| Gene symbol |
Time course plot |
Accession |
Description |
Treatment |
15min |
30min |
1hr |
2hr |
4hr |
8hr |
20hr |
14927508
3 | 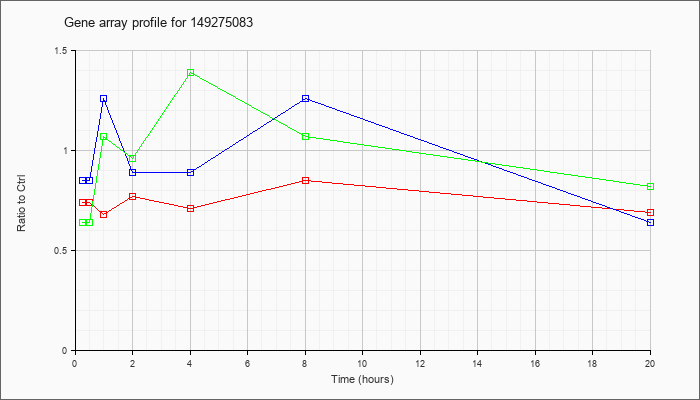 |
149275083 |
Unknown |
KLA | .74 |
.68 |
.68 |
.77 |
.71 |
.85 |
.69 |
| ATP | .85 |
.69 |
1.26 |
.89 |
.89 |
1.26 |
.64 |
| KLA/ATP | .64 |
.63 |
1.07 |
.96 |
1.39 |
1.07 |
.82 |
|
1500003O
03Rik | 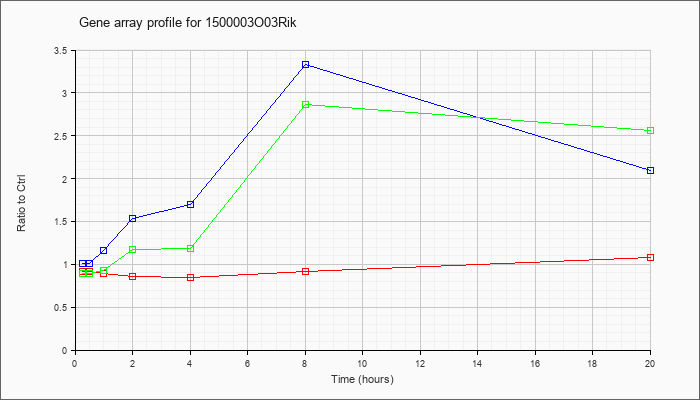 |
NM_019769 |
RIKEN cDNA 1500003O03 gene (1500003O03Rik), mRNA [NM_019769] |
KLA | .92 |
.89 |
.89 |
.86 |
.85 |
.92 |
1.08 |
| ATP | 1.01 |
1.16 |
1.16 |
1.53 |
1.70 |
3.33 |
2.09 |
| KLA/ATP | .89 |
.97 |
.93 |
1.17 |
1.19 |
2.86 |
2.57 |
|
1700009N
14Rik | 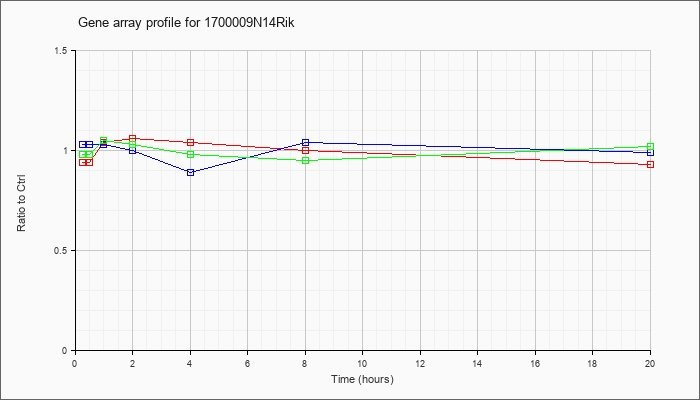 |
NM_001081095 |
RIKEN cDNA 1700009N14 gene (1700009N14Rik), mRNA [NM_001081095] |
KLA | .94 |
.99 |
1.04 |
1.06 |
1.04 |
1.00 |
.93 |
| ATP | 1.03 |
1.12 |
1.03 |
1.00 |
.89 |
1.04 |
.99 |
| KLA/ATP | .98 |
.94 |
1.05 |
1.03 |
.98 |
.95 |
1.02 |
|
2010110P
09Rik | 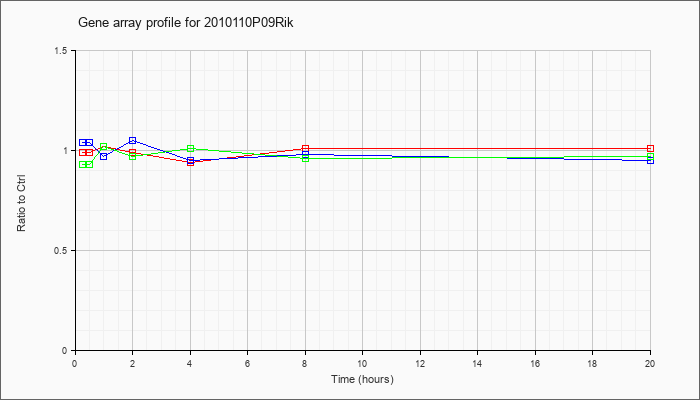 |
NM_027363 |
RIKEN cDNA 2010110P09 gene (2010110P09Rik), mRNA [NM_027363] |
KLA | .99 |
.98 |
1.02 |
.99 |
.94 |
1.01 |
1.01 |
| ATP | 1.04 |
1.05 |
.97 |
1.05 |
.95 |
.98 |
.95 |
| KLA/ATP | .93 |
1.00 |
1.02 |
.97 |
1.01 |
.96 |
.97 |
|
AK019932
|  |
AK019932 |
adult male pituitary gland cDNA, RIKEN full-length enriched library, clone:5330434N09 product:inferred: transmembrane receptor frizzled 5 {Mus musculus}, full insert sequence [AK019932] |
KLA | .99 |
.94 |
.97 |
.98 |
.96 |
.99 |
1.03 |
| ATP | 1.02 |
1.07 |
1.02 |
.97 |
1.00 |
.98 |
.93 |
| KLA/ATP | .93 |
.99 |
.91 |
1.04 |
1.02 |
1.01 |
1.00 |
|
AK036998
|  |
AK036998 |
adult female vagina cDNA, RIKEN full-length enriched library, clone:9930034M05 product:unclassifiable, full insert sequence. [AK036998] |
KLA | 1.43 |
1.09 |
.89 |
.79 |
.96 |
.97 |
.71 |
| ATP | 1.04 |
1.04 |
.71 |
.64 |
.90 |
.82 |
.82 |
| KLA/ATP | 1.34 |
1.23 |
1.01 |
.89 |
1.10 |
.82 |
.75 |
|
AK045358
| 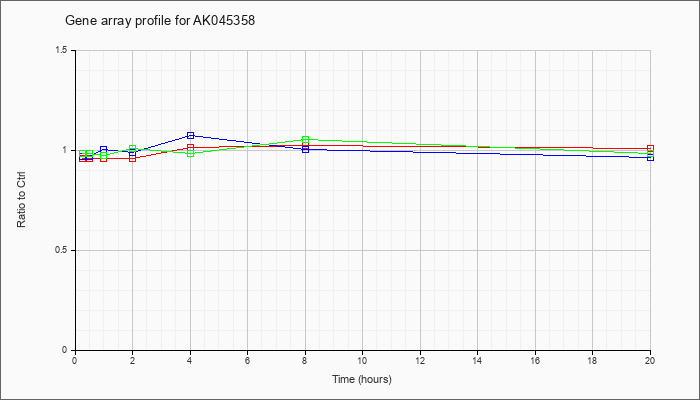 |
AK045358 |
adult male corpora quadrigemina cDNA, RIKEN full-length enriched library, clone:B230105H23 product:voltage-dependent anion channel 3, full insert sequence [AK045358] |
KLA | .96 |
.98 |
.96 |
.96 |
1.02 |
1.03 |
1.01 |
| ATP | .97 |
.95 |
1.01 |
.99 |
1.08 |
1.01 |
.97 |
| KLA/ATP | .99 |
.99 |
.98 |
1.01 |
.99 |
1.06 |
.99 |
|
| Adcy1 | 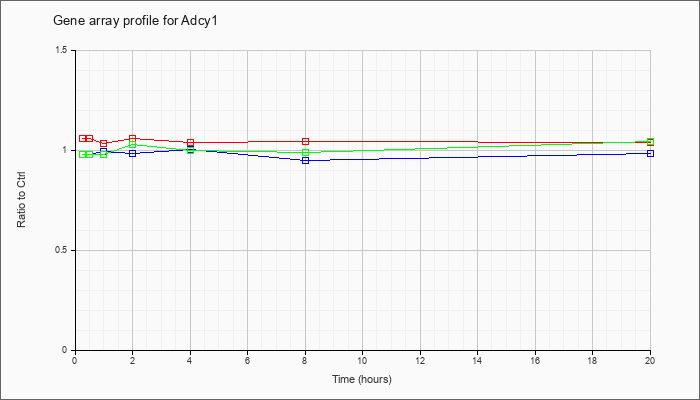 |
BC050125 |
adenylate cyclase 1, mRNA (cDNA clone IMAGE:30015692). [BC050125] |
KLA | 1.05 |
1.03 |
.98 |
1.04 |
1.05 |
1.09 |
1.10 |
| ATP | .90 |
1.02 |
.99 |
.97 |
1.03 |
.95 |
1.01 |
| KLA/ATP | .99 |
1.04 |
.90 |
.98 |
.98 |
1.00 |
1.03 |
|
| Adcy1 |  |
NM_009622 |
adenylate cyclase 1 (Adcy1), mRNA [NM_009622] |
KLA | 1.07 |
1.02 |
1.06 |
1.07 |
1.03 |
1.02 |
1.01 |
| ATP | 1.02 |
1.04 |
1.00 |
.99 |
.99 |
.95 |
.97 |
| KLA/ATP | .97 |
1.05 |
1.02 |
1.05 |
1.01 |
.98 |
1.05 |
|
| Adcy2 | 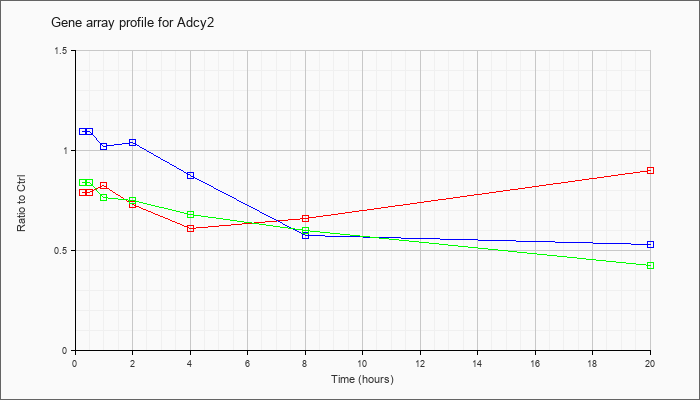 |
NM_153534 |
adenylate cyclase 2 (Adcy2), mRNA [NM_153534] |
KLA | .79 |
.82 |
.83 |
.73 |
.61 |
.66 |
.90 |
| ATP | 1.10 |
1.09 |
1.02 |
1.04 |
.88 |
.58 |
.53 |
| KLA/ATP | .84 |
.87 |
.77 |
.75 |
.68 |
.60 |
.43 |
|
| Adcy3 | 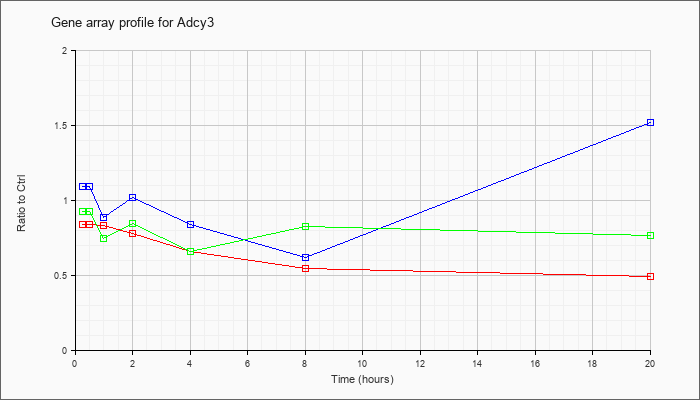 |
NM_138305 |
adenylate cyclase 3 (Adcy3), mRNA [NM_138305] |
KLA | .84 |
.84 |
.83 |
.78 |
.66 |
.55 |
.49 |
| ATP | 1.09 |
1.05 |
.89 |
1.02 |
.84 |
.62 |
1.52 |
| KLA/ATP | .93 |
.87 |
.75 |
.85 |
.66 |
.83 |
.77 |
|
| Adcy4 | 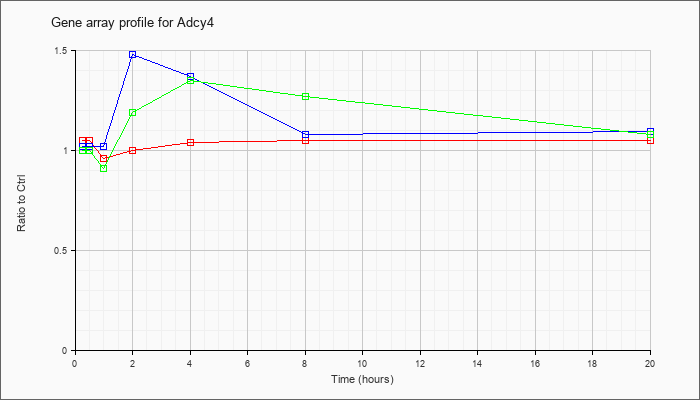 |
NM_080435 |
adenylate cyclase 4 (Adcy4), mRNA [NM_080435] |
KLA | 1.05 |
1.07 |
.96 |
1.00 |
1.04 |
1.05 |
1.05 |
| ATP | 1.02 |
1.05 |
1.02 |
1.48 |
1.37 |
1.08 |
1.09 |
| KLA/ATP | 1.00 |
1.05 |
.91 |
1.19 |
1.35 |
1.27 |
1.08 |
|
| Adcy5 |  |
NM_001012765 |
adenylate cyclase 5 (Adcy5), mRNA [NM_001012765] |
KLA | 1.06 |
1.00 |
.97 |
1.01 |
1.03 |
.95 |
.93 |
| ATP | .95 |
1.08 |
1.08 |
1.07 |
.97 |
1.03 |
.97 |
| KLA/ATP | .97 |
.92 |
1.04 |
.99 |
.91 |
.96 |
1.04 |
|
| Adcy6 | 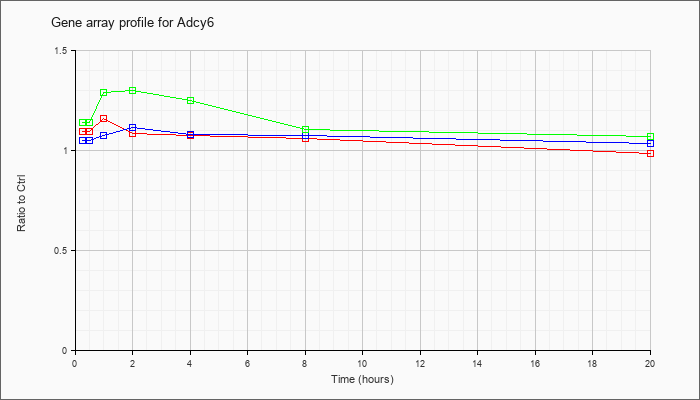 |
NM_007405 |
adenylate cyclase 6 (Adcy6), mRNA [NM_007405] |
KLA | 1.10 |
1.16 |
1.16 |
1.09 |
1.08 |
1.06 |
.99 |
| ATP | 1.05 |
.97 |
1.08 |
1.12 |
1.08 |
1.08 |
1.04 |
| KLA/ATP | 1.14 |
1.20 |
1.29 |
1.30 |
1.25 |
1.11 |
1.07 |
|
| Adcy7 |  |
NM_001037724 |
adenylate cyclase 7 (Adcy7), transcript variant 3, mRNA [NM_001037724] |
KLA | .43 |
.39 |
.35 |
.32 |
.45 |
.72 |
.84 |
| ATP | .90 |
.84 |
.62 |
.41 |
.53 |
.42 |
.94 |
| KLA/ATP | .39 |
.34 |
.23 |
.17 |
.41 |
.34 |
.67 |
|
| Adcy7 |  |
NM_007406 |
adenylate cyclase 7 (Adcy7), transcript variant 1, mRNA [NM_007406] |
KLA | .51 |
.52 |
.48 |
.41 |
.59 |
.81 |
.99 |
| ATP | 1.01 |
1.02 |
.62 |
.52 |
.53 |
.38 |
.98 |
| KLA/ATP | .53 |
.48 |
.30 |
.27 |
.42 |
.36 |
.65 |
|
| Adcy8 | 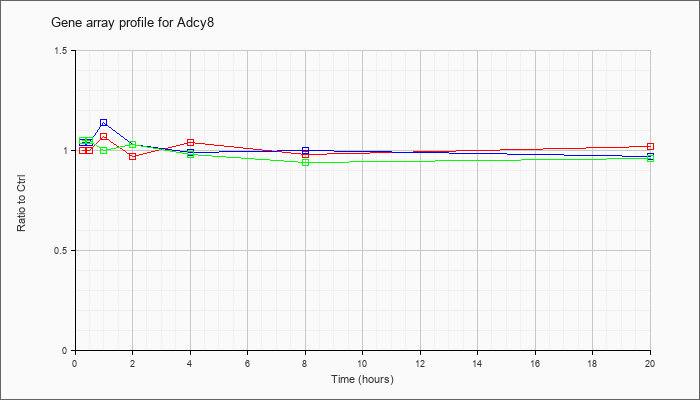 |
NM_009623 |
adenylate cyclase 8 (Adcy8), mRNA [NM_009623] |
KLA | 1.00 |
1.00 |
1.07 |
.97 |
1.04 |
.98 |
1.02 |
| ATP | 1.04 |
1.05 |
1.14 |
1.03 |
.99 |
1.00 |
.97 |
| KLA/ATP | 1.05 |
1.04 |
1.00 |
1.03 |
.98 |
.94 |
.96 |
|
| Adcy9 | 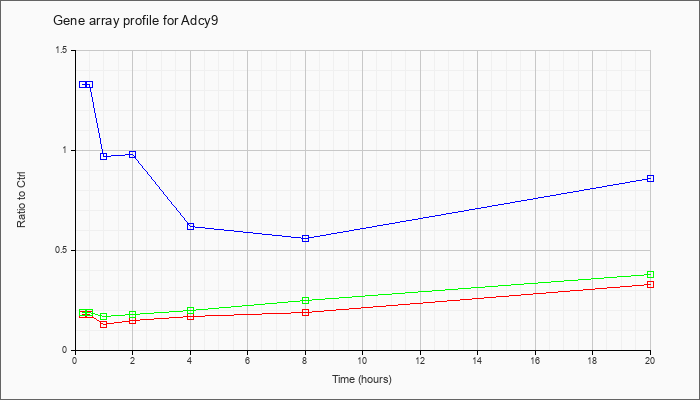 |
NM_009624 |
adenylate cyclase 9 (Adcy9), mRNA [NM_009624] |
KLA | .18 |
.15 |
.13 |
.15 |
.17 |
.19 |
.33 |
| ATP | 1.33 |
1.16 |
.97 |
.98 |
.62 |
.56 |
.86 |
| KLA/ATP | .19 |
.16 |
.17 |
.18 |
.20 |
.25 |
.38 |
|
| Akt1 | 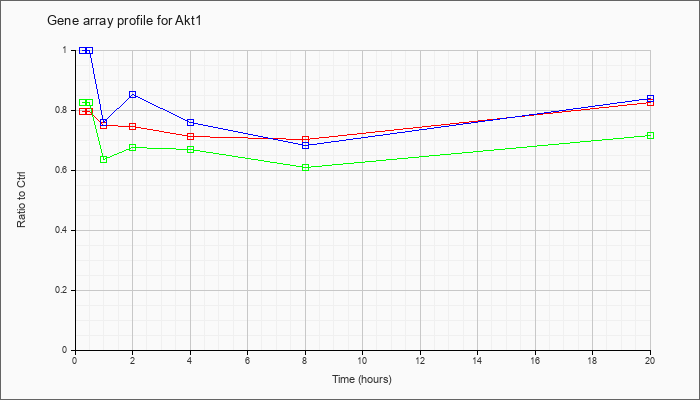 |
NM_009652 |
thymoma viral proto-oncogene 1 (Akt1), mRNA [NM_009652] |
KLA | .80 |
.80 |
.75 |
.75 |
.71 |
.70 |
.82 |
| ATP | 1.00 |
1.02 |
.76 |
.85 |
.76 |
.68 |
.84 |
| KLA/ATP | .82 |
.80 |
.64 |
.68 |
.67 |
.61 |
.72 |
|
| Akt2 | 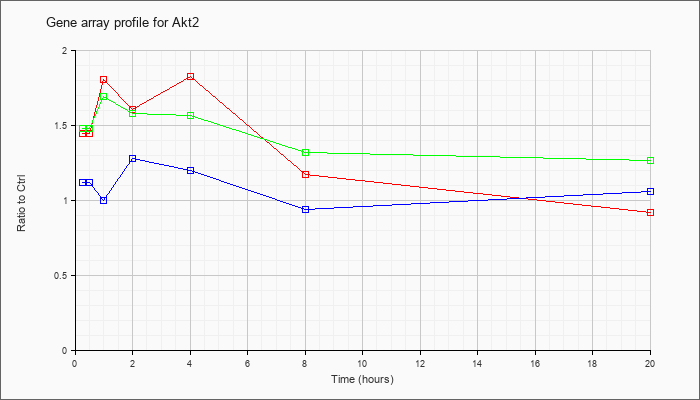 |
NM_007434 |
thymoma viral proto-oncogene 2 (Akt2), transcript variant 2, mRNA [NM_007434] |
KLA | 1.45 |
1.49 |
1.81 |
1.61 |
1.83 |
1.17 |
.92 |
| ATP | 1.12 |
1.12 |
1.00 |
1.28 |
1.20 |
.94 |
1.06 |
| KLA/ATP | 1.48 |
1.70 |
1.69 |
1.58 |
1.57 |
1.32 |
1.27 |
|
| Akt3 | 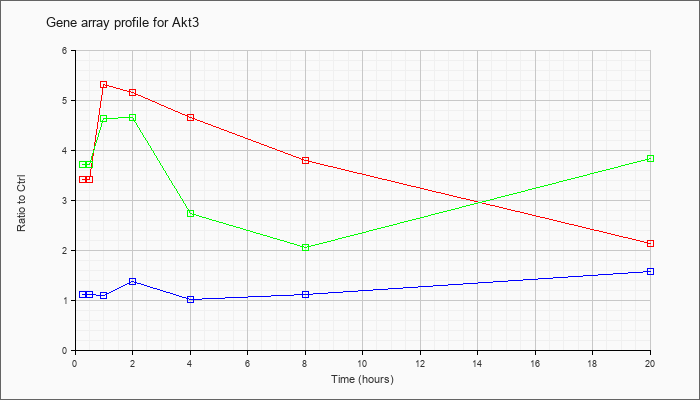 |
NM_011785 |
thymoma viral proto-oncogene 3 (Akt3), mRNA [NM_011785] |
KLA | 3.42 |
3.45 |
5.31 |
5.16 |
4.65 |
3.80 |
2.12 |
| ATP | 1.12 |
1.22 |
1.10 |
1.37 |
1.02 |
1.11 |
1.56 |
| KLA/ATP | 3.72 |
4.17 |
4.62 |
4.64 |
2.74 |
2.05 |
3.82 |
|
| Anapc1 | 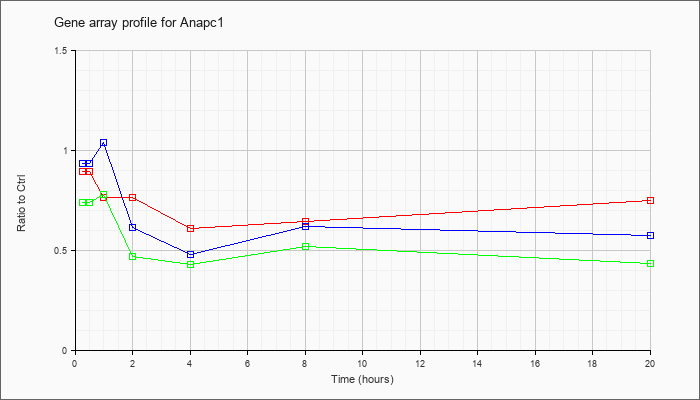 |
AK090134 |
nullipotent stem cell CRL-2070 NE cDNA, RIKEN full-length enriched library, clone:G431003J08 product:meiotic check point regulator, full insert sequence [AK090134] |
KLA | .79 |
.76 |
.72 |
.64 |
.56 |
.54 |
.76 |
| ATP | .99 |
1.04 |
1.11 |
.77 |
.48 |
.31 |
.39 |
| KLA/ATP | .72 |
.75 |
.73 |
.57 |
.42 |
.30 |
.25 |
|
| Anapc1 |  |
NM_008569 |
anaphase promoting complex subunit 1 (Anapc1), mRNA [NM_008569] |
KLA | 1.00 |
.89 |
.81 |
.89 |
.66 |
.75 |
.74 |
| ATP | .88 |
.79 |
.97 |
.46 |
.48 |
.93 |
.76 |
| KLA/ATP | .76 |
.77 |
.83 |
.37 |
.44 |
.74 |
.62 |
|
| Anapc10 | 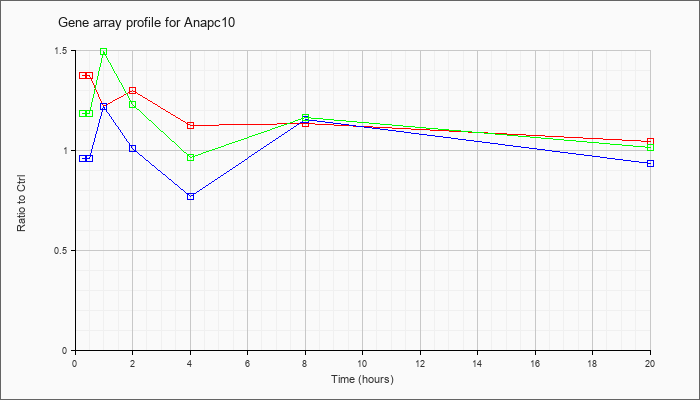 |
NM_026904 |
anaphase promoting complex subunit 10 (Anapc10), mRNA [NM_026904] |
KLA | 1.38 |
1.24 |
1.22 |
1.30 |
1.13 |
1.14 |
1.05 |
| ATP | .96 |
.99 |
1.22 |
1.01 |
.77 |
1.16 |
.94 |
| KLA/ATP | 1.19 |
1.24 |
1.50 |
1.23 |
.97 |
1.17 |
1.02 |
|
| Anapc11 | 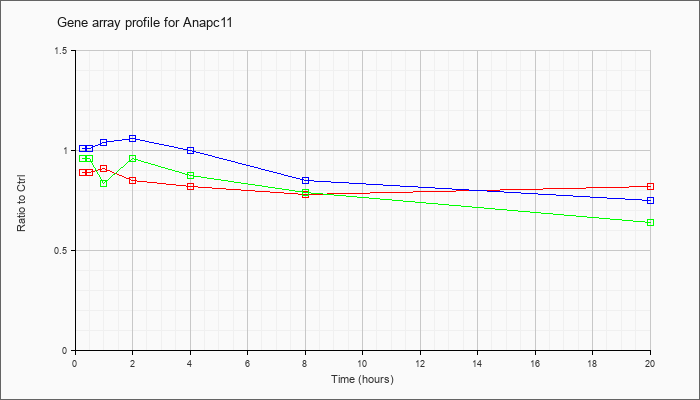 |
NM_001038230 |
anaphase promoting complex subunit 11 homolog (yeast) (Anapc11), transcript variant 1, mRNA [NM_001038230] |
KLA | .89 |
.90 |
.91 |
.85 |
.82 |
.78 |
.82 |
| ATP | 1.01 |
1.06 |
1.04 |
1.06 |
1.00 |
.85 |
.75 |
| KLA/ATP | .96 |
.93 |
.83 |
.96 |
.87 |
.79 |
.64 |
|
| Anapc2 | 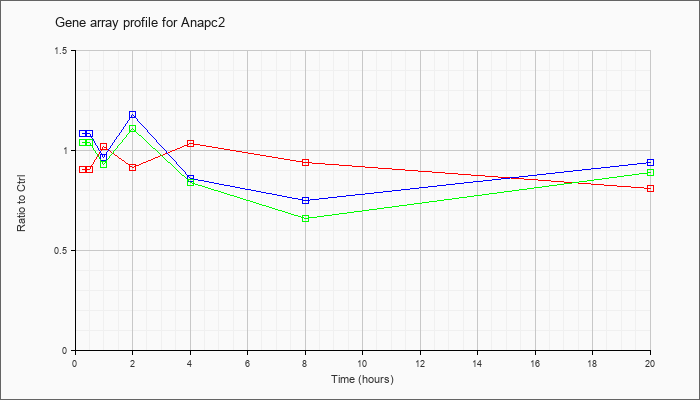 |
NM_175300 |
anaphase promoting complex subunit 2 (Anapc2), mRNA [NM_175300] |
KLA | .90 |
.92 |
1.02 |
.91 |
1.03 |
.94 |
.81 |
| ATP | 1.08 |
1.16 |
.96 |
1.18 |
.86 |
.75 |
.94 |
| KLA/ATP | 1.04 |
1.07 |
.93 |
1.11 |
.84 |
.66 |
.89 |
|
| Anapc4 | 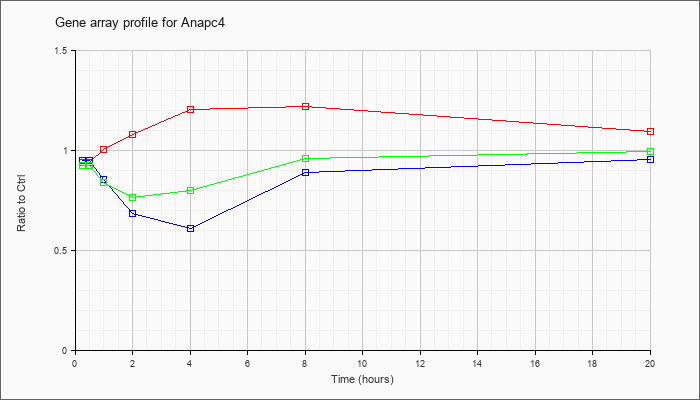 |
NM_024213 |
anaphase promoting complex subunit 4 (Anapc4), mRNA [NM_024213] |
KLA | .94 |
.99 |
1.01 |
1.08 |
1.21 |
1.22 |
1.09 |
| ATP | .95 |
.91 |
.86 |
.69 |
.61 |
.89 |
.96 |
| KLA/ATP | .93 |
.95 |
.84 |
.77 |
.80 |
.96 |
1.00 |
|
| Anapc5 |  |
NM_021505 |
anaphase-promoting complex subunit 5 (Anapc5), transcript variant 1, mRNA [NM_021505] |
KLA | .80 |
.82 |
.83 |
.82 |
.79 |
.84 |
1.10 |
| ATP | .99 |
.99 |
.89 |
.81 |
.79 |
.70 |
.71 |
| KLA/ATP | .82 |
.81 |
.72 |
.62 |
.63 |
.73 |
.68 |
|
| Anapc7 | 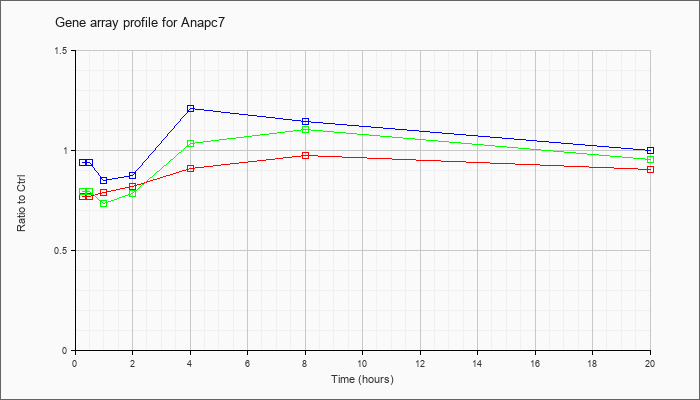 |
AK049968 |
adult male hippocampus cDNA, RIKEN full-length enriched library, clone:C630034H22 product:anaphase-promoting complex subunit 7, full insert sequence. [AK049968] |
KLA | .89 |
.86 |
.88 |
.85 |
.91 |
.95 |
.89 |
| ATP | .92 |
.89 |
.89 |
.86 |
.96 |
.99 |
1.00 |
| KLA/ATP | .89 |
.79 |
.83 |
.85 |
.93 |
.92 |
.87 |
|
| Anapc7 |  |
NM_019805 |
anaphase promoting complex subunit 7 (Anapc7), mRNA [NM_019805] |
KLA | .65 |
.69 |
.70 |
.79 |
.91 |
1.00 |
.92 |
| ATP | .96 |
.99 |
.81 |
.89 |
1.46 |
1.30 |
1.00 |
| KLA/ATP | .70 |
.73 |
.64 |
.72 |
1.14 |
1.28 |
1.04 |
|
| Apc | 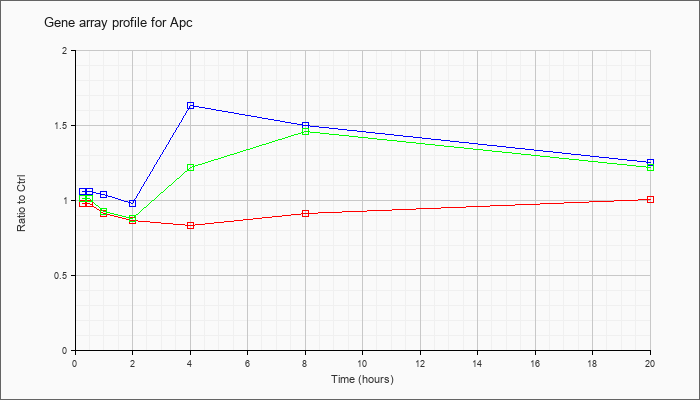 |
NM_007462 |
adenomatosis polyposis coli (Apc), mRNA [NM_007462] |
KLA | .98 |
.94 |
.91 |
.86 |
.83 |
.91 |
1.01 |
| ATP | 1.06 |
1.16 |
1.04 |
.97 |
1.63 |
1.50 |
1.25 |
| KLA/ATP | 1.01 |
1.01 |
.92 |
.88 |
1.22 |
1.46 |
1.21 |
|
| Apc2 | 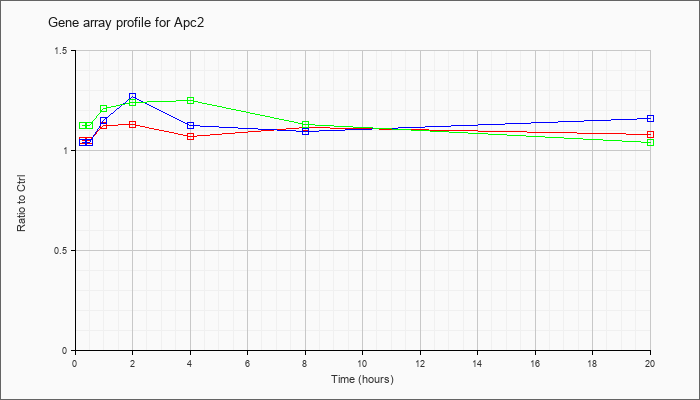 |
NM_011789 |
adenomatosis polyposis coli 2 (Apc2), mRNA [NM_011789] |
KLA | 1.05 |
