1.26 |
.97 |
2.51 |
1.31 |
| KLA/ATP | 2.24 |
2.35 |
2.06 |
2.58 |
1.43 |
1.55 |
4.63 |
|
| Irak4 | 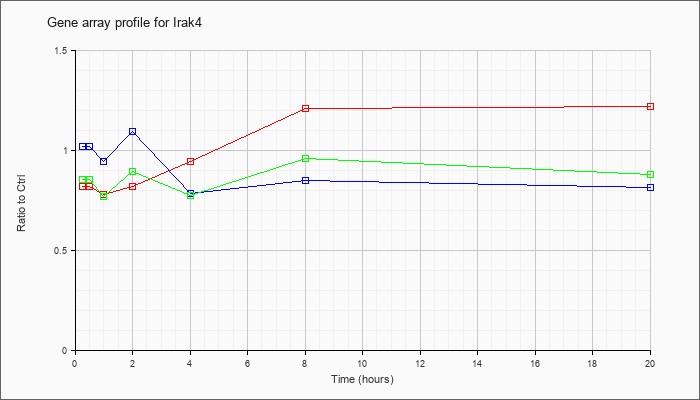 |
NM_029926 |
interleukin-1 receptor-associated kinase 4 (Irak4), mRNA [NM_029926] |
KLA | .82 |
.86 |
.78 |
.82 |
.95 |
1.21 |
1.22 |
| ATP | 1.02 |
1.18 |
.95 |
1.10 |
.79 |
.85 |
.82 |
| KLA/ATP | .86 |
.89 |
.77 |
.90 |
.78 |
.96 |
.88 |
|
| Irs1 | 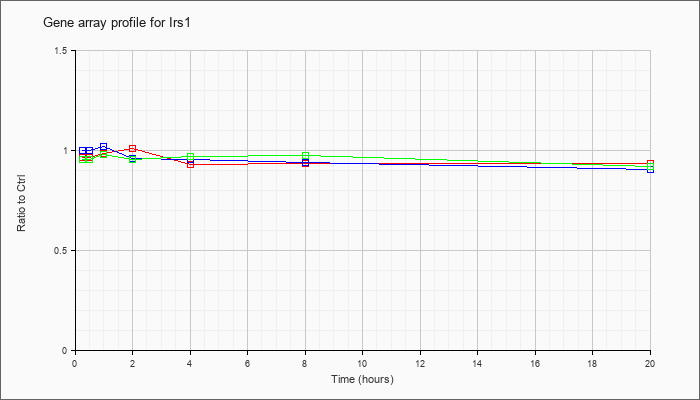 |
NM_010570 |
insulin receptor substrate 1 (Irs1), mRNA [NM_010570] |
KLA | .97 |
.96 |
.99 |
1.01 |
.93 |
.94 |
.94 |
| ATP | 1.00 |
.99 |
1.02 |
.96 |
.96 |
.94 |
.91 |
| KLA/ATP | .96 |
.96 |
.98 |
.96 |
.97 |
.98 |
.92 |
|
| Jun | 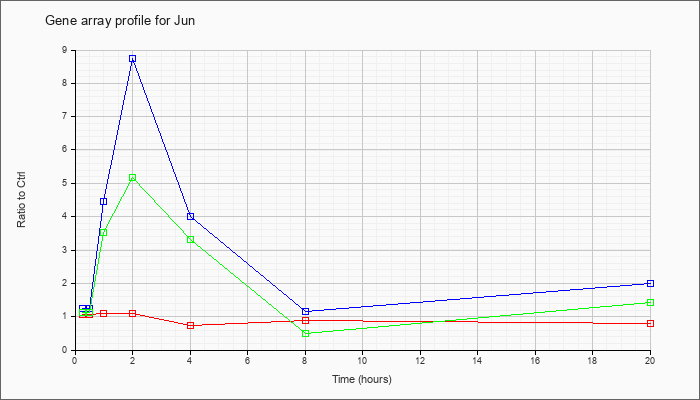 |
NM_010591 |
Jun oncogene (Jun), mRNA [NM_010591] |
KLA | 1.08 |
.96 |
1.09 |
1.09 |
.75 |
.88 |
.81 |
| ATP | 1.26 |
1.48 |
4.45 |
8.73 |
4.02 |
1.16 |
1.99 |
| KLA/ATP | 1.12 |
1.38 |
3.52 |
5.17 |
3.32 |
.49 |
1.42 |
|
| Kras | 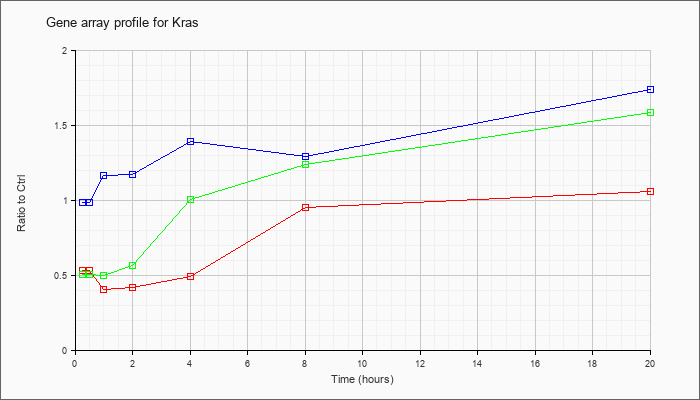 |
NM_021284 |
v-Ki-ras2 Kirsten rat sarcoma viral oncogene homolog (Kras), mRNA [NM_021284] |
KLA | .53 |
.49 |
.40 |
.42 |
.49 |
.95 |
1.05 |
| ATP | .98 |
1.01 |
1.16 |
1.17 |
1.39 |
1.29 |
1.74 |
| KLA/ATP | .50 |
.47 |
.50 |
.56 |
1.00 |
1.24 |
1.58 |
|
| Maged1 | 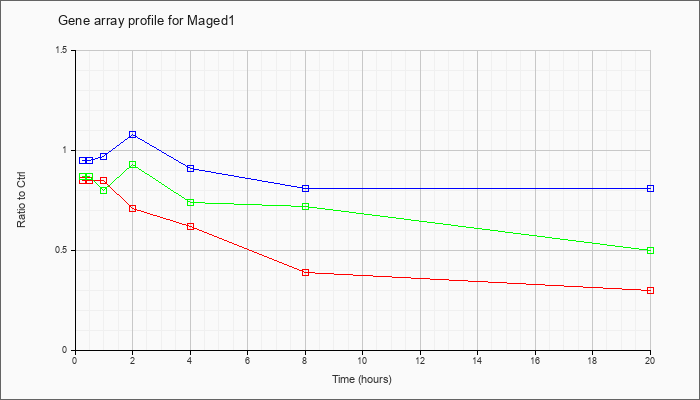 |
NM_019791 |
melanoma antigen, family D, 1 (Maged1), mRNA [NM_019791] |
KLA | .85 |
.87 |
.85 |
.71 |
.62 |
.39 |
.30 |
| ATP | .95 |
1.01 |
.97 |
1.08 |
.91 |
.81 |
.81 |
| KLA/ATP | .87 |
.91 |
.80 |
.93 |
.74 |
.72 |
.50 |
|
| Map2k1 | 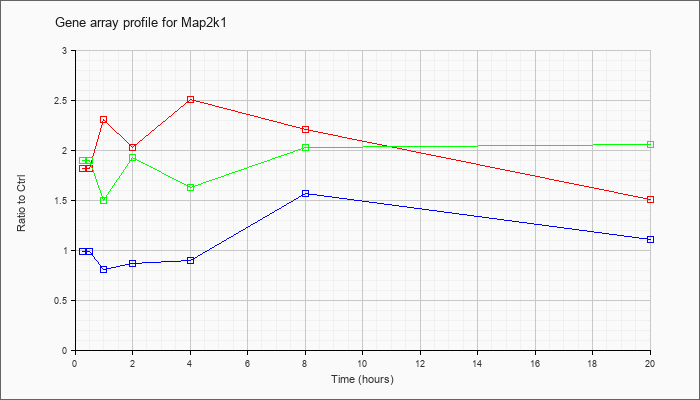 |
NM_008927 |
mitogen-activated protein kinase kinase 1 (Map2k1), mRNA [NM_008927] |
KLA | 1.82 |
2.00 |
2.31 |
2.03 |
2.51 |
2.20 |
1.51 |
| ATP | .99 |
1.14 |
.81 |
.87 |
.90 |
1.57 |
1.11 |
| KLA/ATP | 1.90 |
2.03 |
1.50 |
1.93 |
1.63 |
2.03 |
2.06 |
|
| Map2k2 | 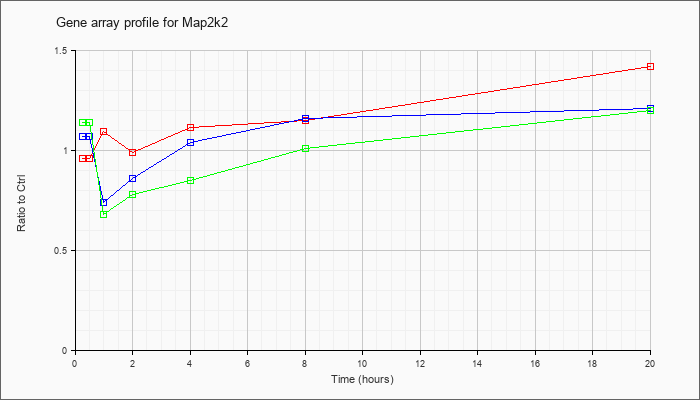 |
NM_023138 |
mitogen-activated protein kinase kinase 2 (Map2k2), mRNA [NM_023138] |
KLA | .96 |
1.01 |
1.09 |
.99 |
1.11 |
1.15 |
1.42 |
| ATP | 1.07 |
1.05 |
.74 |
.86 |
1.04 |
1.16 |
1.21 |
| KLA/ATP | 1.14 |
1.03 |
.68 |
.78 |
.85 |
1.01 |
1.20 |
|
| Map2k5 | 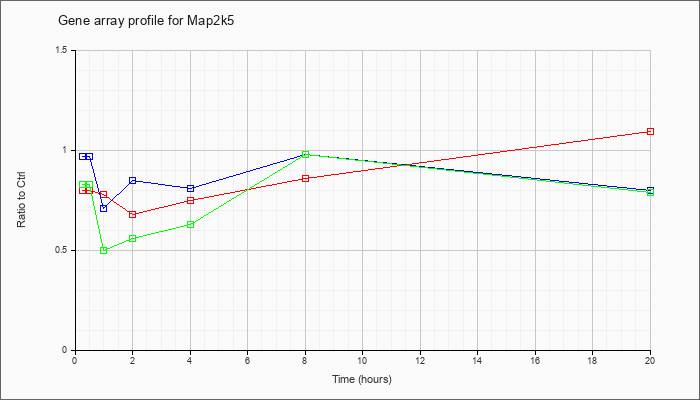 |
NM_011840 |
mitogen-activated protein kinase kinase 5 (Map2k5), mRNA [NM_011840] |
KLA | .80 |
.86 |
.78 |
.68 |
.75 |
.86 |
1.09 |
| ATP | .97 |
.90 |
.71 |
.85 |
.81 |
.98 |
.80 |
| KLA/ATP | .83 |
.73 |
.50 |
.56 |
.63 |
.98 |
.79 |
|
| Map2k7 | 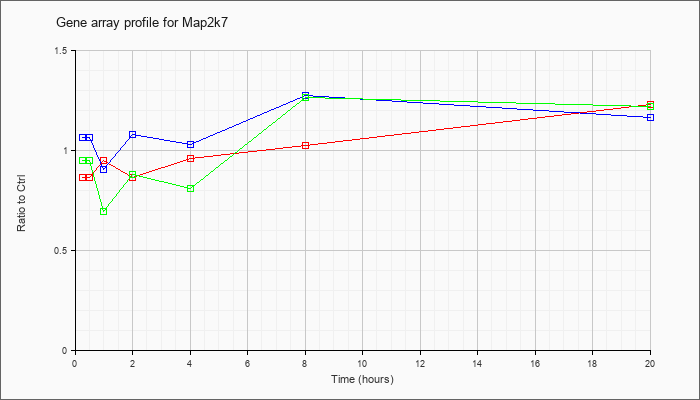 |
BC070467 |
mitogen activated protein kinase kinase 7, mRNA (cDNA clone MGC:96654 IMAGE:30615302), complete cds [BC070467] |
KLA | .77 |
.79 |
.88 |
.77 |
.97 |
.96 |
1.27 |
| ATP | 1.11 |
1.26 |
.93 |
1.19 |
1.13 |
1.45 |
1.16 |
| KLA/ATP | .89 |
.88 |
.59 |
.89 |
.86 |
1.32 |
1.28 |
|
| Map2k7 |  |
NM_001042557 |
mitogen-activated protein kinase kinase 7 (Map2k7), transcript variant 1, mRNA [NM_001042557] |
KLA | .96 |
.95 |
1.02 |
.96 |
.95 |
1.09 |
1.19 |
| ATP | 1.02 |
1.07 |
.88 |
.97 |
.93 |
1.10 |
1.17 |
| KLA/ATP | 1.01 |
.99 |
.80 |
.87 |
.76 |
1.21 |
1.16 |
|
| Map3k1 | 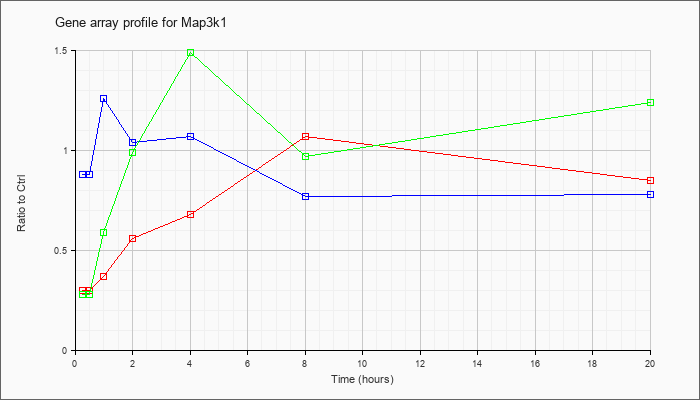 |
NM_011945 |
mitogen-activated protein kinase kinase kinase 1 (Map3k1), mRNA [NM_011945] |
KLA | .30 |
.25 |
.37 |
.56 |
.68 |
1.07 |
.85 |
| ATP | .88 |
.86 |
1.26 |
1.04 |
1.07 |
.77 |
.78 |
| KLA/ATP | .28 |
.24 |
.59 |
.99 |
1.49 |
.97 |
1.24 |
|
| Map3k3 | 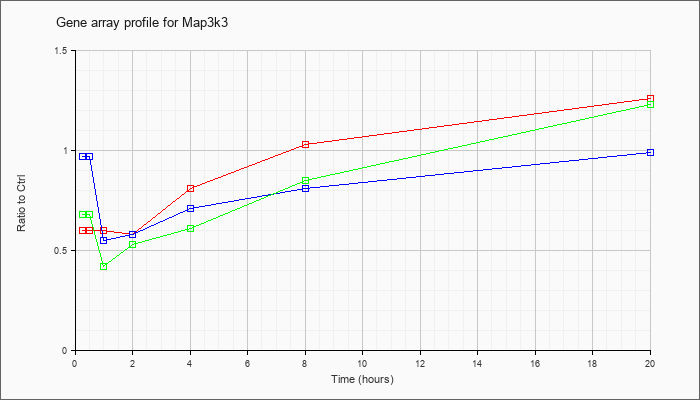 |
NM_011947 |
mitogen-activated protein kinase kinase kinase 3 (Map3k3), mRNA [NM_011947] |
KLA | .60 |
.63 |
.60 |
.58 |
.81 |
1.03 |
1.26 |
| ATP | .97 |
.91 |
.55 |
.58 |
.71 |
.81 |
.99 |
| KLA/ATP | .68 |
.57 |
.42 |
.53 |
.61 |
.85 |
1.23 |
|
| Map3k5 | 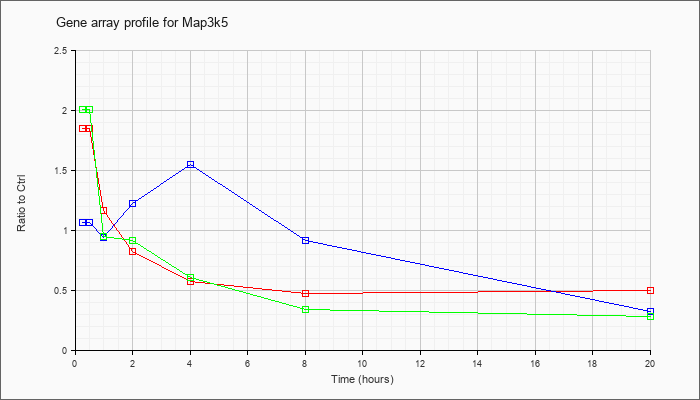 |
NM_008580 |
mitogen-activated protein kinase kinase kinase 5 (Map3k5), mRNA [NM_008580] |
KLA | 1.85 |
1.97 |
1.17 |
.82 |
.57 |
.47 |
.49 |
| ATP | 1.06 |
1.05 |
.94 |
1.22 |
1.55 |
.91 |
.32 |
| KLA/ATP | 2.00 |
1.82 |
.95 |
.91 |
.60 |
.34 |
.28 |
|
| Mapk1 | 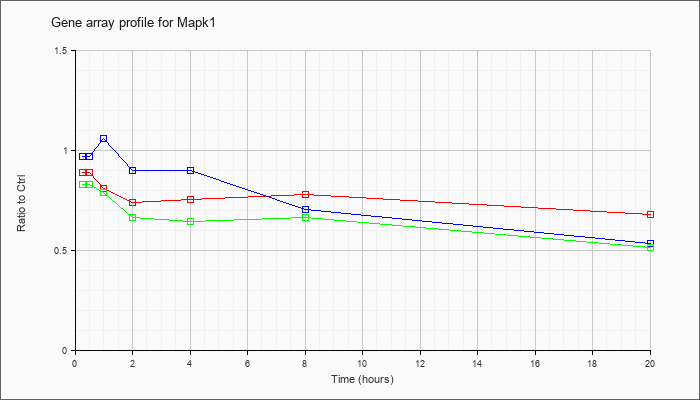 |
NM_001038663 |
mitogen-activated protein kinase 1 (Mapk1), transcript variant 2, mRNA [NM_001038663] |
KLA | .97 |
1.03 |
.93 |
.86 |
.79 |
1.01 |
1.04 |
| ATP | .96 |
1.18 |
1.01 |
1.13 |
.84 |
1.04 |
1.16 |
| KLA/ATP | .99 |
.99 |
.84 |
.87 |
.63 |
.96 |
1.26 |
|
| Mapk1 |  |
NM_011949 |
mitogen-activated protein kinase 1 (Mapk1), transcript variant 1, mRNA [NM_011949] |
KLA | .88 |
.80 |
.80 |
.73 |
.75 |
.76 |
.65 |
| ATP | .97 |
.96 |
1.06 |
.88 |
.90 |
.68 |
.48 |
| KLA/ATP | .82 |
.81 |
.78 |
.65 |
.64 |
.64 |
.45 |
|
| Mapk10 | 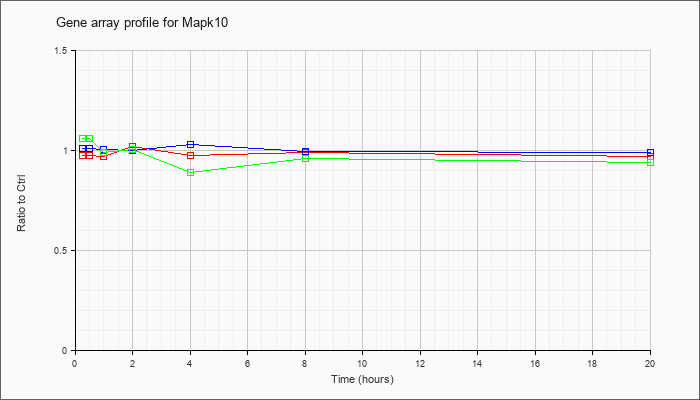 |
AK076990 |
adult male testis cDNA, RIKEN full-length enriched library, clone:4930597C01 product:mitogen activated protein kinase 10, full insert sequence. [AK076990] |
KLA | 1.03 |
1.05 |
.98 |
1.04 |
.94 |
.95 |
1.02 |
| ATP | 1.00 |
.99 |
1.06 |
.97 |
1.01 |
.96 |
.99 |
| KLA/ATP | 1.07 |
1.01 |
.95 |
1.05 |
.82 |
.95 |
.87 |
|
| Mapk10 |  |
NM_009158 |
mitogen-activated protein kinase 10 (Mapk10), transcript variant 1, mRNA [NM_009158] |
KLA | .95 |
1.00 |
.97 |
1.01 |
.99 |
1.01 |
.95 |
| ATP | 1.01 |
1.02 |
.98 |
1.01 |
1.04 |
1.01 |
.99 |
| KLA/ATP | 1.05 |
.99 |
1.01 |
.98 |
.92 |
.97 |
.98 |
|
| Mapk11 | 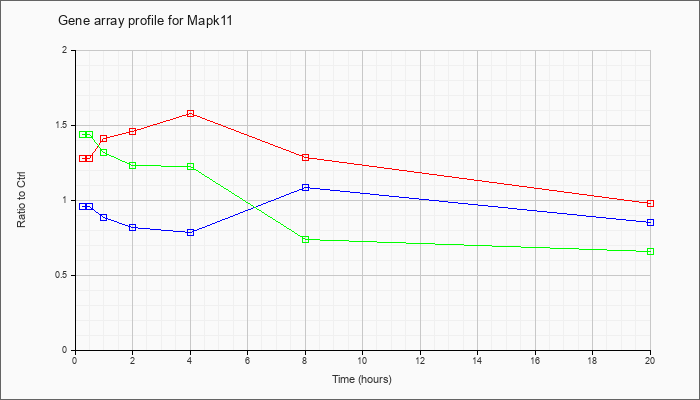 |
NM_011161 |
mitogen-activated protein kinase 11 (Mapk11), mRNA [NM_011161] |
KLA | 1.28 |
1.30 |
1.41 |
1.46 |
1.58 |
1.29 |
.98 |
| ATP | .96 |
.95 |
.89 |
.82 |
.79 |
1.09 |
.85 |
| KLA/ATP | 1.44 |
1.42 |
1.32 |
1.23 |
1.23 |
.74 |
.66 |
|
| Mapk12 | 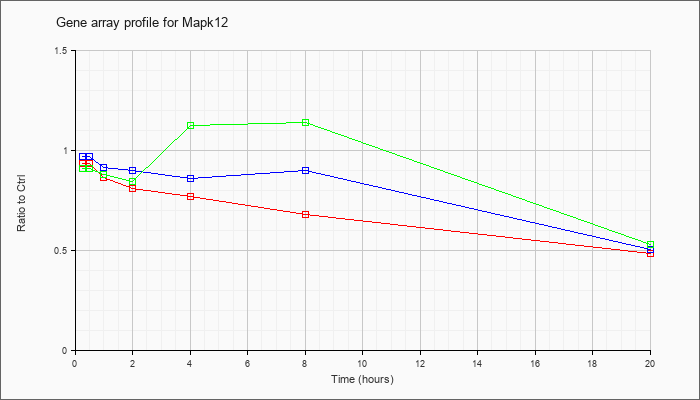 |
NM_013871 |
mitogen-activated protein kinase 12 (Mapk12), mRNA [NM_013871] |
KLA | .94 |
.86 |
.87 |
.81 |
.77 |
.68 |
.49 |
| ATP | .97 |
.83 |
.92 |
.90 |
.86 |
.90 |
.51 |
| KLA/ATP | .91 |
.89 |
.88 |
.85 |
1.12 |
1.14 |
.53 |
|
| Mapk13 | 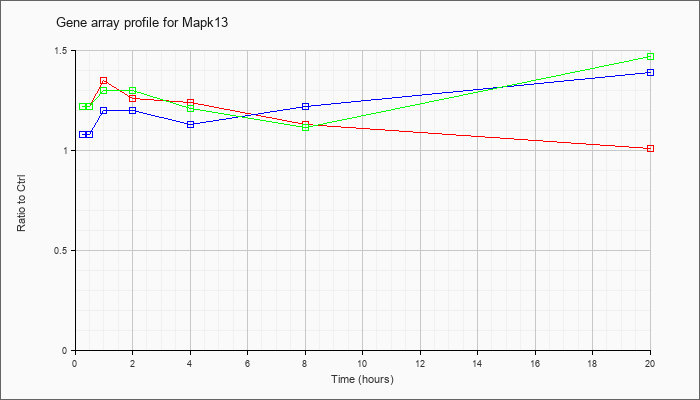 |
NM_011950 |
mitogen-activated protein kinase 13 (Mapk13), mRNA [NM_011950] |
KLA | 1.22 |
1.21 |
1.35 |
1.26 |
1.24 |
1.13 |
1.01 |
| ATP | 1.08 |
1.00 |
1.20 |
1.20 |
1.13 |
1.22 |
1.39 |
| KLA/ATP | 1.22 |
1.27 |
1.30 |
1.30 |
1.21 |
1.11 |
1.47 |
|
| Mapk14 | 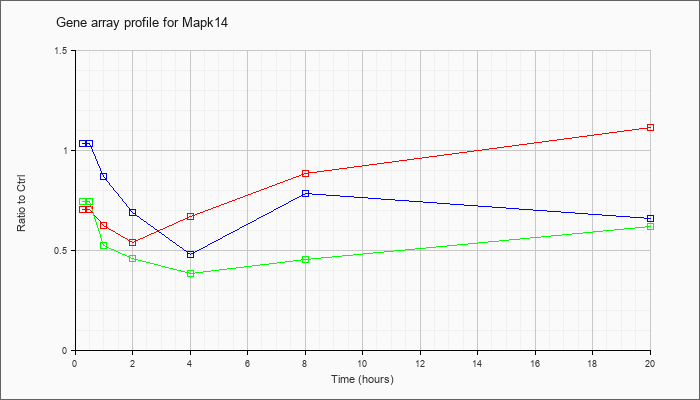 |
NM_011951 |
mitogen-activated protein kinase 14 (Mapk14), mRNA [NM_011951] |
KLA | .70 |
.70 |
.62 |
.54 |
.67 |
.88 |
1.11 |
| ATP | 1.03 |
.98 |
.87 |
.69 |
.48 |
.78 |
.66 |
| KLA/ATP | .74 |
.68 |
.52 |
.46 |
.38 |
.45 |
.62 |
|
| Mapk3 | 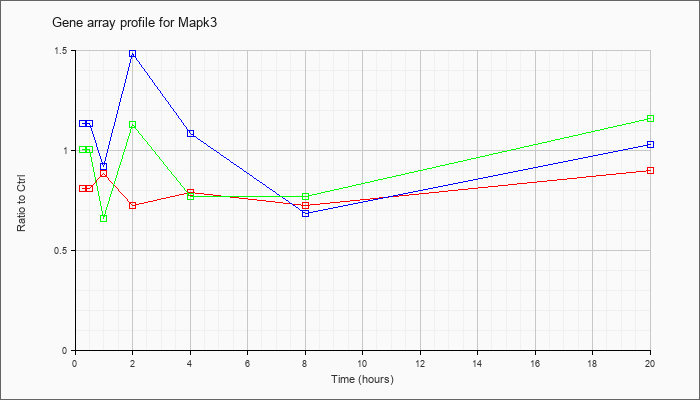 |
NM_011952 |
mitogen-activated protein kinase 3 (Mapk3), mRNA [NM_011952] |
KLA | .81 |
.85 |
.88 |
.72 |
.79 |
.73 |
.90 |
| ATP | 1.13 |
1.30 |
.92 |
1.48 |
1.08 |
.68 |
1.03 |
| KLA/ATP | 1.01 |
1.01 |
.66 |
1.13 |
.77 |
.77 |
1.16 |
|
| Mapk7 | 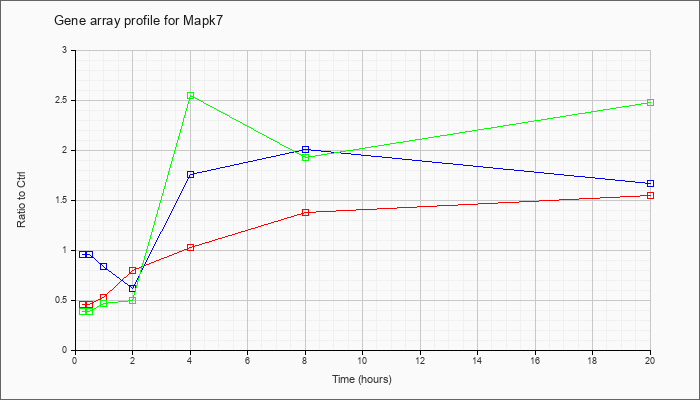 |
NM_011841 |
mitogen-activated protein kinase 7 (Mapk7), mRNA [NM_011841] |
KLA | .46 |
.43 |
.53 |
.80 |
1.03 |
1.38 |
1.55 |
| ATP | .96 |
.80 |
.84 |
.62 |
1.76 |
2.01 |
1.67 |
| KLA/ATP | .39 |
.40 |
.47 |
.50 |
2.55 |
1.93 |
2.48 |
|
| Mapk8 |  |
AK162915 |
adult male spinal cord cDNA, RIKEN full-length enriched library, clone:A330031O19 product:mitogen activated protein kinase 8, full insert sequence [AK162915] |
KLA | .94 |
.86 |
.80 |
.83 |
.77 |
.85 |
.83 |
| ATP | 1.08 |
1.12 |
.94 |
.80 |
.83 |
.71 |
.75 |
| KLA/ATP | .83 |
.81 |
.80 |
.74 |
.74 |
.75 |
.67 |
|
| Mapk8 |  |
AK163829 |
16 days embryo head cDNA, RIKEN full-length enriched library, clone:C130070A06 product:mitogen activated protein kinase 8, full insert sequence [AK163829] |
KLA | .65 |
.61 |
.66 |
.70 |
.90 |
1.02 |
1.03 |
| ATP | .99 |
.94 |
1.68 |
1.44 |
1.75 |
1.84 |
1.10 |
| KLA/ATP | .63 |
.67 |
1.11 |
1.00 |
2.01 |
1.68 |
1.15 |
|
| Mapk8 |  |
NM_016700 |
mitogen-activated protein kinase 8 (Mapk8), mRNA [NM_016700] |
KLA | .96 |
1.00 |
.94 |
.96 |
.94 |
.99 |
1.04 |
| ATP | 1.01 |
1.05 |
1.01 |
.99 |
1.03 |
.98 |
1.02 |
| KLA/ATP | .96 |
.97 |
.96 |
.97 |
.93 |
.94 |
.96 |
|
| Mapk9 | 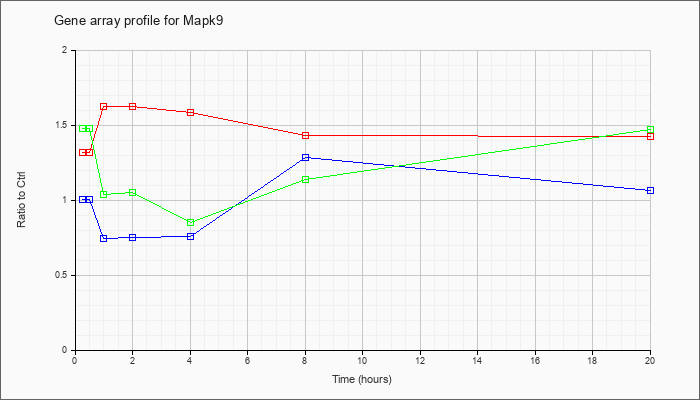 |
NM_016961 |
mitogen-activated protein kinase 9 (Mapk9), transcript variant 2, mRNA [NM_016961] |
KLA | 1.28 |
1.37 |
1.59 |
1.60 |
1.61 |
1.43 |
1.38 |
| ATP | 1.03 |
.99 |
.74 |
.71 |
.79 |
1.20 |
1.01 |
| KLA/ATP | 1.53 |
1.30 |
1.07 |
.99 |
.83 |
1.10 |
1.40 |
|
| Mapk9 |  |
NM_207692 |
mitogen-activated protein kinase 9 (Mapk9), transcript variant 1, mRNA [NM_207692] |
KLA | 1.39 |
1.53 |
1.69 |
1.69 |
1.55 |
1.44 |
1.51 |
| ATP | .95 |