1.15 |
1.12 |
1.13 |
1.07 |
1.11 |
1.08 |
| ATP | 1.04 |
.97 |
1.15 |
1.27 |
1.12 |
1.09 |
1.16 |
| KLA/ATP | 1.12 |
1.20 |
1.21 |
1.24 |
1.25 |
1.13 |
1.04 |
|
| Atf1 | 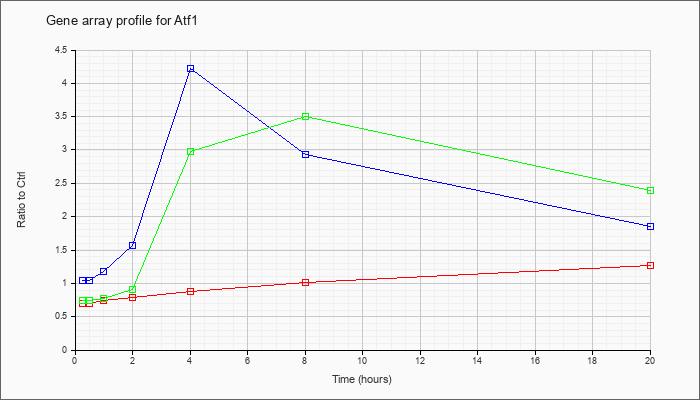 |
NM_007497 |
activating transcription factor 1 (Atf1), mRNA [NM_007497] |
KLA | .71 |
.73 |
.74 |
.79 |
.89 |
1.02 |
1.27 |
| ATP | 1.05 |
1.14 |
1.17 |
1.58 |
4.22 |
2.94 |
1.86 |
| KLA/ATP | .75 |
.78 |
.77 |
.90 |
2.98 |
3.50 |
2.39 |
|
| Atf2 | 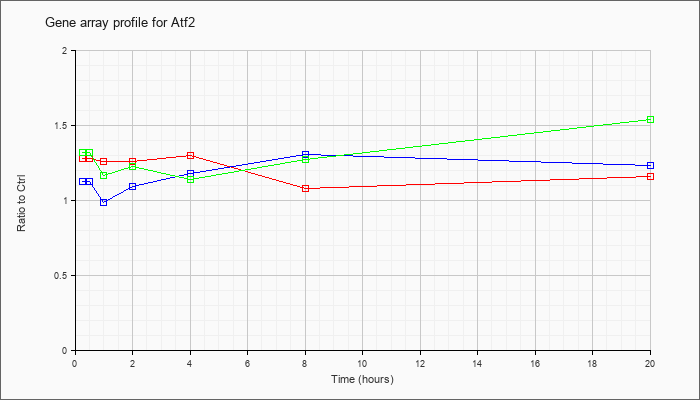 |
AK047405 |
10 days neonate cerebellum cDNA, RIKEN full-length enriched library, clone:B930056P03 product:activating transcription factor 2, full insert sequence. [AK047405] |
KLA | 1.01 |
1.00 |
1.02 |
1.04 |
1.03 |
1.03 |
1.00 |
| ATP | .99 |
1.00 |
.95 |
1.01 |
1.04 |
.98 |
1.03 |
| KLA/ATP | 1.07 |
1.00 |
.98 |
.99 |
1.00 |
1.03 |
.93 |
|
| Atf2 |  |
NM_001025093 |
activating transcription factor 2 (Atf2), transcript variant 1, mRNA [NM_001025093] |
KLA | 1.35 |
1.30 |
1.32 |
1.31 |
1.37 |
1.09 |
1.20 |
| ATP | 1.15 |
1.17 |
.99 |
1.11 |
1.22 |
1.39 |
1.28 |
| KLA/ATP | 1.38 |
1.42 |
1.21 |
1.28 |
1.17 |
1.33 |
1.69 |
|
| Atf3 |  |
NM_007498 |
activating transcription factor 3 (Atf3), mRNA [NM_007498] |
KLA | 1.95 |
2.23 |
1.99 |
2.03 |
1.86 |
2.12 |
1.97 |
| ATP | 1.15 |
2.05 |
4.52 |
7.49 |
3.91 |
1.67 |
1.74 |
| KLA/ATP | 2.02 |
2.85 |
3.90 |
6.24 |
6.00 |
3.32 |
3.79 |
|
| Atf4 | 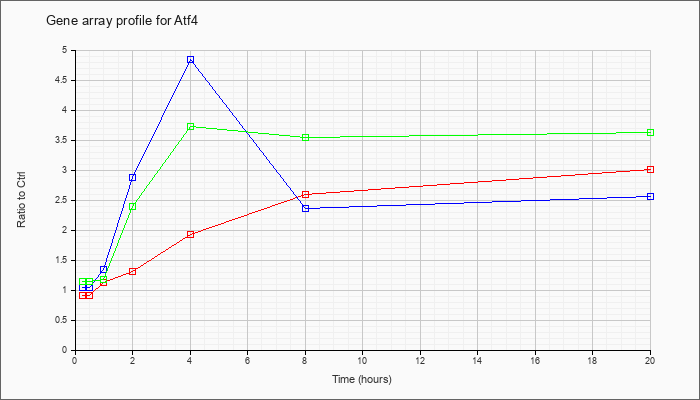 |
NM_009716 |
activating transcription factor 4 (Atf4), mRNA [NM_009716] |
KLA | .91 |
1.06 |
1.13 |
1.31 |
1.92 |
2.59 |
3.01 |
| ATP | 1.05 |
1.14 |
1.34 |
2.88 |
4.84 |
2.36 |
2.56 |
| KLA/ATP | 1.14 |
1.08 |
1.17 |
2.40 |
3.72 |
3.54 |
3.62 |
|
| Atm | 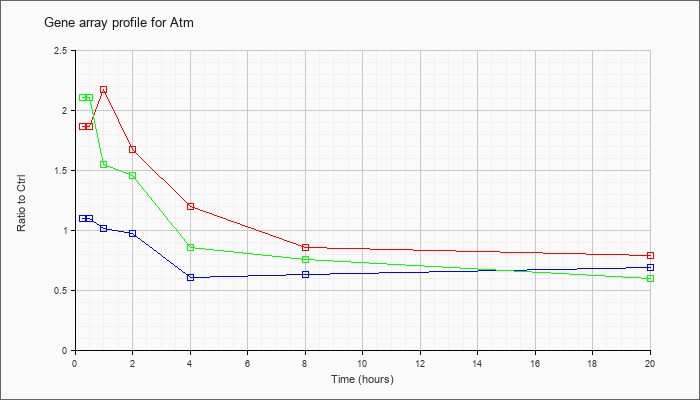 |
NM_007499 |
ataxia telangiectasia mutated homolog (human) (Atm), mRNA [NM_007499] |
KLA | 1.86 |
1.97 |
2.17 |
1.67 |
1.20 |
.86 |
.79 |
| ATP | 1.09 |
1.17 |
1.01 |
.97 |
.61 |
.63 |
.69 |
| KLA/ATP | 2.11 |
2.24 |
1.55 |
1.46 |
.85 |
.75 |
.59 |
|
BB113440
|  |
BB113440 |
gb|BB113440 RIKEN full-length enriched, adult male urinary bladder Mus musculus cDNA clone 9530041B19 3. [BB113440] |
KLA | .94 |
.95 |
.94 |
.95 |
1.05 |
.99 |
1.05 |
| ATP | 1.03 |
1.07 |
1.79 |
1.71 |
1.51 |
1.12 |
1.06 |
| KLA/ATP | .95 |
1.11 |
1.27 |
1.15 |
1.05 |
1.14 |
1.25 |
|
BI646741
| 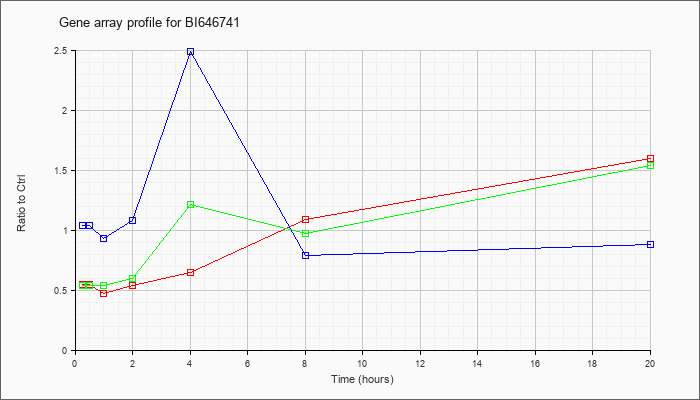 |
BI646741 |
gb|BI646741 603279769F1 NCI_CGAP_Mam3 Mus musculus cDNA clone IMAGE:5320025 5, mRNA sequence [BI646741] |
KLA | .55 |
.54 |
.47 |
.54 |
.65 |
1.09 |
1.60 |
| ATP | 1.04 |
.99 |
.93 |
1.08 |
2.49 |
.79 |
.88 |
| KLA/ATP | .54 |
.49 |
.54 |
.60 |
1.21 |
.97 |
1.54 |
|
| Bax | 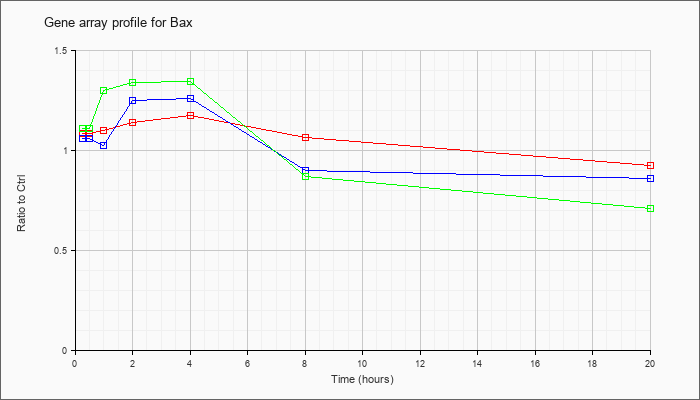 |
NM_007527 |
Bcl2-associated X protein (Bax), mRNA [NM_007527] |
KLA | 1.08 |
1.14 |
1.10 |
1.14 |
1.17 |
1.06 |
.92 |
| ATP | 1.06 |
1.05 |
1.02 |
1.25 |
1.26 |
.90 |
.86 |
| KLA/ATP | 1.11 |
1.19 |
1.30 |
1.34 |
1.34 |
.87 |
.71 |
|
| Bcl2l1 | 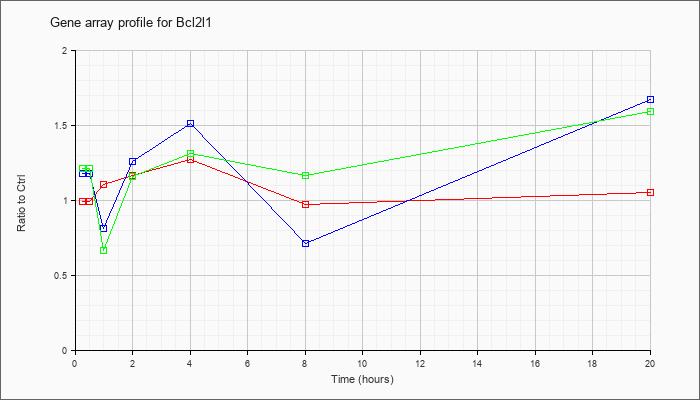 |
NM_009743 |
Bcl2-like 1 (Bcl2l1), nuclear gene encoding mitochondrial protein, mRNA [NM_009743] |
KLA | .99 |
1.02 |
1.10 |
1.16 |
1.27 |
.97 |
1.05 |
| ATP | 1.18 |
1.42 |
.81 |
1.26 |
1.51 |
.71 |
1.67 |
| KLA/ATP | 1.21 |
1.30 |
.66 |
1.15 |
1.31 |
1.17 |
1.59 |
|
| Bub1b | 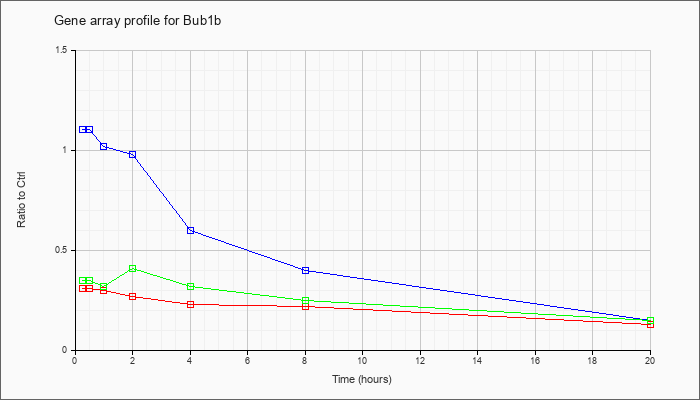 |
NM_009773 |
budding uninhibited by benzimidazoles 1 homolog, beta (S. cerevisiae) (Bub1b), mRNA [NM_009773] |
KLA | .31 |
.30 |
.30 |
.27 |
.23 |
.22 |
.13 |
| ATP | 1.10 |
1.06 |
1.02 |
.98 |
.60 |
.40 |
.15 |
| KLA/ATP | .35 |
.30 |
.32 |
.41 |
.32 |
.25 |
.15 |
|
| Bub3 | 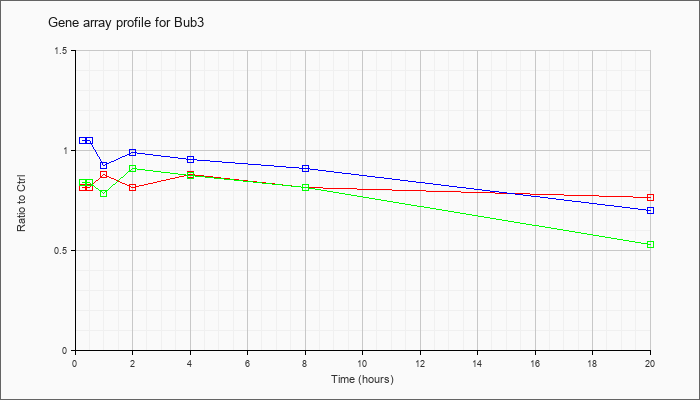 |
AK083742 |
9 days embryo whole body cDNA, RIKEN full-length enriched library, clone:D030073O17 product:unclassifiable, full insert sequence. [AK083742] |
KLA | .77 |
.74 |
.83 |
.76 |
.91 |
.80 |
.78 |
| ATP | 1.02 |
.74 |
.70 |
.69 |
1.08 |
1.05 |
.79 |
| KLA/ATP | .65 |
.65 |
.70 |
.72 |
1.02 |
.88 |
.64 |
|
| Bub3 |  |
NM_009774 |
budding uninhibited by benzimidazoles 3 homolog (S. cerevisiae) (Bub3), mRNA [NM_009774] |
KLA | .84 |
.90 |
.90 |
.84 |
.86 |
.82 |
.76 |
| ATP | 1.07 |
1.10 |
1.04 |
1.14 |
.89 |
.84 |
.65 |
| KLA/ATP | .93 |
.97 |
.83 |
1.00 |
.80 |
.78 |
.48 |
|
| Calr | 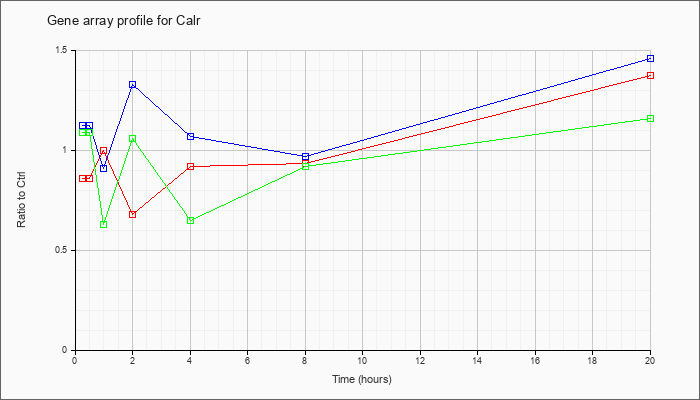 |
NM_007591 |
calreticulin (Calr), mRNA [NM_007591] |
KLA | .86 |
.95 |
1.00 |
.68 |
.92 |
.93 |
1.37 |
| ATP | 1.12 |
1.34 |
.91 |
1.33 |
1.07 |
.97 |
1.46 |
| KLA/ATP | 1.09 |
1.02 |
.63 |
1.06 |
.65 |
.92 |
1.16 |
|
| Canx | 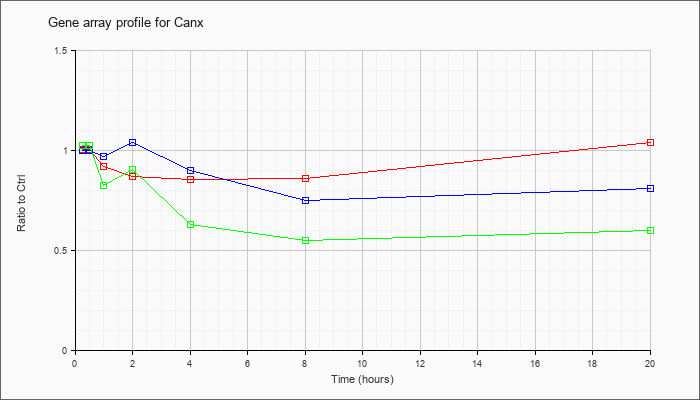 |
NM_007597 |
calnexin (Canx), transcript variant 1, mRNA [NM_007597] |
KLA | 1.00 |
1.01 |
.92 |
.87 |
.85 |
.86 |
1.04 |
| ATP | 1.00 |
1.13 |
.97 |
1.04 |
.90 |
.75 |
.81 |
| KLA/ATP | 1.02 |
1.10 |
.82 |
.90 |
.63 |
.55 |
.60 |
|
| Ccnb2 | 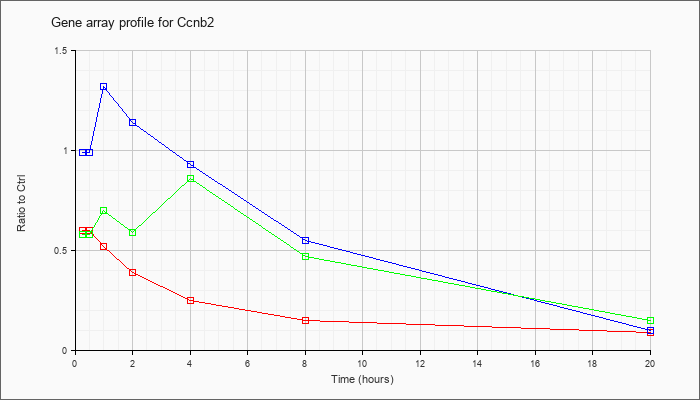 |
NM_007630 |
cyclin B2 (Ccnb2), mRNA [NM_007630] |
KLA | .60 |
.51 |
.52 |
.39 |
.25 |
.15 |
.09 |
| ATP | .99 |
.89 |
1.32 |
1.14 |
.93 |
.55 |
.10 |
| KLA/ATP | .58 |
.52 |
.70 |
.59 |
.86 |
.47 |
.15 |
|
| Ccnd1 | 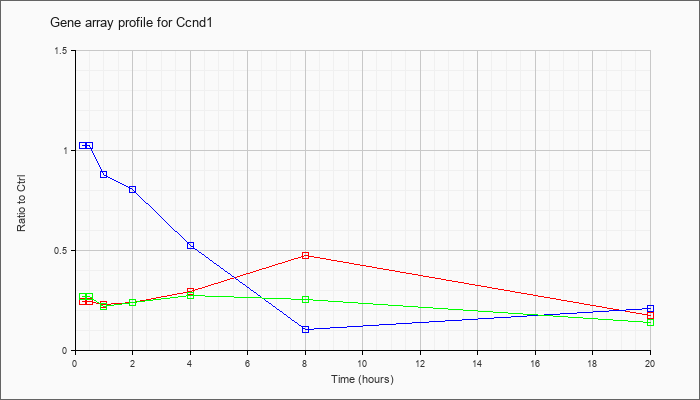 |
S78355 |
gb|Cyl-1=cyclin D1 [mice, BALB/c, brain, mRNA, 3737 nt]. [S78355] |
KLA | .24 |
.23 |
.23 |
.24 |
.29 |
.47 |
.17 |
| ATP | 1.02 |
1.02 |
.88 |
.80 |
.52 |
.10 |
.21 |
| KLA/ATP | .27 |
.23 |
.22 |
.24 |
.27 |
.25 |
.14 |
|
| Ccnd2 | 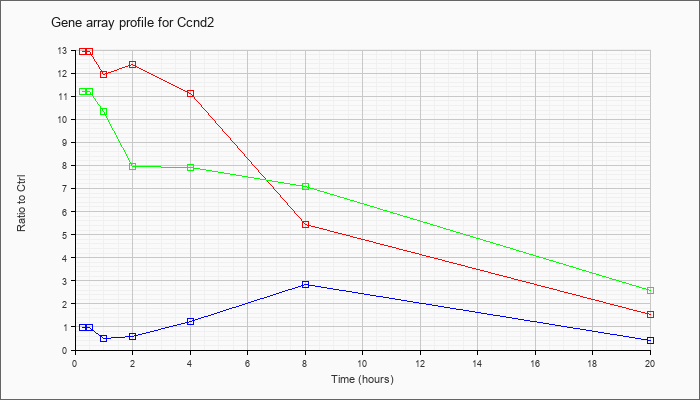 |
NM_009829 |
cyclin D2 (Ccnd2), mRNA [NM_009829] |
KLA | 12.92 |
13.66 |
11.94 |
12.37 |
11.11 |
5.44 |
1.53 |
| ATP | .98 |
.75 |
.50 |
.57 |
1.24 |
2.83 |
.40 |
| KLA/ATP | 11.19 |
11.62 |
10.34 |
7.96 |
7.92 |
7.10 |
2.57 |
|
| Ccnd3 | 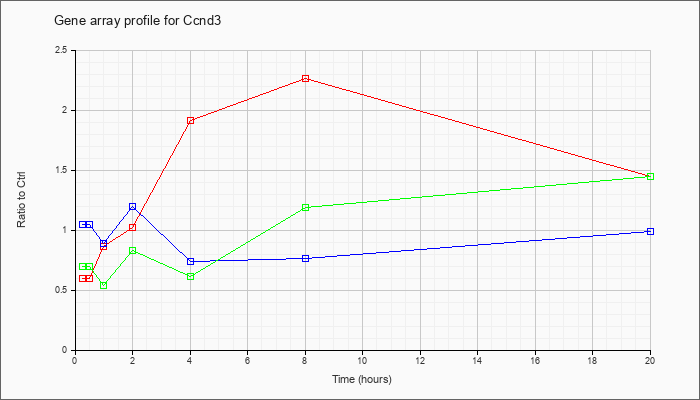 |
NM_007632 |
cyclin D3 (Ccnd3), transcript variant 1, mRNA [NM_007632] |
KLA | .60 |
.65 |
.86 |
1.02 |
1.91 |
2.26 |
1.45 |
| ATP | 1.05 |
1.14 |
.89 |
1.20 |
.74 |
.76 |
.99 |
| KLA/ATP | .70 |
.67 |
.54 |
.83 |
.61 |
1.19 |
1.45 |
|
| Cd3d | 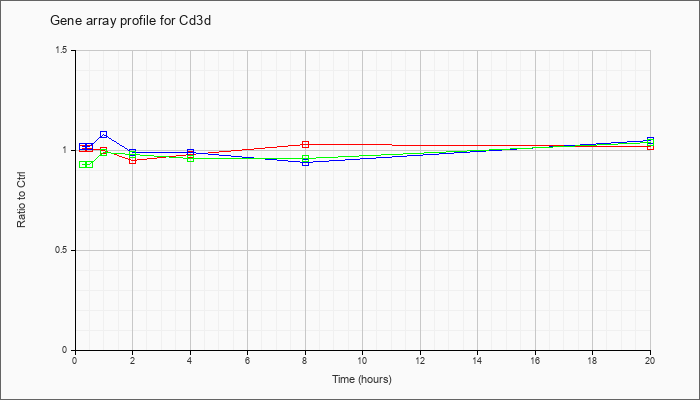 |
NM_013487 |
CD3 antigen, delta polypeptide (Cd3d), mRNA [NM_013487] |
KLA | 1.01 |
1.05 |
1.00 |
.95 |
.98 |
1.03 |
1.02 |
| ATP | 1.02 |
.94 |
1.08 |
.99 |
.99 |
.94 |
1.05 |
| KLA/ATP | .93 |
.91 |
.99 |
.98 |
.96 |
.96 |
1.04 |
|
| Cd3e | 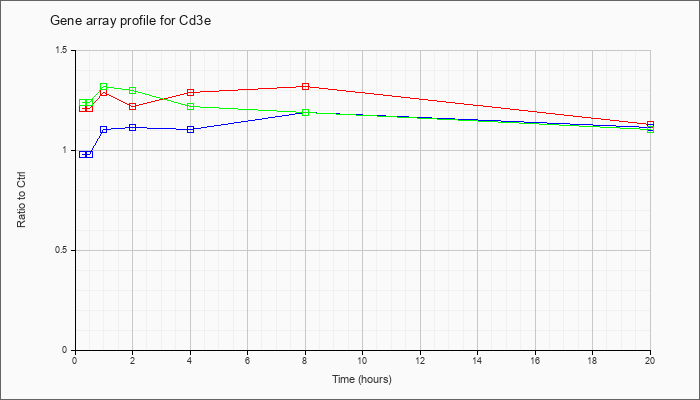 |
NM_007648 |
CD3 antigen, epsilon polypeptide (Cd3e), mRNA [NM_007648] |
KLA | 1.21 |
1.25 |
1.29 |
1.22 |
1.29 |
1.32 |
1.13 |
| ATP | .98 |
1.00 |
1.10 |
1.11 |
1.10 |
1.19 |
1.11 |
| KLA/ATP | 1.24 |
1.30 |
1.32 |
1.30 |
1.22 |
1.19 |
1.10 |
|
| Cd3g | 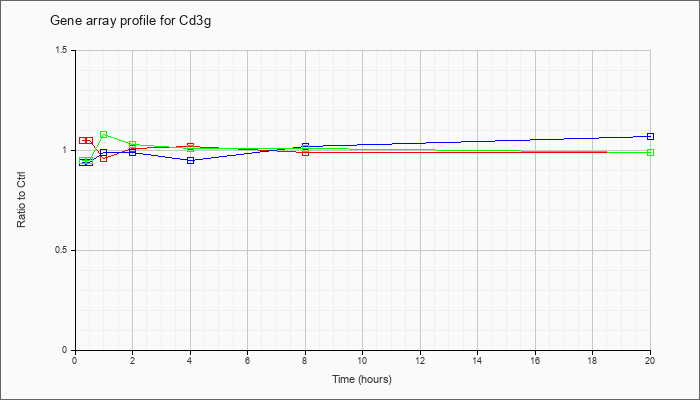 |
NM_009850 |
CD3 antigen, gamma polypeptide (Cd3g), mRNA [NM_009850] |
KLA | 1.05 |
1.01 |
.96 |
1.01 |
1.02 |
.99 |
.99 |
| ATP | .94 |
.97 |
.99 |
.99 |
.95 |
1.02 |
1.07 |
| KLA/ATP | .95 |
.92 |
1.08 |
1.03 |
1.01 |
1.01 |
.99 |
|
| Cd40 | 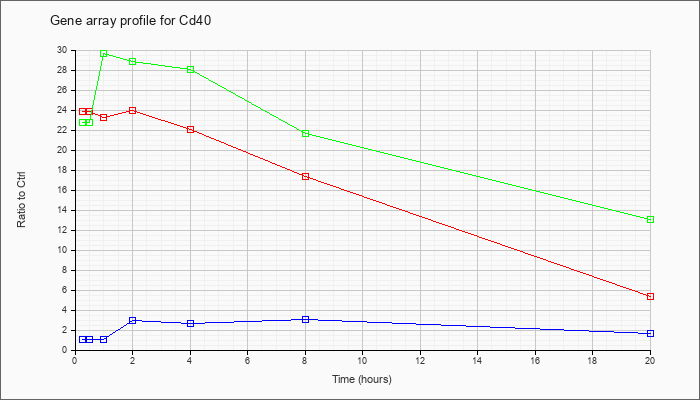 |
NM_170701 |
CD40 antigen (Cd40), transcript variant 3, mRNA [NM_170701] |
KLA | 23.87 |
23.62 |
23.26 |
23.98 |
22.07 |
17.37 |
5.34 |
| ATP | 1.00 |
1.01 |
1.06 |
2.96 |
2.65 |
3.03 |
1.70 |
| KLA/ATP | 22.77 |
23.79 |
29.67 |
28.85 |
28.08 |
21.62 |
13.02 |
|
| Cdc16 | 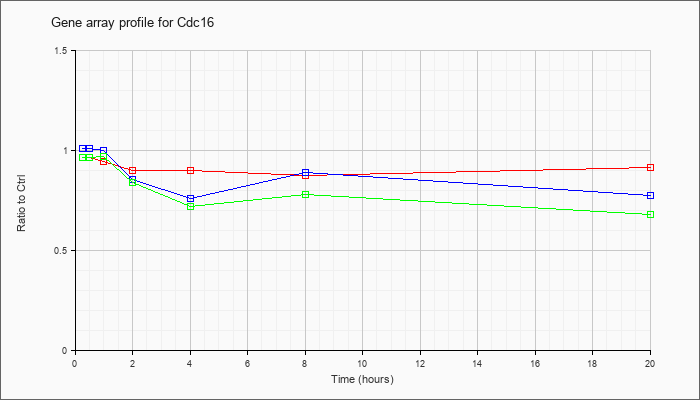 |
AK012511 |
11 days embryo whole body cDNA, RIKEN full-length enriched library, clone:2700071J12 product:unclassifiable, full insert sequence. [AK012511] |
KLA | .97 |
.92 |
.89 |
.90 |
.93 |
.92 |
.99 |
| ATP | .97 |
.94 |
.89 |
.92 |
.89 |
.94 |
.91 |
| KLA/ATP | .98 |
.88 |
.99 |
.93 |
.80 |
.89 |
.82 |
|
| Cdc16 |  |
NM_027276 |
CDC16 cell division cycle 16 homolog (S. cerevisiae) (Cdc16), mRNA [NM_027276] |
KLA | .96 |
.98 |
.97 |
.90 |
.89 |
.85 |
.88 |
| ATP | 1.03 |
1.02 |
1.06 |
.82 |
.70 |
.87 |
.71 |
| KLA/ATP | .96 |
.93 |
.96 |
.79 |
.68 |
.73 |
.61 |
|
| Cdc20 | 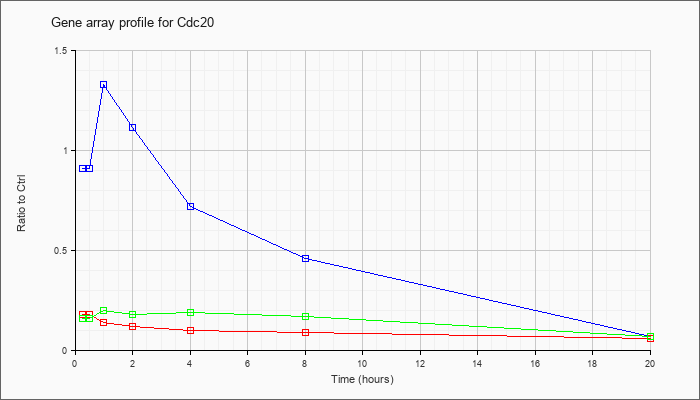 |
NM_023223 |
cell division cycle 20 homolog (S. cerevisiae) (Cdc20), mRNA [NM_023223] |
KLA | .18 |
.15 |
.14 |
.12 |
.10 |
.09 |
.06 |
| ATP | .91 |
.88 |
1.33 |
1.11 |
.72 |
.46 |
.07 |
| KLA/ATP | .16 |
.15 |
.20 |
.18 |
.19 |
.17 |
.07 |
|
| Cdc23 |  |
NM_178347 |
CDC23 (cell division cycle 23, yeast, homolog) (Cdc23), mRNA [NM_178347] |
KLA | .74 |
.76 |
.75 |
.74 |
.72 |
.79 |
.79 |
| ATP | 1.06 |
1.04 |
.91 |
.71 |
.48 |
.60 |
.71 |
| KLA/ATP | .76 |
.77 |
.73 |
.66 |
.46 |
.51 |
.59 |
|
| Cdc26 | 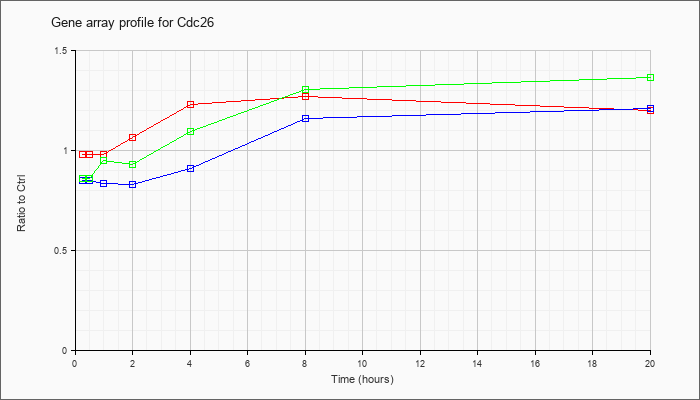 |
AK041840 |
3 days neonate thymus cDNA, RIKEN full-length enriched library, clone:A630041F01 product:unclassifiable, full insert sequence. [AK041840] |
KLA | .92 |
.88 |
.95 |
.88 |
1.08 |
.99 |
1.15 |
| ATP | .72 |
.64 |
.61 |
.75 |
.91 |
.85 |
1.04 |
| KLA/ATP | .84 |
.75 |
.66 |
.87 |
.94 |
.95 |
.99 |
|
| Cdc26 |  |
NM_139291 |
cell division cycle 26 (Cdc26), mRNA [NM_139291] |
KLA | 1.04 |
1.07 |
1.01 |
1.25 |
1.38 |
1.55 |
1.25 |
| ATP | .98 |
.97 |
1.06 |
.91 |
.91 |
1.47 |
1.38 |
| KLA/ATP | .88 |
1.02 |
1.24 |
.99 |
1.25 |
1.66 |
1.74 |
|
| Cdc27 | 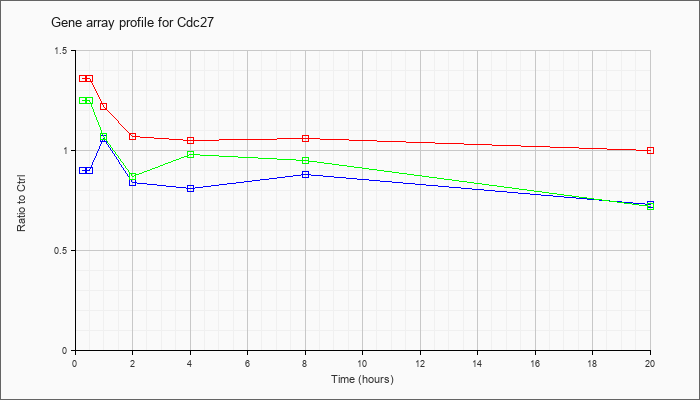 |
NM_145436 |
cell division cycle 27 homolog (S. cerevisiae) (Cdc27), mRNA [NM_145436] |
KLA | 1.36 |
1.25 |
1.22 |
1.07 |
1.05 |
1.06 |
1.00 |
| ATP | .90 |
.97 |
1.06 |
.84 |
.81 |
.88 |
.73 |
| KLA/ATP | 1.25 |
1.22 |
1.07 |
.87 |
.98 |
.95 |
.72 |
|
| Cdk4 | 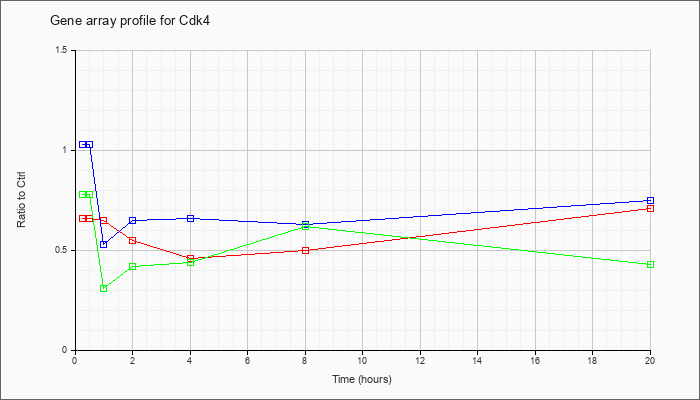 |
NM_009870 |
cyclin-dependent kinase 4 (Cdk4), mRNA [NM_009870] |
KLA | .66 |
.72 |
.65 |
.55 |
.46 |
.50 |
.71 |
| ATP | 1.03 |
1.02 |
.53 |
.65 |
.66 |
.63 |
.75 |
| KLA/ATP | .78 |