.92 |
.75 |
.85 |
.70 |
1.46 |
1.17 |
| KLA/ATP | 1.37 |
1.35 |
.99 |
1.18 |
.91 |
1.22 |
1.62 |
|
Mapkapk2
| 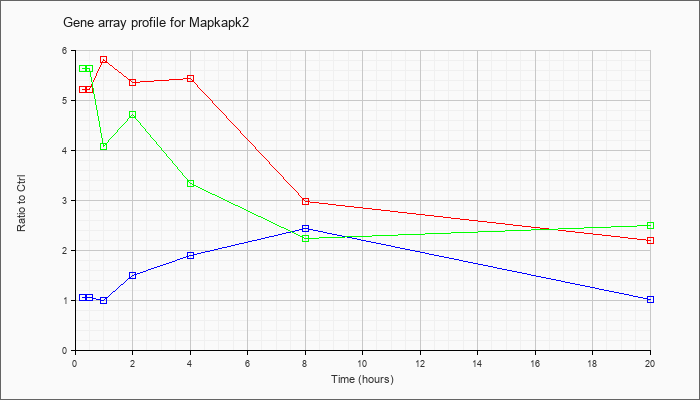 |
NM_008551 |
MAP kinase-activated protein kinase 2 (Mapkapk2), mRNA [NM_008551] |
KLA | 5.20 |
5.49 |
5.81 |
5.35 |
5.43 |
2.98 |
2.19 |
| ATP | 1.04 |
1.12 |
1.00 |
1.50 |
1.88 |
2.43 |
1.01 |
| KLA/ATP | 5.63 |
5.95 |
4.08 |
4.71 |
3.33 |
2.24 |
2.50 |
|
| Matk | 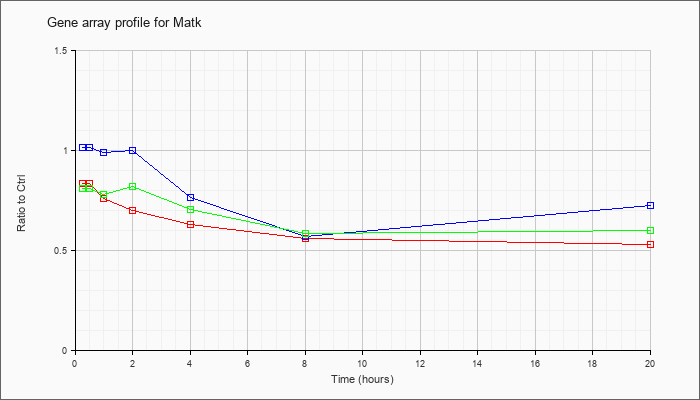 |
NM_010768 |
megakaryocyte-associated tyrosine kinase (Matk), mRNA [NM_010768] |
KLA | .84 |
.77 |
.76 |
.70 |
.63 |
.56 |
.53 |
| ATP | 1.02 |
1.06 |
.99 |
1.00 |
.77 |
.57 |
.73 |
| KLA/ATP | .81 |
.78 |
.78 |
.82 |
.71 |
.59 |
.60 |
|
Mm.13153
0 | 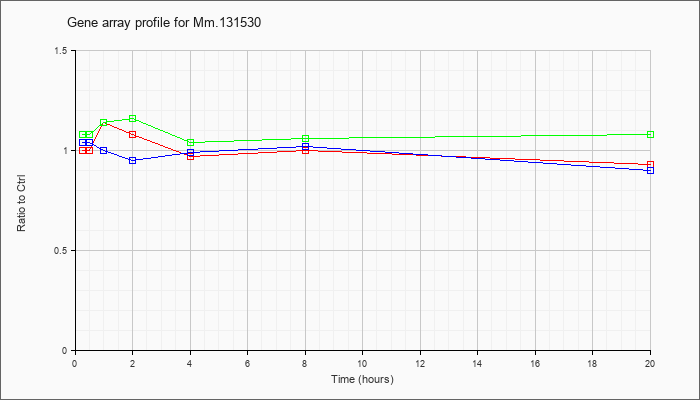 |
161086915 |
Unknown |
KLA | 1.00 |
1.07 |
1.14 |
1.08 |
.97 |
1.00 |
.93 |
| ATP | 1.04 |
.99 |
1.00 |
.95 |
.99 |
1.02 |
.90 |
| KLA/ATP | 1.08 |
1.18 |
1.14 |
1.16 |
1.04 |
1.06 |
1.08 |
|
Mm.21312
8 | 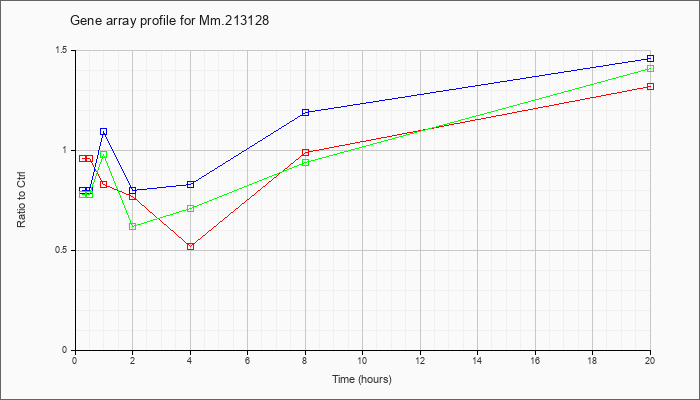 |
141801344 |
Unknown |
KLA | .96 |
.95 |
.83 |
.77 |
.52 |
.99 |
1.32 |
| ATP | .80 |
.82 |
1.09 |
.80 |
.83 |
1.19 |
1.46 |
| KLA/ATP | .78 |
.92 |
.98 |
.62 |
.71 |
.94 |
1.41 |
|
Mm.25381
9 | 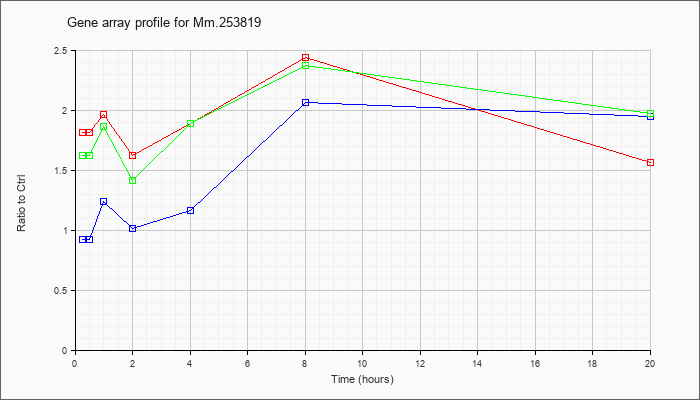 |
118130676 |
Unknown |
KLA | 1.81 |
1.88 |
1.96 |
1.62 |
1.89 |
2.44 |
1.56 |
| ATP | .92 |
.91 |
1.24 |
1.01 |
1.16 |
2.06 |
1.95 |
| KLA/ATP | 1.62 |
1.86 |
1.86 |
1.41 |
1.89 |
2.37 |
1.97 |
|
Mm.27578
| 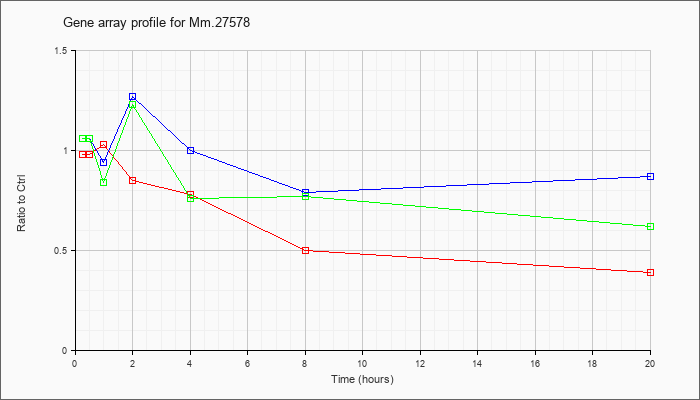 |
118130592 |
Unknown |
KLA | .98 |
1.03 |
1.03 |
.85 |
.78 |
.50 |
.39 |
| ATP | 1.06 |
1.15 |
.94 |
1.27 |
1.00 |
.79 |
.87 |
| KLA/ATP | 1.06 |
1.10 |
.84 |
1.23 |
.76 |
.77 |
.62 |
|
Mm.27970
| 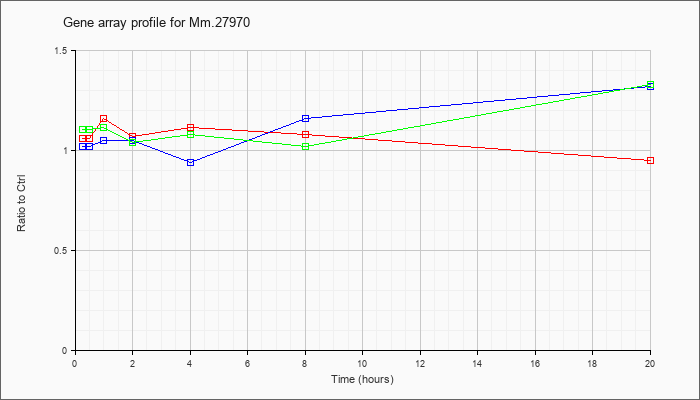 |
13994135 |
Unknown |
KLA | 1.06 |
1.11 |
1.16 |
1.07 |
1.11 |
1.08 |
.95 |
| ATP | 1.02 |
1.09 |
1.05 |
1.05 |
.94 |
1.16 |
1.32 |
| KLA/ATP | 1.10 |
1.19 |
1.11 |
1.04 |
1.08 |
1.02 |
1.33 |
|
Mm.32574
6 | 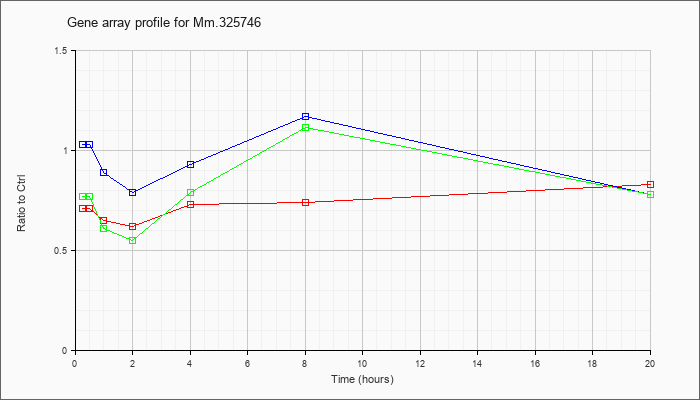 |
118130017 |
Unknown |
KLA | .71 |
.67 |
.65 |
.62 |
.73 |
.74 |
.83 |
| ATP | 1.03 |
.83 |
.89 |
.79 |
.93 |
1.17 |
.78 |
| KLA/ATP | .77 |
.62 |
.61 |
.55 |
.79 |
1.11 |
.78 |
|
Mm.33085
0 | 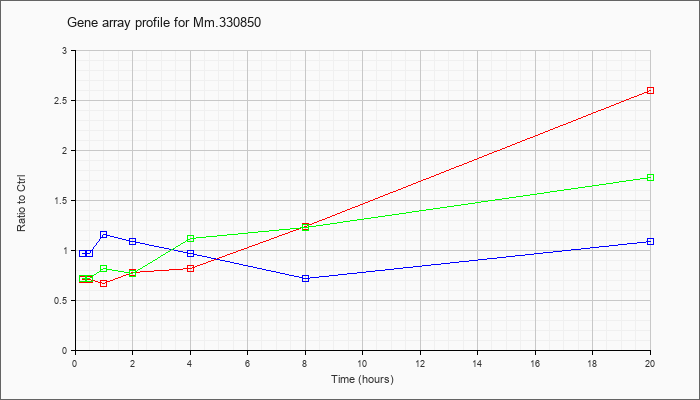 |
190684662 |
Unknown |
KLA | .71 |
.70 |
.67 |
.78 |
.82 |
1.24 |
2.60 |
| ATP | .97 |
.98 |
1.16 |
1.09 |
.97 |
.72 |
1.09 |
| KLA/ATP | .72 |
.73 |
.82 |
.77 |
1.12 |
1.23 |
1.73 |
|
| Mm.4387 |  |
133892666 |
Unknown |
KLA | .85 |
.87 |
.85 |
.89 |
.77 |
.91 |
1.18 |
| ATP | 1.01 |
1.01 |
.87 |
1.07 |
.75 |
.86 |
.93 |
| KLA/ATP | .88 |
.86 |
.77 |
.81 |
.72 |
.91 |
.84 |
|
Mm.44463
|  |
118129851 |
Unknown |
KLA | 1.01 |
.98 |
.97 |
1.02 |
.93 |
1.06 |
1.05 |
| ATP | .97 |
1.01 |
1.04 |
1.06 |
1.02 |
1.08 |
.97 |
| KLA/ATP | 1.02 |
1.06 |
.95 |
1.04 |
1.09 |
1.14 |
1.03 |
|
Mm.46002
4 | 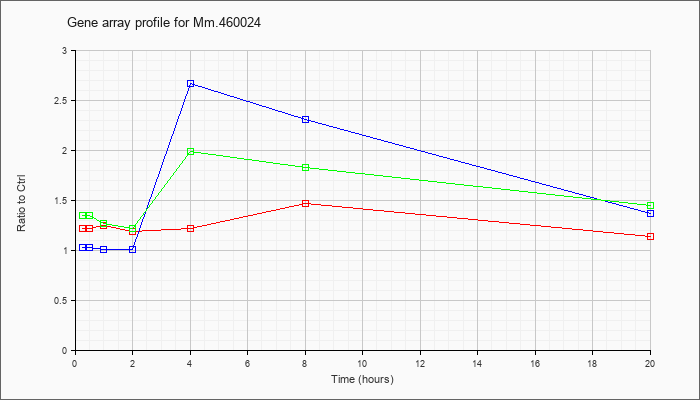 |
38348245 |
Unknown |
KLA | 1.22 |
1.25 |
1.25 |
1.19 |
1.22 |
1.47 |
1.14 |
| ATP | 1.03 |
.94 |
1.01 |
1.01 |
2.67 |
2.31 |
1.37 |
| KLA/ATP | 1.35 |
1.19 |
1.27 |
1.22 |
1.99 |
1.83 |
1.45 |
|
Mm.80682
| 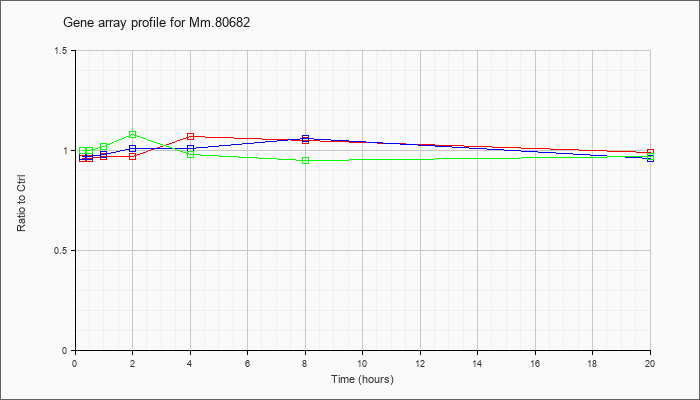 |
94369627 |
Unknown |
KLA | .96 |
1.00 |
.97 |
.97 |
1.07 |
1.05 |
.99 |
| ATP | .97 |
1.02 |
.98 |
1.01 |
1.01 |
1.06 |
.96 |
| KLA/ATP | 1.00 |
.93 |
1.02 |
1.08 |
.98 |
.95 |
.97 |
|
| Mm.998 |  |
118130584 |
Unknown |
KLA | 1.14 |
1.17 |
1.21 |
1.06 |
1.22 |
1.16 |
1.13 |
| ATP | 1.02 |
1.07 |
1.06 |
.97 |
.92 |
1.01 |
.98 |
| KLA/ATP | 1.05 |
1.17 |
1.22 |
1.24 |
1.13 |
1.07 |
1.12 |
|
| Nfkb1 | 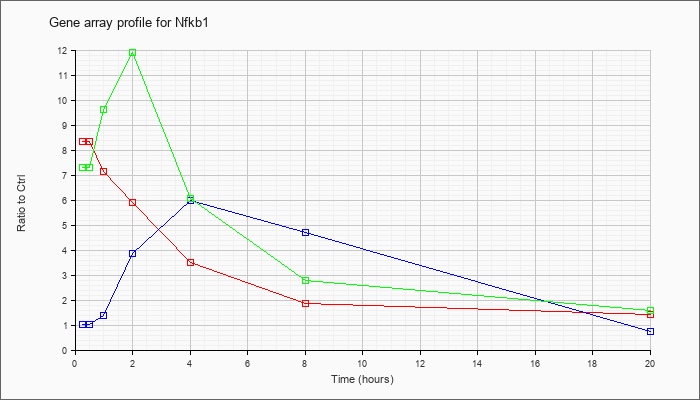 |
BC050841 |
nuclear factor of kappa light chain gene enhancer in B-cells 1, p105, mRNA (cDNA clone IMAGE:6392793), complete cds. [BC050841] |
KLA | 5.81 |
4.96 |
4.77 |
3.21 |
3.37 |
2.00 |
1.71 |
| ATP | 1.39 |
1.52 |
3.25 |
12.63 |
13.64 |
4.80 |
1.08 |
| KLA/ATP | 7.21 |
8.69 |
19.53 |
23.93 |
11.70 |
2.85 |
2.01 |
|
| Nfkb1 |  |
M57999 |
gb|Mouse transcription factor NF-kappa-B DNA binding subunit mRNA, complete cds. [M57999] |
KLA | 8.55 |
7.87 |
7.35 |
6.13 |
3.51 |
1.85 |
1.38 |
| ATP | .98 |
.96 |
1.21 |
3.06 |
5.29 |
4.67 |
.69 |
| KLA/ATP | 7.32 |
7.20 |
8.71 |
10.82 |
5.55 |
2.77 |
1.54 |
|
| Nfkbia | 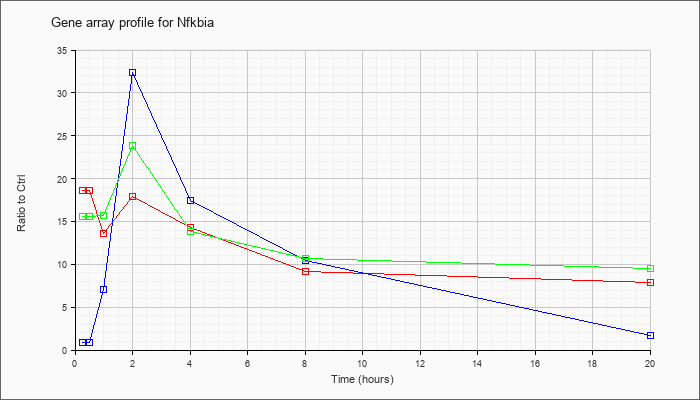 |
NM_010907 |
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha (Nfkbia), mRNA [NM_010907] |
KLA | 18.65 |
18.46 |
13.63 |
17.88 |
14.25 |
9.17 |
7.92 |
| ATP | .91 |
1.09 |
7.11 |
32.39 |
17.44 |
10.49 |
1.72 |
| KLA/ATP | 15.57 |
17.66 |
15.73 |
23.91 |
13.83 |
10.73 |
9.48 |
|
| Nfkbib | 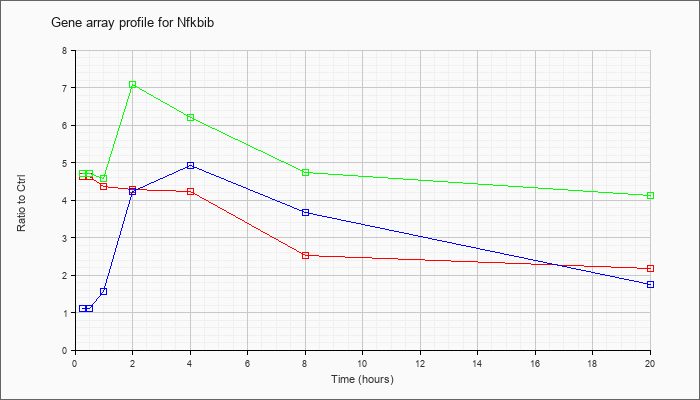 |
NM_010908 |
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, beta (Nfkbib), mRNA [NM_010908] |
KLA | 4.63 |
4.69 |
4.36 |
4.27 |
4.23 |
2.52 |
2.17 |
| ATP | 1.11 |
1.22 |
1.55 |
4.23 |
4.93 |
3.66 |
1.76 |
| KLA/ATP | 4.71 |
5.41 |
4.58 |
7.08 |
6.20 |
4.73 |
4.11 |
|
| Nfkbie | 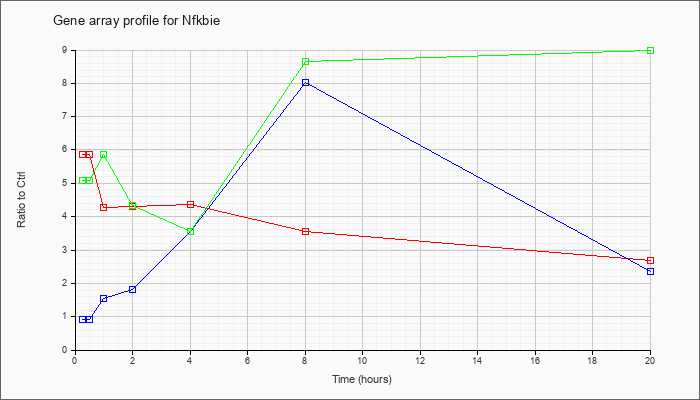 |
NM_008690 |
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, epsilon (Nfkbie), mRNA [NM_008690] |
KLA | 5.87 |
5.26 |
4.28 |
4.30 |
4.38 |
3.56 |
2.69 |
| ATP | .92 |
1.04 |
1.55 |
1.83 |
3.57 |
8.04 |
2.35 |
| KLA/ATP | 5.07 |
5.19 |
5.86 |
4.34 |
3.56 |
8.64 |
9.00 |
|
| Ngf | 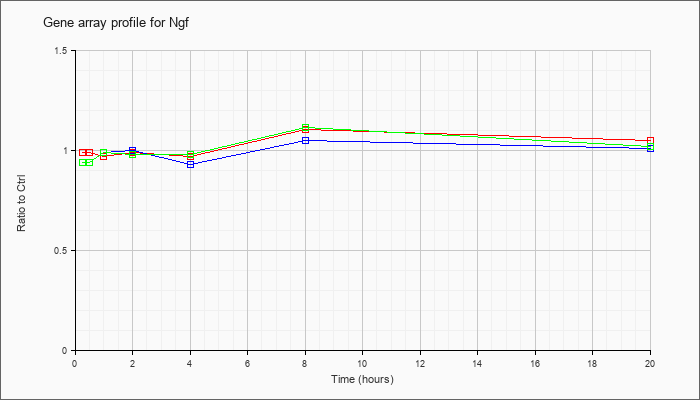 |
NM_013609 |
nerve growth factor (Ngf), transcript variant A, mRNA [NM_013609] |
KLA | .99 |
.98 |
.97 |
.99 |
.97 |
1.10 |
1.05 |
| ATP | .94 |
1.03 |
.99 |
1.00 |
.93 |
1.05 |
1.01 |
| KLA/ATP | .94 |
1.01 |
.99 |
.98 |
.98 |
1.11 |
1.02 |
|
| Ngfr | 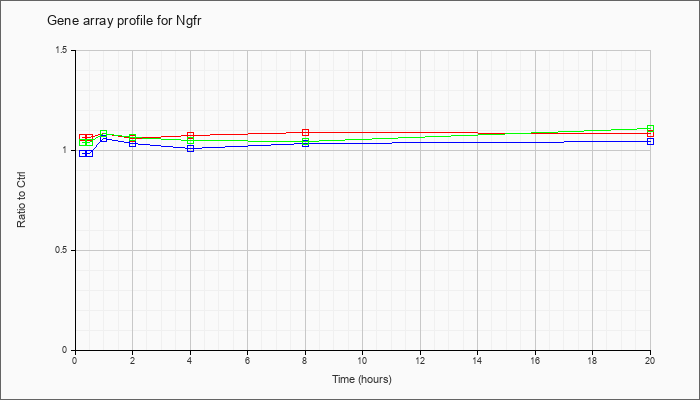 |
NM_033217 |
nerve growth factor receptor (TNFR superfamily, member 16) (Ngfr), mRNA [NM_033217] |
KLA | 1.06 |
1.08 |
1.08 |
1.06 |
1.07 |
1.09 |
1.08 |
| ATP | .99 |
1.01 |
1.06 |
1.03 |
1.01 |
1.04 |
1.04 |
| KLA/ATP | 1.04 |
1.07 |
1.08 |
1.06 |
1.05 |
1.04 |
1.11 |
|
| Ngfrap1 | 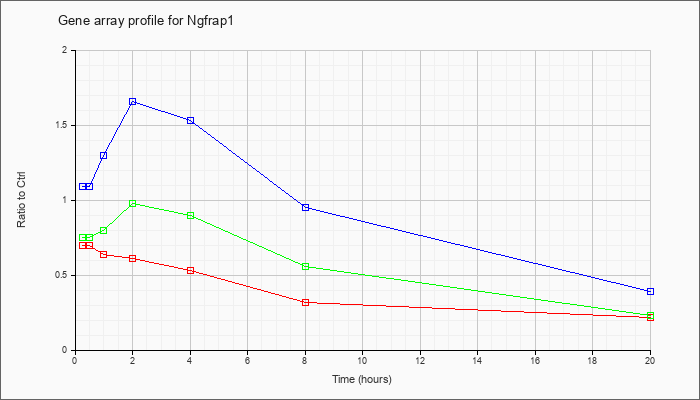 |
NM_009750 |
nerve growth factor receptor (TNFRSF16) associated protein 1 (Ngfrap1), transcript variant 1, mRNA [NM_009750] |
KLA | .70 |
.74 |
.64 |
.61 |
.53 |
.32 |
.22 |
| ATP | 1.09 |
1.15 |
1.30 |
1.66 |
1.53 |
.95 |
.39 |
| KLA/ATP | .75 |
.71 |
.80 |
.98 |
.90 |
.56 |
.23 |
|
| Nras | 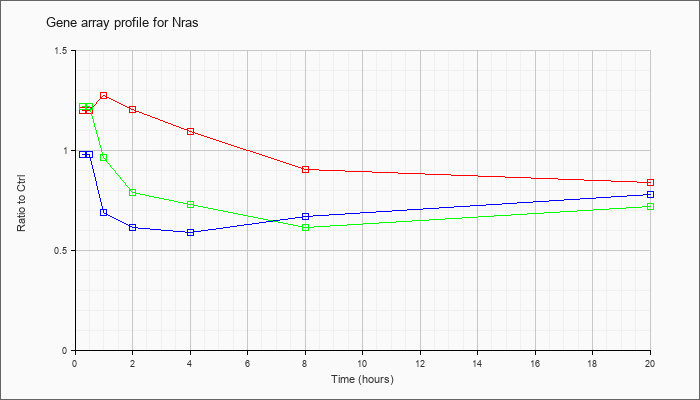 |
NM_010937 |
neuroblastoma ras oncogene (Nras), mRNA [NM_010937] |
KLA | 1.20 |
1.22 |
1.27 |
1.20 |
1.09 |
.91 |
.84 |
| ATP | .98 |
.91 |
.69 |
.61 |
.59 |
.67 |
.78 |
| KLA/ATP | 1.22 |
1.14 |
.97 |
.79 |
.73 |
.61 |
.72 |
|
| Ntf3 | 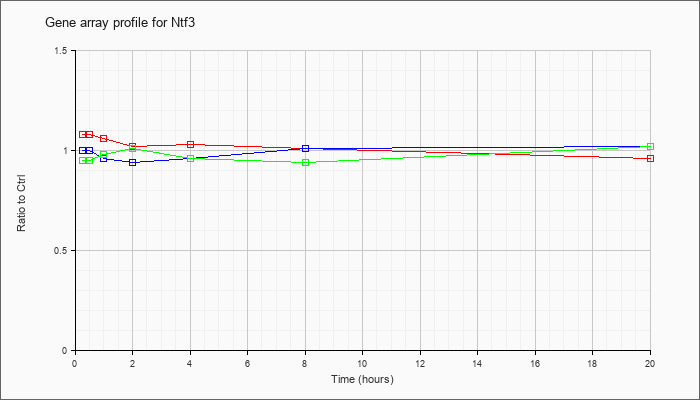 |
NM_008742 |
neurotrophin 3 (Ntf3), mRNA [NM_008742] |
KLA | 1.08 |
.93 |
1.06 |
1.02 |
1.03 |
1.01 |
.96 |
| ATP | 1.00 |
1.03 |
.96 |
.94 |
.96 |
1.01 |
1.02 |
| KLA/ATP | .95 |
1.00 |
.98 |
1.01 |
.96 |
.94 |
1.02 |
|
| Ntf5 | 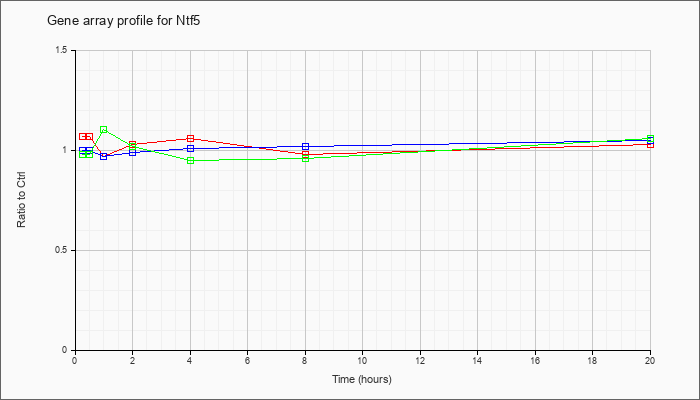 |
NM_198190 |
neurotrophin 5 (Ntf5), mRNA [NM_198190] |
KLA | 1.07 |
1.02 |
.97 |
1.03 |
1.06 |
.98 |
1.03 |
| ATP | 1.00 |
1.05 |
.97 |
.99 |
1.01 |
1.02 |
1.05 |
| KLA/ATP | .98 |
1.00 |
1.10 |
1.02 |
.95 |
.96 |
1.06 |
|
| Ntrk1 | 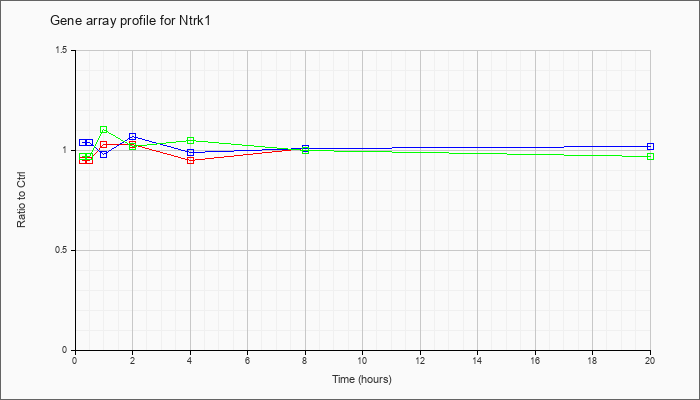 |
NM_001033124 |
neurotrophic tyrosine kinase, receptor, type 1 (Ntrk1), mRNA [NM_001033124] |
KLA | .95 |
.96 |
1.03 |
1.03 |
.95 |
1.01 |
1.02 |
| ATP | 1.04 |
.98 |
.98 |
1.07 |
.99 |
1.01 |
1.02 |
| KLA/ATP | .97 |
1.03 |
1.10 |
1.02 |
1.05 |
1.00 |
.97 |
|
| Ntrk2 |  |
AK018789 |
adult male cerebellum cDNA, RIKEN full-length enriched library, clone:1500040I13 product:neurotrophic tyrosine kinase, receptor, type 2, full insert sequence. [AK018789] |
KLA | 1.02 |
.94 |
1.05 |
1.03 |
1.03 |
1.03 |
1.04 |
| ATP | .99 |
1.01 |
1.09 |
.95 |
1.02 |
.99 |
1.04 |
| KLA/ATP | 1.02 |
.96 |
1.00 |
1.04 |
1.13 |
.99 |
1.03 |
|
| Ntrk2 |  |
NM_001025074 |
neurotrophic tyrosine kinase, receptor, type 2 (Ntrk2), transcript variant 1, mRNA [NM_001025074] |
KLA | .99 |
.99 |
1.02 |
1.02 |
1.02 |
1.00 |
1.03 |
| ATP | .99 |
.99 |
.99 |
.99 |
1.00 |
1.01 |
1.03 |
| KLA/ATP | .97 |
1.00 |
1.01 |
1.00 |
.99 |
.98 |
1.00 |
|
| Ntrk2 |  |
NM_008745 |
neurotrophic tyrosine kinase, receptor, type 2 (Ntrk2), transcript variant 2, mRNA [NM_008745] |
KLA | .99 |
1.00 |
.99 |
1.00 |
.99 |
1.00 |
1.02 |
| ATP | 1.01 |
1.00 |
1.01 |
.97 |
1.01 |
1.01 |
.99 |
| KLA/ATP | .99 |
1.01 |
.99 |
1.01 |
.99 |
.99 |
1.01 |
|
| Ntrk3 |  |
NM_182809 |
neurotrophic tyrosine kinase, receptor, type 3 (Ntrk3), transcript variant 2, mRNA [NM_182809] |
KLA | 1.09 |
1.12 |
1.11 |
1.12 |
1.05 |
1.17 |
1.06 |
| ATP | 1.03 |
1.02 |