.61 |
.31 |
.42 |
.44 |
.62 |
.43 |
|
| Cdkn1a | 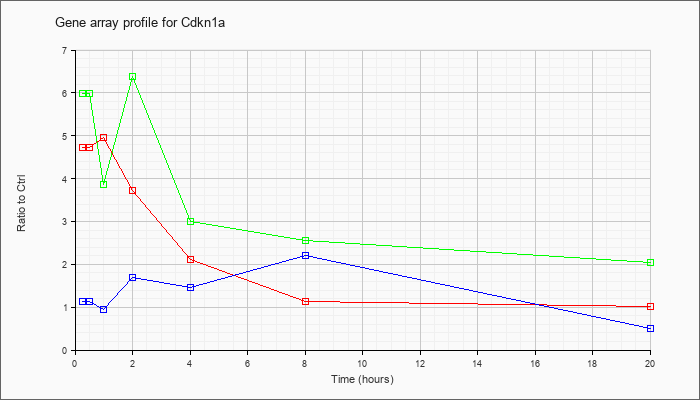 |
NM_007669 |
cyclin-dependent kinase inhibitor 1A (P21) (Cdkn1a), transcript variant 1, mRNA [NM_007669] |
KLA | 4.72 |
4.90 |
4.96 |
3.73 |
2.11 |
1.14 |
1.02 |
| ATP | 1.14 |
1.52 |
.95 |
1.69 |
1.47 |
2.21 |
.50 |
| KLA/ATP | 5.99 |
7.08 |
3.86 |
6.37 |
3.00 |
2.55 |
2.05 |
|
| Cdkn2a | 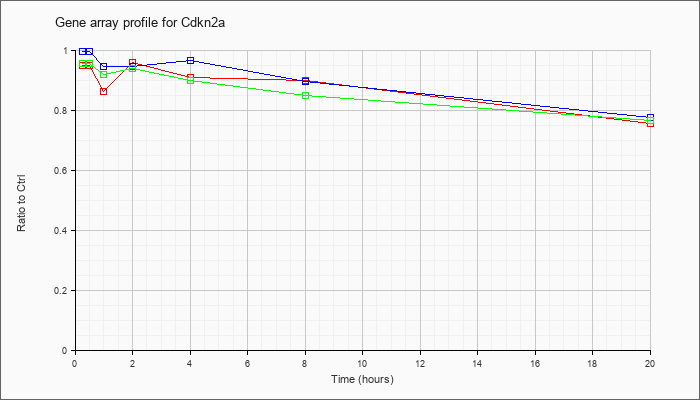 |
NM_009877 |
cyclin-dependent kinase inhibitor 2A (Cdkn2a), transcript variant 1, mRNA [NM_009877] |
KLA | .95 |
.95 |
.86 |
.96 |
.91 |
.90 |
.75 |
| ATP | 1.00 |
.99 |
.95 |
.95 |
.97 |
.89 |
.77 |
| KLA/ATP | .95 |
.96 |
.92 |
.94 |
.90 |
.85 |
.76 |
|
| Cdkn2b | 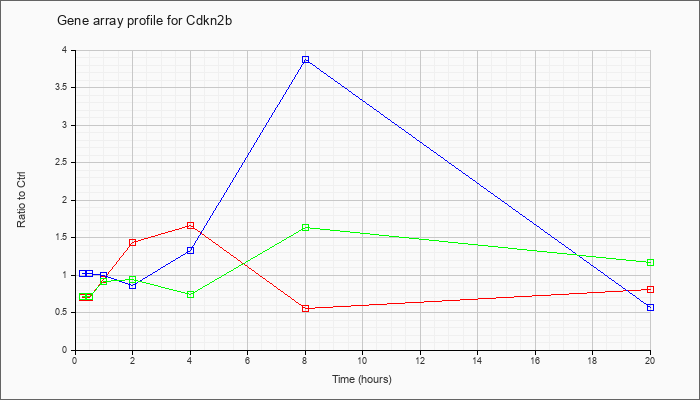 |
NM_007670 |
cyclin-dependent kinase inhibitor 2B (p15, inhibits CDK4) (Cdkn2b), mRNA [NM_007670] |
KLA | .70 |
.75 |
.94 |
1.44 |
1.66 |
.55 |
.81 |
| ATP | 1.02 |
.89 |
1.00 |
.86 |
1.33 |
3.87 |
.57 |
| KLA/ATP | .71 |
.77 |
.91 |
.94 |
.74 |
1.64 |
1.17 |
|
| Cdkn2c | 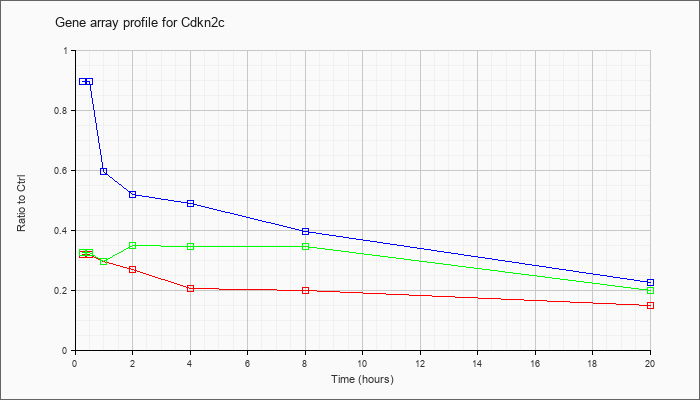 |
NM_007671 |
cyclin-dependent kinase inhibitor 2C (p18, inhibits CDK4) (Cdkn2c), mRNA [NM_007671] |
KLA | .32 |
.29 |
.29 |
.27 |
.20 |
.20 |
.15 |
| ATP | .88 |
.87 |
.62 |
.52 |
.49 |
.39 |
.23 |
| KLA/ATP | .30 |
.29 |
.30 |
.34 |
.34 |
.34 |
.20 |
|
| Cdkn2c |  |
U19596 |
Cdk4 and Cdk6 inhibitor p18 protein mRNA, complete cds. [U19596] |
KLA | .32 |
.32 |
.30 |
.27 |
.21 |
.20 |
.15 |
| ATP | .91 |
.90 |
.57 |
.52 |
.49 |
.40 |
.22 |
| KLA/ATP | .35 |
.31 |
.29 |
.36 |
.35 |
.35 |
.20 |
|
| Chek1 | 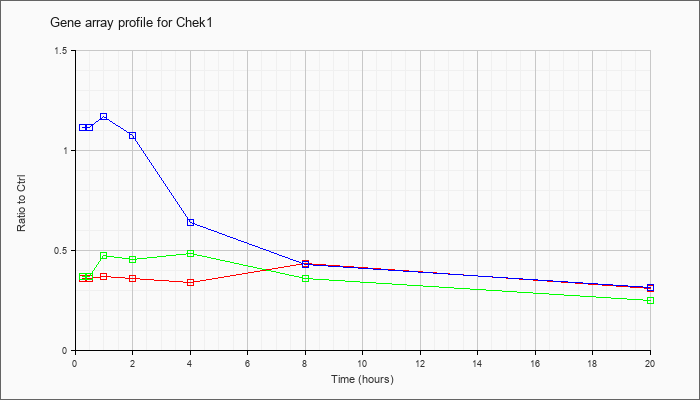 |
NM_007691 |
checkpoint kinase 1 homolog (S. pombe) (Chek1), mRNA [NM_007691] |
KLA | .36 |
.35 |
.37 |
.36 |
.34 |
.44 |
.31 |
| ATP | 1.12 |
1.14 |
1.17 |
1.08 |
.64 |
.43 |
.32 |
| KLA/ATP | .37 |
.36 |
.48 |
.46 |
.49 |
.36 |
.25 |
|
| Chek2 | 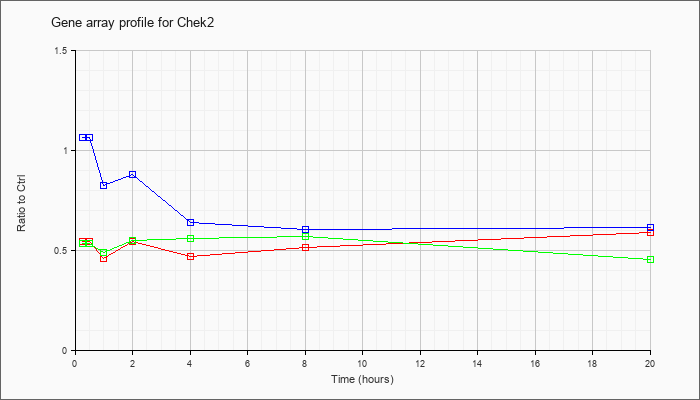 |
NM_016681 |
CHK2 checkpoint homolog (S. pombe) (Chek2), mRNA [NM_016681] |
KLA | .55 |
.54 |
.46 |
.55 |
.47 |
.52 |
.59 |
| ATP | 1.07 |
1.10 |
.83 |
.88 |
.64 |
.61 |
.62 |
| KLA/ATP | .54 |
.51 |
.49 |
.55 |
.56 |
.57 |
.46 |
|
| Chuk | 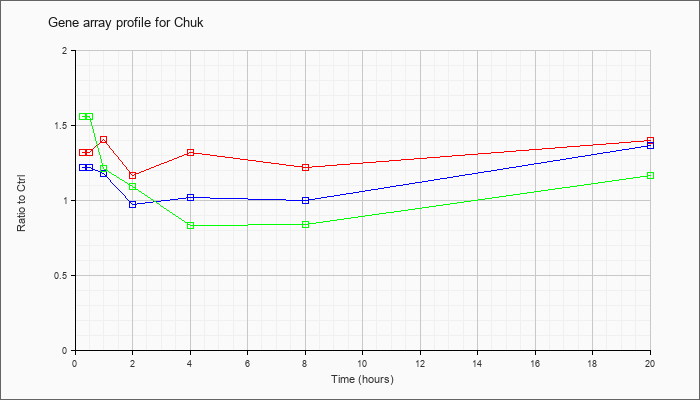 |
NM_007700 |
conserved helix-loop-helix ubiquitous kinase (Chuk), mRNA [NM_007700] |
KLA | 1.32 |
1.42 |
1.41 |
1.17 |
1.32 |
1.22 |
1.40 |
| ATP | 1.22 |
1.30 |
1.18 |
.97 |
1.02 |
1.00 |
1.37 |
| KLA/ATP | 1.56 |
1.61 |
1.21 |
1.09 |
.83 |
.84 |
1.17 |
|
| Creb1 | 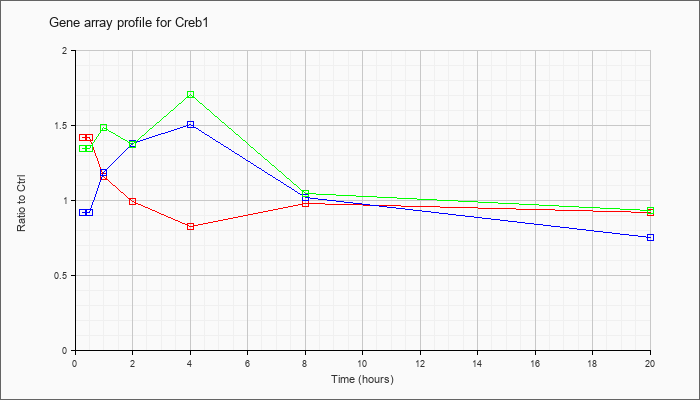 |
NM_009952 |
cAMP responsive element binding protein 1 (Creb1), transcript variant B, mRNA [NM_009952] |
KLA | 1.42 |
1.33 |
1.16 |
.99 |
.83 |
.97 |
.92 |
| ATP | .92 |
.91 |
1.18 |
1.38 |
1.51 |
1.02 |
.75 |
| KLA/ATP | 1.34 |
1.31 |
1.49 |
1.37 |
1.71 |
1.05 |
.93 |
|
| Crebbp | 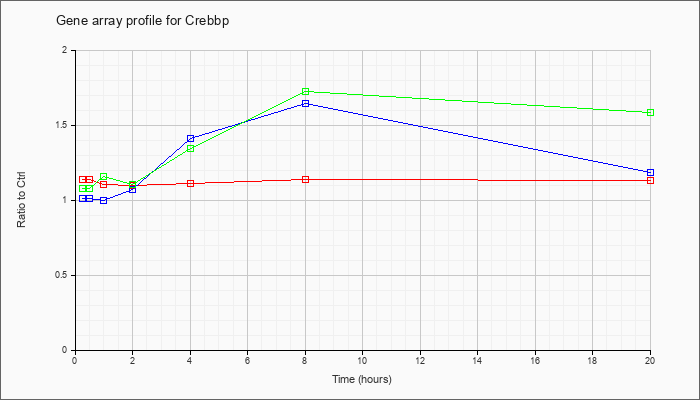 |
AK048818 |
0 day neonate cerebellum cDNA, RIKEN full-length enriched library, clone:C230072N21 product:hypothetical protein, full insert sequence [AK048818] |
KLA | 1.44 |
1.19 |
1.26 |
1.25 |
1.25 |
1.36 |
1.24 |
| ATP | .98 |
.90 |
1.04 |
1.11 |
1.95 |
2.72 |
1.51 |
| KLA/ATP | 1.17 |
1.27 |
1.45 |
1.28 |
1.98 |
2.80 |
2.52 |
|
| Crebbp |  |
NM_001025432 |
CREB binding protein (Crebbp), mRNA [NM_001025432] |
KLA | .97 |
1.01 |
.98 |
1.06 |
1.05 |
.99 |
1.07 |
| ATP | 1.07 |
1.05 |
.96 |
1.01 |
1.15 |
1.14 |
1.00 |
| KLA/ATP | 1.01 |
.97 |
.97 |
1.00 |
1.03 |
1.23 |
1.13 |
|
| Crebbp |  |
S66385 |
gb|CREB-binding protein [mice, brain, mRNA Partial, 7326 nt]. [S66385] |
KLA | 1.00 |
.98 |
1.08 |
.98 |
1.04 |
1.07 |
1.08 |
| ATP | .99 |
.98 |
.99 |
1.09 |
1.13 |
1.08 |
1.05 |
| KLA/ATP | 1.06 |
1.00 |
1.05 |
1.03 |
1.02 |
1.14 |
1.11 |
|
| Crem | 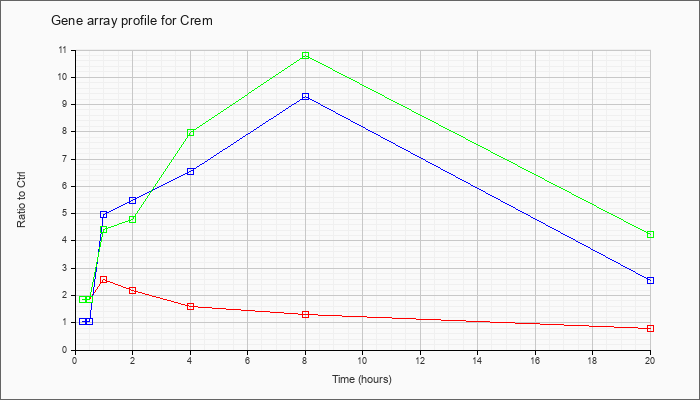 |
NM_013498 |
cAMP responsive element modulator (Crem), transcript variant 3, mRNA [NM_013498] |
KLA | 1.85 |
2.02 |
2.58 |
2.17 |
1.59 |
1.31 |
.79 |
| ATP | 1.04 |
1.44 |
4.98 |
5.47 |
6.53 |
9.31 |
2.55 |
| KLA/ATP | 1.84 |
2.15 |
4.41 |
4.79 |
7.98 |
10.80 |
4.25 |
|
| Crtc2 | 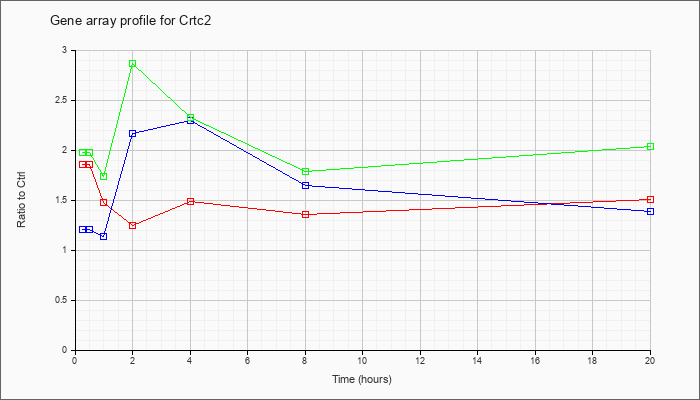 |
NM_028881 |
CREB regulated transcription coactivator 2 (Crtc2), mRNA [NM_028881] |
KLA | 1.86 |
1.66 |
1.48 |
1.25 |
1.49 |
1.36 |
1.51 |
| ATP | 1.21 |
1.19 |
1.14 |
2.17 |
2.30 |
1.65 |
1.39 |
| KLA/ATP | 1.98 |
1.95 |
1.74 |
2.87 |
2.33 |
1.79 |
2.04 |
|
| Crtc3 |  |
NM_173863 |
CREB regulated transcription coactivator 3 (Crtc3), mRNA [NM_173863] |
KLA | .83 |
.94 |
.88 |
.94 |
1.00 |
1.06 |
1.05 |
| ATP | 1.11 |
1.12 |
1.11 |
1.27 |
1.34 |
1.27 |
1.15 |
| KLA/ATP | .95 |
.94 |
.99 |
1.00 |
1.26 |
1.47 |
1.45 |
|
| Csf2 | 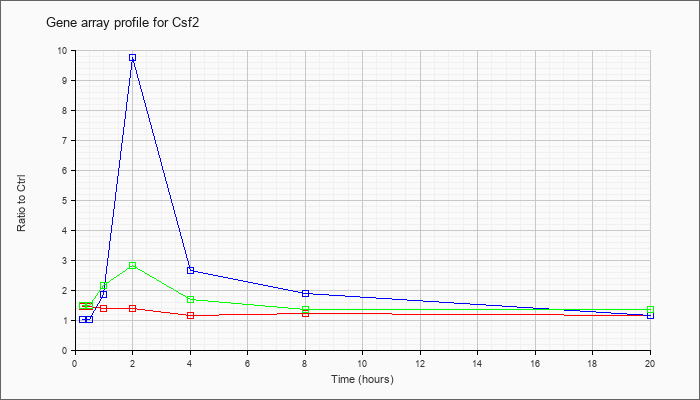 |
NM_009969 |
colony stimulating factor 2 (granulocyte-macrophage) (Csf2), mRNA [NM_009969] |
KLA | 1.44 |
1.53 |
1.38 |
1.37 |
1.15 |
1.21 |
1.14 |
| ATP | 1.01 |
1.02 |
1.85 |
9.73 |
2.64 |
1.90 |
1.14 |
| KLA/ATP | 1.49 |
1.84 |
2.14 |
2.80 |
1.67 |
1.36 |
1.35 |
|
| Ctnnb1 | 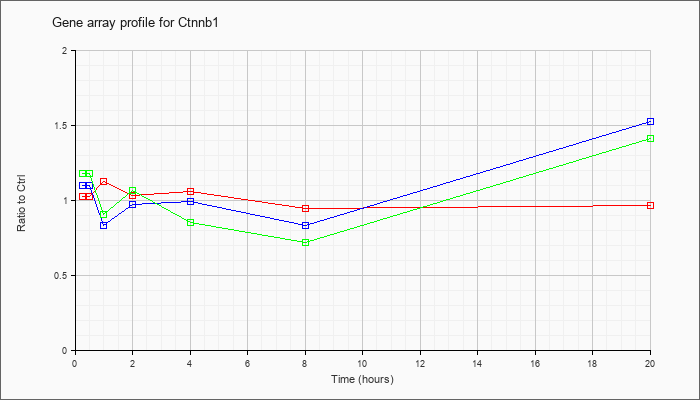 |
AK020013 |
adult male thymus cDNA, RIKEN full-length enriched library, clone:5830416C23 product:catenin beta, full insert sequence. [AK020013] |
KLA | 1.06 |
1.15 |
1.00 |
1.06 |
1.27 |
1.14 |
1.03 |
| ATP | .85 |
.57 |
1.74 |
3.36 |
4.13 |
2.19 |
1.55 |
| KLA/ATP | 1.12 |
.87 |
2.47 |
3.61 |
3.79 |
2.21 |
2.06 |
|
| Ctnnb1 |  |
NM_007614 |
catenin (cadherin associated protein), beta 1 (Ctnnb1), mRNA [NM_007614] |
KLA | 1.02 |
1.07 |
1.13 |
1.03 |
1.04 |
.93 |
.96 |
| ATP | 1.12 |
1.14 |
.75 |
.77 |
.73 |
.72 |
1.52 |
| KLA/ATP | 1.18 |
1.17 |
.77 |
.85 |
.60 |
.59 |
1.35 |
|
| Dlg1 | 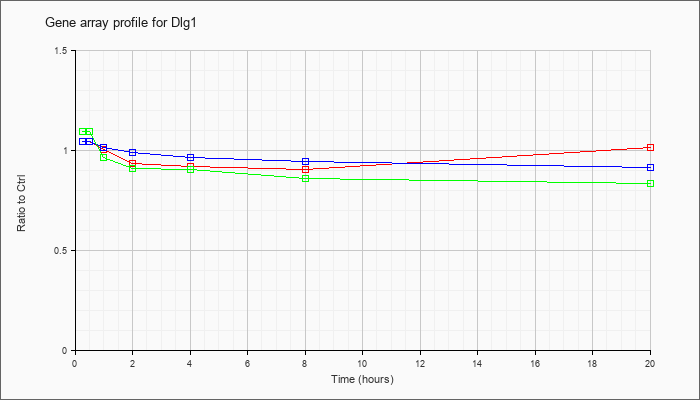 |
AK031364 |
13 days embryo male testis cDNA, RIKEN full-length enriched library, clone:6030413K05 product:discs, large homolog 1 (Drosophila), full insert sequence. [AK031364] |
KLA | .92 |
.85 |
.88 |
.82 |
.95 |
.91 |
.98 |
| ATP | 1.11 |
1.01 |
.90 |
.87 |
1.03 |
.93 |
.92 |
| KLA/ATP | .94 |
.93 |
.80 |
.81 |
1.01 |
.90 |
.87 |
|
| Dlg1 |  |
NM_007862 |
discs, large homolog 1 (Drosophila) (Dlg1), mRNA [NM_007862] |
KLA | 1.17 |
1.11 |
1.13 |
1.05 |
.89 |
.91 |
1.05 |
| ATP | .98 |
.97 |
1.13 |
1.12 |
.90 |
.96 |
.91 |
| KLA/ATP | 1.24 |
1.11 |
1.14 |
1.02 |
.80 |
.83 |
.80 |
|
| Dvl1 | 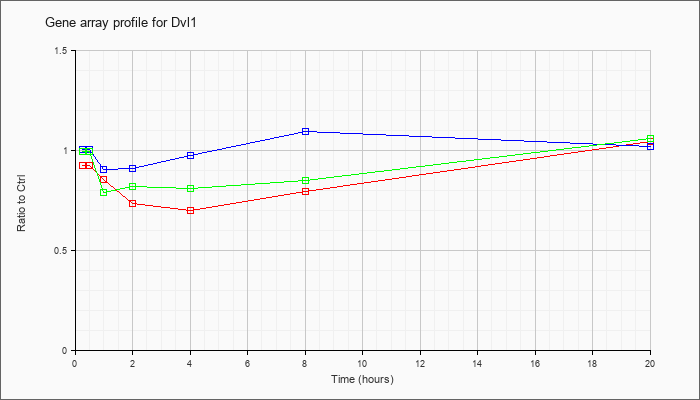 |
NM_010091 |
dishevelled, dsh homolog 1 (Drosophila) (Dvl1), mRNA [NM_010091] |
KLA | .93 |
.91 |
.86 |
.74 |
.70 |
.80 |
1.05 |
| ATP | 1.01 |
1.01 |
.91 |
.91 |
.98 |
1.10 |
1.02 |
| KLA/ATP | 1.00 |
.93 |
.79 |
.82 |
.81 |
.85 |
1.06 |
|
| Dvl2 | 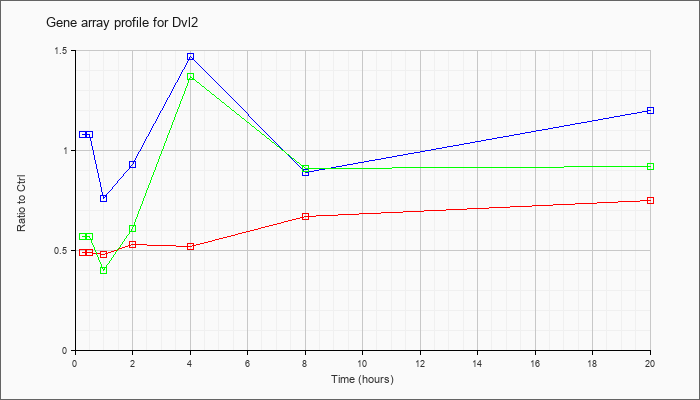 |
NM_007888 |
dishevelled 2, dsh homolog (Drosophila) (Dvl2), mRNA [NM_007888] |
KLA | .49 |
.52 |
.48 |
.53 |
.52 |
.67 |
.75 |
| ATP | 1.08 |
1.08 |
.76 |
.93 |
1.47 |
.89 |
1.20 |
| KLA/ATP | .57 |
.47 |
.40 |
.61 |
1.37 |
.91 |
.92 |
|
| Dvl3 | 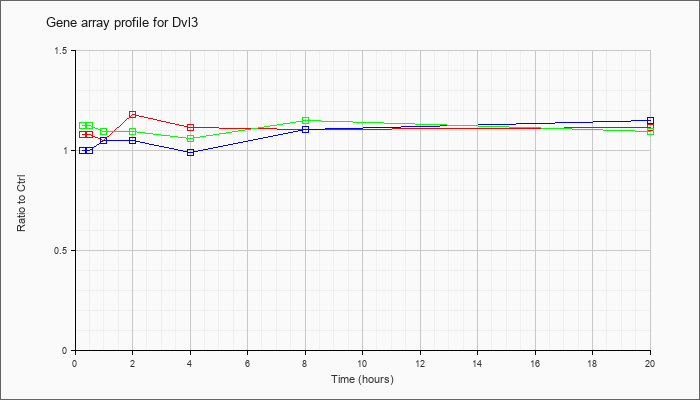 |
NM_007889 |
dishevelled 3, dsh homolog (Drosophila) (Dvl3), mRNA [NM_007889] |
KLA | 1.08 |
1.09 |
1.05 |
1.18 |
1.11 |
1.10 |
1.11 |
| ATP | 1.00 |
1.01 |
1.05 |
1.05 |
.99 |
1.10 |
1.15 |
| KLA/ATP | 1.12 |
1.07 |
1.09 |
1.09 |
1.06 |
1.15 |
1.09 |
|
| E2f1 | 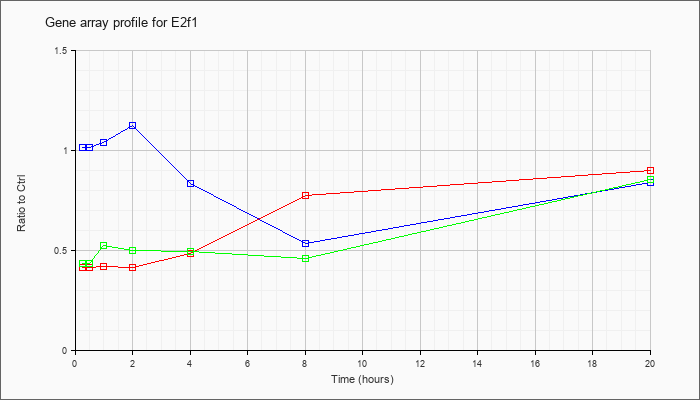 |
NM_007891 |
E2F transcription factor 1 (E2f1), mRNA [NM_007891] |
KLA | .41 |
.44 |
.42 |
.41 |
.48 |
.77 |
.90 |
| ATP | 1.01 |
1.09 |
1.04 |
1.13 |
.83 |
.53 |
.84 |
| KLA/ATP | .44 |
.44 |
.52 |
.50 |
.49 |
.46 |
.85 |
|
| E2f2 | 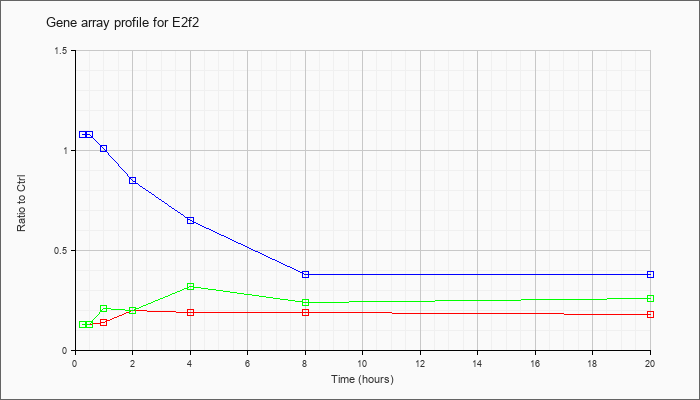 |
NM_177733 |
E2F transcription factor 2 (E2f2), mRNA [NM_177733] |
KLA | .13 |
.13 |
.14 |
.20 |
.19 |
.19 |
.18 |
| ATP | 1.08 |
.92 |
1.01 |
.85 |
.65 |
.38 |
.38 |
| KLA/ATP | .13 |
.13 |
.21 |
.20 |
.32 |
.24 |
.26 |
|
| E2f3 | 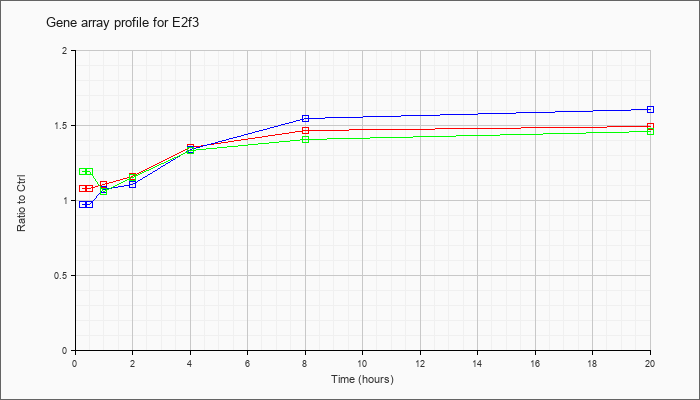 |
AK085296 |
0 day neonate kidney cDNA, RIKEN full-length enriched library, clone:D630007K21 product:E2F transcription factor 3, full insert sequence. [AK085296] |
KLA | .98 |
.97 |
1.02 |
1.00 |
.99 |
.99 |
.99 |
| ATP | .97 |
1.06 |
1.03 |
.98 |
1.01 |
.99 |
1.00 |
| KLA/ATP | 1.12 |
1.14 |
1.03 |
.99 |
.96 |
.93 |
.96 |
|
| E2f3 |  |
NM_010093 |
E2F transcription factor 3 (E2f3), mRNA [NM_010093] |
KLA | 1.11 |
1.09 |
1.13 |
1.21 |
1.47 |
1.62 |
1.65 |
| ATP | .97 |
.98 |
1.08 |
1.15 |
1.45 |
1.73 |
1.80 |
| KLA/ATP | 1.21 |
1.13 |
1.07 |
1.20 |
1.45 |
1.56 |
1.62 |
|
| Egr1 | 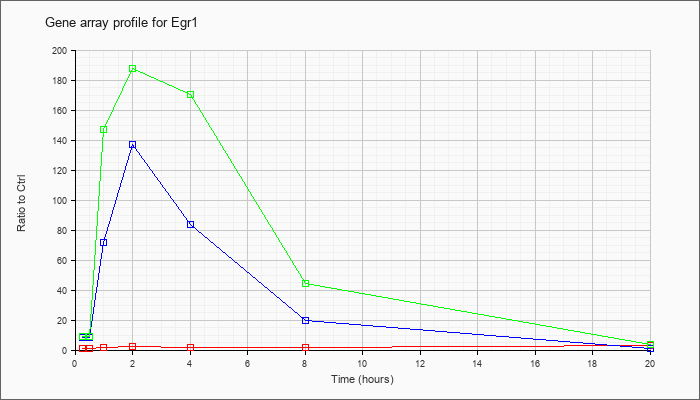 |
NM_007913 |
early growth response 1 (Egr1), mRNA [NM_007913] |
KLA | 1.00 |
.56 |
1.72 |
2.17 |
1.88 |
1.59 |
2.89 |
| ATP | 8.02 |
5.01 |
71.67 |
137.15 |
83.52 |
19.89 |
1.19 |
| KLA/ATP | 8.95 |
15.55 |
146.78 |
187.64 |
170.49 |
44.28 |
3.89 |
|
| Egr2 | 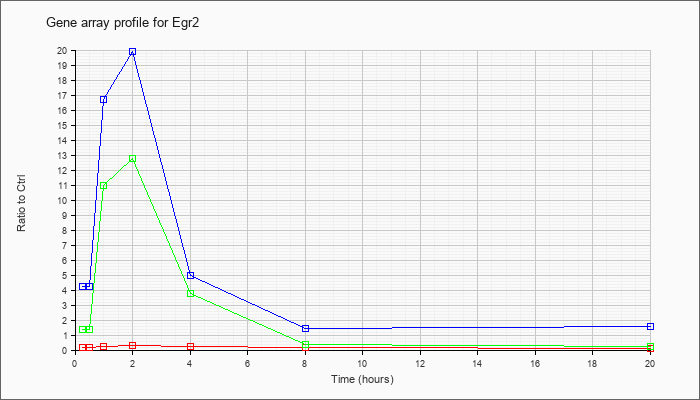 |
NM_010118 |
early growth response 2 (Egr2), mRNA [NM_010118] |
KLA | .20 |
.24 |
.23 |
.28 |
.25 |
.15 |
.08 |
| ATP | 4.23 |
6.98 |
16.71 |
19.88 |
4.99 |
1.41 |
1.59 |
| KLA/ATP | 1.38 |
4.50 |
10.98 |
12.78 |
3.75 |
.37 |
.22 |
|
| Elk1 | 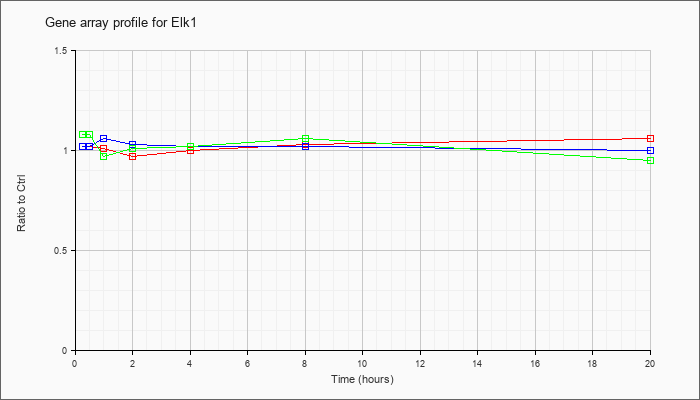 |
AK082260 |
0 day neonate cerebellum cDNA, RIKEN full-length enriched library, clone:C230030F12 product:ELK1, member of ETS oncogene family, full insert sequence. [AK082260] |
KLA | 1.02 |
1.01 |
1.01 |
.97 |
1.00 |
1.03 |
1.06 |
| ATP | 1.02 |
1.07 |
1.06 |
1.03 |
1.02 |
1.02 |
1.00 |
| KLA/ATP | 1.08 |
.99 |
.97 |
1.01 |
1.02 |
1.06 |
.95 |
|
| Elk4 |  |
AK037559 |
16 days neonate thymus cDNA, RIKEN full-length enriched library, clone:A130026I01 product:unclassifiable, full insert sequence. [AK037559] |
KLA | 1.04 |
.93 |
.96 |
.94 |
.99 |
1.06 |
.93 |
| ATP | .95 |
.98 |
.89 |
.92 |
.96 |
1.00 |
.96 |
| KLA/ATP | .94 |
.94 |
.99 |
.92 |
.93 |
.99 |
.96 |
|
| Elk4 |  |
NM_007923 |
ELK4, member of ETS oncogene family (Elk4), mRNA [NM_007923] |
KLA | .96 |
1.04 |
1.07 |
1.04 |
1.05 |
1.01 |
1.09 |
| ATP | .93 |
1.01 |
1.02 |
1.01 |
1.00 |
1.10 |
1.01 |
| KLA/ATP | .98 |
1.01 |
.94 |
1.02 |
1.03 |
1.04 |
1.04 |
|
| Ep300 | 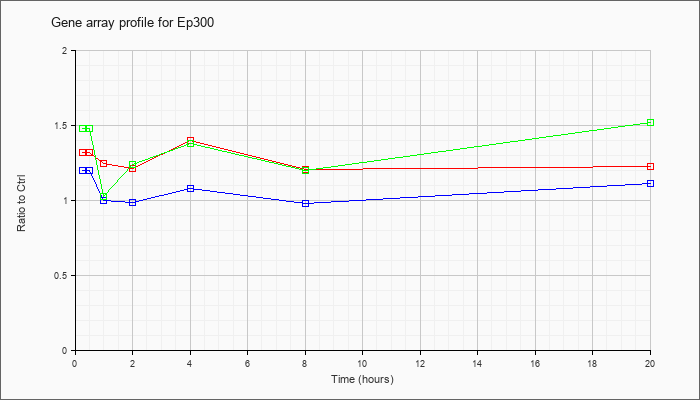 |
AK042627 |
7 days neonate cerebellum cDNA, RIKEN full-length enriched library, clone:A730011L11 product:P300 TRANSCRIPTION COACTIVATOR (FRAGMENT) homolog [Mus sp], full insert sequence. [AK042627] |
KLA | 1.02 |
.95 |
.92 |
1.04 |
1.01 |
1.00 |
1.05 |
| ATP | 1.06 |
.89 |
.88 |
.89 |
1.00 |
.90 |
1.07 |
| KLA/ATP | 1.10 |
1.02 |
.87 |
.91 |
1.04 |
.97 |
.98 |
|
| Ep300 |  |
NM_177821 |
E1A binding protein p300 (Ep300), mRNA [NM_177821] |
KLA | 1.61 |
1.26 |
1.57 |
1.38 |
1.78 |
1.41 |
1.40 |
| ATP | 1.34 |
1.60 |
1.11 |
1.08 |
1.16 |
1.05 |
1.15 |
| KLA/ATP | 1.86 |
1.94 |
1.18 |
1.56 |
1.71 |
1.42 |
2.05 |
|
| Ets1 | 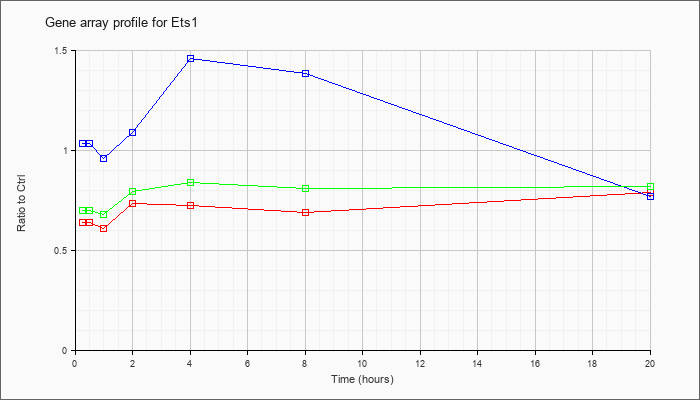 |
NM_011808 |
E26 avian leukemia oncogene 1, 5 domain (Ets1), transcript variant 1, mRNA [NM_011808] |
KLA | .55 |
.54 |
.49 |
.66 |
.62 |
.58 |
.67 |
| ATP | 1.01 |
1.14 |
.98 |
1.11 |
1.59 |
1.48 |
.67 |
| KLA/ATP | .58 |
.53 |
.58 |
.70 |
.80 |
.76 |
.76 |
|
| Ets1 |  |
X55787 |
gb|M.musculus c-ets-1 gene. [X55787] |
KLA | .83 |
.93 |
.85 |
.89 |
.94 |
.90 |
1.03 |
| ATP | 1.08 |
1.11 |
.92 |
1.04 |
1.19 |
1.19 |
.98 |
| KLA/ATP | .94 |
.90 |
.88 |
.98 |
.91 |
.92 |
.95 |
|
| Ets2 | 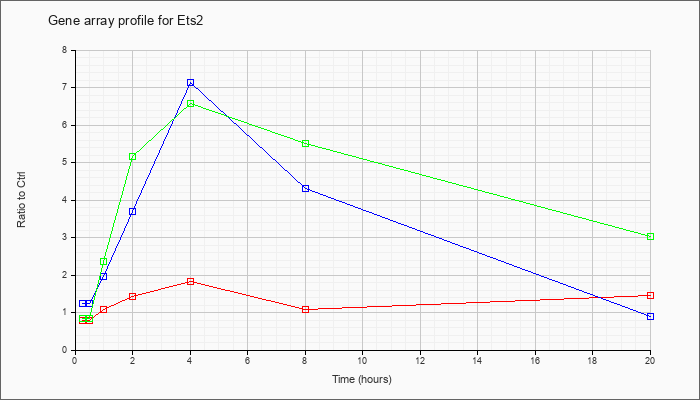 |
NM_011809 |
E26 avian leukemia oncogene 2, 3 domain (Ets2), mRNA [NM_011809] |
KLA | .79 |
.82 |
1.09 |
1.44 |
1.83 |
1.08 |
1.46 |
| ATP | 1.23 |
1.59 |
1.97 |
3.69 |
7.13 |
4.30 |
.90 |
| KLA/ATP | .84 |
1.33 |
2.37 |
5.17 |
6.58 |
5.50 |
3.04 |
|
| Fdps | 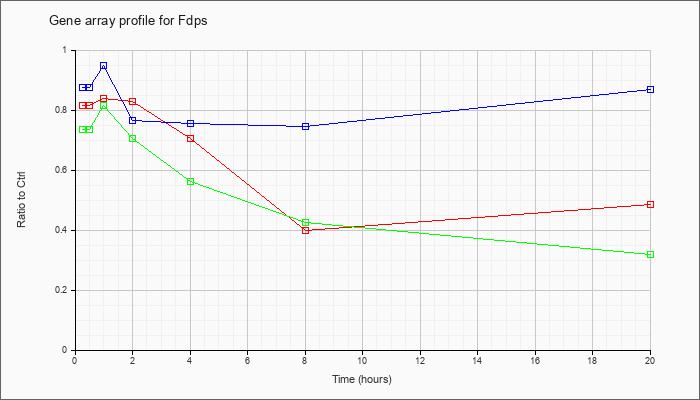 |
AK077979 |
13 days embryo male testis cDNA, RIKEN full-length enriched library, clone:6030492I17 product:farnesyl diphosphate synthetase, full insert sequence. [AK077979] |
KLA | .76 |
.79 |
.79 |
.69 |
.65 |
.37 |
.49 |
| ATP | .63 |
.70 |
.85 |
.30 |
.46 |
.89 |
.86 |
| KLA/ATP | .54 |
.52 |
.63 |
.39 |
.42 |
.40 |
.31 |
|
| Fdps |  |
NM_134469 |
farnesyl diphosphate synthetase (Fdps), mRNA [NM_134469] |
KLA | .87 |
.93 |
.89 |
.97 |
.76 |
.43 |
.48 |
| ATP | 1.12 |
1.10 |
1.05 |
1.23 |
1.05 |
.60 |
.88 |
| KLA/ATP | .93 |
.99 |
1.00 |
1.02 |
.70 |
.45 |
.33 |
|
| Fos | 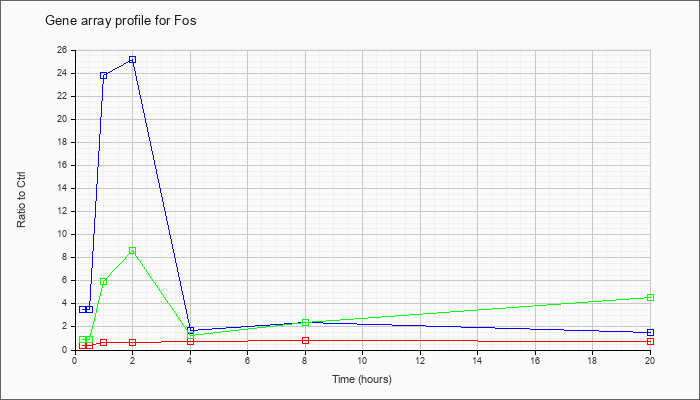 |
NM_010234 |
FBJ osteosarcoma oncogene (Fos), mRNA [NM_010234] |
KLA | .42 |
.33 |
.64 |
.61 |
.71 |
.85 |
.71 |
| ATP | 3.48 |
5.46 |
23.78 |
25.22 |
1.71 |
2.36 |
1.50 |
| KLA/ATP | .90 |
1.74 |
5.97 |
8.62 |
1.28 |
2.42 |
4.52 |
|
| Fosl1 | 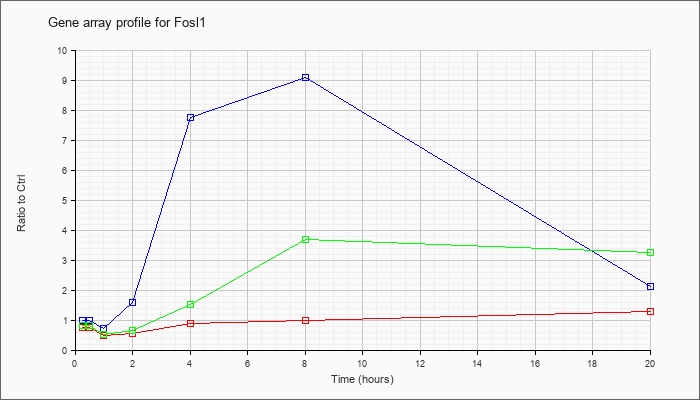 |
NM_010235 |
fos-like antigen 1 (Fosl1), mRNA [NM_010235] |
KLA | .75 |
.72 |
.50 |
.55 |
.87 |
.97 |
1.29 |
| ATP | 1.00 |
.99 |
.72 |
1.57 |
7.76 |
9.10 |
2.11 |
| KLA/ATP | .83 |
.75 |
.53 |
.65 |
1.53 |
3.70 |
3.26 |
|
| Fzd1 | 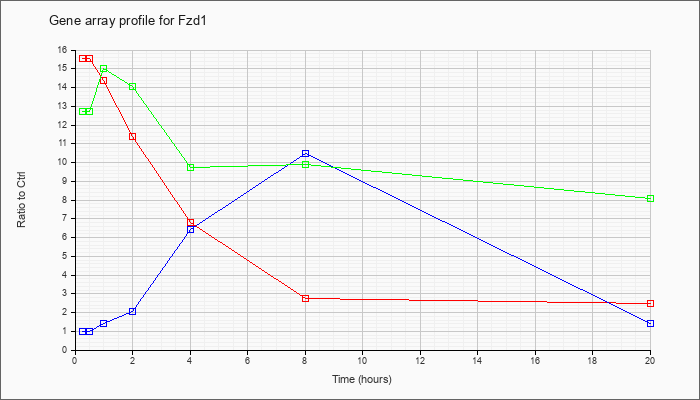 |
NM_021457 |
frizzled homolog 1 (Drosophila) (Fzd1), mRNA [NM_021457] |
KLA | 15.53 |
16.47 |
14.36 |
11.40 |
6.80 |
2.73 |
2.49 |
| ATP | 1.01 |
1.02 |
1.39 |
2.05 |
6.43 |
10.47 |
1.41 |
| KLA/ATP | 12.71 |
14.30 |
14.99 |
14.06 |
9.72 |
9.90 |
8.10 |
|
| Fzd10 | 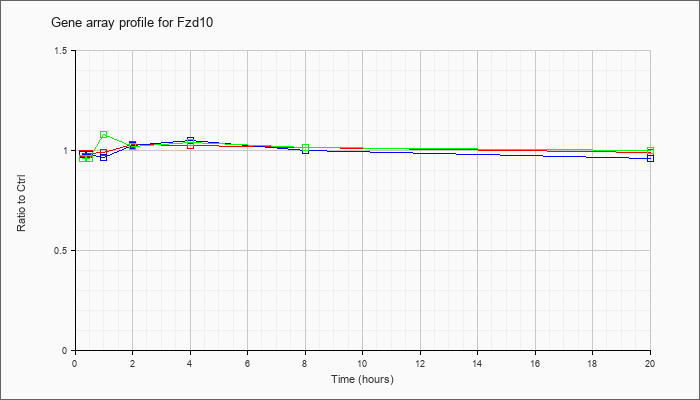 |
NM_175284 |
frizzled homolog 10 (Drosophila) (Fzd10), mRNA [NM_175284] |
KLA | .99 |
.98 |
.99 |
1.03 |
1.03 |
1.02 |
.99 |
| ATP | .98 |
.98 |
.97 |
1.03 |
1.05 |
1.00 |
.96 |
| KLA/ATP | .96 |
.98 |
1.08 |
1.02 |
1.04 |
1.02 |
1.00 |
|
| Fzd2 | 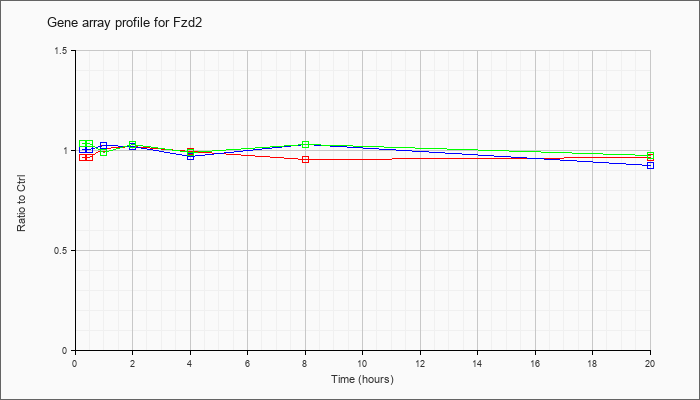 |
NM_020510 |
frizzled homolog 2 (Drosophila) (Fzd2), mRNA [NM_020510] |
KLA | .97 |
1.01 |
1.01 |
1.02 |
1.00 |
.96 |
.97 |
| ATP | 1.01 |
1.02 |
1.03 |
1.02 |
.97 |
1.03 |
.93 |
| KLA/ATP | 1.04 |
.98 |
.99 |
1.03 |
.99 |
1.03 |
.98 |
|
| Fzd3 | 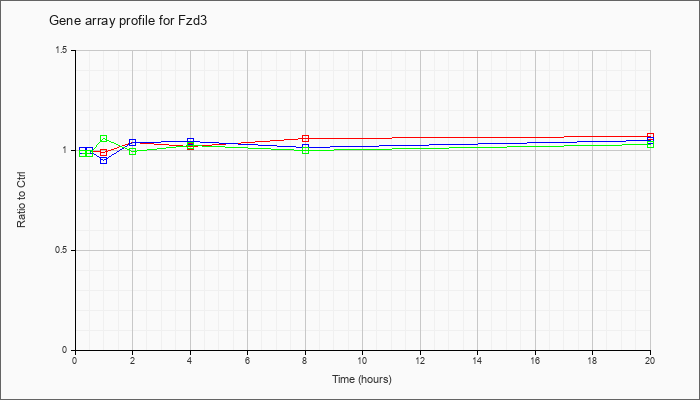 |
NM_021458 |
frizzled homolog 3 (Drosophila) (Fzd3), mRNA [NM_021458] |
KLA | 1.00 |
1.00 |
.99 |
1.04 |
1.02 |
1.06 |
1.07 |
| ATP | 1.00 |
1.00 |
.95 |
1.04 |
1.04 |
1.01 |
1.05 |
| KLA/ATP | .98 |
1.05 |
1.06 |
.99 |
1.02 |
1.00 |
1.03 |
|
| Fzd4 | 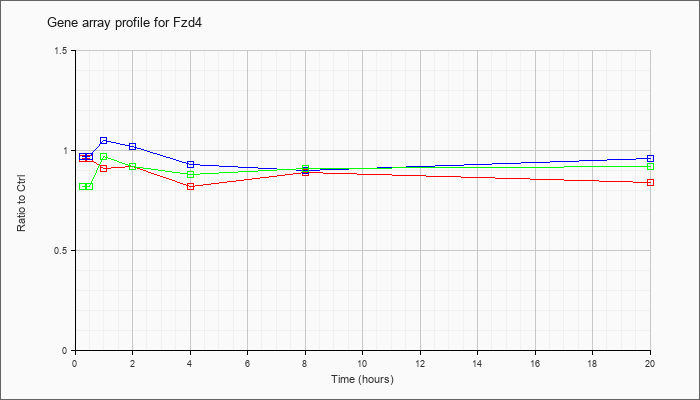 |
NM_008055 |
frizzled homolog 4 (Drosophila) (Fzd4), mRNA [NM_008055] |
KLA | .96 |
.88 |
.91 |
.92 |
.82 |
.89 |
.84 |
| ATP | .97 |
.98 |
1.05 |
1.02 |
.93 |
.90 |
.96 |
| KLA/ATP | .82 |
.86 |
.97 |
.92 |
.88 |
.91 |
.92 |
|
| Fzd5 |  |
NM_022721 |
frizzled homolog 5 (Drosophila) (Fzd5), transcript variant 1, mRNA [NM_022721] |
KLA | 1.63 |
1.69 |
1.56 |
1.23 |
.91 |
1.01 |
.92 |
| ATP | 1.03 |
1.32 |
4.46 |
9.23 |
7.72 |
2.14 |
1.11 |
| KLA/ATP | 1.63 |
1.92 |
4.20 |
8.67 |
8.29 |
3.07 |
1.12 |
|
| Fzd6 |  |
NM_008056 |
frizzled homolog 6 (Drosophila) (Fzd6), mRNA [NM_008056] |
KLA | .97 |
1.01 |
.96 |
.90 |
.87 |
.94 |
.98 |
| ATP | .98 |
.97 |
.98 |
.94 |
.95 |
1.09 |
.96 |
| KLA/ATP | .97 |
1.02 |
.94 |
.92 |
.90 |
1.02 |
1.00 |
|
| Fzd7 |  |
NM_008057 |
frizzled homolog 7 (Drosophila) (Fzd7), mRNA [NM_008057] |
KLA | 1.50 |
1.68 |
1.53 |
1.31 |
1.13 |
1.00 |
.94 |
| ATP | 1.04 |
1.12 |
1.60 |
3.45 |
2.56 |
1.72 |
1.02 |
| KLA/ATP | 1.89 |
1.92 |
1.96 |
4.61 |
3.75 |
3.38 |
1.88 |
|
| Fzd8 |  |
AK034561 |
12 days embryo embryonic body between diaphragm region and neck cDNA, RIKEN full-length enriched library, clone:9430006K02 product:inferred: transmembrane receptor {Mus musculus}; frizzled homolog 8 (Drosophila) homolog {GNF expression}, |
KLA | 1.00 |
.98 |
1.05 |
.98 |
.96 |
1.03 |
.94 |
| ATP | 1.00 |
.92 |
1.06 |
1.05 |
1.02 |
1.01 |
.98 |
| KLA/ATP | .99 |
1.06 |
.99 |
.94 |
.95 |
.99 |
1.03 |
|
| Fzd8 |  |
NM_008058 |
frizzled homolog 8 (Drosophila) (Fzd8), mRNA [NM_008058] |
KLA | 1.09 |
1.15 |
1.07 |
1.05 |
1.03 |
1.05 |
.98 |
| ATP | 1.05 |
1.08 |
2.01 |
3.38 |
2.89 |
1.20 |
1.03 |
| KLA/ATP | 1.04 |
1.17 |
1.65 |
2.57 |
2.62 |
1.40 |
1.04 |
|
| Fzd9 | 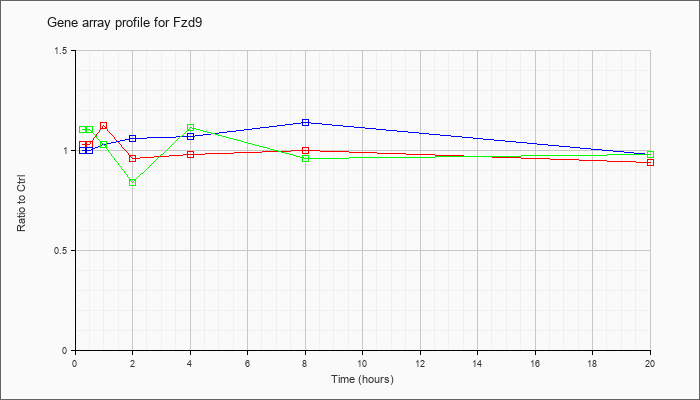 |
NM_010246 |
frizzled homolog 9 (Drosophila) (Fzd9), mRNA [NM_010246] |
KLA | 1.03 |
1.10 |
1.12 |
.96 |
.98 |
1.00 |
.94 |
| ATP | 1.00 |
1.00 |
1.03 |
1.06 |
1.07 |
1.14 |
.98 |
| KLA/ATP | 1.10 |
1.07 |
1.03 |
.84 |
1.11 |
.96 |
.98 |
|
| Gcn5l2 |  |
NM_020004 |
GCN5 general control of amino acid synthesis-like 2 (yeast) (Gcn5l2), transcript variant 1, mRNA [NM_020004] |
KLA | .86 |
.86 |
.77 |
.82 |
.91 |
.97 |
1.10 |
| ATP | 1.13 |
1.27 |
1.06 |
1.44 |
.73 |
.63 |
.90 |
| KLA/ATP | 1.02 |
1.05 |
.82 |
1.05 |
.54 |
.62 |
.69 |
|
| Gps2 | 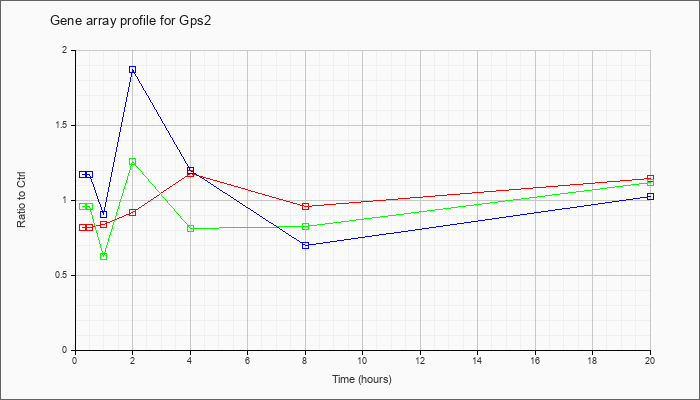 |
NM_019726 |
G protein pathway suppressor 2 (Gps2), mRNA [NM_019726] |
KLA | .82 |
.88 |
.84 |
.92 |
1.18 |
.96 |
1.15 |
| ATP | 1.17 |
1.28 |
.91 |
1.87 |
1.20 |
.70 |
1.03 |
| KLA/ATP | .96 |
1.02 |
.63 |
1.26 |
.81 |
.83 |
1.12 |
|
| Gsk3b | 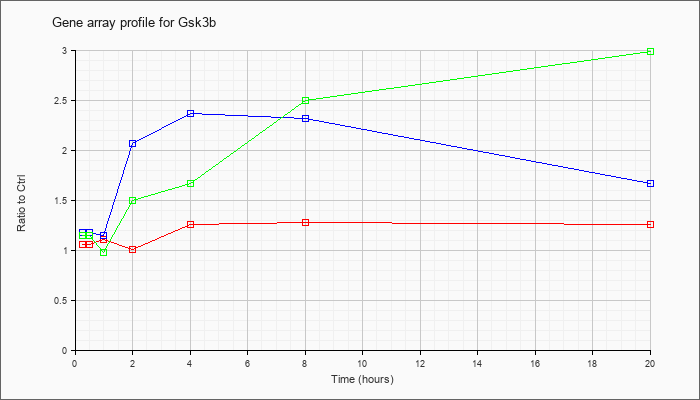 |
NM_019827 |
glycogen synthase kinase 3 beta (Gsk3b), mRNA [NM_019827] |
KLA | 1.06 |
1.03 |
1.11 |
1.01 |
1.26 |
1.28 |
1.26 |
| ATP | 1.18 |
1.27 |
1.15 |
2.07 |
2.37 |
2.32 |
1.67 |
| KLA/ATP | 1.15 |
1.21 |
.98 |
1.50 |
1.67 |
2.50 |
2.99 |
|
| H2-Aa | 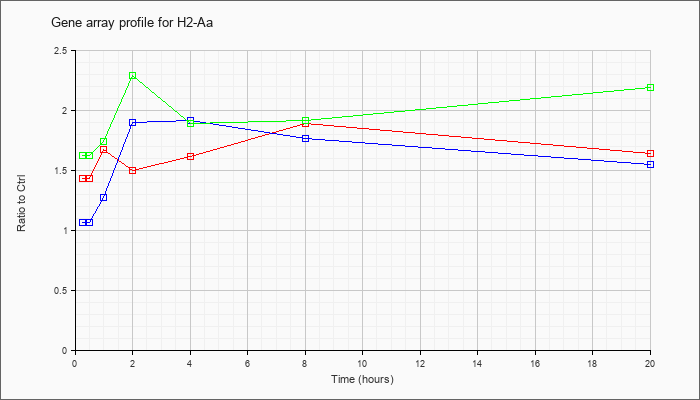 |
NM_010378 |
histocompatibility 2, class II antigen A, alpha (H2-Aa), mRNA [NM_010378] |
KLA | 1.43 |
1.59 |
1.68 |
1.50 |
1.62 |
1.89 |
1.64 |
| ATP | 1.07 |
1.15 |
1.27 |
1.90 |
1.92 |
1.77 |
1.55 |
| KLA/ATP | 1.62 |
1.81 |
1.74 |
2.29 |
1.89 |
1.92 |
2.19 |
|
| H2-Ab1 |  |
BC010322 |
histocompatibility 2, class II antigen A, beta 1, mRNA (cDNA clone MGC:6297 IMAGE:2651058), complete cds [BC010322] |
KLA | 1.73 |
1.76 |
1.77 |
1.55 |
1.95 |
2.10 |
2.27 |
| ATP | 1.03 |
1.18 |
1.32 |
1.99 |
1.79 |
1.56 |
1.61 |
| KLA/ATP | 1.70 |
2.01 |
1.74 |
2.52 |
1.58 |
2.01 |
2.75 |
|
| H2-Ab1 |  |
NM_207105 |
histocompatibility 2, class II antigen A, beta 1 (H2-Ab1), mRNA [NM_207105] |
KLA | 2.02 |
2.15 |
2.07 |
2.09 |
2.32 |
2.78 |
3.08 |
| ATP | 1.11 |
1.20 |
1.42 |
2.33 |
2.33 |
2.25 |
1.87 |
| KLA/ATP | 2.11 |
2.32 |
2.52 |
3.62 |
2.67 |
2.96 |
5.04 |
|
| H2-Bl | 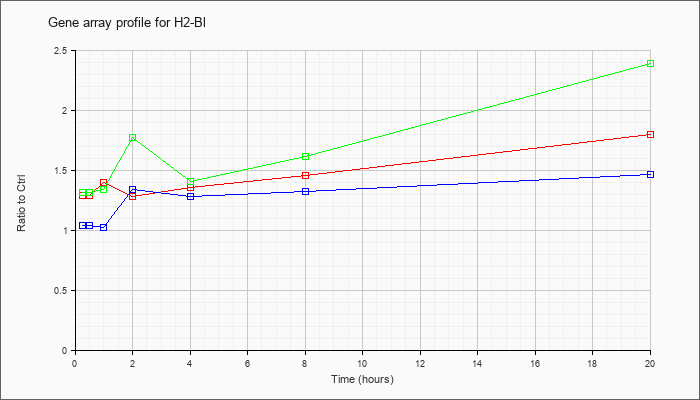 |
NM_008199 |
histocompatibility 2, blastocyst (H2-Bl), mRNA [NM_008199] |
KLA | 1.29 |
1.32 |
1.40 |
1.28 |
1.36 |
1.46 |
1.80 |
| ATP | 1.04 |
1.11 |
1.02 |
1.34 |
1.28 |
1.32 |
1.47 |
| KLA/ATP | 1.32 |
1.39 |
1.34 |
1.77 |
1.41 |
1.61 |
2.39 |
|
| H2-D1 | 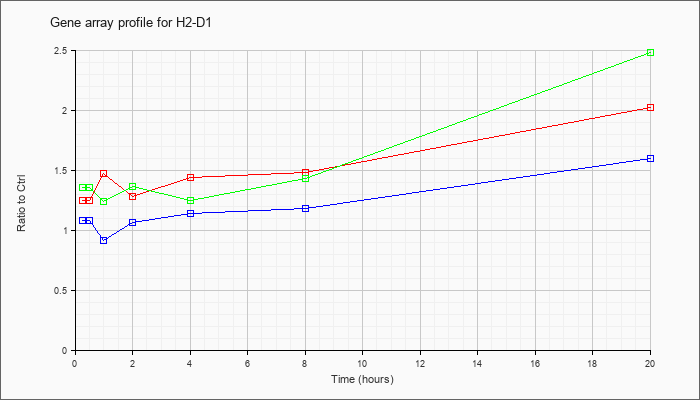 |
AK169750 |
2 days neonate thymus thymic cells cDNA, RIKEN full-length enriched library, clone:E430019E13 product:histocompatibility 2, D region locus 1, full insert sequence [AK169750] |
KLA | .99 |
.98 |
.96 |
.99 |
.93 |
.91 |
.94 |
| ATP | .96 |
1.01 |
1.02 |
.95 |
1.05 |
1.00 |
.98 |
| KLA/ATP | .92 |
.94 |
.97 |
.97 |
.98 |
.92 |
1.03 |
|
| H2-D1 |  |
NM_010380 |
histocompatibility 2, D region locus 1 (H2-D1), mRNA [NM_010380] |
KLA | 1.38 |
1.45 |
1.73 |
1.42 |
1.70 |
1.76 |
2.56 |
| ATP | 1.15 |
1.11 |
.86 |
1.12 |
1.19 |
1.28 |
1.90 |
| KLA/ATP | 1.58 |
1.72 |
1.38 |
1.56 |
1.38 |
1.69 |
3.21 |
|
| H2-DMa | 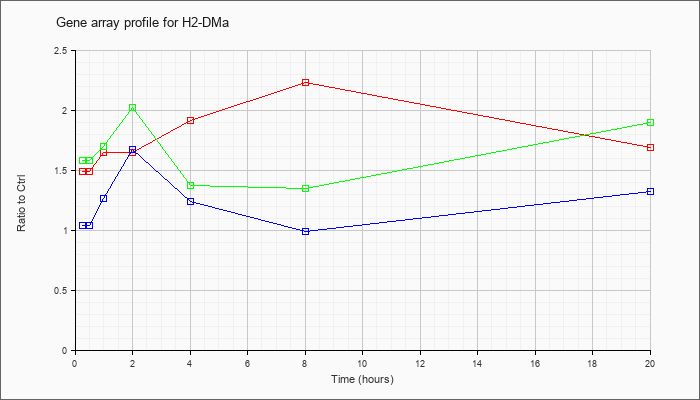 |
NM_010386 |
histocompatibility 2, class II, locus DMa (H2-DMa), mRNA [NM_010386] |
KLA | 1.49 |
1.50 |
1.65 |
1.65 |
1.91 |
2.23 |
1.69 |
| ATP | 1.04 |
1.12 |
1.26 |
1.67 |
1.24 |
.99 |
1.32 |
| KLA/ATP | 1.58 |
1.68 |
1.70 |
2.02 |
1.37 |
1.35 |
1.90 |
|