1.06 |
1.07 |
1.02 |
1.01 |
1.02 |
| KLA/ATP | 1.07 |
1.07 |
1.26 |
1.01 |
1.13 |
1.08 |
1.07 |
|
| Pdk1 | 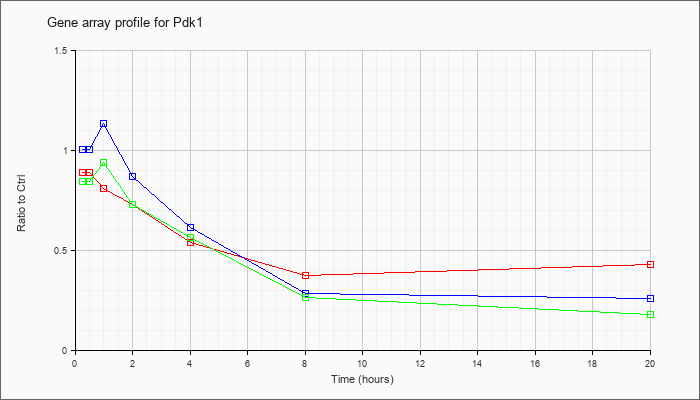 |
AK165090 |
2 days pregnant adult female ovary cDNA, RIKEN full-length enriched library, clone:E330005M21 product:hypothetical protein, full insert sequence. [AK165090] |
KLA | .86 |
.77 |
.71 |
.67 |
.45 |
.29 |
.35 |
| ATP | 1.06 |
.88 |
1.21 |
.72 |
.60 |
.26 |
.15 |
| KLA/ATP | .81 |
.74 |
.97 |
.57 |
.51 |
.22 |
.09 |
|
| Pdk1 |  |
NM_172665 |
pyruvate dehydrogenase kinase, isoenzyme 1 (Pdk1), nuclear gene encoding mitochondrial protein, mRNA [NM_172665] |
KLA | .92 |
.94 |
.91 |
.79 |
.63 |
.46 |
.51 |
| ATP | .95 |
1.03 |
1.06 |
1.02 |
.63 |
.31 |
.37 |
| KLA/ATP | .88 |
.91 |
.91 |
.89 |
.62 |
.31 |
.27 |
|
| Pik3ca | 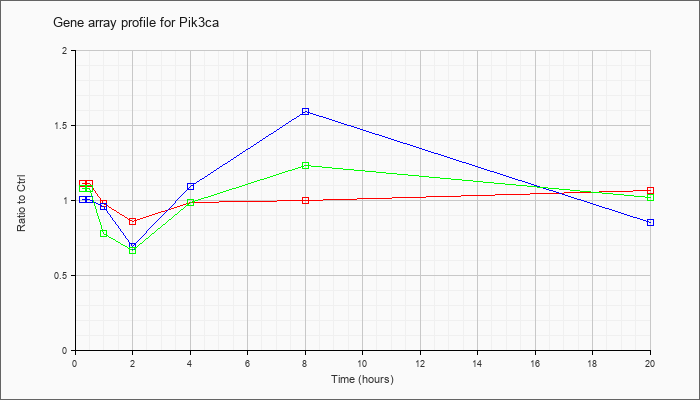 |
ENSMUST00000108243 |
ens|Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha isoform (EC 2.7.1.153) (PI3-kinase p110 subunit alpha) (PtdIns-3- kinase p110) (PI3K). [Source:Uniprot/SWISSPROT;Acc:P42337] [ENSMUST00000108243] |
KLA | 1.07 |
1.01 |
.91 |
.90 |
.98 |
.96 |
.86 |
| ATP | 1.03 |
.87 |
1.12 |
.63 |
1.22 |
1.86 |
.60 |
| KLA/ATP | 1.05 |
.82 |
.91 |
.61 |
1.17 |
1.26 |
.63 |
|
| Pik3ca |  |
NM_008839 |
phosphatidylinositol 3-kinase, catalytic, alpha polypeptide (Pik3ca), mRNA [NM_008839] |
KLA | 1.15 |
1.11 |
1.05 |
.82 |
.99 |
1.03 |
1.27 |
| ATP | .98 |
1.00 |
.79 |
.75 |
.96 |
1.32 |
1.10 |
| KLA/ATP | 1.10 |
.98 |
.64 |
.72 |
.80 |
1.20 |
1.41 |
|
| Pik3cb | 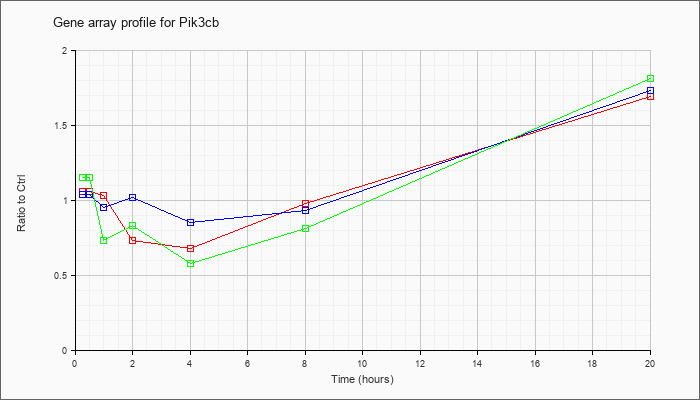 |
NM_029094 |
phosphatidylinositol 3-kinase, catalytic, beta polypeptide (Pik3cb), mRNA [NM_029094] |
KLA | 1.06 |
1.06 |
1.03 |
.73 |
.68 |
.98 |
1.69 |
| ATP | 1.04 |
1.21 |
.95 |
1.02 |
.85 |
.93 |
1.73 |
| KLA/ATP | 1.15 |
1.27 |
.73 |
.83 |
.58 |
.81 |
1.81 |
|
| Pik3cd | 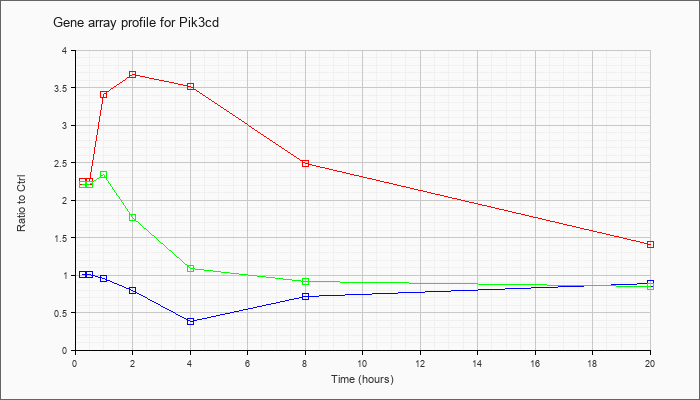 |
NM_008840 |
phosphatidylinositol 3-kinase catalytic delta polypeptide (Pik3cd), transcript variant 1, mRNA [NM_008840] |
KLA | 2.28 |
2.42 |
3.10 |
4.17 |
3.35 |
2.31 |
1.34 |
| ATP | 1.02 |
.91 |
.99 |
.63 |
.30 |
.67 |
.80 |
| KLA/ATP | 2.14 |
2.23 |
2.70 |
1.51 |
1.17 |
.85 |
.72 |
|
| Pik3cd |  |
U86587 |
phosphatidylinositol 3-kinase catalytic subunit p110 delta mRNA, complete cds. [U86587] |
KLA | 2.21 |
2.24 |
3.71 |
3.17 |
3.67 |
2.66 |
1.48 |
| ATP | 1.00 |
1.03 |
.93 |
.96 |
.46 |
.75 |
.97 |
| KLA/ATP | 2.27 |
2.32 |
1.97 |
2.02 |
1.01 |
.99 |
.98 |
|
| Pik3cg | 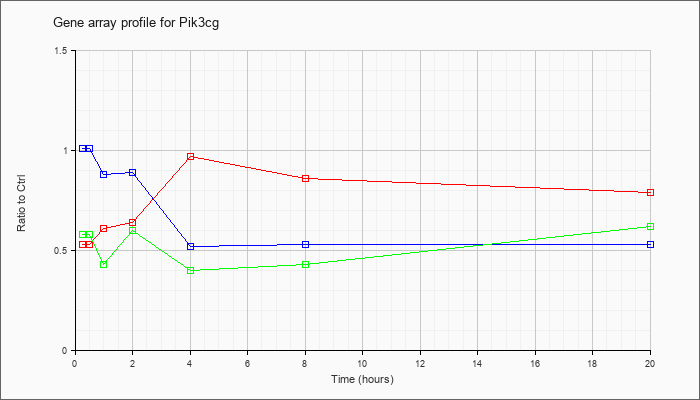 |
NM_020272 |
phosphoinositide-3-kinase, catalytic, gamma polypeptide (Pik3cg), mRNA [NM_020272] |
KLA | .53 |
.53 |
.61 |
.64 |
.97 |
.86 |
.79 |
| ATP | 1.01 |
1.13 |
.88 |
.89 |
.52 |
.53 |
.53 |
| KLA/ATP | .58 |
.53 |
.43 |
.60 |
.40 |
.43 |
.62 |
|
| Pik3r1 | 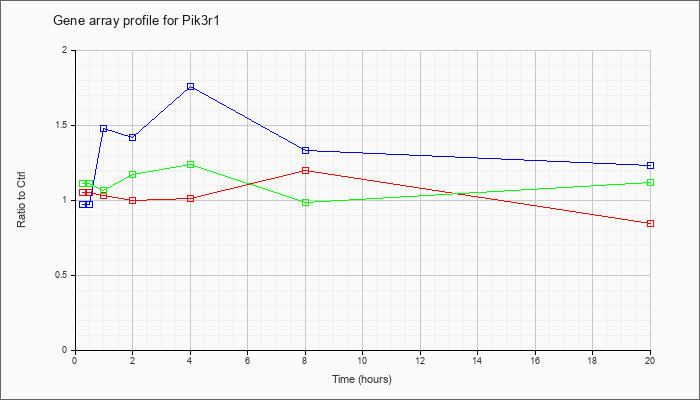 |
NM_001077495 |
phosphatidylinositol 3-kinase, regulatory subunit, polypeptide 1 (p85 alpha) (Pik3r1), transcript variant 2, mRNA [NM_001077495] |
KLA | 1.05 |
1.07 |
1.03 |
1.00 |
1.01 |
1.20 |
.85 |
| ATP | .97 |
1.02 |
1.48 |
1.42 |
1.76 |
1.33 |
1.23 |
| KLA/ATP | 1.11 |
1.12 |
1.06 |
1.17 |
1.24 |
.99 |
1.12 |
|
| Pik3r2 | 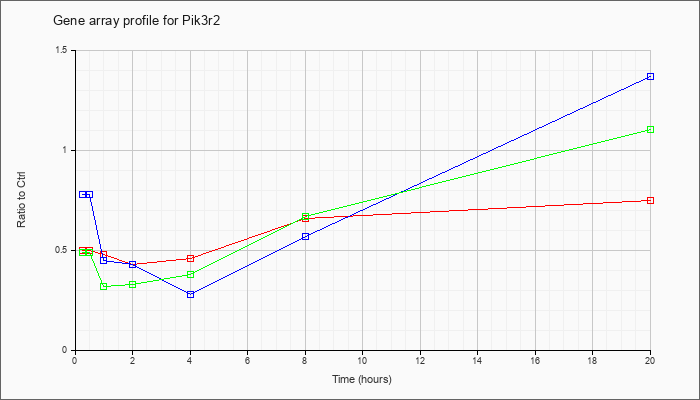 |
NM_008841 |
phosphatidylinositol 3-kinase, regulatory subunit, polypeptide 2 (p85 beta) (Pik3r2), mRNA [NM_008841] |
KLA | .50 |
.50 |
.48 |
.43 |
.46 |
.66 |
.75 |
| ATP | .78 |
.68 |
.45 |
.43 |
.28 |
.57 |
1.37 |
| KLA/ATP | .49 |
.38 |
.32 |
.33 |
.38 |
.67 |
1.10 |
|
| Pik3r3 | 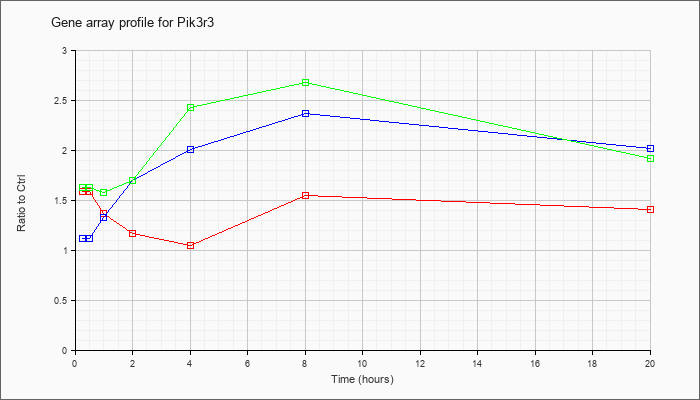 |
NM_181585 |
phosphatidylinositol 3 kinase, regulatory subunit, polypeptide 3 (p55) (Pik3r3), mRNA [NM_181585] |
KLA | 1.59 |
1.49 |
1.37 |
1.17 |
1.05 |
1.55 |
1.41 |
| ATP | 1.12 |
1.00 |
1.33 |
1.70 |
2.01 |
2.37 |
2.02 |
| KLA/ATP | 1.63 |
1.54 |
1.58 |
1.70 |
2.43 |
2.68 |
1.92 |
|
| Pik3r5 | 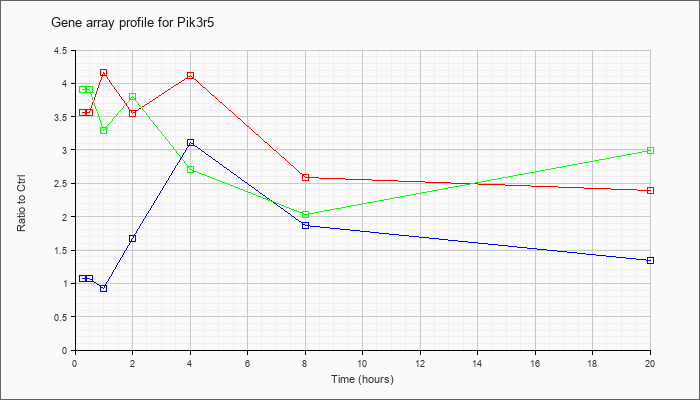 |
NM_177320 |
phosphoinositide-3-kinase, regulatory subunit 5, p101 (Pik3r5), mRNA [NM_177320] |
KLA | 3.57 |
3.58 |
4.16 |
3.54 |
4.11 |
2.59 |
2.39 |
| ATP | 1.08 |
1.08 |
.93 |
1.67 |
3.11 |
1.88 |
1.34 |
| KLA/ATP | 3.91 |
3.70 |
3.30 |
3.80 |
2.70 |
2.04 |
3.00 |
|
| Plcg2 | 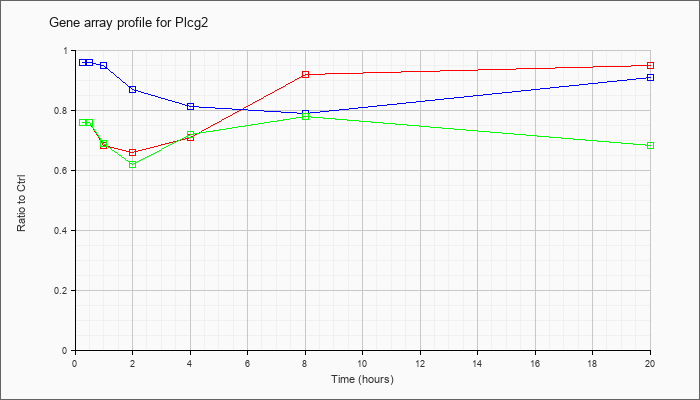 |
NM_172285 |
phospholipase C, gamma 2 (Plcg2), mRNA [NM_172285] |
KLA | .76 |
.74 |
.68 |
.66 |
.71 |
.92 |
.95 |
| ATP | .96 |
.90 |
.95 |
.87 |
.81 |
.79 |
.91 |
| KLA/ATP | .76 |
.71 |
.69 |
.62 |
.72 |
.78 |
.68 |
|
| Prdm4 | 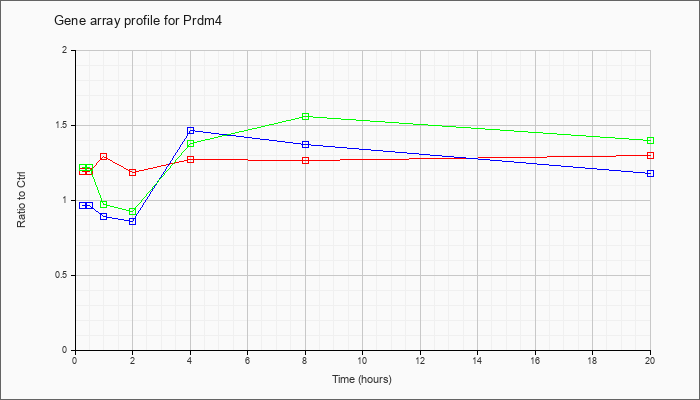 |
NM_181650 |
PR domain containing 4 (Prdm4), mRNA [NM_181650] |
KLA | 1.19 |
1.15 |
1.29 |
1.19 |
1.27 |
1.27 |
1.30 |
| ATP | .97 |
1.08 |
.89 |
.86 |
1.47 |
1.37 |
1.18 |
| KLA/ATP | 1.22 |
1.16 |
.97 |
.93 |
1.38 |
1.56 |
1.40 |
|
| Prkcd | 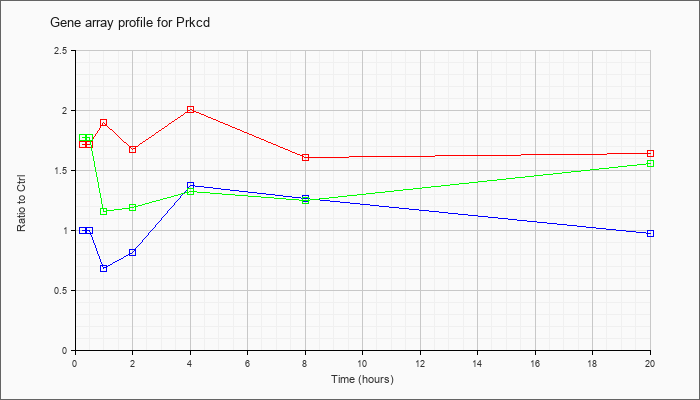 |
NM_011103 |
protein kinase C, delta (Prkcd), mRNA [NM_011103] |
KLA | 1.72 |
1.84 |
1.90 |
1.68 |
2.01 |
1.61 |
1.64 |
| ATP | 1.00 |
1.03 |
.68 |
.81 |
1.38 |
1.26 |
.98 |
| KLA/ATP | 1.77 |
1.79 |
1.16 |
1.19 |
1.33 |
1.25 |
1.56 |
|
| Psen1 | 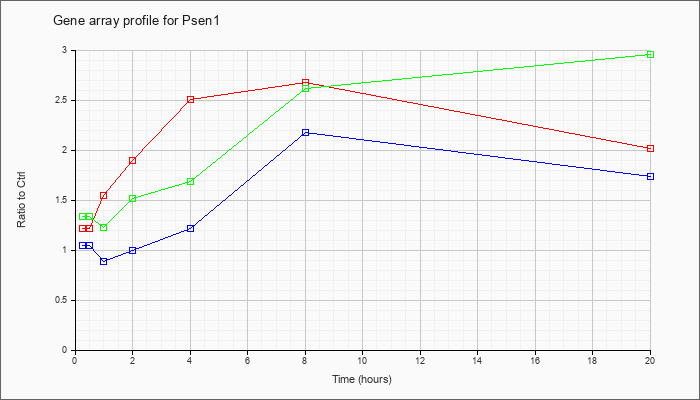 |
NM_008943 |
presenilin 1 (Psen1), mRNA [NM_008943] |
KLA | 1.22 |
1.31 |
1.54 |
1.90 |
2.50 |
2.67 |
2.01 |
| ATP | 1.04 |
1.10 |
.89 |
1.00 |
1.21 |
2.18 |
1.73 |
| KLA/ATP | 1.33 |
1.37 |
1.22 |
1.52 |
1.69 |
2.61 |
2.95 |
|
| Psen2 | 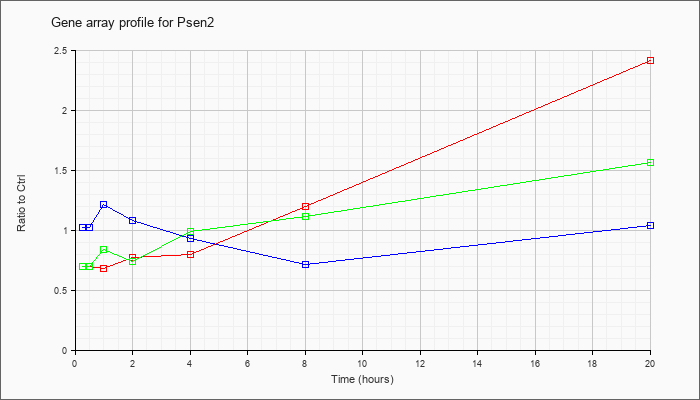 |
NM_011183 |
presenilin 2 (Psen2), mRNA [NM_011183] |
KLA | .70 |
.68 |
.68 |
.77 |
.80 |
1.20 |
2.41 |
| ATP | 1.02 |
1.04 |
1.21 |
1.08 |
.93 |
.71 |
1.04 |
| KLA/ATP | .70 |
.71 |
.84 |
.74 |
.99 |
1.11 |
1.56 |
|
| Ptpn11 | 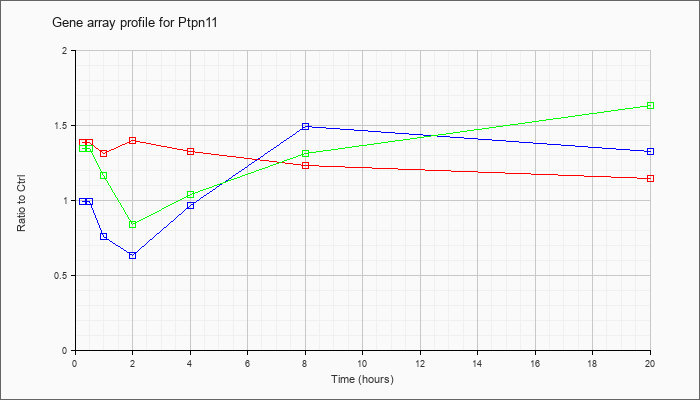 |
NM_001109992 |
protein tyrosine phosphatase, non-receptor type 11 (Ptpn11), transcript variant 2, mRNA [NM_001109992] |
KLA | 1.52 |
1.71 |
1.59 |
1.78 |
1.62 |
1.57 |
1.35 |
| ATP | 1.04 |
.93 |
.86 |
.79 |
1.09 |
1.95 |
1.90 |
| KLA/ATP | 1.59 |
1.61 |
1.45 |
1.11 |
1.18 |
1.82 |
3.02 |
|
| Ptpn11 |  |
NM_011202 |
protein tyrosine phosphatase, non-receptor type 11 (Ptpn11), transcript variant 1, mRNA [NM_011202] |
KLA | 1.32 |
1.28 |
1.17 |
1.21 |
1.18 |
1.06 |
1.05 |
| ATP | .97 |
.77 |
.71 |
.56 |
.90 |
1.27 |
1.04 |
| KLA/ATP | 1.22 |
1.21 |
1.02 |
.70 |
.96 |
1.06 |
.94 |
|
| Rac1 | 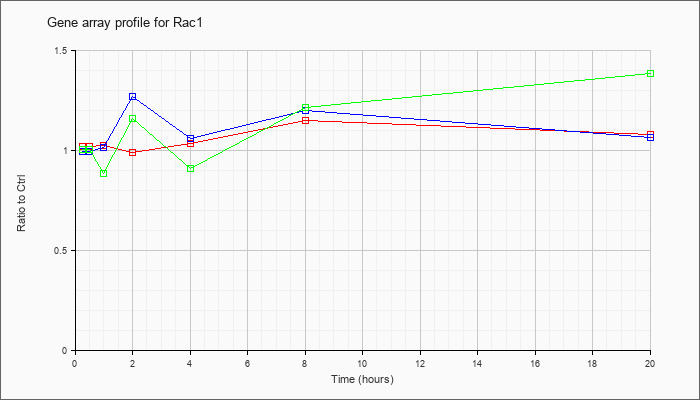 |
NM_009007 |
RAS-related C3 botulinum substrate 1 (Rac1), mRNA [NM_009007] |
KLA | 1.02 |
1.01 |
1.02 |
.99 |
1.03 |
1.15 |
1.08 |
| ATP | .99 |
1.15 |
1.01 |
1.27 |
1.06 |
1.20 |
1.06 |
| KLA/ATP | 1.00 |
1.04 |
.88 |
1.16 |
.91 |
1.21 |
1.38 |
|
| Raf1 | 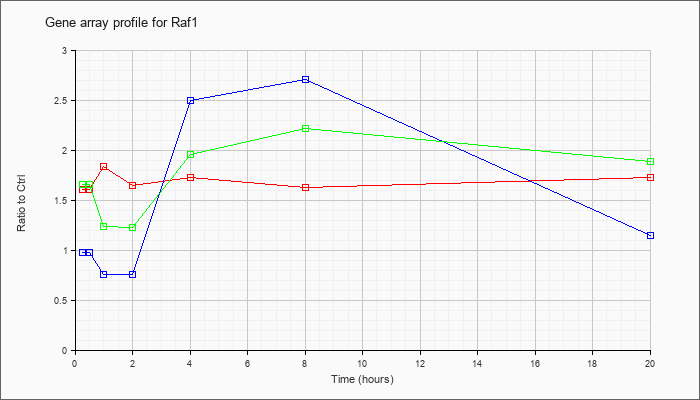 |
AK036317 |
16 days neonate cerebellum cDNA, RIKEN full-length enriched library, clone:9630056E02 product:v-raf-1 leukemia viral oncogene 1, full insert sequence. [AK036317] |
KLA | 1.47 |
1.34 |
1.55 |
1.29 |
1.59 |
1.32 |
1.46 |
| ATP | 1.05 |
.98 |
.54 |
.63 |
3.32 |
2.07 |
1.01 |
| KLA/ATP | 1.57 |
1.33 |
.79 |
.96 |
2.26 |
1.38 |
1.35 |
|
| Raf1 |  |
NM_029780 |
v-raf-leukemia viral oncogene 1 (Raf1), mRNA [NM_029780] |
KLA | 1.65 |
1.64 |
1.93 |
1.77 |
1.77 |
1.72 |
1.81 |
| ATP | .95 |
.96 |
.83 |
.80 |
2.22 |
2.92 |
1.19 |
| KLA/ATP | 1.68 |
1.64 |
1.39 |
1.32 |
1.85 |
2.50 |
2.06 |
|
| Rap1a | 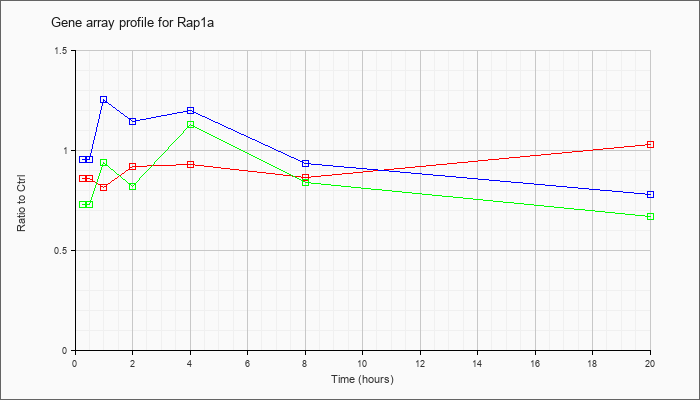 |
AK030897 |
adult male thymus cDNA, RIKEN full-length enriched library, clone:5830450G12 product:RAS-related protein-1a, full insert sequence [AK030897] |
KLA | .83 |
.80 |
.92 |
1.04 |
1.06 |
.91 |
1.06 |
| ATP | 1.04 |
.92 |
1.11 |
1.03 |
1.05 |
.73 |
.72 |
| KLA/ATP | .83 |
.81 |
.97 |
.75 |
1.14 |
.71 |
.53 |
|
| Rap1a |  |
NM_145541 |
RAS-related protein-1a (Rap1a), mRNA [NM_145541] |
KLA | .87 |
.78 |
.76 |
.86 |
.86 |
.84 |
1.01 |
| ATP | .91 |
.95 |
1.33 |
1.20 |
1.27 |
1.04 |
.81 |
| KLA/ATP | .68 |
.75 |
.92 |
.85 |
1.13 |
.90 |
.74 |
|
| Rap1b | 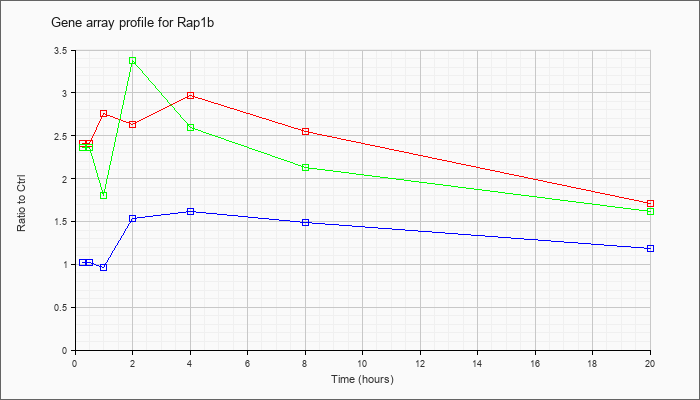 |
M79314 |
GTP-binding protein RAP1b mRNA, partial cds. [M79314] |
KLA | 2.61 |
2.95 |
2.81 |
2.76 |
3.07 |
2.67 |
1.88 |
| ATP | 1.00 |
1.10 |
1.00 |
1.94 |
1.94 |
1.72 |
1.31 |
| KLA/ATP | 2.49 |
2.45 |
2.03 |
4.23 |
3.03 |
2.60 |
1.86 |
|
| Rap1b |  |
NM_024457 |
RAS related protein 1b (Rap1b), mRNA [NM_024457] |
KLA | 2.21 |
2.35 |
2.71 |
2.51 |
2.86 |
2.44 |
1.54 |
| ATP | 1.05 |
.92 |
.93 |
1.14 |
1.30 |
1.26 |
1.07 |
| KLA/ATP | 2.23 |
2.15 |
1.57 |
2.53 |
2.16 |
1.67 |
1.38 |
|
| Rapgef1 | 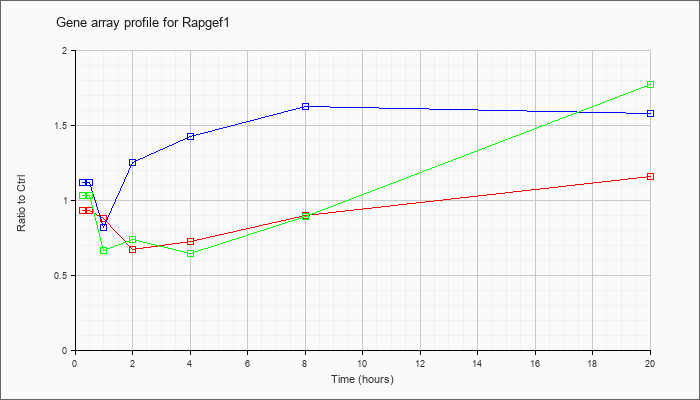 |
NM_001039087 |
Rap guanine nucleotide exchange factor (GEF) 1 (Rapgef1), transcript variant 1, mRNA [NM_001039087] |