| H2-DMb1 | 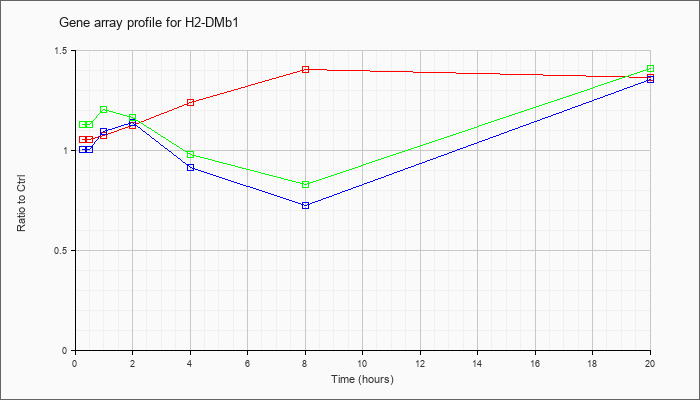 |
NM_010387 |
histocompatibility 2, class II, locus Mb1 (H2-DMb1), mRNA [NM_010387] |
KLA | 1.06 |
1.14 |
1.08 |
1.13 |
1.24 |
1.41 |
1.37 |
| ATP | 1.01 |
1.02 |
1.09 |
1.14 |
.92 |
.73 |
1.36 |
| KLA/ATP | 1.13 |
1.05 |
1.21 |
1.17 |
.98 |
.83 |
1.41 |
|
| H2-DMb2 | 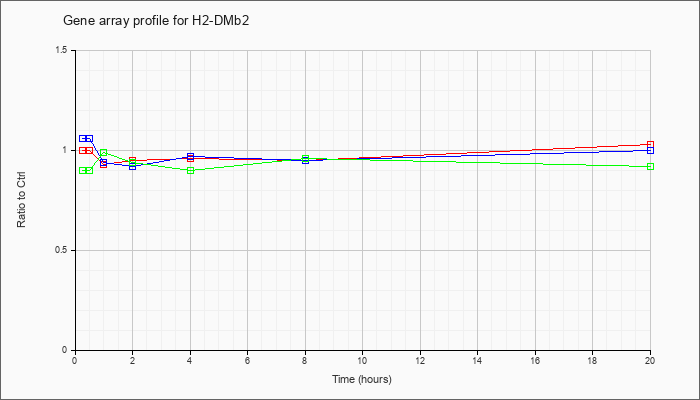 |
NM_010388 |
histocompatibility 2, class II, locus Mb2 (H2-DMb2), mRNA [NM_010388] |
KLA | 1.00 |
1.04 |
.93 |
.95 |
.96 |
.95 |
1.03 |
| ATP | 1.06 |
1.04 |
.94 |
.92 |
.97 |
.95 |
1.00 |
| KLA/ATP | .90 |
.94 |
.99 |
.94 |
.90 |
.96 |
.92 |
|
| H2-Eb1 | 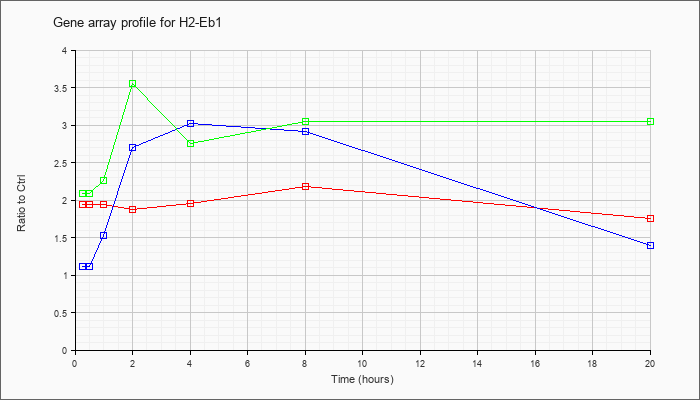 |
X52641 |
gb|Mouse mRNA for I-A beta NOD. [X52641] |
KLA | 1.94 |
2.06 |
1.94 |
1.87 |
1.96 |
2.18 |
1.76 |
| ATP | 1.11 |
1.20 |
1.53 |
2.70 |
3.02 |
2.91 |
1.40 |
| KLA/ATP | 2.09 |
2.11 |
2.26 |
3.55 |
2.76 |
3.05 |
3.05 |
|
| H2-K1 | 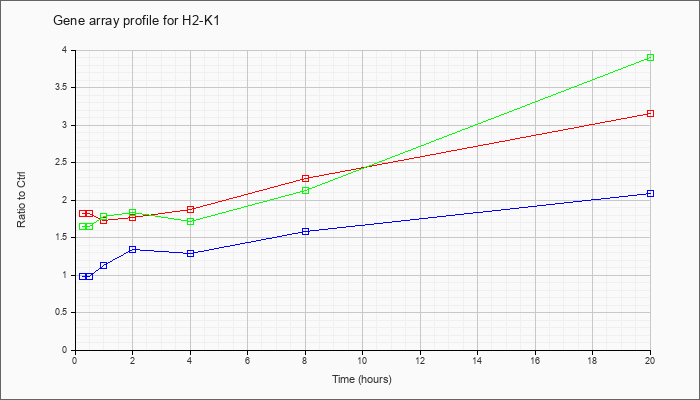 |
M69070 |
mRNA, complete cds. [M69070] |
KLA | 1.76 |
1.66 |
1.78 |
1.78 |
1.87 |
2.23 |
2.99 |
| ATP | .98 |
.97 |
1.19 |
1.45 |
1.36 |
1.68 |
2.17 |
| KLA/ATP | 1.68 |
1.70 |
1.87 |
1.96 |
1.87 |
2.13 |
3.82 |
|
| H2-K1 |  |
NM_001001892 |
histocompatibility 2, K1, K region (H2-K1), transcript variant 1, mRNA [NM_001001892] |
KLA | 1.82 |
1.66 |
1.73 |
1.76 |
1.87 |
2.29 |
3.17 |
| ATP | .98 |
.97 |
1.13 |
1.32 |
1.29 |
1.57 |
2.09 |
| KLA/ATP | 1.64 |
1.71 |
1.77 |
1.82 |
1.71 |
2.12 |
3.91 |
|
| H2-M1 | 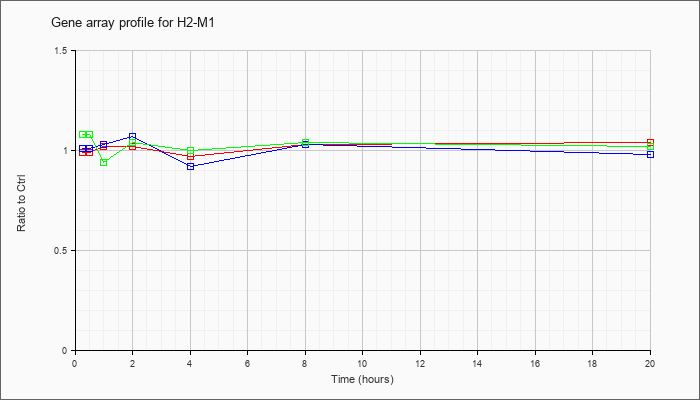 |
NM_177636 |
histocompatibility 2, M region locus 1 (H2-M1), mRNA [NM_177636] |
KLA | .99 |
1.01 |
1.02 |
1.02 |
.97 |
1.03 |
1.04 |
| ATP | 1.01 |
.99 |
1.03 |
1.07 |
.92 |
1.03 |
.98 |
| KLA/ATP | 1.08 |
.97 |
.94 |
1.04 |
1.00 |
1.04 |
1.02 |
|
H2-M10.1
| 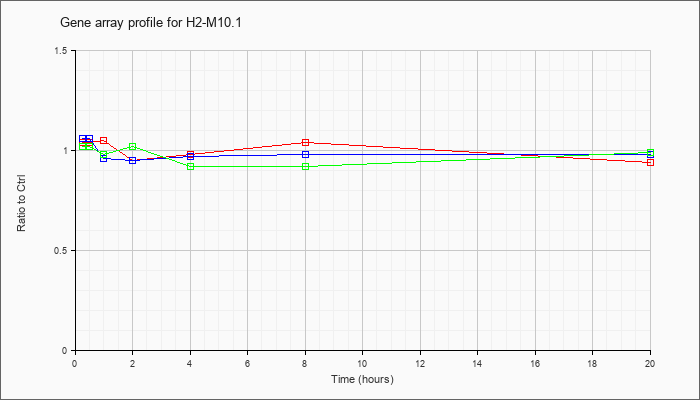 |
NM_013544 |
histocompatibility 2, M region locus 10.1 (H2-M10.1), mRNA [NM_013544] |
KLA | 1.04 |
1.00 |
1.05 |
.95 |
.98 |
1.04 |
.94 |
| ATP | 1.06 |
1.04 |
.96 |
.95 |
.97 |
.98 |
.98 |
| KLA/ATP | 1.02 |
1.07 |
.98 |
1.02 |
.92 |
.92 |
.99 |
|
H2-M10.2
| 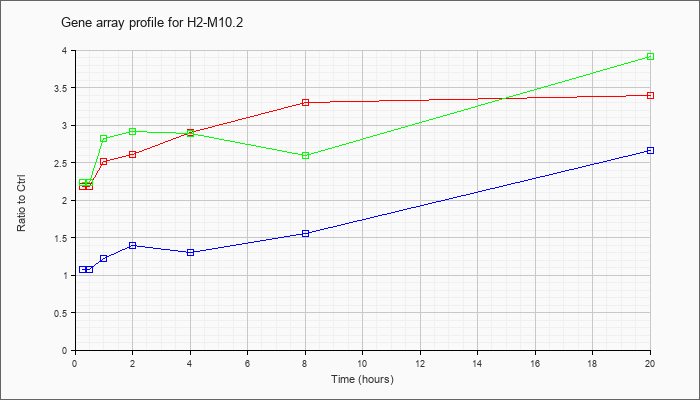 |
NM_177923 |
histocompatibility 2, M region locus 10.2 (H2-M10.2), mRNA [NM_177923] |
KLA | 2.18 |
2.23 |
2.51 |
2.61 |
2.90 |
3.30 |
3.40 |
| ATP | 1.08 |
1.02 |
1.22 |
1.39 |
1.30 |
1.56 |
2.66 |
| KLA/ATP | 2.23 |
2.53 |
2.82 |
2.91 |
2.89 |
2.60 |
3.92 |
|
H2-M10.3
| 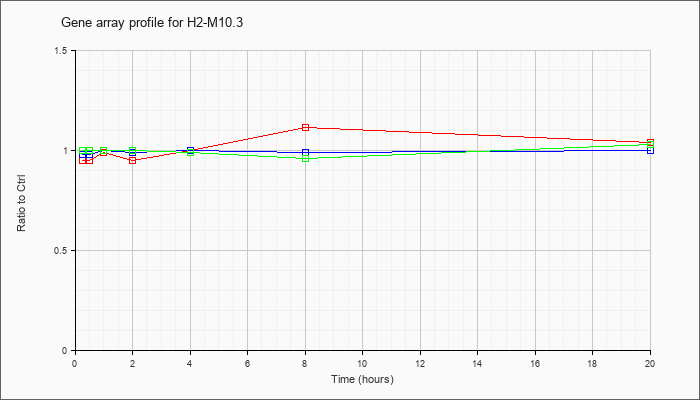 |
AY211080 |
major histocompatibility complex class Ib M10.3 mRNA, complete cds [AY211080] |
KLA | .95 |
1.08 |
.99 |
.95 |
1.00 |
1.11 |
1.04 |
| ATP | .98 |
.96 |
1.00 |
.99 |
1.00 |
.99 |
1.00 |
| KLA/ATP | 1.00 |
.99 |
1.00 |
1.00 |
.99 |
.96 |
1.03 |
|
H2-M10.4
| 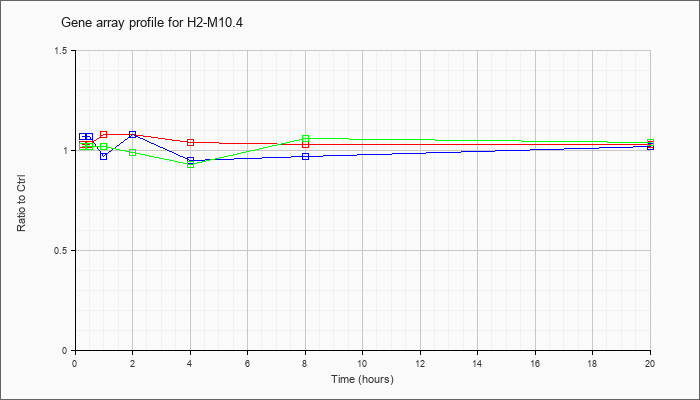 |
NM_177634 |
histocompatibility 2, M region locus 10.4 (H2-M10.4), mRNA [NM_177634] |
KLA | 1.03 |
1.04 |
1.08 |
1.08 |
1.04 |
1.03 |
1.03 |
| ATP | 1.07 |
1.05 |
.97 |
1.08 |
.95 |
.97 |
1.02 |
| KLA/ATP | 1.02 |
.97 |
1.02 |
.99 |
.93 |
1.06 |
1.04 |
|
H2-M10.5
| 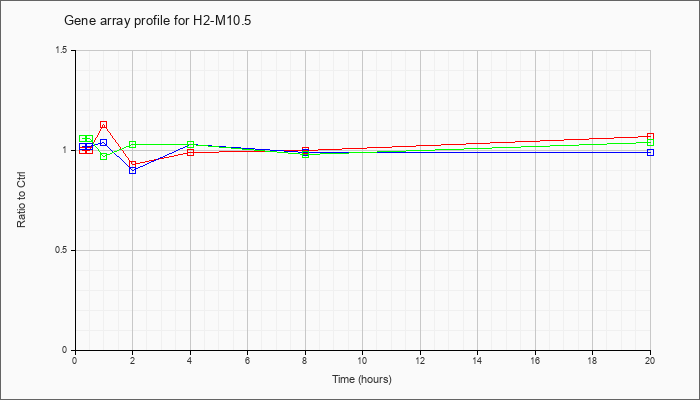 |
NM_177637 |
histocompatibility 2, M region locus 10.5 (H2-M10.5), mRNA [NM_177637] |
KLA | 1.00 |
1.04 |
1.13 |
.93 |
.99 |
1.00 |
1.07 |
| ATP | 1.02 |
1.02 |
1.04 |
.90 |
1.03 |
.99 |
.99 |
| KLA/ATP | 1.06 |
1.03 |
.97 |
1.03 |
1.03 |
.98 |
1.04 |
|
H2-M10.6
| 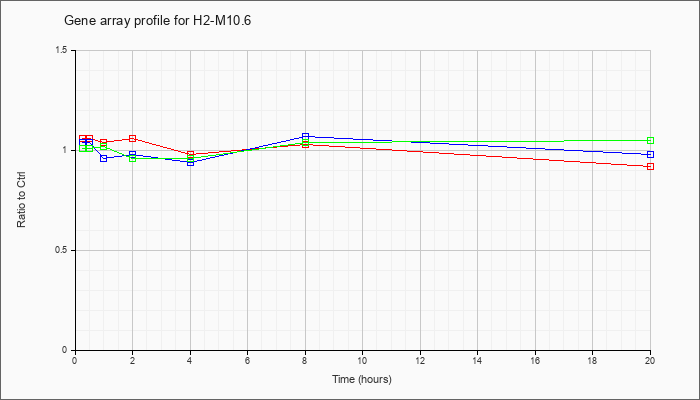 |
NM_201611 |
histocompatibility 2, M region locus 10.6 (H2-M10.6), mRNA [NM_201611] |
KLA | 1.06 |
.99 |
1.04 |
1.06 |
.98 |
1.03 |
.92 |
| ATP | 1.04 |
1.09 |
.96 |
.98 |
.94 |
1.07 |
.98 |
| KLA/ATP | 1.01 |
1.06 |
1.02 |
.96 |
.96 |
1.04 |
1.05 |
|
| H2-M11 | 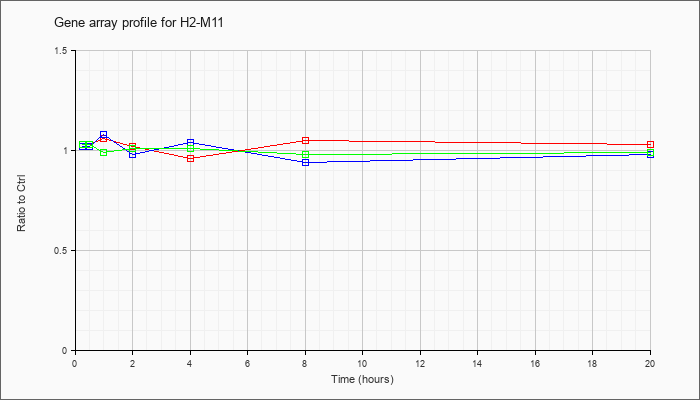 |
NM_177635 |
histocompatibility 2, M region locus 11 (H2-M11), mRNA [NM_177635] |
KLA | 1.03 |
1.07 |
1.06 |
1.02 |
.96 |
1.05 |
1.03 |
| ATP | 1.02 |
.96 |
1.08 |
.98 |
1.04 |
.94 |
.98 |
| KLA/ATP | 1.03 |
1.03 |
.99 |
1.01 |
1.01 |
.98 |
.99 |
|
| H2-M3 | 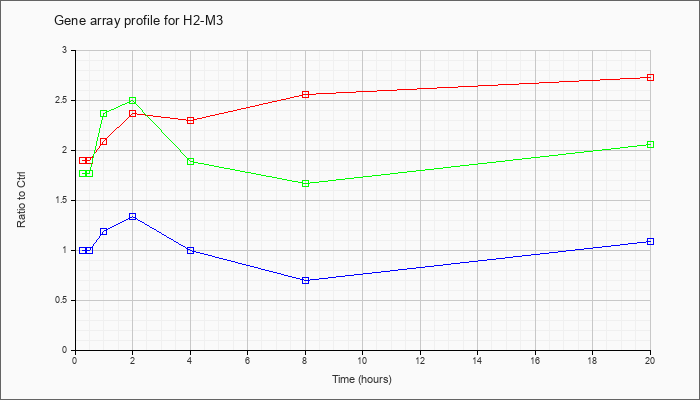 |
NM_013819 |
histocompatibility 2, M region locus 3 (H2-M3), mRNA [NM_013819] |
KLA | 1.90 |
2.04 |
2.09 |
2.37 |
2.30 |
2.56 |
2.73 |
| ATP | 1.00 |
1.00 |
1.19 |
1.34 |
1.00 |
.70 |
1.09 |
| KLA/ATP | 1.77 |
1.94 |
2.37 |
2.49 |
1.89 |
1.67 |
2.06 |
|
| H2-M9 | 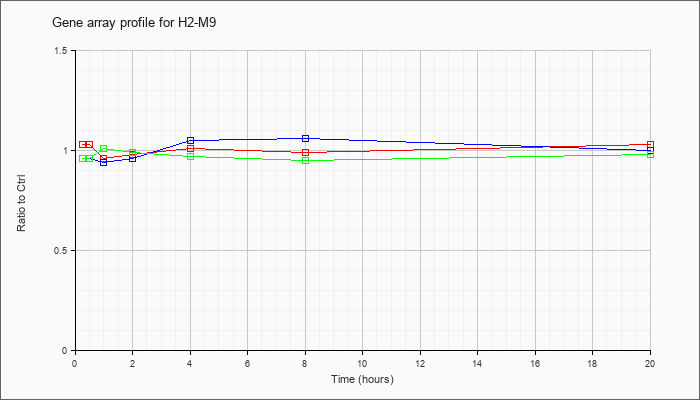 |
NM_008205 |
histocompatibility 2, M region locus 9 (H2-M9), mRNA [NM_008205] |
KLA | 1.03 |
.99 |
.96 |
.98 |
1.01 |
.99 |
1.03 |
| ATP | .96 |
1.06 |
.94 |
.96 |
1.05 |
1.06 |
1.00 |
| KLA/ATP | .96 |
1.01 |
1.01 |
.99 |
.97 |
.95 |
.98 |
|
| H2-Oa | 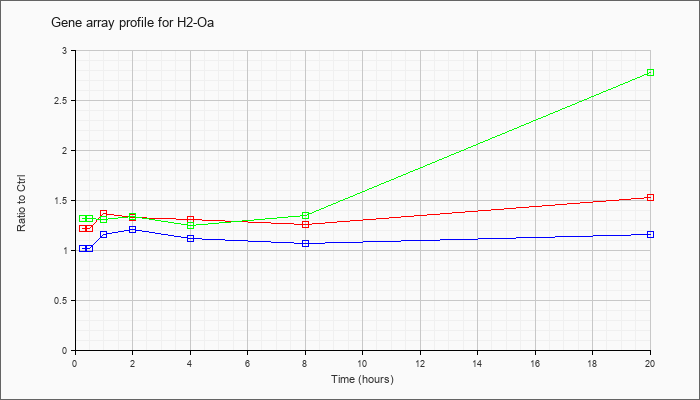 |
NM_008206 |
histocompatibility 2, O region alpha locus (H2-Oa), mRNA [NM_008206] |
KLA | 1.22 |
1.32 |
1.37 |
1.33 |
1.31 |
1.26 |
1.53 |
| ATP | 1.02 |
1.05 |
1.16 |
1.21 |
1.12 |
1.07 |
1.16 |
| KLA/ATP | 1.32 |
1.34 |
1.31 |
1.34 |
1.25 |
1.35 |
2.78 |
|
| H2-Ob | 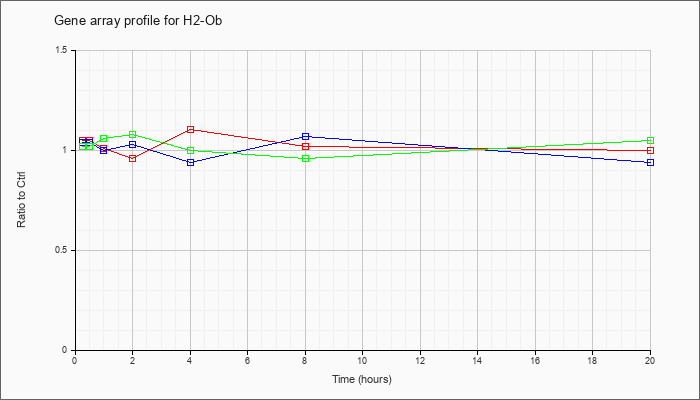 |
NM_010389 |
histocompatibility 2, O region beta locus (H2-Ob), mRNA [NM_010389] |
KLA | 1.05 |
1.07 |
1.01 |
.96 |
1.10 |
1.02 |
1.00 |
| ATP | 1.04 |
.99 |
1.00 |
1.03 |
.94 |
1.07 |
.94 |
| KLA/ATP | 1.02 |
.97 |
1.06 |
1.08 |
1.00 |
.96 |
1.05 |
|
| H2-Q1 | 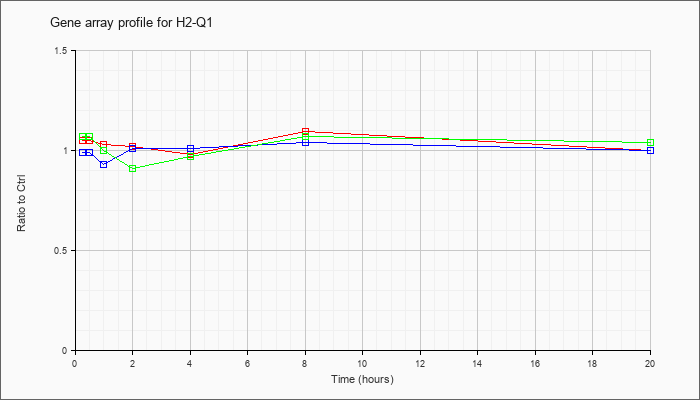 |
NM_010390 |
histocompatibility 2, Q region locus 1 (H2-Q1), mRNA [NM_010390] |
KLA | 1.05 |
1.08 |
1.03 |
1.02 |
.98 |
1.09 |
1.00 |
| ATP | .99 |
.92 |
.93 |
1.01 |
1.01 |
1.04 |
1.00 |
| KLA/ATP | 1.07 |
1.00 |
1.00 |
.91 |
.97 |
1.07 |
1.04 |
|
| H2-Q10 | 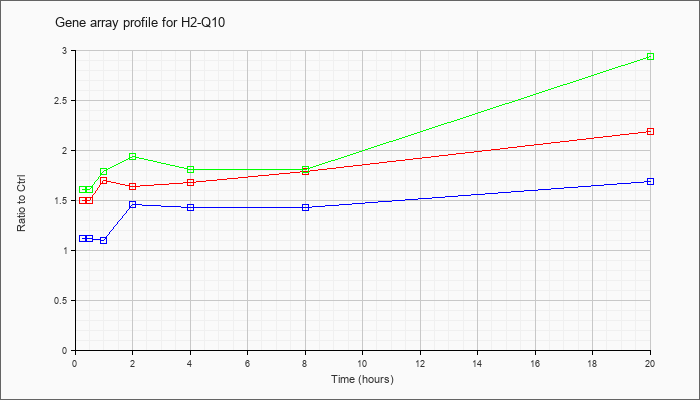 |
NM_010391 |
histocompatibility 2, Q region locus 10 (H2-Q10), mRNA [NM_010391] |
KLA | 1.50 |
1.50 |
1.70 |
1.64 |
1.67 |
1.79 |
2.18 |
| ATP | 1.12 |
1.11 |
1.09 |
1.46 |
1.42 |
1.43 |
1.69 |
| KLA/ATP | 1.61 |
1.72 |
1.79 |
1.94 |
1.81 |
1.81 |
2.93 |
|
| H2-Q2 | 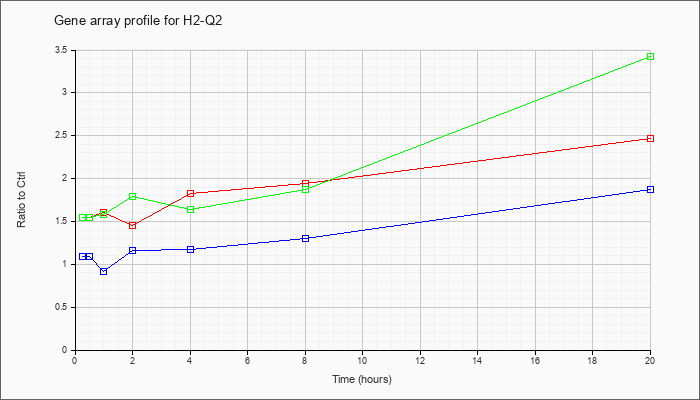 |
NM_010392 |
histocompatibility 2, Q region locus 2 (H2-Q2), mRNA [NM_010392] |
KLA | 1.55 |
1.52 |
1.61 |
1.45 |
1.83 |
1.94 |
2.47 |
| ATP | 1.09 |
1.13 |
.92 |
1.16 |
1.17 |
1.30 |
1.88 |
| KLA/ATP | 1.55 |
1.70 |
1.58 |
1.80 |
1.64 |
1.87 |
3.43 |
|
| H2-Q7 | 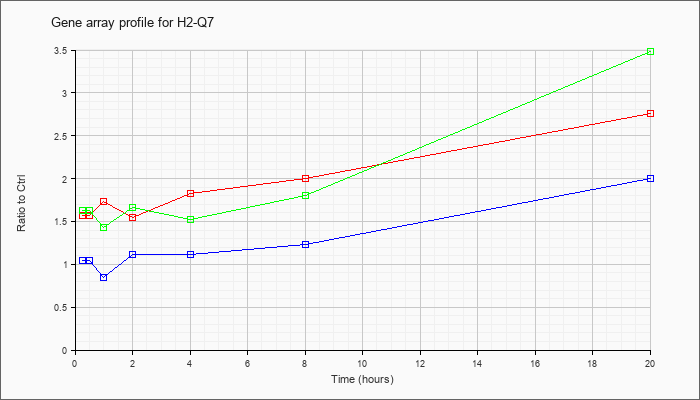 |
NM_010394 |
histocompatibility 2, Q region locus 7 (H2-Q7), mRNA [NM_010394] |
KLA | 1.57 |
1.58 |
1.74 |
1.55 |
1.82 |
2.00 |
2.76 |
| ATP | 1.05 |
1.09 |
.84 |
1.12 |
1.12 |
1.23 |
2.00 |
| KLA/ATP | 1.63 |
1.75 |
1.43 |
1.67 |
1.53 |
1.80 |
3.48 |
|
| H2-Q8 | 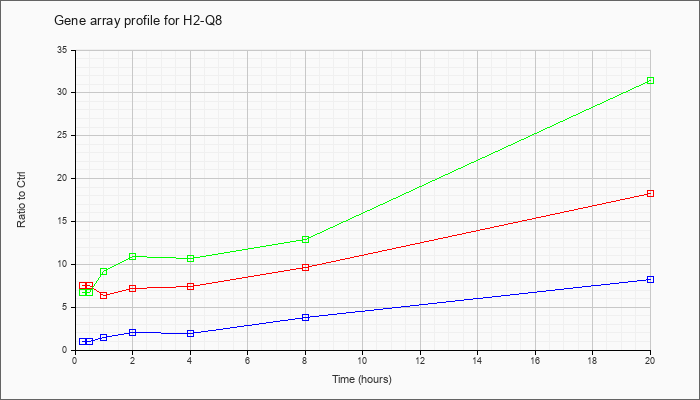 |
NM_023124 |
histocompatibility 2, Q region locus 8 (H2-Q8), mRNA [NM_023124] |
KLA | 7.48 |
6.46 |
6.41 |
7.15 |
7.46 |
9.60 |
18.27 |
| ATP | 1.01 |
.98 |
1.48 |
2.05 |
1.93 |
3.83 |
8.26 |
| KLA/ATP | 6.72 |
7.10 |
9.11 |
10.95 |
10.71 |
12.88 |
31.43 |
|
| H2-T22 | 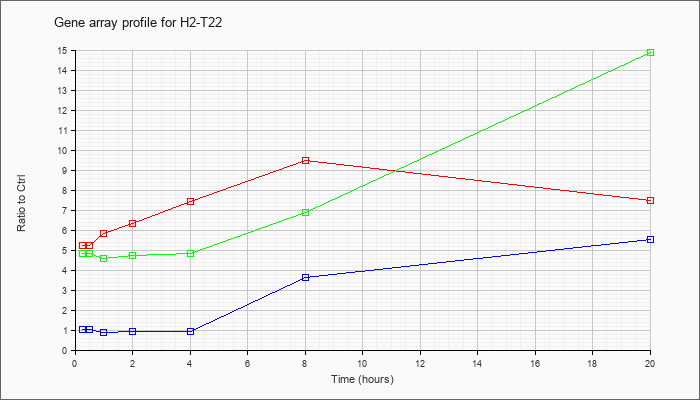 |
NM_010397 |
histocompatibility 2, T region locus 22 (H2-T22), mRNA [NM_010397] |
KLA | 5.21 |
5.40 |
5.81 |
6.31 |
7.42 |
9.49 |
7.50 |
| ATP | 1.01 |
.95 |
.88 |
.92 |
.93 |
3.65 |
5.53 |
| KLA/ATP | 4.83 |
5.12 |
4.58 |
4.74 |
4.81 |
6.89 |
14.89 |
|
| H2-T23 | 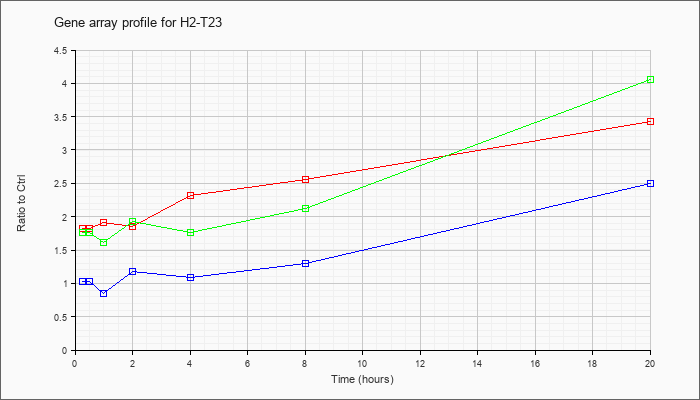 |
NM_010398 |
histocompatibility 2, T region locus 23 (H2-T23), mRNA [NM_010398] |
KLA | 1.83 |
1.80 |
1.92 |
1.86 |
2.32 |
2.56 |
3.43 |
| ATP | 1.03 |
1.08 |
.85 |
1.18 |
1.09 |
1.30 |
2.49 |
| KLA/ATP | 1.76 |
1.92 |
1.62 |
1.93 |
1.76 |
2.13 |
4.06 |
|
| H2-T24 | 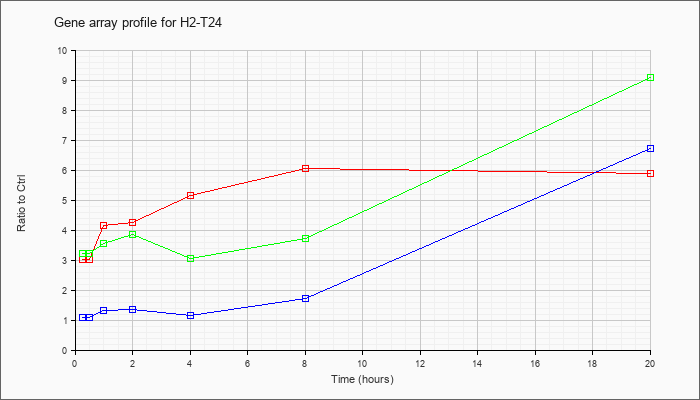 |
NM_008207 |
histocompatibility 2, T region locus 24 (H2-T24), mRNA [NM_008207] |
KLA | 3.02 |
3.26 |
4.14 |
4.26 |
5.15 |
6.06 |
5.89 |
| ATP | 1.08 |
1.09 |
1.32 |
1.35 |
1.17 |
1.71 |
6.71 |
| KLA/ATP | 3.23 |
3.31 |
3.54 |
3.85 |
3.07 |
3.73 |
9.10 |
|
| H2-T3 | 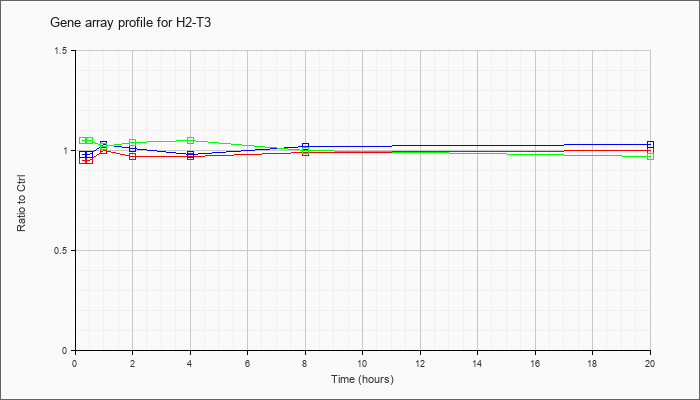 |
NM_008208 |
histocompatibility 2, T region locus 3 (H2-T3), mRNA [NM_008208] |
KLA | .95 |
1.04 |
1.00 |
.97 |
.97 |
.99 |
1.00 |
| ATP | .98 |
1.08 |
1.03 |
1.01 |
.98 |
1.02 |
1.03 |
| KLA/ATP | 1.05 |
.96 |
1.02 |
1.04 |
1.05 |
1.00 |
.97 |
|
| Hras1 | 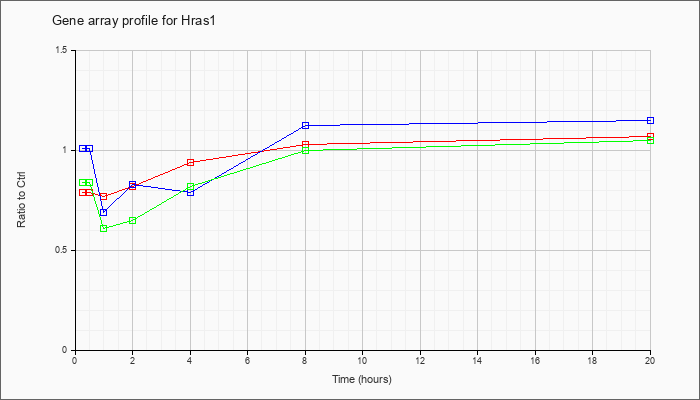 |
NM_008284 |
Harvey rat sarcoma virus oncogene 1 (Hras1), mRNA [NM_008284] |
KLA | .79 |
.82 |
.77 |
.82 |
.94 |
1.03 |
1.07 |
| ATP | 1.01 |
.95 |
.69 |
.83 |
.79 |
1.12 |
1.15 |
| KLA/ATP | .84 |
.77 |
.61 |
.65 |
.82 |
1.00 |
1.05 |
|
| Htatip | 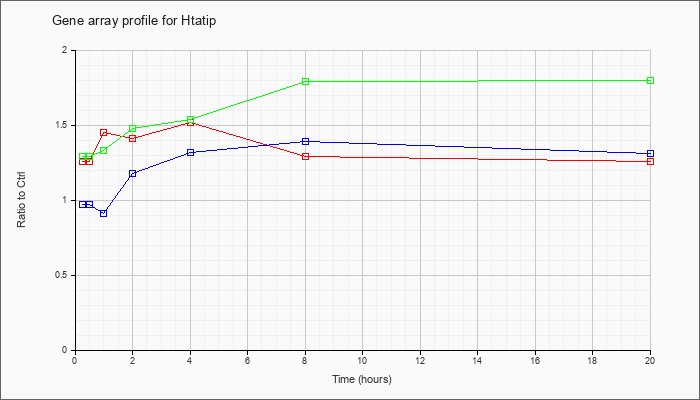 |
NM_178637 |
HIV-1 tat interactive protein, homolog (human) (Htatip), mRNA [NM_178637] |
KLA | 1.26 |
1.40 |
1.45 |
1.41 |
1.52 |
1.29 |
1.26 |
| ATP | .97 |
.96 |
.91 |
1.18 |
1.32 |
1.39 |
1.31 |
| KLA/ATP | 1.29 |
1.32 |
1.33 |
1.48 |
1.54 |
1.79 |
1.80 |
|
| Icam1 | 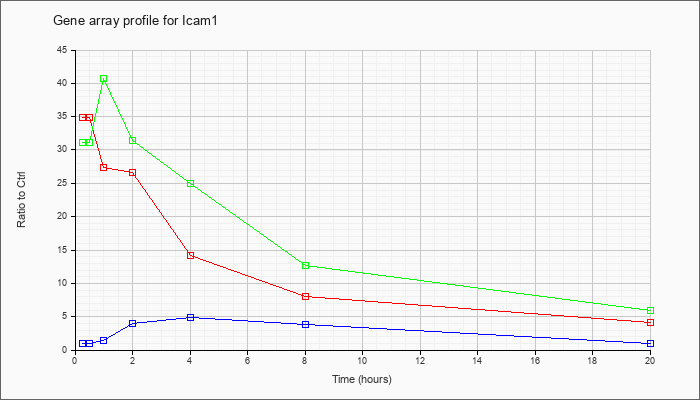 |
BC008626 |
intercellular adhesion molecule, mRNA (cDNA clone MGC:6195 IMAGE:3588949), complete cds [BC008626] |
KLA | 36.67 |
32.93 |
28.63 |
27.91 |
14.65 |
8.16 |
4.15 |
| ATP | .96 |
.88 |
1.50 |
3.88 |
4.98 |
3.90 |
1.01 |
| KLA/ATP | 32.58 |
31.60 |
42.74 |
32.60 |
26.00 |
13.02 |
6.01 |
|
| Icam1 |  |
NM_010493 |
intercellular adhesion molecule 1 (Icam1), mRNA [NM_010493] |
KLA | 15.66 |
15.20 |
12.96 |
12.31 |
9.39 |
6.17 |
3.81 |
| ATP | .96 |
1.06 |
1.21 |
4.19 |
4.33 |
3.16 |
1.06 |
| KLA/ATP | 14.31 |
15.46 |
17.84 |
17.60 |
13.77 |
8.36 |
5.42 |
|
| Ikbkb | 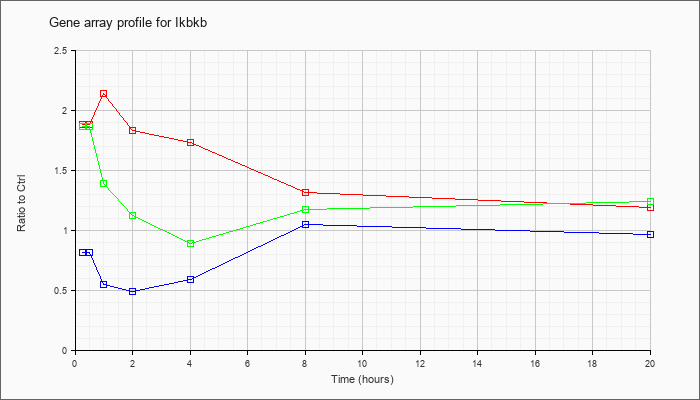 |
NM_010546 |
inhibitor of kappaB kinase beta (Ikbkb), mRNA [NM_010546] |
KLA | 1.88 |
2.10 |
2.14 |
1.83 |
1.73 |
1.31 |
1.19 |
| ATP | .81 |
.74 |
.55 |
.49 |
.59 |
1.05 |
.96 |
| KLA/ATP | 1.86 |
1.73 |
1.39 |
1.12 |
.89 |
1.17 |
1.24 |
|
| Ikbkg | 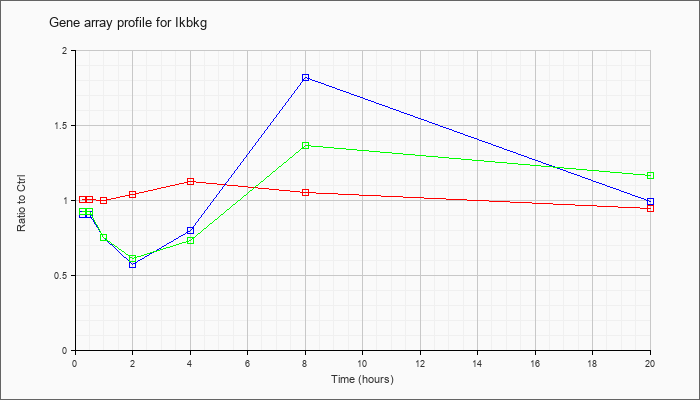 |
AK042138 |
3 days neonate thymus cDNA, RIKEN full-length enriched library, clone:A630062P04 product:unclassifiable, full insert sequence. [AK042138] |
KLA | .92 |
.87 |
.78 |
.78 |
.87 |
.88 |
.62 |
| ATP | .84 |
.71 |
.80 |
.50 |
1.15 |
2.33 |
.61 |
| KLA/ATP | .72 |
.69 |
.74 |
.48 |
1.06 |
1.97 |
.68 |
|
| Ikbkg |  |
NM_010547 |
inhibitor of kappaB kinase gamma (Ikbkg), transcript variant 1, mRNA [NM_010547] |
KLA | 1.51 |
1.63 |
1.74 |
1.83 |
1.83 |
1.73 |
1.72 |
| ATP | .80 |
.79 |
.82 |
.75 |
.66 |
2.58 |
2.37 |
| KLA/ATP | 1.43 |
1.42 |
1.17 |
1.04 |
.71 |
1.54 |
3.17 |
|
| Ikbkg |  |
NM_178590 |
inhibitor of kappaB kinase gamma (Ikbkg), transcript variant 2, mRNA [NM_178590] |
KLA | .84 |
.93 |
.86 |
.91 |
1.03 |
.89 |
.88 |
| ATP | 1.03 |
.98 |
.68 |
.56 |
.51 |
.93 |
.68 |
| KLA/ATP | .89 |
.81 |
.55 |
.52 |
.41 |
.67 |
.65 |
|
| Il15 | 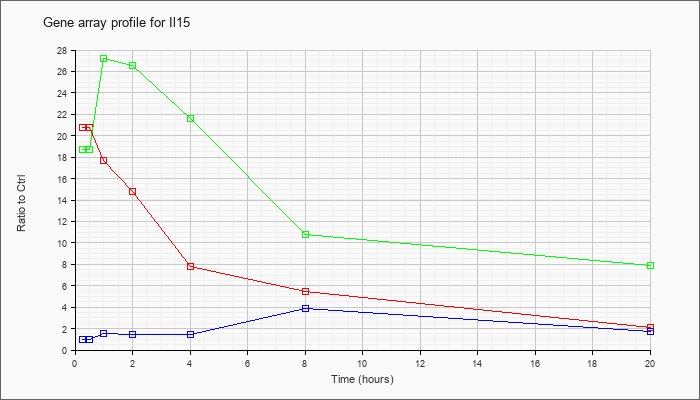 |
AB022307 |
mRNA for shorter isoform of interleukin 15, partial cds. [AB022307] |
KLA | 13.11 |
12.14 |
12.05 |
8.72 |
4.50 |
3.62 |
1.70 |
| ATP | 1.04 |
1.05 |
1.94 |
1.72 |
1.74 |
3.48 |
1.69 |
| KLA/ATP | 13.23 |
13.71 |
21.01 |
21.43 |
21.51 |
10.85 |
4.55 |
|
| Il15 |  |
NM_008357 |
interleukin 15 (Il15), mRNA [NM_008357] |
KLA | 28.44 |
24.75 |
23.34 |
20.83 |
11.02 |
7.27 |
2.51 |
| ATP | 1.00 |
.96 |
1.22 |
1.21 |
1.24 |
4.27 |
1.85 |
| KLA/ATP | 24.24 |
23.93 |
33.47 |
31.76 |
21.73 |
10.79 |
11.23 |
|
| Il15ra |  |
NM_008358 |
interleukin 15 receptor, alpha chain (Il15ra), transcript variant 1, mRNA [NM_008358] |
KLA | 8.74 |
9.57 |
10.15 |
8.79 |
7.56 |
7.63 |
3.77 |
| ATP | .96 |
.93 |
1.02 |
1.07 |
2.02 |
3.73 |
1.46 |
| KLA/ATP | 9.56 |
10.41 |
10.94 |
10.62 |
11.08 |
9.82 |
8.07 |
|
| Il1r1 |  |
NM_008362 |
interleukin 1 receptor, type I (Il1r1), transcript variant 1, mRNA [NM_008362] |
KLA | 1.20 |
1.08 |
1.22 |
1.52 |
2.19 |
2.32 |
1.61 |
| ATP | 1.10 |
.96 |
1.18 |
1.56 |
8.14 |
9.56 |
7.54 |
| KLA/ATP | 1.22 |
1.13 |
1.37 |
1.85 |
4.54 |
6.85 |
8.96 |
|
| Il1r2 | 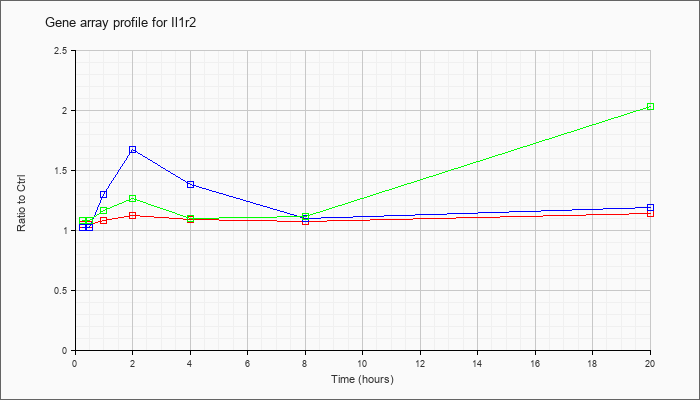 |
NM_010555 |
interleukin 1 receptor, type II (Il1r2), mRNA [NM_010555] |
KLA | 1.05 |
1.03 |
1.08 |
1.13 |
1.09 |
1.08 |
1.14 |
| ATP | 1.03 |
1.08 |
1.30 |
1.68 |
1.38 |
1.10 |
1.19 |
| KLA/ATP | 1.08 |
1.14 |
1.17 |
1.27 |
1.10 |
1.11 |
2.03 |
|
| Il2 | 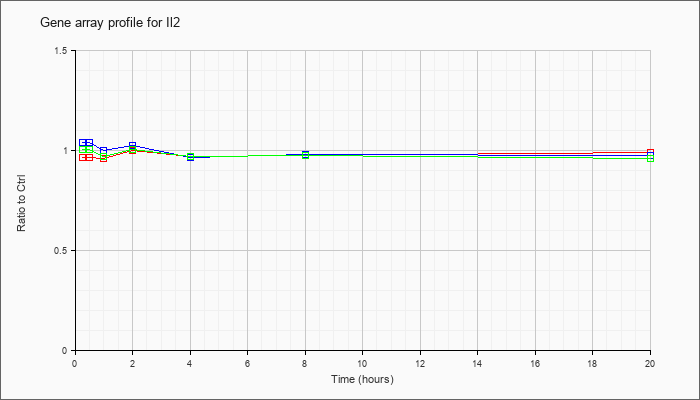 |
NM_008366 |
interleukin 2 (Il2), mRNA [NM_008366] |
KLA | .96 |
.97 |
.96 |
1.00 |
.97 |
.97 |
.99 |
| ATP | 1.04 |
1.02 |
1.00 |
1.03 |
.97 |
.98 |
.97 |
| KLA/ATP | 1.01 |
.98 |
.97 |
1.00 |
.97 |
.97 |
.96 |
|
| Il2ra | 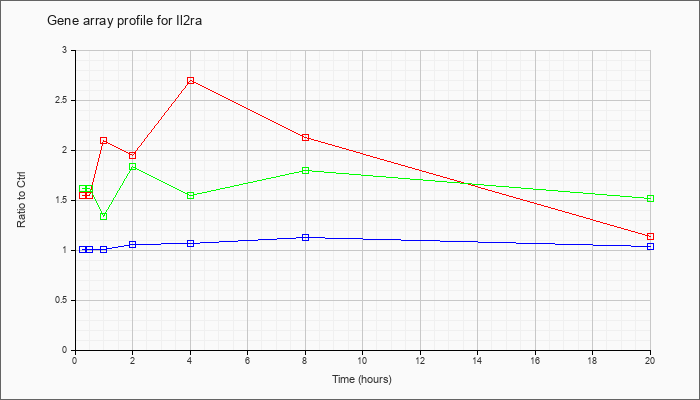 |
AF054581 |
interleukin 2 receptor mRNA, 3UTR, partial sequence. [AF054581] |
KLA | 1.76 |
2.05 |
2.73 |
2.46 |
3.58 |
2.68 |
1.13 |
| ATP | .99 |
.99 |
1.00 |
1.09 |
1.13 |
1.14 |
1.03 |
| KLA/ATP | 1.93 |
2.11 |
1.32 |
2.27 |
1.87 |
2.13 |
1.37 |
|
| Il2ra |  |
NM_008367 |
interleukin 2 receptor, alpha chain (Il2ra), mRNA [NM_008367] |
KLA | 1.33 |
1.29 |
1.46 |
1.43 |
1.82 |
1.57 |
1.15 |
| ATP | 1.02 |
1.03 |
1.01 |
1.02 |
1.01 |
1.12 |
1.05 |
| KLA/ATP | 1.30 |
1.46 |
1.36 |
1.41 |
1.22 |
1.47 |
1.66 |
|
| Il2rb | 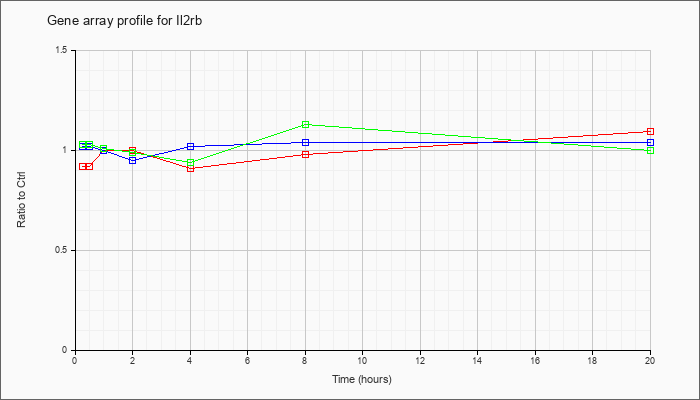 |
NM_008368 |
interleukin 2 receptor, beta chain (Il2rb), mRNA [NM_008368] |
KLA | .92 |
1.00 |
1.00 |
1.00 |
.91 |
.98 |
1.09 |
| ATP | 1.02 |
.98 |
1.00 |
.95 |
1.02 |
1.04 |
1.04 |
| KLA/ATP | 1.03 |
.98 |
1.01 |
.99 |
.94 |
1.13 |
1.00 |
|
| Il2rg | 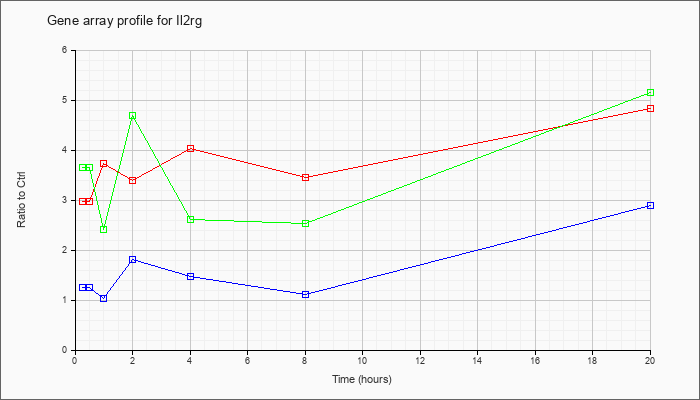 |
NM_013563 |
interleukin 2 receptor, gamma chain (Il2rg), mRNA [NM_013563] |
KLA | 2.97 |
2.92 |
3.73 |
3.39 |
4.04 |
3.45 |
4.83 |
| ATP | 1.25 |
1.41 |
1.04 |
1.82 |
1.48 |
1.12 |
2.89 |
| KLA/ATP | 3.65 |
4.34 |
2.42 |
4.70 |
2.62 |
2.54 |
5.15 |
|
| Il6 | 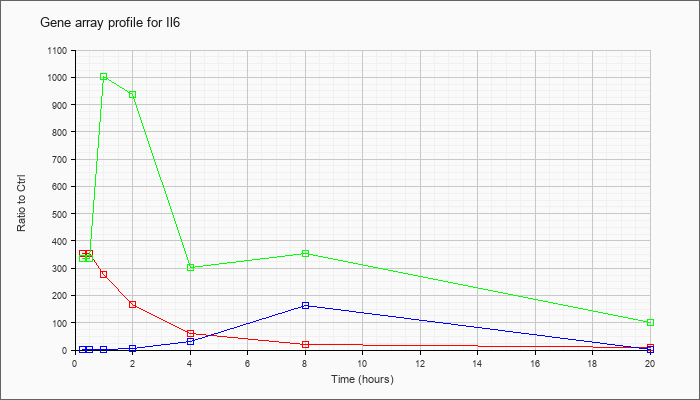 |
NM_031168 |
interleukin 6 (Il6), mRNA [NM_031168] |
KLA | 352.99 |
378.70 |
276.81 |
168.47 |
59.99 |
18.66 |
8.27 |
| ATP | 1.04 |
1.08 |
2.28 |
3.69 |
30.40 |
161.93 |
2.11 |
| KLA/ATP | 334.28 |
528.66 |
###.## |
935.86 |
302.45 |
355.36 |
99.65 |
|
| Itgal | 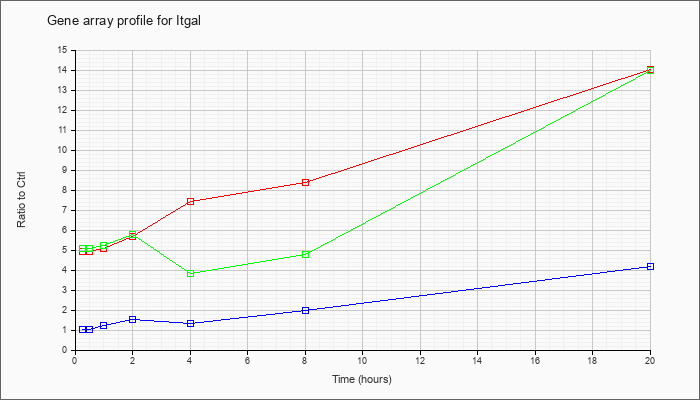 |
AK156251 |
activated spleen cDNA, RIKEN full-length enriched library, clone:F830013L17 product:integrin alpha L, full insert sequence [AK156251] |
KLA | 5.36 |
5.78 |
4.86 |
5.55 |
7.02 |
8.63 |
11.12 |
| ATP | .99 |
1.17 |
1.22 |
1.57 |
1.35 |
1.95 |
3.79 |
| KLA/ATP | 5.42 |
5.89 |
4.95 |
5.91 |
3.96 |
4.93 |
10.15 |
|
| Itgal |  |
NM_008400 |
integrin alpha L (Itgal), mRNA [NM_008400] |
KLA | 4.48 |
5.17 |
5.28 |
5.82 |
7.82 |
8.09 |
16.96 |
| ATP | 1.03 |
1.06 |
1.24 |
1.44 |
1.34 |
2.02 |
4.59 |
| KLA/ATP | 4.71 |
5.63 |
5.49 |
5.60 |
3.66 |
4.65 |
17.76 |
|
| Itgb2 | 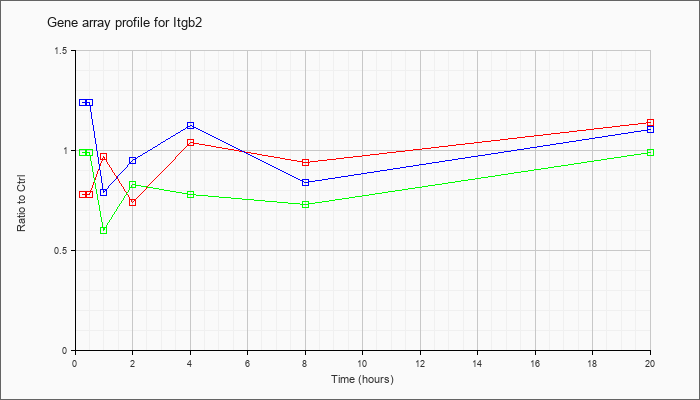 |
NM_008404 |
integrin beta 2 (Itgb2), mRNA [NM_008404] |
KLA | .78 |
.78 |
.97 |
.74 |
1.04 |
.94 |
1.14 |
| ATP | 1.24 |
1.23 |
.79 |
.95 |
1.12 |
.84 |
1.10 |
| KLA/ATP | .99 |
1.01 |
.60 |
.83 |
.78 |
.73 |
.99 |
|
| Itgb2l | 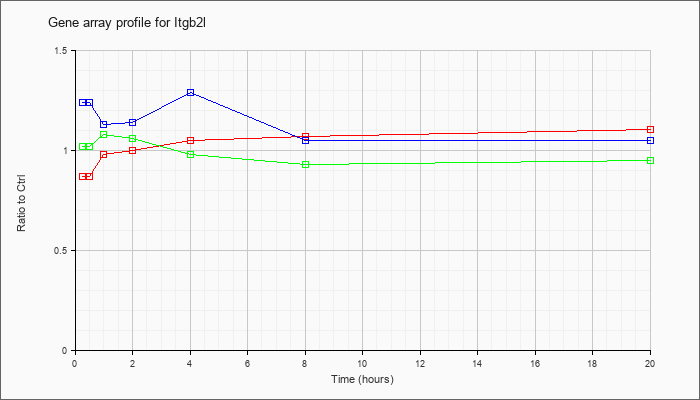 |
NM_008405 |
integrin beta 2-like (Itgb2l), mRNA [NM_008405] |
KLA | .87 |
.98 |
.98 |
1.00 |
1.05 |
1.07 |
1.10 |
| ATP | 1.24 |
1.23 |
1.13 |
1.14 |
1.29 |
1.05 |
1.05 |
| KLA/ATP | 1.02 |
1.08 |
1.08 |
1.06 |
.98 |
.93 |
.95 |
|
| Jak1 | 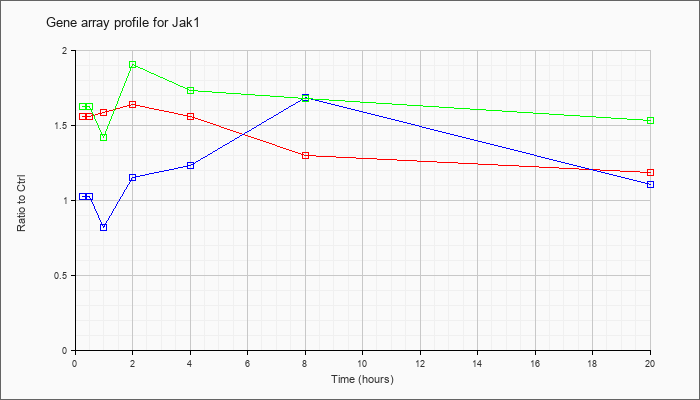 |
AK029232 |
0 day neonate head cDNA, RIKEN full-length enriched library, clone:4831433J01 product:Janus kinase 1, full insert sequence. [AK029232] |
KLA | 1.25 |
1.13 |
1.19 |
1.31 |
1.24 |
1.17 |
1.14 |
| ATP | 1.01 |
1.11 |
.96 |
1.38 |
1.06 |
1.26 |
1.05 |
| KLA/ATP | 1.19 |
1.29 |
1.27 |
1.59 |
1.18 |
1.38 |
1.24 |
|
| Jak1 |  |
NM_146145 |
Janus kinase 1 (Jak1), mRNA [NM_146145] |
KLA | 1.71 |
1.75 |
1.79 |
1.81 |
1.72 |
1.37 |
1.21 |
| ATP | 1.03 |
1.01 |
.75 |
1.04 |
1.32 |
1.90 |
1.14 |
| KLA/ATP | 1.84 |
1.73 |
1.49 |
2.07 |
2.01 |
1.83 |
1.68 |
|
| Jak3 |  |
NM_010589 |
Janus kinase 3 (Jak3), mRNA [NM_010589] |
KLA | 1.46 |
1.56 |
1.79 |
1.62 |
2.05 |
1.72 |
1.65 |
| ATP | 1.11 |
1.19 |
.84 |
1.25 |
.82 |
.61 |
.71 |
| KLA/ATP | 1.76 |
1.82 |
1.10 |
1.74 |
.93 |
.83 |
1.01 |
|
| Jun | 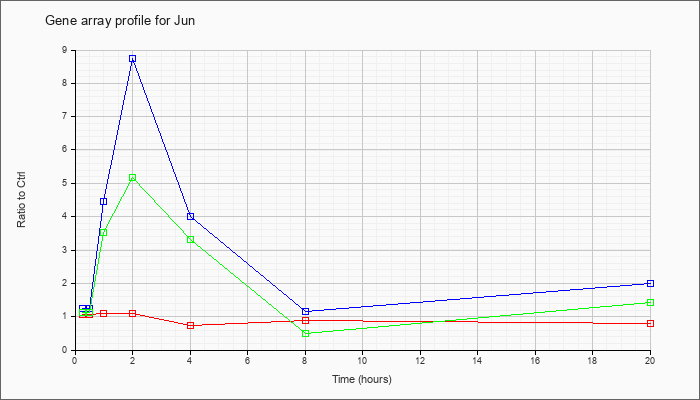 |
NM_010591 |
Jun oncogene (Jun), mRNA [NM_010591] |
KLA | 1.08 |
.96 |
1.09 |
1.09 |
.75 |
.88 |
.81 |
| ATP | 1.26 |
1.48 |
4.45 |
8.73 |
4.02 |
1.16 |
1.99 |
| KLA/ATP | 1.12 |
1.38 |
3.52 |
5.17 |
3.32 |
.49 |
1.42 |
|
| Kras | 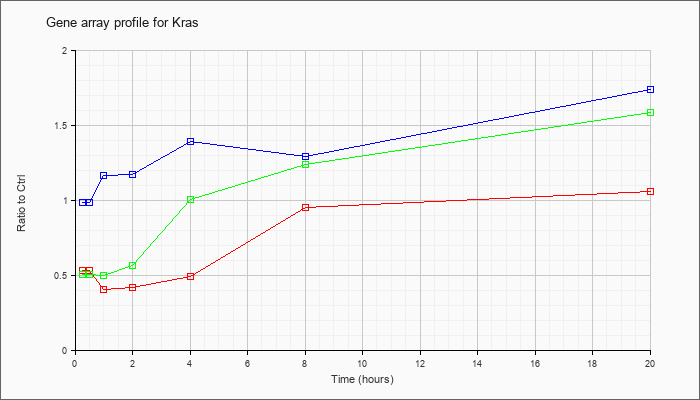 |
NM_021284 |
v-Ki-ras2 Kirsten rat sarcoma viral oncogene homolog (Kras), mRNA [NM_021284] |
KLA | .53 |
.49 |
.40 |
.42 |
.49 |
.95 |
1.05 |
| ATP | .98 |
1.01 |
1.16 |
1.17 |
1.39 |
1.29 |
1.74 |
| KLA/ATP | .50 |
.47 |
.50 |
.56 |
1.00 |
1.24 |
1.58 |
|
LOC67670
8 | 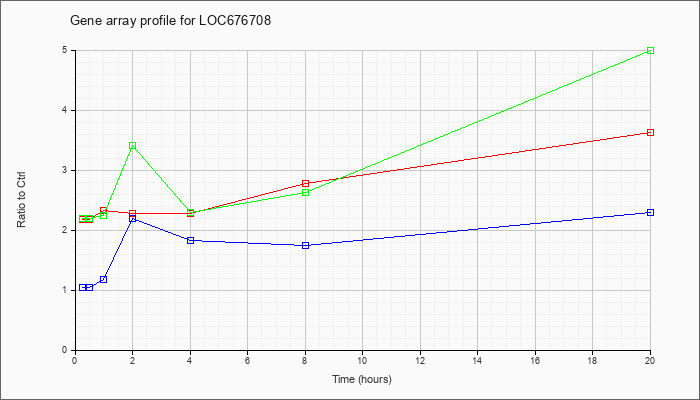 |
XM_992322 |
ref|PREDICTED: Mus musculus similar to class I (Qa) Q2k antigen (LOC676708), mRNA [XM_992322] |
KLA | 2.18 |
2.22 |
2.32 |
2.27 |
2.28 |
2.77 |
3.62 |
| ATP | 1.04 |
1.17 |
1.17 |
2.19 |
1.83 |
1.75 |
2.30 |
| KLA/ATP | 2.20 |
2.44 |
2.25 |
3.41 |
2.30 |
2.63 |
5.00 |
|
| Lck | 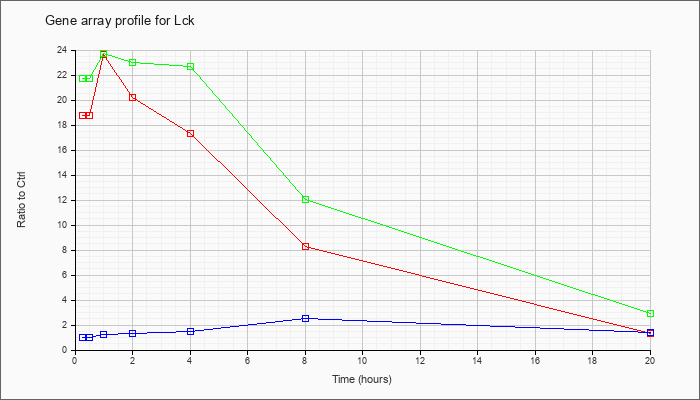 |
NM_010693 |
lymphocyte protein tyrosine kinase (Lck), mRNA [NM_010693] |
KLA | 18.74 |
17.97 |
23.64 |
20.20 |
17.34 |
8.29 |
1.29 |
| ATP | 1.00 |
1.04 |
1.28 |
1.30 |
1.48 |
2.55 |
1.44 |
| KLA/ATP | 21.76 |
24.22 |
23.73 |
23.04 |
22.66 |
12.06 |
2.90 |
|
| Lta | 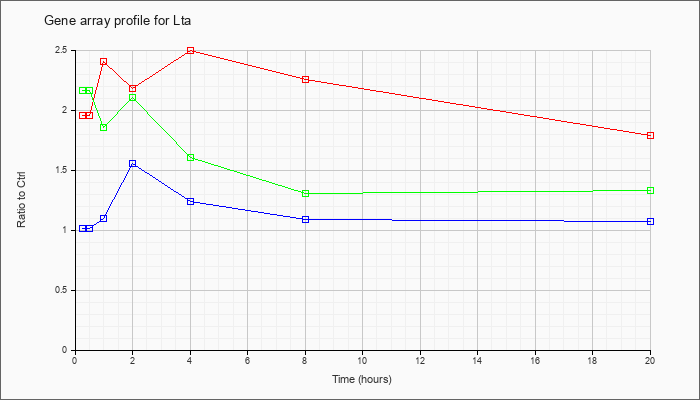 |
NM_010735 |
lymphotoxin A (Lta), mRNA [NM_010735] |
KLA | 1.95 |
2.18 |
2.41 |
2.18 |
2.50 |
2.26 |
1.79 |
| ATP | 1.01 |
1.00 |
1.10 |
1.55 |
1.24 |
1.09 |
1.07 |
| KLA/ATP | 2.16 |
2.23 |
1.85 |
2.11 |
1.60 |
1.31 |
1.33 |
|
| Ltbr | 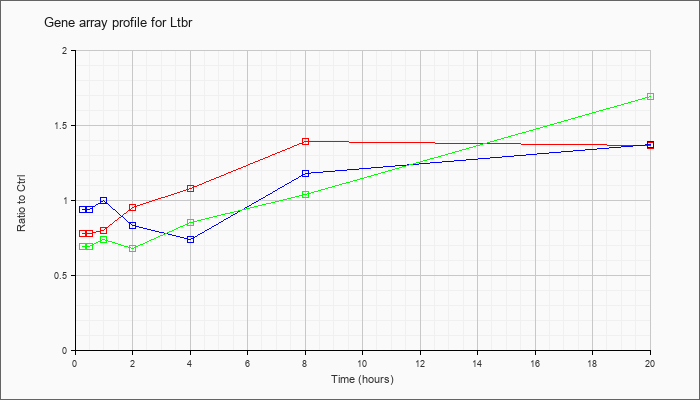 |
NM_010736 |
lymphotoxin B receptor (Ltbr), mRNA [NM_010736] |
KLA | .78 |
.77 |
.80 |
.95 |
1.08 |
1.39 |
1.36 |
| ATP | .94 |
.86 |
1.00 |
.83 |
.74 |
1.18 |
1.37 |
| KLA/ATP | .69 |
.71 |
.74 |
.68 |
.85 |
1.04 |
1.69 |
|
| Mad2l1 | 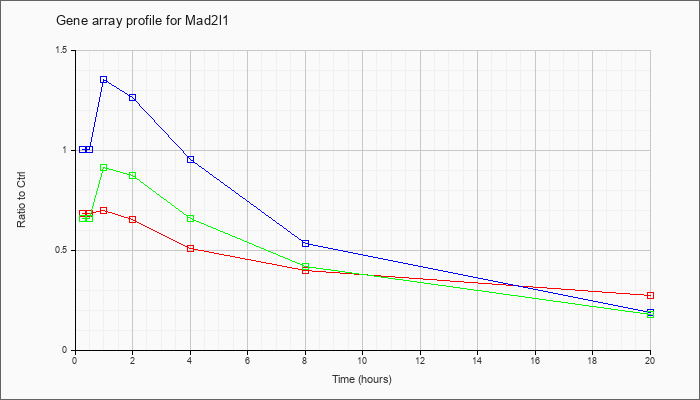 |
NM_019499 |
MAD2 (mitotic arrest deficient, homolog)-like 1 (yeast) (Mad2l1), mRNA [NM_019499] |
KLA | .70 |
.68 |
.73 |
.71 |
.52 |
.41 |
.28 |
| ATP | 1.06 |
1.02 |
1.43 |
1.25 |
1.03 |
.53 |
.19 |
| KLA/ATP | .70 |
.69 |
.99 |
.84 |
.70 |
.43 |
.18 |
|
| Mad2l1 |  |
U83902 |
mitotic checkpoint component Mad2 mRNA, complete cds. [U83902] |
KLA | .67 |
.68 |
.67 |
.60 |
.50 |
.39 |
.27 |
| ATP | .95 |
.97 |
1.28 |
1.28 |
.88 |
.54 |
.19 |
| KLA/ATP | .62 |
.66 |
.84 |
.91 |
.62 |
.41 |
.18 |
|
| Map2k4 | 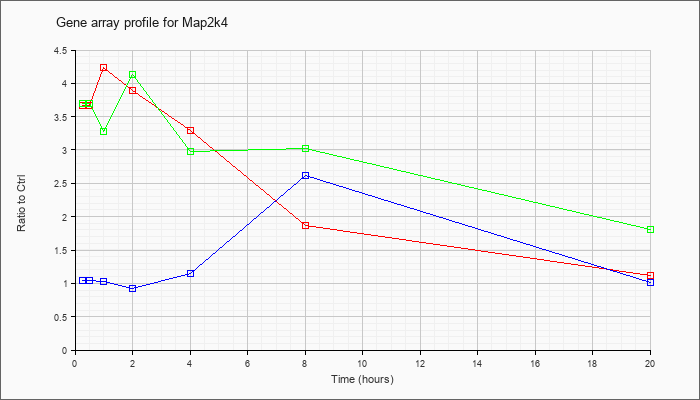 |
NM_009157 |
mitogen-activated protein kinase kinase 4 (Map2k4), mRNA [NM_009157] |
KLA | 3.67 |
3.75 |
4.23 |
3.90 |
3.30 |
1.87 |
1.12 |
| ATP | 1.05 |
1.11 |
1.04 |
.93 |
1.16 |
2.62 |
1.02 |
| KLA/ATP | 3.70 |
4.13 |
3.27 |
4.13 |
2.98 |
3.03 |
1.82 |
|
| Map3k1 | 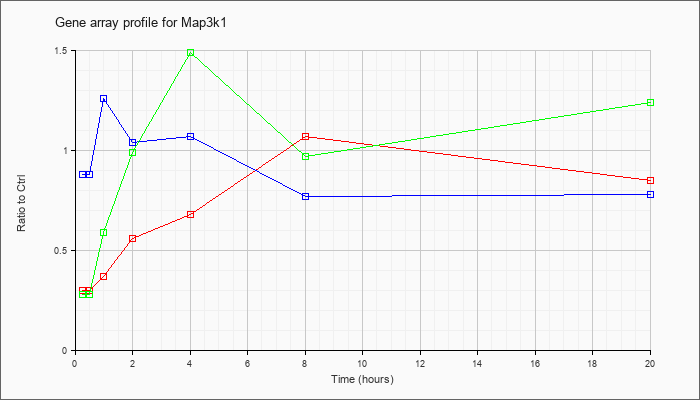 |
NM_011945 |
mitogen-activated protein kinase kinase kinase 1 (Map3k1), mRNA [NM_011945] |
KLA | .30 |
.25 |
.37 |
.56 |
.68 |
1.07 |
.85 |
| ATP | .88 |
.86 |
1.26 |
1.04 |
1.07 |
.77 |
.78 |
| KLA/ATP | .28 |
.24 |
.59 |
.99 |
1.49 |
.97 |
1.24 |
|
| Map3k14 | 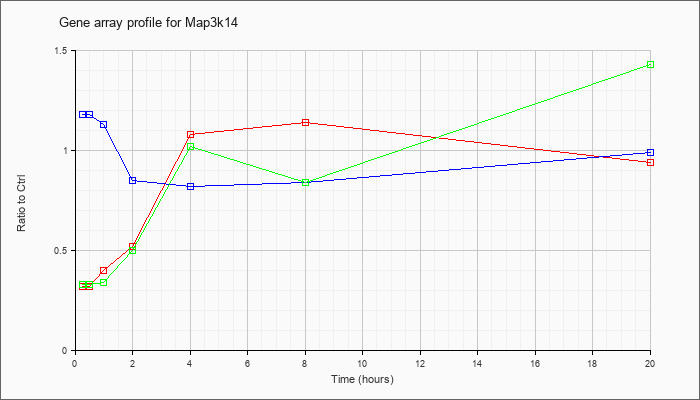 |
NM_016896 |
mitogen-activated protein kinase kinase kinase 14 (Map3k14), mRNA [NM_016896] |
KLA | .32 |
.30 |
.40 |
.52 |
1.08 |
1.14 |
.94 |
| ATP | 1.18 |
1.24 |
1.13 |
.85 |
.82 |
.84 |
.99 |
| KLA/ATP | .33 |
.30 |
.34 |
.50 |
1.02 |
.84 |
1.43 |
|
| Map3k3 | 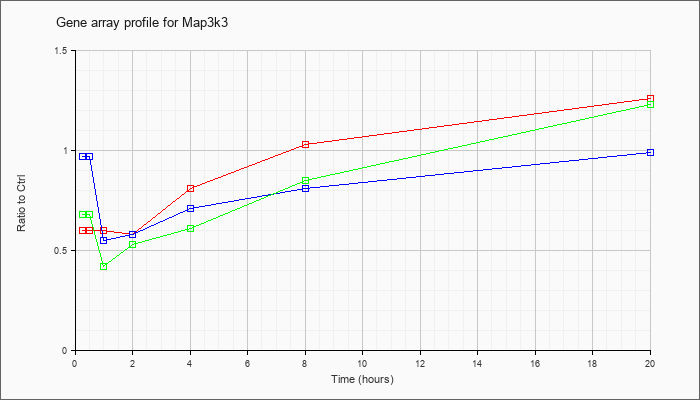 |