KLA | .93 |
.89 |
.88 |
.67 |
.73 |
.90 |
1.16 |
| ATP | 1.12 |
1.09 |
.82 |
1.25 |
1.43 |
1.63 |
1.58 |
| KLA/ATP | 1.03 |
.95 |
.67 |
.74 |
.65 |
.89 |
1.77 |
|
| Rela | 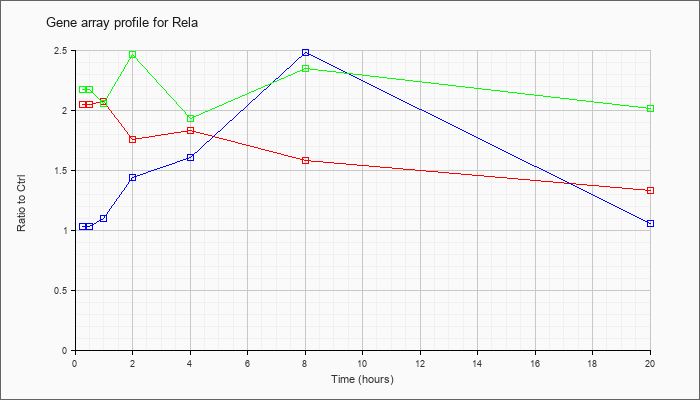 |
NM_009045 |
v-rel reticuloendotheliosis viral oncogene homolog A (avian) (Rela), mRNA [NM_009045] |
KLA | 2.05 |
2.05 |
2.07 |
1.75 |
1.83 |
1.58 |
1.33 |
| ATP | 1.03 |
1.08 |
1.10 |
1.44 |
1.61 |
2.48 |
1.05 |
| KLA/ATP | 2.17 |
2.27 |
2.05 |
2.47 |
1.93 |
2.35 |
2.02 |
|
| Rhoa | 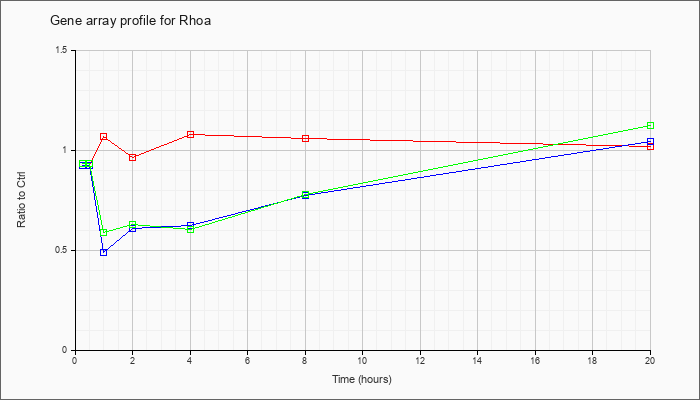 |
NM_016802 |
ras homolog gene family, member A (Rhoa), mRNA [NM_016802] |
KLA | .93 |
1.00 |
1.07 |
.97 |
1.08 |
1.06 |
1.02 |
| ATP | .93 |
.88 |
.49 |
.61 |
.63 |
.78 |
1.05 |
| KLA/ATP | .94 |
.92 |
.59 |
.63 |
.61 |
.78 |
1.12 |
|
| Ripk2 |  |
NM_138952 |
receptor (TNFRSF)-interacting serine-threonine kinase 2 (Ripk2), mRNA [NM_138952] |
KLA | 19.51 |
18.67 |
18.82 |
10.27 |
5.08 |
3.42 |
2.58 |
| ATP | 1.05 |
1.02 |
1.41 |
3.66 |
9.06 |
6.82 |
1.68 |
| KLA/ATP | 19.33 |
20.48 |
15.93 |
16.05 |
8.62 |
9.36 |
5.23 |
|
| Rps6ka1 | 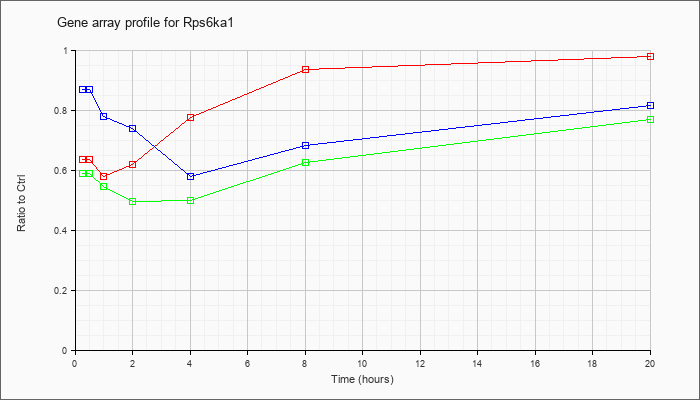 |
NM_009097 |
ribosomal protein S6 kinase polypeptide 1 (Rps6ka1), mRNA [NM_009097] |
KLA | .64 |
.65 |
.58 |
.62 |
.78 |
.94 |
.98 |
| ATP | .87 |
.85 |
.78 |
.74 |
.58 |
.68 |
.82 |
| KLA/ATP | .59 |
.52 |
.55 |
.50 |
.50 |
.63 |
.77 |
|
| Rps6ka2 | 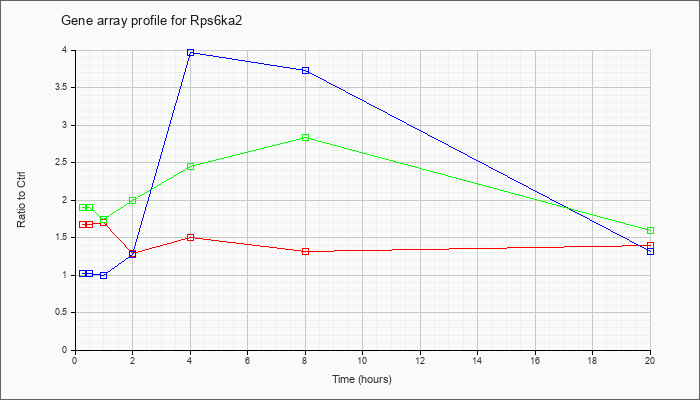 |
NM_011299 |
ribosomal protein S6 kinase, polypeptide 2 (Rps6ka2), mRNA [NM_011299] |
KLA | 1.67 |
1.69 |
1.70 |
1.29 |
1.50 |
1.32 |
1.40 |
| ATP | 1.02 |
1.02 |
.99 |
1.28 |
3.97 |
3.73 |
1.32 |
| KLA/ATP | 1.90 |
1.63 |
1.74 |
1.99 |
2.45 |
2.84 |
1.59 |
|
| Rps6ka3 | 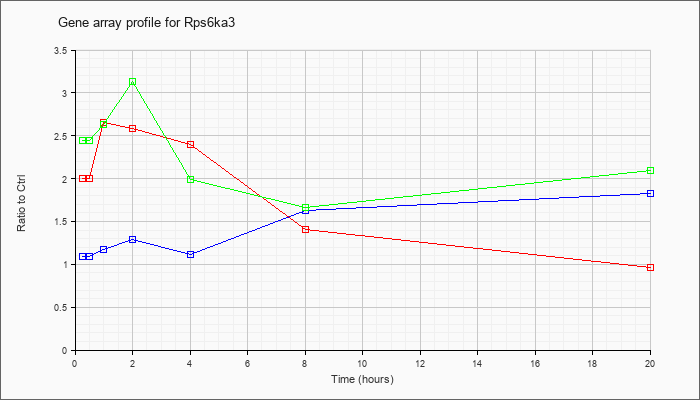 |
NM_148945 |
ribosomal protein S6 kinase polypeptide 3 (Rps6ka3), mRNA [NM_148945] |
KLA | 2.00 |
2.37 |
2.65 |
2.59 |
2.40 |
1.41 |
.96 |
| ATP | 1.09 |
1.09 |
1.17 |
1.29 |
1.11 |
1.63 |
1.83 |
| KLA/ATP | 2.44 |
2.44 |
2.63 |
3.13 |
1.99 |
1.66 |
2.09 |
|
| Rps6ka5 | 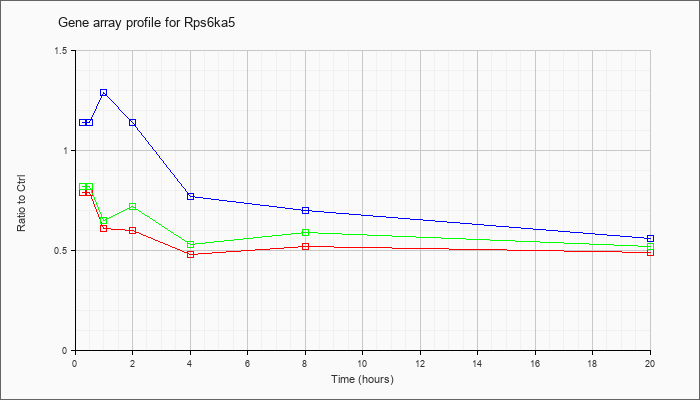 |
NM_153587 |
ribosomal protein S6 kinase, polypeptide 5 (Rps6ka5), mRNA [NM_153587] |
KLA | .79 |
.69 |
.61 |
.60 |
.48 |
.52 |
.49 |
| ATP | 1.14 |
1.25 |
1.29 |
1.14 |
.77 |
.70 |
.56 |
| KLA/ATP | .82 |
.73 |
.65 |
.72 |
.53 |
.59 |
.52 |
|
| Rps6ka6 | 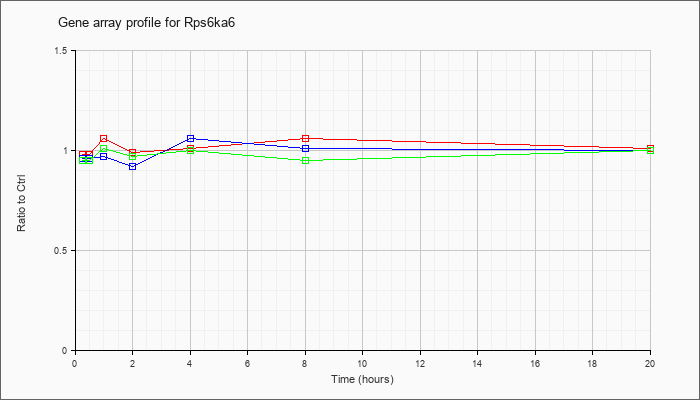 |
NM_025949 |
ribosomal protein S6 kinase polypeptide 6 (Rps6ka6), mRNA [NM_025949] |
KLA | .98 |
1.03 |
1.06 |
.99 |
1.01 |
1.06 |
1.01 |
| ATP | .96 |
1.00 |
.97 |
.92 |
1.06 |
1.01 |
1.00 |
| KLA/ATP | .95 |
1.01 |
1.01 |
.97 |
1.00 |
.95 |
1.00 |
|
| Sh2b1 | 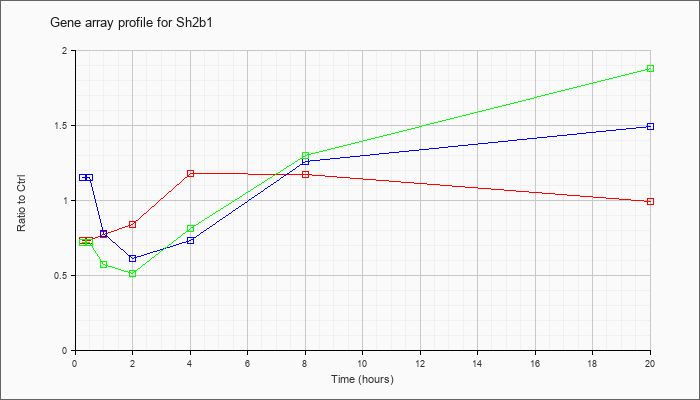 |
NM_011363 |
SH2B adaptor protein 1 (Sh2b1), transcript variant 2, mRNA [NM_011363] |
KLA | .73 |
.65 |
.77 |
.84 |
1.18 |
1.17 |
.99 |
| ATP | 1.15 |
1.16 |
.78 |
.61 |
.73 |
1.26 |
1.49 |
| KLA/ATP | .72 |
.70 |
.57 |
.51 |
.81 |
1.30 |
1.88 |
|
| Sh2b2 | 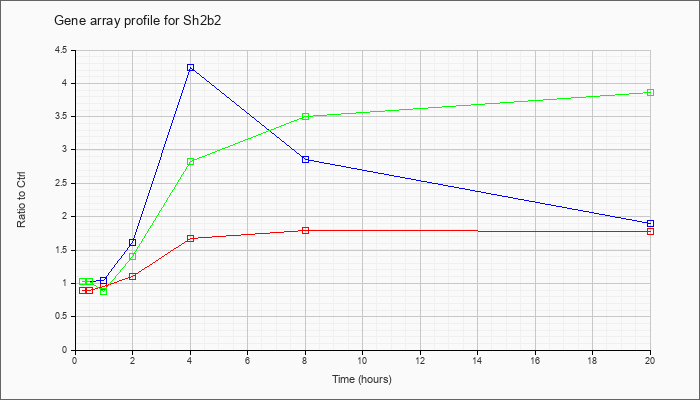 |
NM_018825 |
SH2B adaptor protein 2 (Sh2b2), mRNA [NM_018825] |
KLA | .89 |
.94 |
.95 |
1.10 |
1.67 |
1.80 |
1.78 |
| ATP | 1.03 |
1.15 |
1.05 |
1.62 |
4.24 |
2.85 |
1.90 |
| KLA/ATP | 1.03 |
.97 |
.88 |
1.41 |
2.83 |
3.50 |
3.87 |
|
| Sh2b3 | 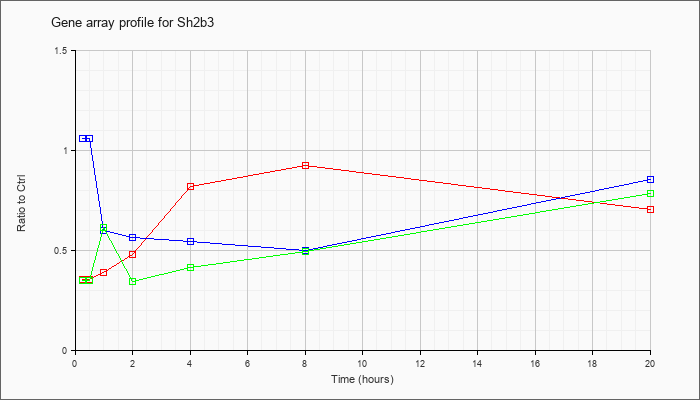 |
AK156499 |
activated spleen cDNA, RIKEN full-length enriched library, clone:F830027H19 product:linker of T-cell receptor pathways, full insert sequence [AK156499] |
KLA | .26 |
.25 |
.32 |
.39 |
.84 |
.84 |
.63 |
| ATP | 1.12 |
1.03 |
.52 |
.47 |
.55 |
.40 |
.65 |
| KLA/ATP | .25 |
.25 |
.18 |
.22 |
.34 |
.41 |
.64 |
|
| Sh2b3 |  |
NM_008507 |
SH2B adaptor protein 3 (Sh2b3), mRNA [NM_008507] |
KLA | .45 |
.43 |
.46 |
.57 |
.80 |
1.01 |
.78 |
| ATP | 1.00 |
.94 |
.68 |
.66 |
.54 |
.60 |
1.06 |
| KLA/ATP | .45 |
.41 |
1.05 |
.47 |
.49 |
.58 |
.93 |
|
| Shc1 | 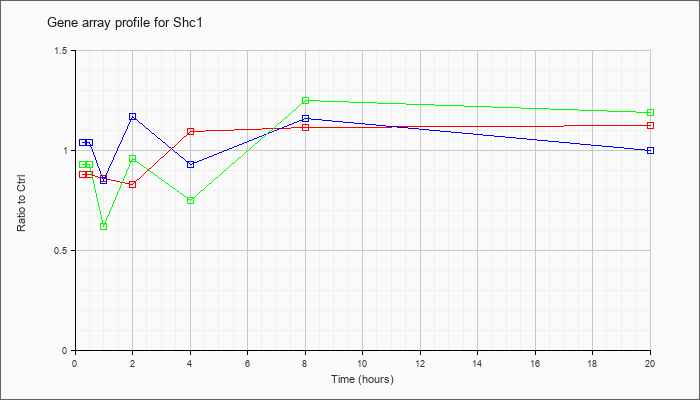 |
NM_011368 |
src homology 2 domain-containing transforming protein C1 (Shc1), transcript variant 2, mRNA [NM_011368] |
KLA | .88 |
.87 |
.86 |
.83 |
1.09 |
1.11 |
1.12 |
| ATP | 1.04 |
1.25 |
.85 |
1.17 |
.93 |
1.16 |
1.00 |
| KLA/ATP | .93 |
.96 |
.62 |
.96 |
.75 |
1.25 |
1.19 |
|
| Shc2 | 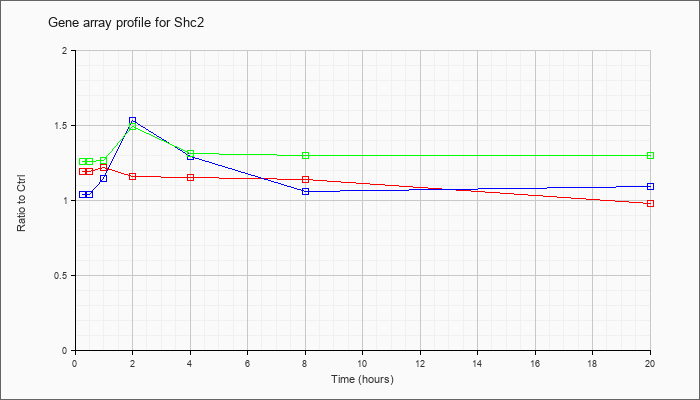 |
NM_001024539 |
src homology 2 domain-containing transforming protein C2 (Shc2), mRNA [NM_001024539] |
KLA | 1.19 |
1.25 |
1.22 |
1.16 |
1.15 |
1.14 |
.98 |
| ATP | 1.04 |
1.14 |
1.15 |
1.53 |
1.29 |
1.06 |
1.09 |
| KLA/ATP | 1.26 |
1.24 |
1.26 |
1.49 |
1.31 |
1.30 |
1.30 |
|
| Shc3 | 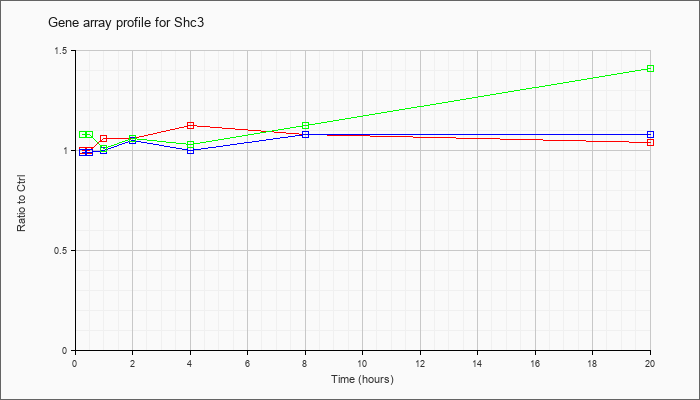 |
NM_009167 |
src homology 2 domain-containing transforming protein C3 (Shc3), mRNA [NM_009167] |
KLA | 1.00 |
.98 |
1.06 |
1.06 |
1.12 |
1.08 |
1.04 |
| ATP | .99 |
1.02 |
1.00 |
1.05 |
1.00 |
1.08 |
1.08 |
| KLA/ATP | 1.08 |
1.06 |
1.01 |
1.06 |
1.03 |
1.12 |
1.41 |
|
| Shc4 | 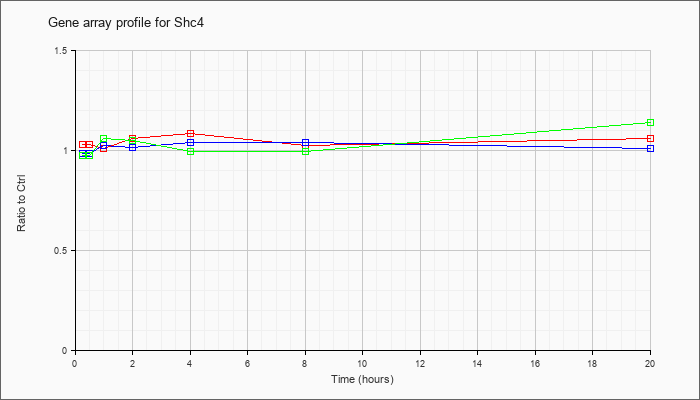 |
NM_199022 |
SHC (Src homology 2 domain containing) family, member 4 (Shc4), mRNA [NM_199022] |
KLA | 1.03 |
1.03 |
1.01 |
1.06 |
1.09 |
1.03 |
1.06 |
| ATP | .99 |
1.00 |
1.03 |
1.02 |
1.04 |
1.04 |
1.01 |
| KLA/ATP | .98 |
.97 |
1.06 |
1.05 |
1.00 |
1.00 |
1.14 |
|
| Sort1 | 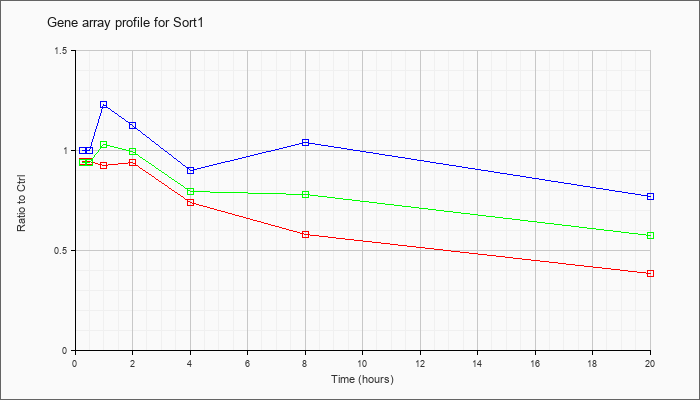 |
NM_019972 |
sortilin 1 (Sort1), mRNA [NM_019972] |
KLA | .94 |
.96 |
.92 |
.94 |
.74 |
.58 |
.38 |
| ATP | 1.00 |
1.01 |
1.23 |
1.12 |
.90 |
1.04 |
.77 |
| KLA/ATP | .94 |
.96 |
1.03 |
.99 |
.79 |
.78 |
.57 |
|
| Sos1 | 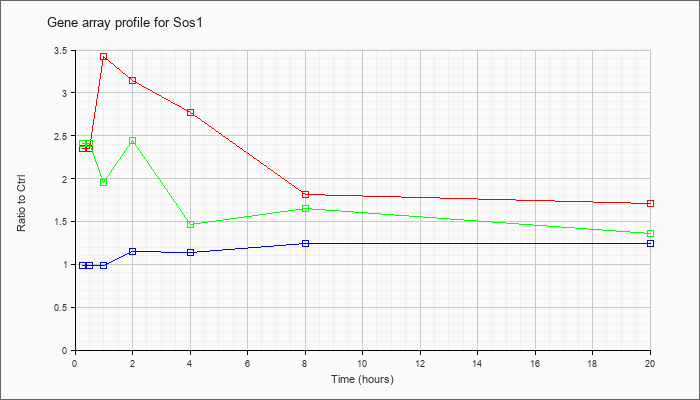 |
NM_009231 |
Son of sevenless homolog 1 (Drosophila) (Sos1), mRNA [NM_009231] |
KLA | 2.36 |
2.40 |
3.42 |
3.15 |
2.78 |
1.81 |
1.71 |
| ATP | .99 |
1.00 |
.99 |
1.15 |
1.14 |
1.25 |
1.24 |
| KLA/ATP | 2.41 |
2.26 |
1.95 |
2.44 |
1.47 |
1.66 |
1.37 |
|
| Sos2 | 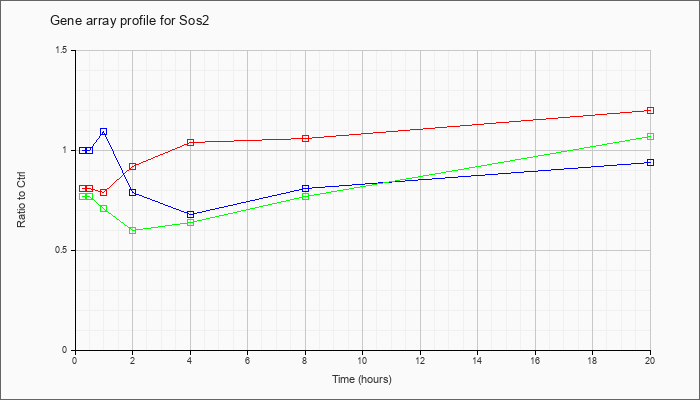 |
XM_127051 |
gb|PREDICTED: Mus musculus Son of sevenless homolog 2 (Drosophila) (Sos2), mRNA [XM_127051] |
KLA | .81 |
.77 |
.79 |
.92 |
1.04 |
1.06 |
1.20 |
| ATP | 1.00 |
1.00 |
1.09 |
.79 |
.68 |
.81 |
.94 |
| KLA/ATP | .77 |
.69 |
.71 |
.60 |
.64 |
.77 |
1.07 |
|
| Traf6 | 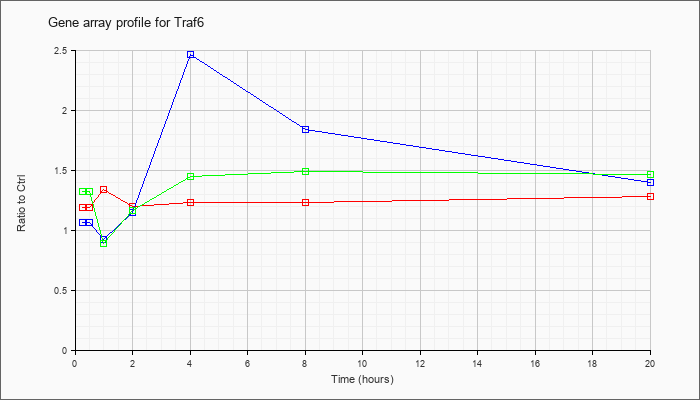 |
NM_009424 |
Tnf receptor-associated factor 6 (Traf6), mRNA [NM_009424] |
KLA | 1.19 |
1.28 |
1.34 |
1.20 |
1.23 |
1.23 |
1.28 |
| ATP | 1.06 |
1.12 |
.92 |
1.15 |
2.46 |
1.84 |
1.40 |
| KLA/ATP | 1.32 |
1.37 |
.89 |
1.16 |
1.45 |
1.49 |
1.46 |
|
| Trp53 | 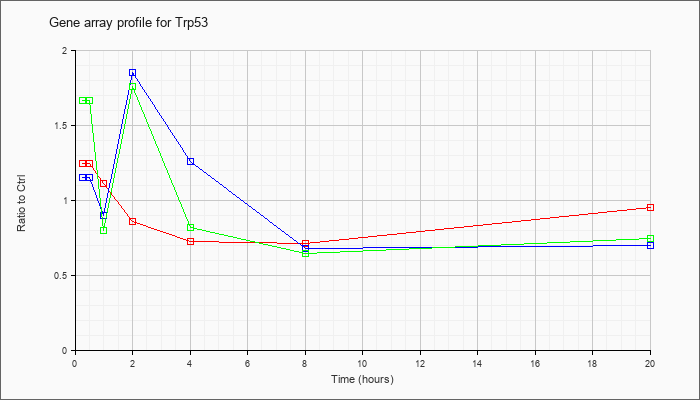 |
NM_011640 |
transformation related protein 53 (Trp53), mRNA [NM_011640] |
KLA | 1.25 |
1.37 |
1.11 |
.86 |
.72 |
.71 |
.95 |
| ATP | 1.15 |
1.32 |
.90 |
1.85 |
1.26 |
.68 |
.70 |
| KLA/ATP | 1.66 |
1.49 |
.80 |
1.76 |
.81 |
.65 |
.74 |
|
| Trp73 | 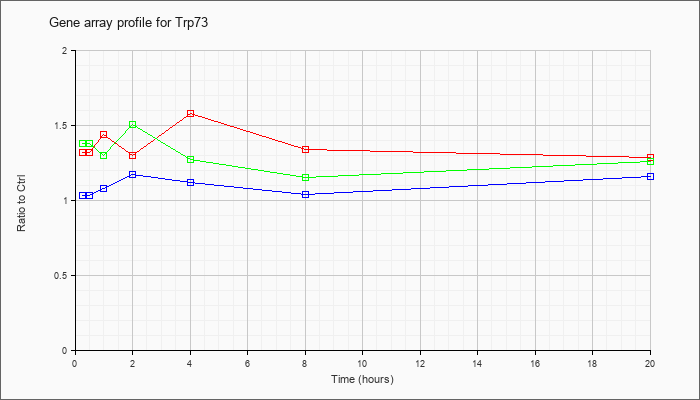 |
BC069182 |
transformation related protein 73, mRNA (cDNA clone IMAGE:6826464) [BC069182] |
KLA | .98 |
1.00 |
.95 |
.99 |
1.04 |
1.03 |
1.00 |
| ATP | 1.01 |
.97 |
1.01 |
1.06 |
1.04 |
1.01 |
1.08 |
| KLA/ATP | 1.02 |
.96 |
1.00 |
1.07 |
1.06 |
1.07 |
1.00 |
|
| Trp73 |  |
NM_011642 |
transformation related protein 73 (Trp73), mRNA [NM_011642] |
KLA | 1.66 |
1.63 |
1.93 |
1.60 |
2.12 |
1.64 |
1.57 |
| ATP | 1.05 |
1.07 |
1.14 |
1.28 |
1.19 |
1.07 |
1.24 |
| KLA/ATP | 1.73 |
1.86 |
1.60 |
1.94 |
1.48 |
1.23 |
1.52 |
|
| Ywhae | 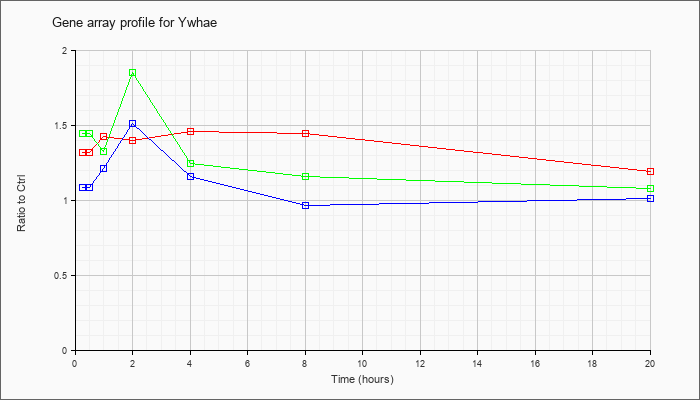 |
NM_009536 |
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, epsilon polypeptide (Ywhae), mRNA [NM_009536] |
KLA | 1.32 |
1.35 |
1.43 |
1.40 |
1.46 |
1.45 |
1.19 |
| ATP | 1.09 |
1.30 |
1.21 |
1.51 |
1.16 |
.97 |
1.01 |
| KLA/ATP | 1.45 |
1.50 |
1.33 |
1.85 |
1.25 |
1.16 |
1.08 |
|
| Zfp110 | 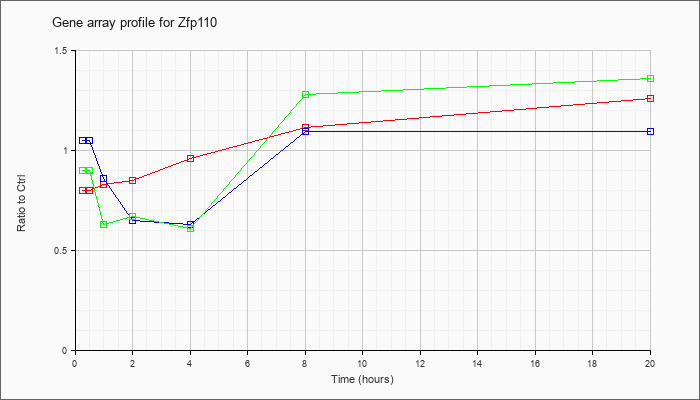 |
NM_022981 |
zinc finger protein 110 (Zfp110), mRNA [NM_022981] |
KLA | .80 |
.85 |
.83 |
.85 |
.96 |
1.11 |
1.26 |
| ATP | 1.05 |
.95 |
.86 |
.65 |
.63 |
1.09 |
1.09 |
| KLA/ATP | .90 |
.81 |
.63 |
.67 |
.61 |
1.28 |
1.36 |
|
| Zfp369 | 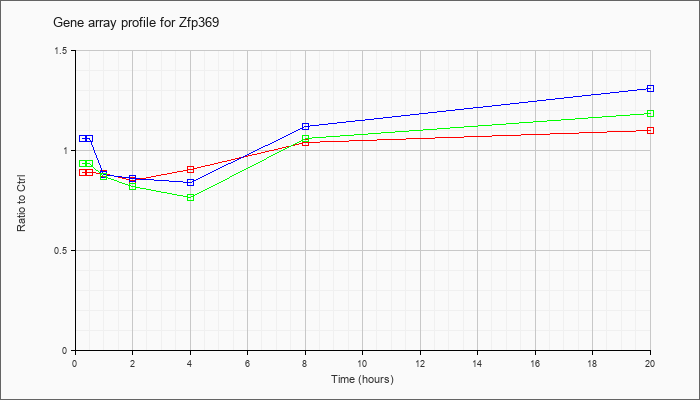 |
AK082535 |
0 day neonate cerebellum cDNA, RIKEN full-length enriched library, clone:C230060D12 product:hypothetical KRAB box containing protein, full insert sequence [AK082535] |
KLA | .86 |
.77 |
.83 |
.86 |
.79 |
1.00 |
.87 |
| ATP | 1.04 |
.81 |
.79 |
.82 |
.95 |
.98 |
.94 |
| KLA/ATP | .89 |
.82 |
.91 |
.84 |
.84 |
.87 |
.81 |
|
| Zfp369 |  |
NM_178364 |
zinc finger protein 369 (Zfp369), mRNA [NM_178364] |
KLA | .91 |
.93 |
.91 |
.85 |
.96 |
1.06 |
1.21 |
| ATP | 1.07 |
1.03 |
.92 |
.88 |
.78 |
1.19 |
1.50 |
| KLA/ATP | .96 |
.92 |
.85 |
.81 |
.73 |
1.16 |
1.37 |
|