NM_011947 |
mitogen-activated protein kinase kinase kinase 3 (Map3k3), mRNA [NM_011947] |
KLA | .60 |
.63 |
.60 |
.58 |
.81 |
1.03 |
1.26 |
| ATP | .97 |
.91 |
.55 |
.58 |
.71 |
.81 |
.99 |
| KLA/ATP | .68 |
.57 |
.42 |
.53 |
.61 |
.85 |
1.23 |
|
| Mapk10 | 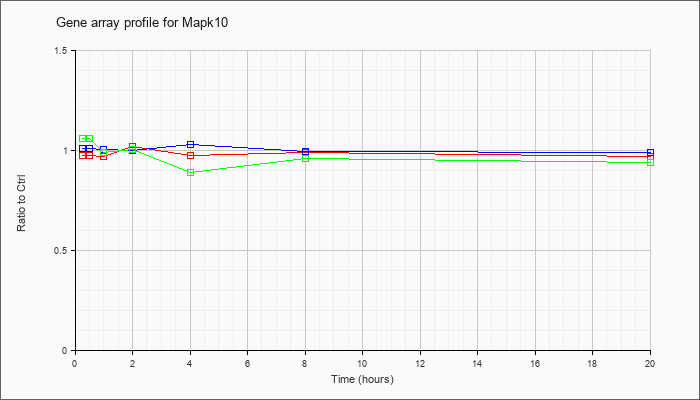 |
AK076990 |
adult male testis cDNA, RIKEN full-length enriched library, clone:4930597C01 product:mitogen activated protein kinase 10, full insert sequence. [AK076990] |
KLA | 1.03 |
1.05 |
.98 |
1.04 |
.94 |
.95 |
1.02 |
| ATP | 1.00 |
.99 |
1.06 |
.97 |
1.01 |
.96 |
.99 |
| KLA/ATP | 1.07 |
1.01 |
.95 |
1.05 |
.82 |
.95 |
.87 |
|
| Mapk10 |  |
NM_009158 |
mitogen-activated protein kinase 10 (Mapk10), transcript variant 1, mRNA [NM_009158] |
KLA | .95 |
1.00 |
.97 |
1.01 |
.99 |
1.01 |
.95 |
| ATP | 1.01 |
1.02 |
.98 |
1.01 |
1.04 |
1.01 |
.99 |
| KLA/ATP | 1.05 |
.99 |
1.01 |
.98 |
.92 |
.97 |
.98 |
|
| Mapk8 |  |
AK162915 |
adult male spinal cord cDNA, RIKEN full-length enriched library, clone:A330031O19 product:mitogen activated protein kinase 8, full insert sequence [AK162915] |
KLA | .94 |
.86 |
.80 |
.83 |
.77 |
.85 |
.83 |
| ATP | 1.08 |
1.12 |
.94 |
.80 |
.83 |
.71 |
.75 |
| KLA/ATP | .83 |
.81 |
.80 |
.74 |
.74 |
.75 |
.67 |
|
| Mapk8 |  |
AK163829 |
16 days embryo head cDNA, RIKEN full-length enriched library, clone:C130070A06 product:mitogen activated protein kinase 8, full insert sequence [AK163829] |
KLA | .65 |
.61 |
.66 |
.70 |
.90 |
1.02 |
1.03 |
| ATP | .99 |
.94 |
1.68 |
1.44 |
1.75 |
1.84 |
1.10 |
| KLA/ATP | .63 |
.67 |
1.11 |
1.00 |
2.01 |
1.68 |
1.15 |
|
| Mapk8 |  |
NM_016700 |
mitogen-activated protein kinase 8 (Mapk8), mRNA [NM_016700] |
KLA | .96 |
1.00 |
.94 |
.96 |
.94 |
.99 |
1.04 |
| ATP | 1.01 |
1.05 |
1.01 |
.99 |
1.03 |
.98 |
1.02 |
| KLA/ATP | .96 |
.97 |
.96 |
.97 |
.93 |
.94 |
.96 |
|
| Mapk9 | 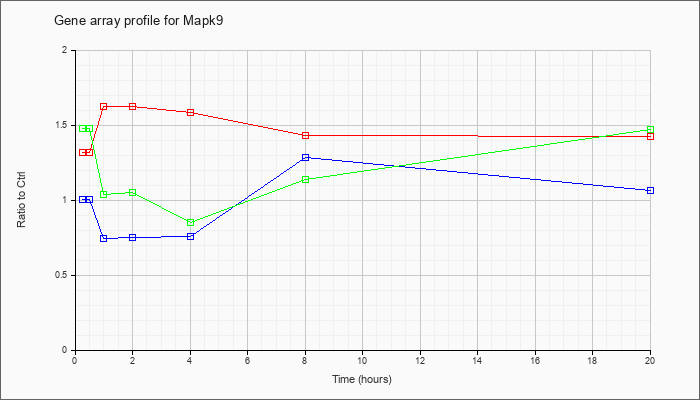 |
NM_016961 |
mitogen-activated protein kinase 9 (Mapk9), transcript variant 2, mRNA [NM_016961] |
KLA | 1.28 |
1.37 |
1.59 |
1.60 |
1.61 |
1.43 |
1.38 |
| ATP | 1.03 |
.99 |
.74 |
.71 |
.79 |
1.20 |
1.01 |
| KLA/ATP | 1.53 |
1.30 |
1.07 |
.99 |
.83 |
1.10 |
1.40 |
|
| Mapk9 |  |
NM_207692 |
mitogen-activated protein kinase 9 (Mapk9), transcript variant 1, mRNA [NM_207692] |
KLA | 1.39 |
1.53 |
1.69 |
1.69 |
1.55 |
1.44 |
1.51 |
| ATP | .95 |
.92 |
.75 |
.85 |
.70 |
1.46 |
1.17 |
| KLA/ATP | 1.37 |
1.35 |
.99 |
1.18 |
.91 |
1.22 |
1.62 |
|
Mm.15898
1 | 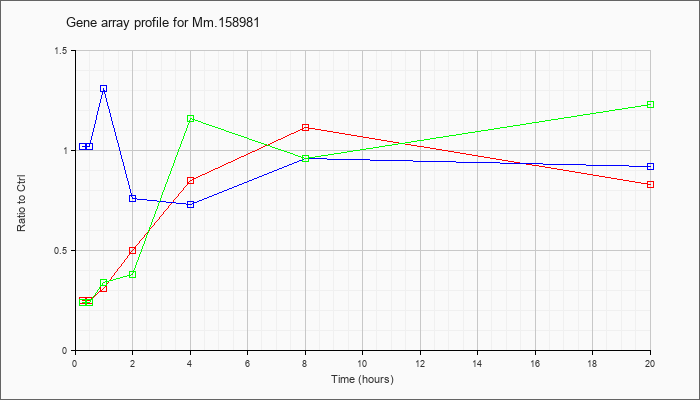 |
142388182 |
Unknown |
KLA | .25 |
.22 |
.31 |
.50 |
.85 |
1.11 |
.83 |
| ATP | 1.02 |
.94 |
1.31 |
.76 |
.73 |
.96 |
.92 |
| KLA/ATP | .24 |
.22 |
.34 |
.38 |
1.16 |
.96 |
1.23 |
|
| Mm.1741 | 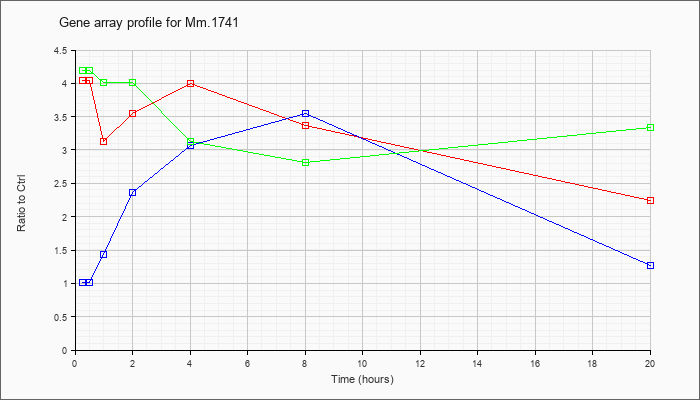 |
31982052 |
Unknown |
KLA | 4.05 |
3.90 |
3.12 |
3.55 |
4.00 |
3.37 |
2.25 |
| ATP | 1.01 |
1.01 |
1.44 |
2.36 |
3.07 |
3.54 |
1.27 |
| KLA/ATP | 4.19 |
3.74 |
4.02 |
4.02 |
3.13 |
2.82 |
3.33 |
|
Mm.19506
1 | 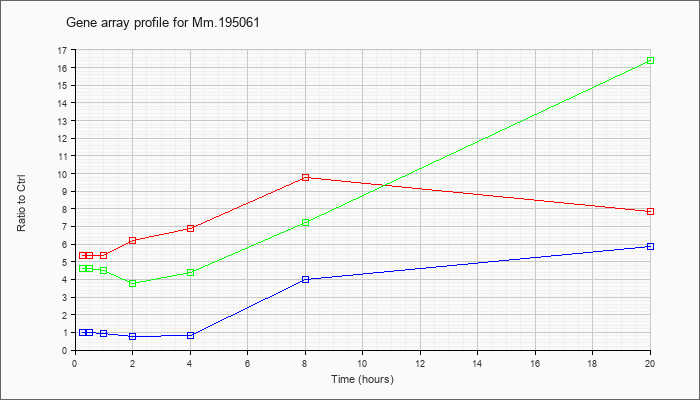 |
76253707 |
Unknown |
KLA | 5.35 |
4.83 |
5.36 |
6.20 |
6.87 |
9.80 |
7.83 |
| ATP | .97 |
.90 |
.96 |
.77 |
.80 |
3.98 |
5.87 |
| KLA/ATP | 4.63 |
4.51 |
4.51 |
3.75 |
4.39 |
7.21 |
16.38 |
|
Mm.21036
1 | 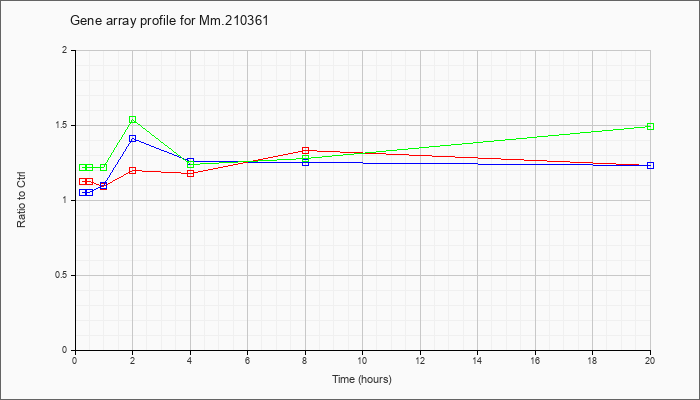 |
158508719 |
Unknown |
KLA | 1.12 |
1.15 |
1.09 |
1.20 |
1.18 |
1.33 |
1.23 |
| ATP | 1.05 |
1.18 |
1.10 |
1.41 |
1.26 |
1.25 |
1.23 |
| KLA/ATP | 1.22 |
1.40 |
1.22 |
1.54 |
1.24 |
1.28 |
1.49 |
|
Mm.21312
8 | 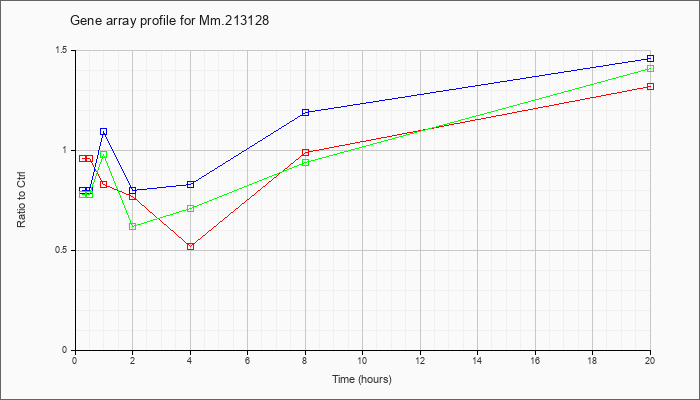 |
141801344 |
Unknown |
KLA | .96 |
.95 |
.83 |
.77 |
.52 |
.99 |
1.32 |
| ATP | .80 |
.82 |
1.09 |
.80 |
.83 |
1.19 |
1.46 |
| KLA/ATP | .78 |
.92 |
.98 |
.62 |
.71 |
.94 |
1.41 |
|
Mm.21516
1 | 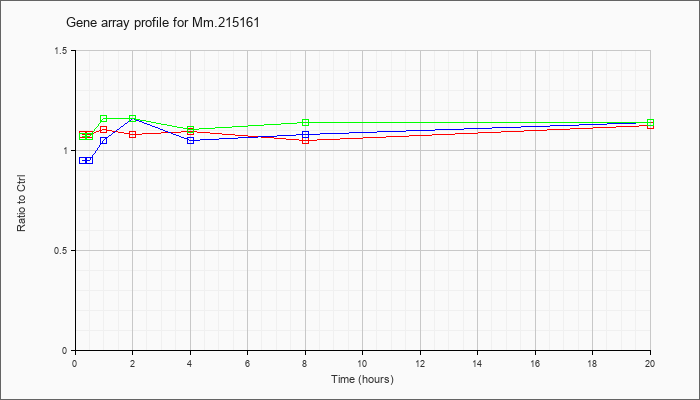 |
141801906 |
Unknown |
KLA | 1.08 |
1.09 |
1.10 |
1.08 |
1.09 |
1.05 |
1.12 |
| ATP | .95 |
.94 |
1.05 |
1.16 |
1.05 |
1.08 |
1.14 |
| KLA/ATP | 1.07 |
1.15 |
1.16 |
1.16 |
1.10 |
1.14 |
1.14 |
|
Mm.23522
| 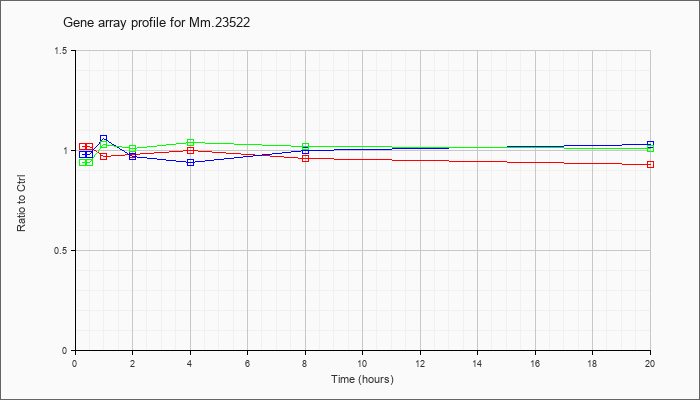 |
124487357 |
Unknown |
KLA | 1.02 |
1.00 |
.97 |
.98 |
1.00 |
.96 |
.93 |
| ATP | .98 |
.99 |
1.06 |
.97 |
.94 |
1.00 |
1.03 |
| KLA/ATP | .94 |
1.01 |
1.03 |
1.01 |
1.04 |
1.02 |
1.01 |
|
Mm.24004
7 | 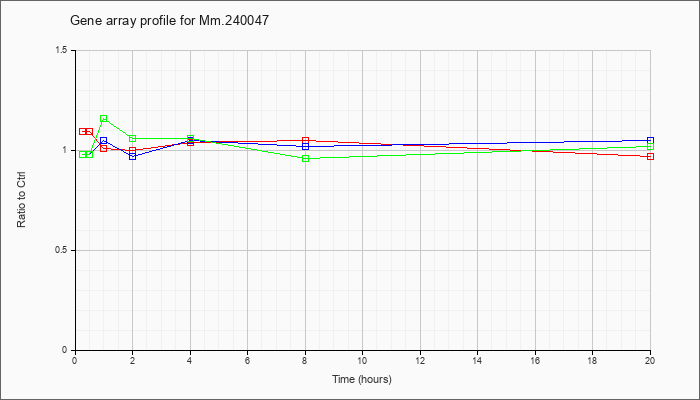 |
118130430 |
Unknown |
KLA | 1.09 |
1.07 |
1.01 |
1.00 |
1.04 |
1.05 |
.97 |
| ATP | .98 |
1.00 |
1.05 |
.97 |
1.05 |
1.02 |
1.05 |
| KLA/ATP | .98 |
1.04 |
1.16 |
1.06 |
1.06 |
.96 |
1.02 |
|
Mm.24600
3 | 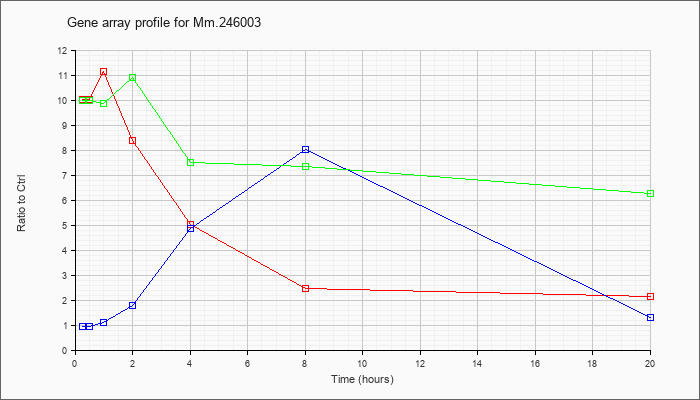 |
161086914 |
Unknown |
KLA | 10.04 |
10.23 |
11.14 |
8.37 |
5.02 |
2.48 |
2.15 |
| ATP | .96 |
1.08 |
1.11 |
1.79 |
4.88 |
8.01 |
1.29 |
| KLA/ATP | 9.96 |
11.13 |
9.87 |
10.92 |
7.51 |
7.35 |
6.26 |
|
Mm.25381
9 | 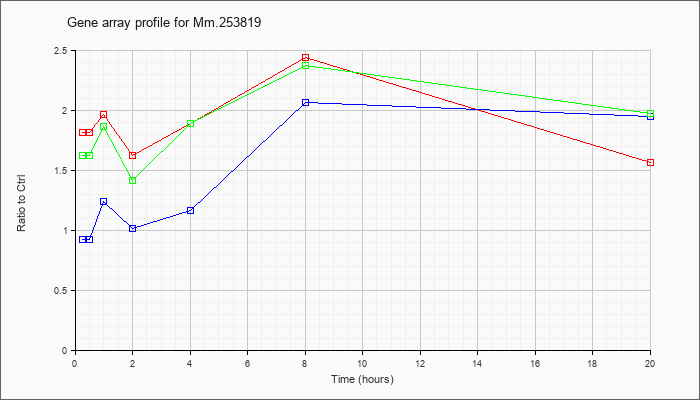 |
118130676 |
Unknown |
KLA | 1.81 |
1.88 |
1.96 |
1.62 |
1.89 |
2.44 |
1.56 |
| ATP | .92 |
.91 |
1.24 |
1.01 |
1.16 |
2.06 |
1.95 |
| KLA/ATP | 1.62 |
1.86 |
1.86 |
1.41 |
1.89 |
2.37 |
1.97 |
|
Mm.26232
7 |  |
118130063 |
Unknown |
KLA | .91 |
.88 |
.82 |
.99 |
.84 |
.98 |
.82 |
| ATP | .84 |
.72 |
.77 |
.54 |
.54 |
.50 |
.72 |
| KLA/ATP | .74 |
.71 |
.74 |
.50 |
.71 |
.60 |
.39 |
|
Mm.27304
9 | 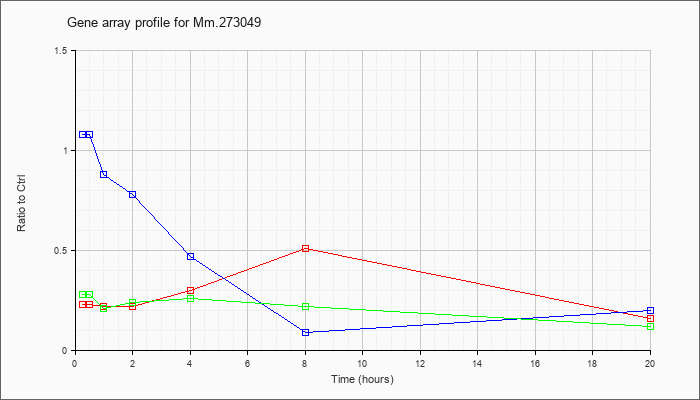 |
119672895 |
Unknown |
KLA | .23 |
.22 |
.22 |
.22 |
.30 |
.51 |
.16 |
| ATP | 1.08 |
1.05 |
.88 |
.78 |
.47 |
.09 |
.20 |
| KLA/ATP | .28 |
.22 |
.21 |
.24 |
.26 |
.22 |
.12 |
|
Mm.30793
2 |  |
160333363 |
Unknown |
KLA | .13 |
.12 |
.14 |
.16 |
.15 |
.18 |
.15 |
| ATP | .92 |
.94 |
1.12 |
.93 |
.58 |
.29 |
.31 |
| KLA/ATP | .11 |
.11 |
.18 |
.20 |
.28 |
.20 |
.19 |
|
Mm.33138
9 | 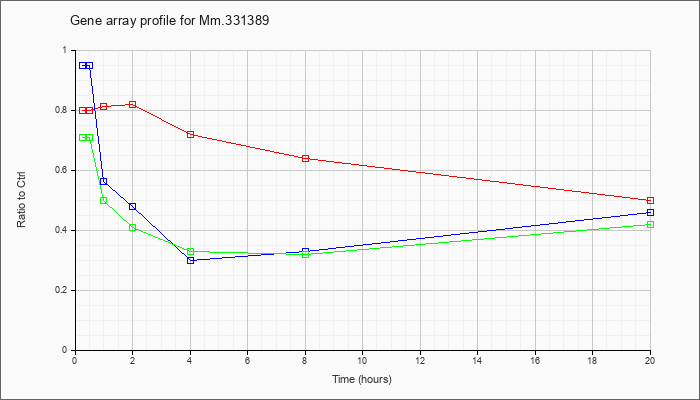 |
158508469 |
Unknown |
KLA | .80 |
.79 |
.81 |
.82 |
.72 |
.64 |
.50 |
| ATP | .95 |
.85 |
.56 |
.48 |
.30 |
.33 |
.46 |
| KLA/ATP | .71 |
.64 |
.50 |
.41 |
.33 |
.32 |
.42 |
|
Mm.41137
| 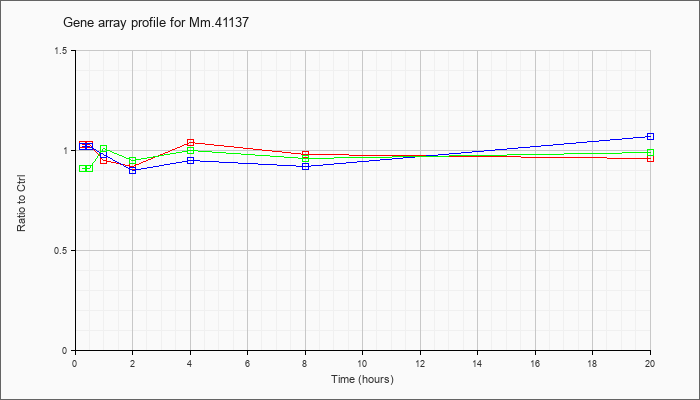 |
148747308 |
Unknown |
KLA | 1.03 |
1.00 |
.95 |
.92 |
1.04 |
.98 |
.96 |
| ATP | 1.02 |
.98 |
.98 |
.90 |
.95 |
.92 |
1.07 |
| KLA/ATP | .91 |
.89 |
1.01 |
.95 |
1.00 |
.96 |
.99 |
|
Mm.41292
2 | 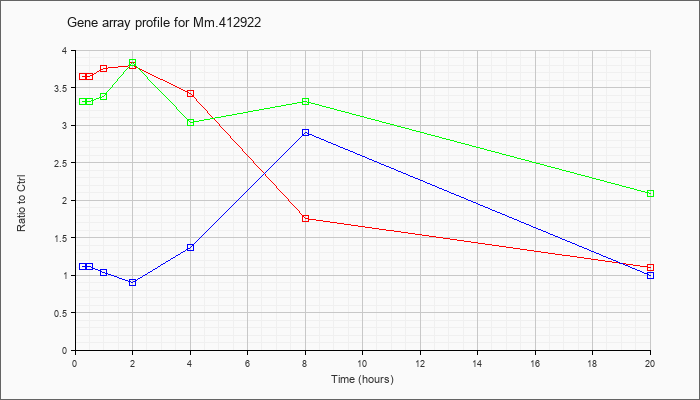 |
145966829 |
Unknown |
KLA | 3.65 |
3.76 |
3.75 |
3.79 |
3.42 |
1.75 |
1.10 |
| ATP | 1.11 |
1.17 |
1.04 |
.90 |
1.37 |
2.90 |
1.00 |
| KLA/ATP | 3.31 |
3.97 |
3.38 |
3.84 |
3.04 |
3.31 |
2.09 |
|
Mm.43967
5 | 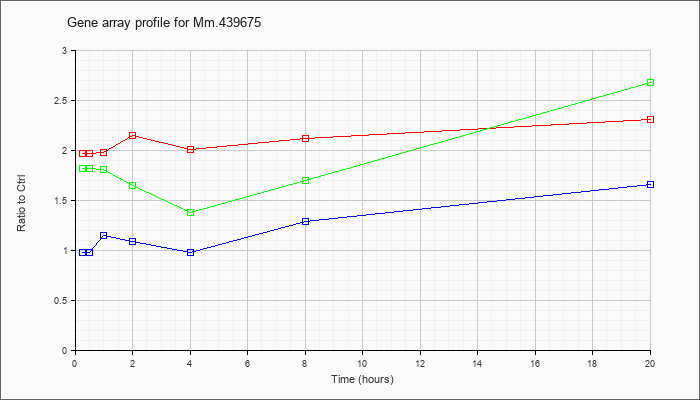 |
133778954 |
Unknown |
KLA | 1.76 |
1.74 |
1.70 |
1.72 |
1.75 |
2.14 |
3.19 |
| ATP | 1.00 |
1.01 |
1.18 |
1.28 |
1.25 |
1.48 |
1.84 |
| KLA/ATP | 1.67 |
1.64 |
1.70 |
1.79 |
1.63 |
2.04 |
3.36 |
|
Mm.43967
5 |  |
149275074 |
Unknown |
KLA | 2.17 |
2.23 |
2.26 |
2.58 |
2.26 |
2.10 |
1.43 |
| ATP | .96 |
.91 |
1.12 |
.90 |
.70 |
1.10 |
1.47 |
| KLA/ATP | 1.96 |
2.10 |
1.92 |
1.51 |
1.12 |
1.35 |
1.99 |
|
Mm.44165
1 |  |
160333403 |
Unknown |
KLA | 5.45 |
5.27 |
5.02 |
6.34 |
6.63 |
10.05 |
8.10 |
| ATP | .91 |
.84 |
.96 |
.75 |
.78 |
3.91 |
6.03 |
| KLA/ATP | 4.29 |
4.38 |
4.50 |
3.55 |
4.22 |
7.22 |
17.63 |
|
Mm.45044
|  |
133892959 |
Unknown |
KLA | .51 |
.47 |
.52 |
.83 |
1.25 |
1.16 |
.60 |
| ATP | 1.09 |
.78 |
.80 |
.63 |
.83 |
.89 |
.41 |
| KLA/ATP | .49 |
.42 |
.44 |
.39 |
.89 |
1.04 |
.41 |
|
Mm.45825
9 | 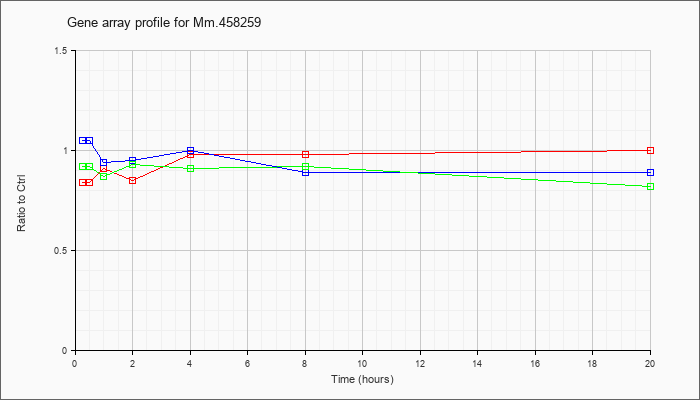 |
116292181 |
Unknown |
KLA | .84 |
.86 |
.91 |
.85 |
.98 |
.98 |
1.00 |
| ATP | 1.05 |
1.06 |
.94 |
.95 |
1.00 |
.89 |
.89 |
| KLA/ATP | .92 |
.84 |
.87 |
.93 |
.91 |
.92 |
.82 |
|
Mm.46781
1 | 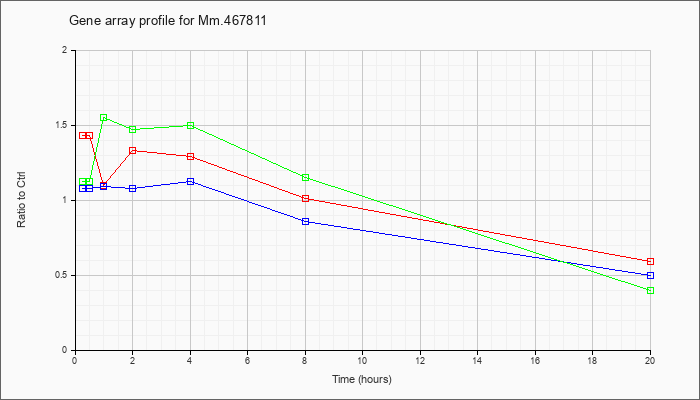 |
153792000 |
Unknown |
KLA | 1.43 |
1.25 |
1.10 |
1.33 |
1.29 |
1.01 |
.59 |
| ATP | 1.08 |
1.15 |
1.09 |
1.08 |
1.12 |
.86 |
.50 |
| KLA/ATP | 1.12 |
1.24 |
1.55 |
1.47 |
1.50 |
1.15 |
.40 |
|
Mm.47124
2 | 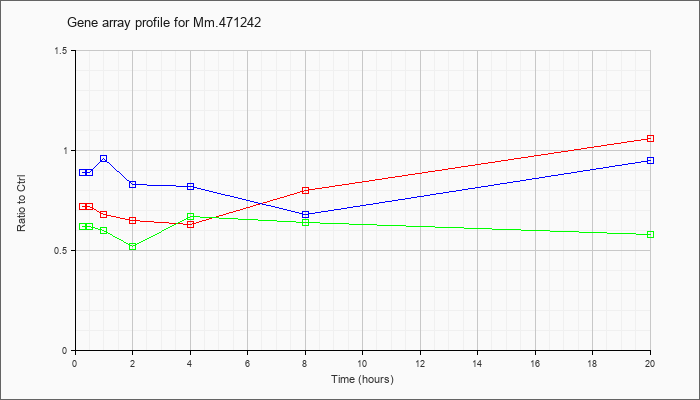 |
31542487 |
Unknown |
KLA | .72 |
.67 |
.68 |
.65 |
.63 |
.80 |
1.06 |
| ATP | .89 |
.87 |
.96 |
.83 |
.82 |
.68 |
.95 |
| KLA/ATP | .62 |
.60 |
.60 |
.52 |
.67 |
.64 |
.58 |
|
Mm.47351
0 | 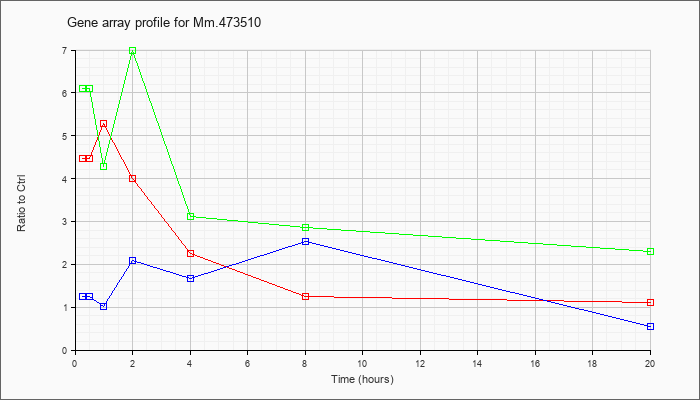 |
162287332 |
Unknown |
KLA | 4.46 |
4.82 |
5.29 |
4.00 |
2.26 |
1.25 |
1.10 |
| ATP | 1.24 |
1.72 |
1.02 |
2.08 |
1.66 |
2.53 |
.55 |
| KLA/ATP | 6.11 |
7.67 |
4.29 |
6.99 |
3.11 |
2.86 |
2.30 |
|
| Mm.4769 | 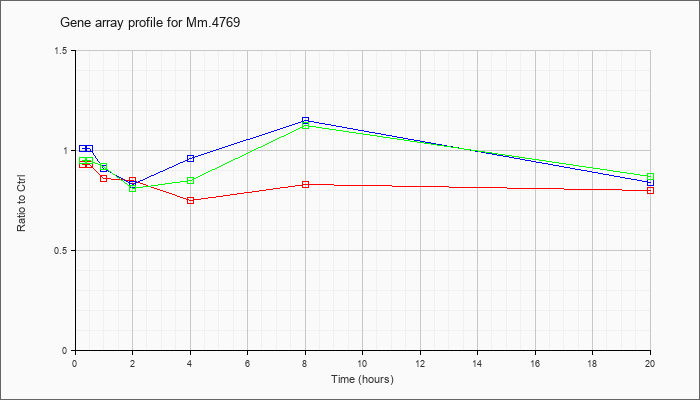 |
31542830 |
Unknown |
KLA | .93 |
.99 |
.86 |
.85 |
.75 |
.83 |
.80 |
| ATP | 1.01 |
1.00 |
.91 |
.83 |
.96 |
1.15 |
.84 |
| KLA/ATP | .95 |
.95 |
.92 |
.81 |
.85 |
1.12 |
.87 |
|
Mm.56964
| 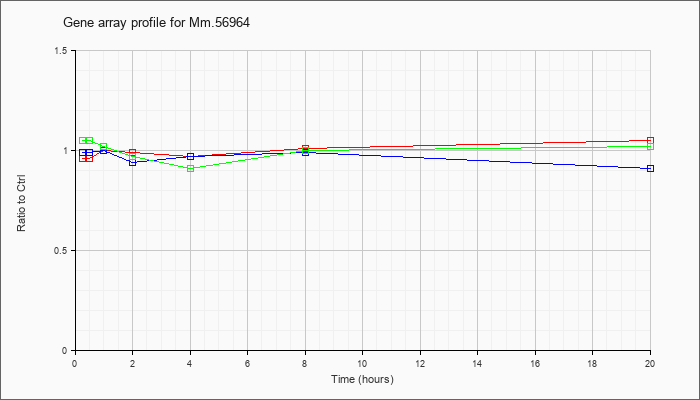 |
144227223 |
Unknown |
KLA | .96 |
.99 |
1.00 |
.99 |
.97 |
1.01 |
1.05 |
| ATP | .99 |
.94 |
1.00 |
.94 |
.97 |
.99 |
.91 |
| KLA/ATP | 1.05 |
.97 |
1.02 |
.97 |
.91 |
1.00 |
1.02 |
|
| Mras |  |
NM_008624 |
muscle and microspikes RAS (Mras), mRNA [NM_008624] |
KLA | .77 |
.76 |
.75 |
.84 |
.92 |
.87 |
1.35 |
| ATP | 1.02 |
1.00 |
1.11 |
1.14 |
.87 |
.79 |
.80 |
| KLA/ATP | .77 |
.77 |
.86 |
.78 |
.69 |
.63 |
.61 |
|
| Msx1 | 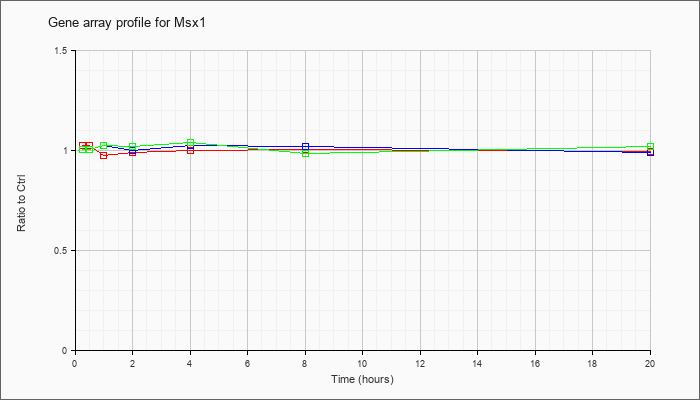 |
NM_010835 |
homeobox, msh-like 1 (Msx1), mRNA [NM_010835] |
KLA | 1.11 |
1.06 |
.85 |
1.00 |
.93 |
.96 |
1.04 |
| ATP | 1.09 |
1.07 |
1.02 |
1.26 |
1.03 |
.96 |
1.03 |
| KLA/ATP | 1.02 |
1.04 |
.93 |
1.11 |
1.06 |
1.03 |
1.17 |
|
| Msx1 |  |
X59251 |
gb|Mouse Hox-7 mRNA for homeobox protein, homolog of Drosophila Msh gene. [X59251] |
KLA | 1.01 |
1.00 |
.98 |
.99 |
1.00 |
1.01 |
.99 |
| ATP | .99 |
1.01 |
1.03 |
.98 |
1.02 |
1.02 |
.99 |
| KLA/ATP | 1.00 |
1.03 |
1.03 |
1.01 |
1.04 |
.98 |
1.01 |
|
| Msx2 | 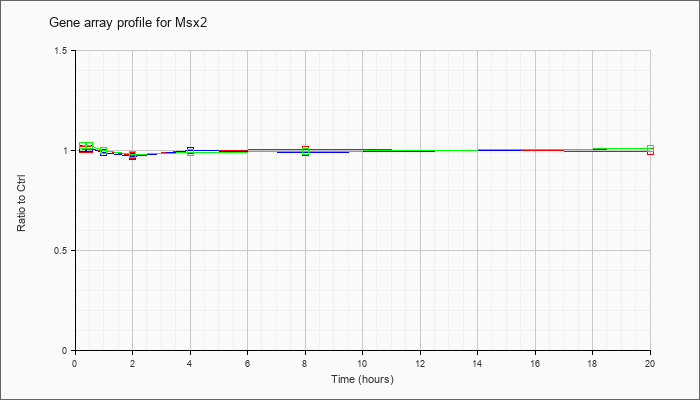 |
NM_013601 |
homeobox, msh-like 2 (Msx2), mRNA [NM_013601] |
KLA | 1.00 |
1.00 |
1.00 |
.97 |
1.00 |
1.00 |
.99 |
| ATP | 1.01 |
.97 |
.99 |
.97 |
1.00 |
.99 |
1.01 |
| KLA/ATP | 1.02 |
.99 |
1.00 |
.98 |
.99 |
.99 |
1.01 |
|
| Msx3 |  |
NM_010836 |
homeobox, msh-like 3 (Msx3), mRNA [NM_010836] |
KLA | 1.00 |
.98 |
.95 |
.99 |
1.02 |
.99 |
.98 |
| ATP | .98 |
.99 |
.97 |
.93 |
.96 |
1.03 |
1.01 |
| KLA/ATP | 1.01 |
.98 |
1.03 |
.97 |
.95 |
.96 |
1.04 |
|
| Myb |  |
AK036518 |
adult male bone cDNA, RIKEN full-length enriched library, clone:9830125P17 product:myeloblastosis oncogene, full insert sequence. [AK036518] |
KLA | 1.02 |
.96 |
1.03 |
1.00 |
.96 |
1.07 |
.99 |
| ATP | 1.07 |
.96 |
1.03 |
.99 |
.96 |
1.03 |
1.00 |
| KLA/ATP | 1.04 |
1.07 |
.83 |
.93 |
1.06 |
1.03 |
1.00 |
|
| Myb |  |
M16449 |
gb|Mouse myb mRNA, complete cds [M16449] |
KLA | .97 |
1.07 |
1.09 |
1.01 |
1.01 |
1.02 |
.98 |
| ATP | 1.06 |
.98 |
1.05 |
1.06 |
.91 |
.91 |
.84 |
| KLA/ATP | 1.08 |
1.07 |
.80 |
1.03 |
1.01 |
.92 |
.90 |
|
| Myb |  |
NM_010848 |
myeloblastosis oncogene (Myb), mRNA [NM_010848] |
KLA | 1.07 |
1.05 |
1.11 |
1.09 |
1.06 |
.99 |
.91 |
| ATP | .99 |
1.05 |
1.05 |
.95 |
.94 |
.93 |
.82 |
| KLA/ATP | 1.04 |
1.09 |
1.07 |
.97 |
.94 |
.95 |
.82 |
|
| Mybl1 | 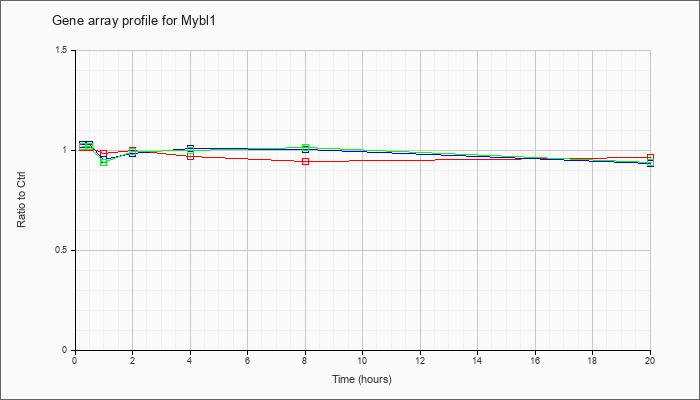 |
NM_008651 |
myeloblastosis oncogene-like 1 (Mybl1), mRNA [NM_008651] |
KLA | 1.04 |
1.01 |
.98 |
1.02 |
.97 |
.96 |
.90 |
| ATP | 1.04 |
1.01 |
.97 |
1.07 |
1.02 |
1.03 |
.94 |
| KLA/ATP | 1.06 |
.97 |
.96 |
1.06 |
1.03 |
1.01 |
.96 |
|
| Mybl1 |  |
X82327 |
gb|M.musculus mRNA for A-Myb protein. [X82327] |
KLA | .99 |
.93 |
.99 |
.98 |
.97 |
.93 |
1.03 |
| ATP | 1.02 |
1.01 |
.94 |
.90 |
1.00 |
.98 |
.93 |
| KLA/ATP | .98 |
1.06 |
.92 |
.93 |
.97 |
1.02 |
.92 |
|
| Mybl2 | 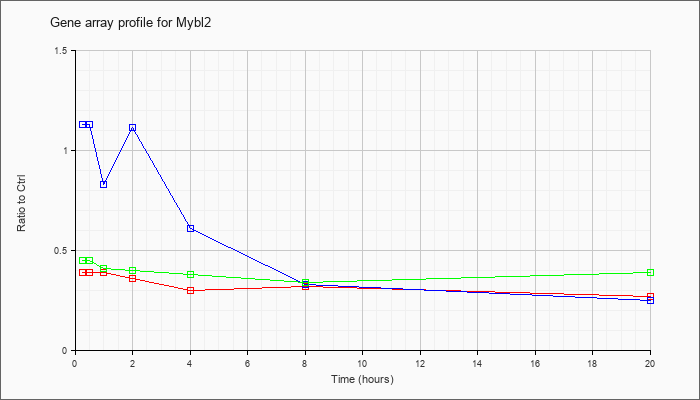 |
NM_008652 |
myeloblastosis oncogene-like 2 (Mybl2), mRNA [NM_008652] |
KLA | .39 |
.38 |
.39 |
.36 |
.30 |
.32 |
.27 |
| ATP | 1.13 |
1.13 |
.83 |
1.11 |
.61 |
.33 |
.25 |
| KLA/ATP | .45 |
.39 |
.41 |
.40 |
.38 |
.34 |
.39 |
|
| Myc |  |
NM_010849 |
myelocytomatosis oncogene (Myc), mRNA [NM_010849] |
KLA | .38 |
.31 |
.45 |
.45 |
.49 |
.35 |
.33 |
| ATP | 1.05 |
1.09 |
1.72 |
4.41 |
1.28 |
.45 |
.76 |
| KLA/ATP | .37 |
.30 |
.44 |
1.23 |
.34 |
.17 |
.19 |
|
| Nfat5 | 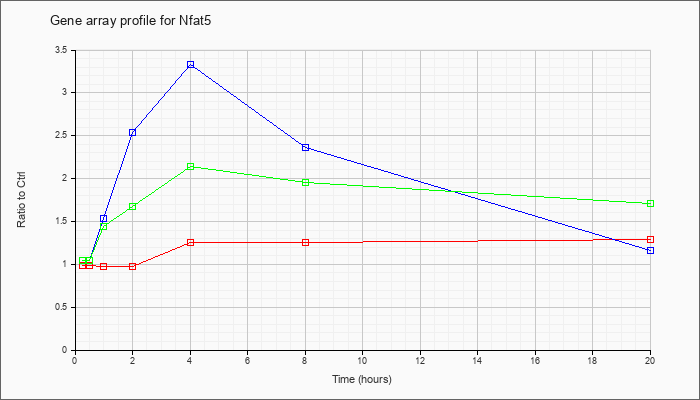 |
AK172011 |
activated spleen cDNA, RIKEN full-length enriched library, clone:F830029B05 product:nuclear factor of activated T-cells 5, full insert sequence. [AK172011] |
KLA | .94 |
.90 |
.93 |
.93 |
1.28 |
1.27 |
1.16 |
| ATP | .95 |
.76 |
1.13 |
1.27 |
1.93 |
2.21 |
1.18 |
| KLA/ATP | .85 |
.84 |
1.10 |
.92 |
1.53 |
1.63 |
1.66 |
|
| Nfat5 |  |
NM_018823 |
nuclear factor of activated T-cells 5 (Nfat5), transcript variant b, mRNA [NM_018823] |
KLA | .95 |
1.01 |
.99 |
1.03 |
1.08 |
1.13 |
1.19 |
| ATP | 1.05 |
1.22 |
1.67 |
1.98 |
1.74 |
1.38 |
1.17 |
| KLA/ATP | 1.03 |
1.07 |
1.34 |
1.38 |
1.30 |
1.19 |
1.40 |
|
| Nfat5 |  |
NM_133957 |
nuclear factor of activated T-cells 5 (Nfat5), transcript variant a, mRNA [NM_133957] |
KLA | 1.07 |
1.07 |
1.00 |
.96 |
1.42 |
1.36 |
1.51 |
| ATP | 1.15 |
1.19 |
1.81 |
4.36 |
6.34 |
3.49 |
1.12 |
| KLA/ATP | 1.27 |
1.08 |
1.87 |
2.72 |
3.58 |
3.05 |
2.07 |
|
| Nfatc1 |  |
NM_016791 |
nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 1 (Nfatc1), transcript variant 1, mRNA [NM_016791] |
KLA | .71 |
.66 |
.68 |
.62 |
.63 |
.67 |
.81 |
| ATP | 1.12 |
1.29 |
1.15 |
1.21 |
.83 |
.63 |
1.25 |
| KLA/ATP | .78 |
.79 |
.65 |
.74 |
.59 |
.45 |
.92 |
|
| Nfatc1 |  |
NM_198429 |
nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 1 (Nfatc1), transcript variant 2, mRNA [NM_198429] |
KLA | .49 |
.41 |
.50 |
.43 |
.50 |
.78 |
.69 |
| ATP | 1.05 |
.81 |
.84 |
.54 |
.60 |
.38 |
.69 |
| KLA/ATP | .46 |
.37 |
.45 |
.32 |
.62 |
.39 |
.61 |
|
| Nfatc2 | 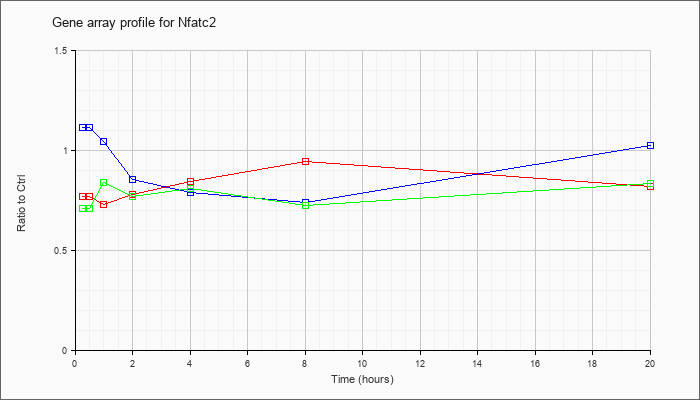 |
AK081853 |
16 days embryo head cDNA, RIKEN full-length enriched library, clone:C130082H17 product:nuclear factor of activated T-cells, cytoplasmic 2, full insert sequence. [AK081853] |
KLA | .56 |
.52 |
.59 |
.61 |
.78 |
.78 |
.66 |
| ATP | 1.19 |
1.27 |
.67 |
.65 |
.62 |
.53 |
.79 |
| KLA/ATP | .52 |
.49 |
.71 |
.57 |
.65 |
.52 |
.53 |
|
| Nfatc2 |  |
NM_001037177 |
nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 2 (Nfatc2), transcript variant 2, mRNA [NM_001037177] |
KLA | .84 |
.79 |
.74 |
.82 |
.89 |
.98 |
.80 |
| ATP | 1.06 |
1.13 |
1.01 |
.87 |
.86 |
.83 |
1.10 |
| KLA/ATP | .76 |
.72 |
.89 |
.84 |
.84 |
.81 |
.84 |
|
| Nfatc2 |  |
NM_010899 |
nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 2 (Nfatc2), transcript variant 1, mRNA [NM_010899] |
KLA | .90 |
.91 |
.86 |
.91 |
.86 |
1.07 |
1.00 |
| ATP | 1.08 |
1.11 |
1.45 |
1.04 |
.88 |
.85 |
1.18 |
| KLA/ATP | .85 |
.89 |
.92 |
.89 |
.93 |
.84 |
1.13 |
|
| Nfatc3 | 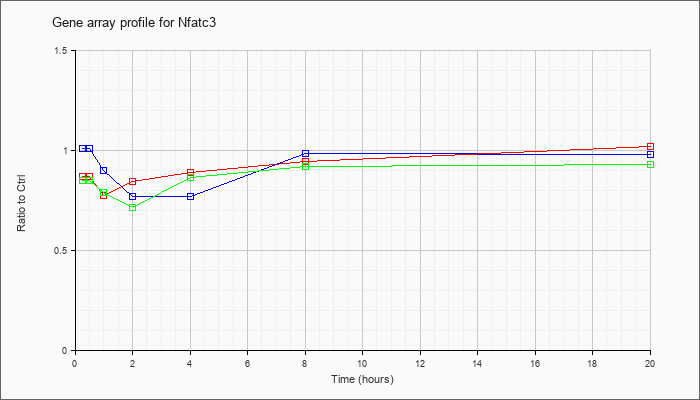 |
AK084848 |
13 days embryo lung cDNA, RIKEN full-length enriched library, clone:D430001G08 product:nuclear factor of activated T-cells, cytoplasmic 3, full insert sequence. [AK084848] |
KLA | 1.08 |
.96 |
.98 |
1.00 |
.98 |
1.03 |
.98 |
| ATP | 1.00 |
1.03 |
.93 |
.92 |
.92 |
1.11 |
.98 |
| KLA/ATP | 1.01 |
.87 |
1.10 |
.97 |
.96 |
.96 |
.99 |
|
| Nfatc3 |  |
NM_010901 |
nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 3 (Nfatc3), mRNA [NM_010901] |
KLA | .77 |
.68 |
.67 |
.77 |
.85 |
.90 |
1.04 |
| ATP | 1.01 |
.95 |
.89 |
.70 |
.70 |
.92 |
.98 |
| KLA/ATP | .77 |
.66 |
.64 |
.59 |
.82 |
.90 |
.90 |
|
| Nfatc4 | 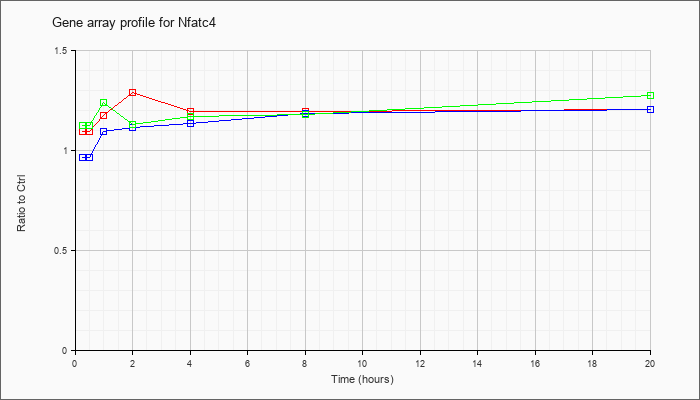 |
AK014164 |
13 days embryo head cDNA, RIKEN full-length enriched library, clone:3110041H08 product:nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 4, full insert sequence. [AK014164] |
KLA | .90 |
.92 |
.91 |
1.00 |
.99 |
.98 |
.95 |
| ATP | .90 |
1.03 |
.96 |
.98 |
.95 |
.97 |
1.00 |
| KLA/ATP | .95 |
.87 |
1.00 |
.98 |
.96 |
.96 |
.92 |
|
| Nfatc4 |  |
NM_023699 |
nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 4 (Nfatc4), mRNA [NM_023699] |
KLA | 1.28 |
1.38 |
1.44 |
1.58 |
1.40 |
1.41 |
1.46 |
| ATP | 1.03 |
1.06 |
1.23 |
1.24 |
1.32 |
1.40 |
1.41 |
| KLA/ATP | 1.30 |
1.38 |
1.47 |
1.28 |
1.38 |
1.40 |
1.63 |
|
| Nfkb1 | 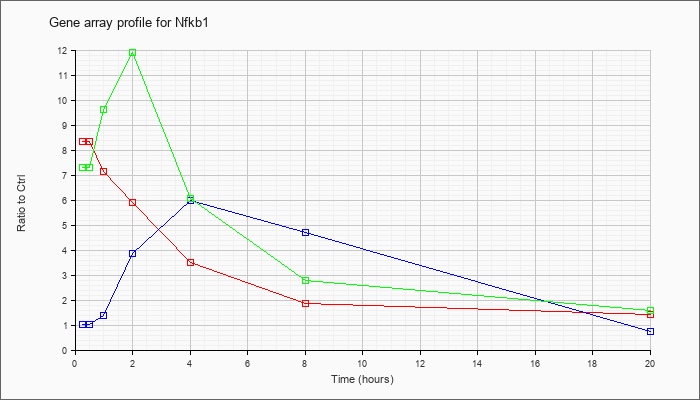 |
BC050841 |
nuclear factor of kappa light chain gene enhancer in B-cells 1, p105, mRNA (cDNA clone IMAGE:6392793), complete cds. [BC050841] |
KLA | 5.81 |
4.96 |
4.77 |
3.21 |
3.37 |
2.00 |
1.71 |
| ATP | 1.39 |
1.52 |
3.25 |
12.63 |
13.64 |
4.80 |
1.08 |
| KLA/ATP | 7.21 |
8.69 |
19.53 |
23.93 |
11.70 |
2.85 |
2.01 |
|
| Nfkb1 |  |
M57999 |
gb|Mouse transcription factor NF-kappa-B DNA binding subunit mRNA, complete cds. [M57999] |
KLA | 8.55 |
7.87 |
7.35 |
6.13 |
3.51 |
1.85 |
1.38 |
| ATP | .98 |
.96 |
1.21 |
3.06 |
5.29 |
4.67 |
.69 |
| KLA/ATP | 7.32 |
7.20 |
8.71 |
10.82 |
5.55 |
2.77 |
1.54 |
|
| Nfkb2 |  |
NM_019408 |
nuclear factor of kappa light polypeptide gene enhancer in B-cells 2, p49/p100 (Nfkb2), mRNA [NM_019408] |
KLA | 7.31 |
7.92 |
7.95 |
6.42 |
6.41 |
4.42 |
2.81 |
| ATP | .99 |
.98 |
.72 |
1.48 |
2.80 |
5.31 |
1.44 |
| KLA/ATP | 8.25 |
7.93 |
6.51 |
7.68 |
5.08 |
4.38 |
4.86 |
|
| Nfkbia | 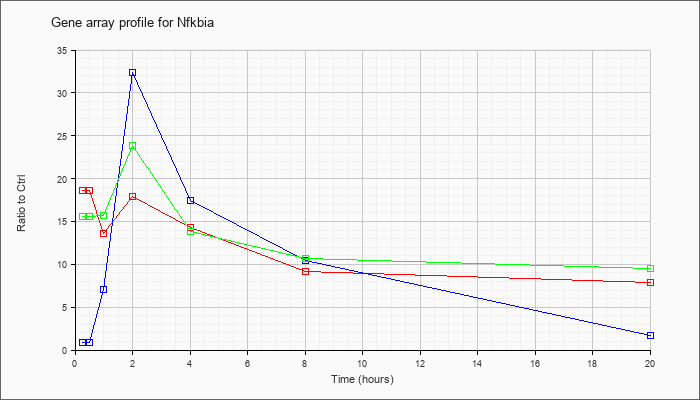 |
NM_010907 |
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha (Nfkbia), mRNA [NM_010907] |
KLA | 18.65 |
18.46 |
13.63 |
17.88 |
14.25 |
9.17 |
7.92 |
| ATP | .91 |
1.09 |
7.11 |
32.39 |
17.44 |
10.49 |
1.72 |
| KLA/ATP | 15.57 |
17.66 |
15.73 |
23.91 |
13.83 |
10.73 |
9.48 |
|
| Nfyb | 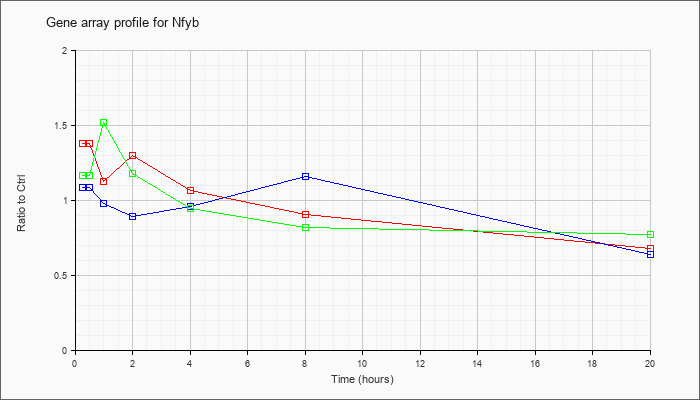 |
NM_010914 |
nuclear transcription factor-Y beta (Nfyb), mRNA [NM_010914] |
KLA | 1.38 |
1.22 |
1.12 |
1.30 |
1.07 |
.91 |
.68 |
| ATP | 1.09 |
1.01 |
.98 |
.89 |
.96 |
1.16 |
.64 |
| KLA/ATP | 1.17 |
1.21 |
1.52 |
1.18 |
.95 |
.82 |
.77 |
|
| Nras | 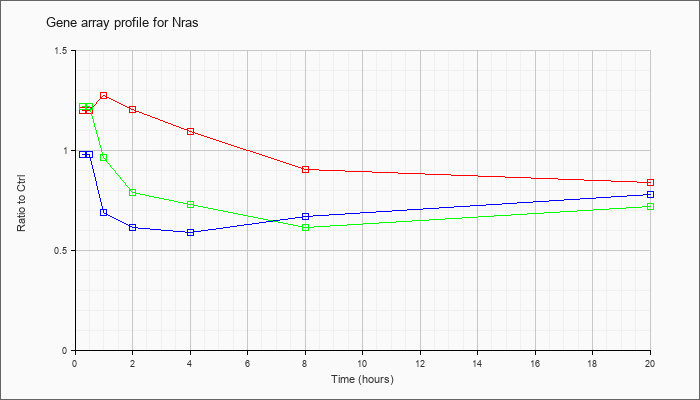 |
NM_010937 |
neuroblastoma ras oncogene (Nras), mRNA [NM_010937] |
KLA | 1.20 |
1.22 |
1.27 |
1.20 |
1.09 |
.91 |
.84 |
| ATP | .98 |
.91 |
.69 |
.61 |
.59 |
.67 |
.78 |
| KLA/ATP | 1.22 |
1.14 |
.97 |
.79 |
.73 |
.61 |
.72 |
|
| Nrp1 |  |
AK040447 |
0 day neonate thymus cDNA, RIKEN full-length enriched library, clone:A430096P05 product:neuropilin, full insert sequence. [AK040447] |
KLA | .33 |
.32 |
.34 |
.36 |
.34 |
.50 |
.45 |
| ATP | .92 |
.85 |
1.03 |
.87 |
.56 |
.82 |
.82 |
| KLA/ATP | .31 |
.33 |
.44 |
.37 |
.41 |
.56 |
.58 |
|
| Nrp1 |  |
NM_008737 |
neuropilin 1 (Nrp1), mRNA [NM_008737] |
KLA | .53 |
.52 |
.45 |
.39 |
.25 |
.25 |
.25 |
| ATP | 1.14 |
1.11 |
.84 |
.98 |
.64 |
.66 |
.66 |
| KLA/ATP | .64 |
.54 |
.39 |
.48 |
.36 |
.40 |
.36 |
|
| Pcaf | 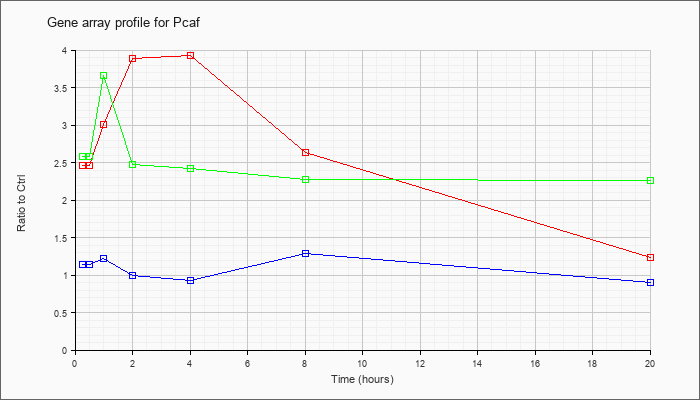 |
NM_020005 |
p300/CBP-associated factor (Pcaf), mRNA [NM_020005] |
KLA | 2.47 |
2.38 |
3.01 |
3.89 |
3.93 |
2.64 |
1.24 |
| ATP | 1.14 |
1.21 |
1.23 |
.99 |
.93 |
1.29 |
.91 |
| KLA/ATP | 2.58 |
2.82 |
3.67 |
2.48 |
2.42 |
2.28 |
2.26 |
|
| Pcna | 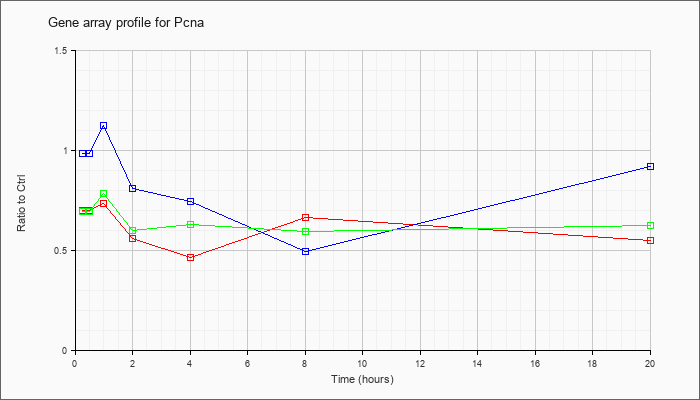 |
NM_011045 |
proliferating cell nuclear antigen (Pcna), mRNA [NM_011045] |
KLA | .70 |
.69 |
.73 |
.56 |
.46 |
.66 |
.55 |
| ATP | .98 |
.95 |
1.12 |
.81 |
.74 |
.49 |
.92 |
| KLA/ATP | .69 |
.66 |
.78 |
.60 |
.63 |
.59 |
.62 |
|
| Pdgfa | 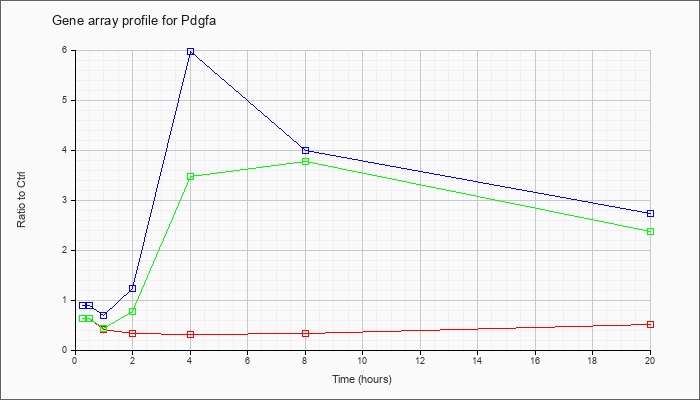 |
NM_008808 |
platelet derived growth factor, alpha (Pdgfa), mRNA [NM_008808] |
KLA | .63 |
.56 |
.42 |
.34 |
.31 |
.33 |
.52 |
| ATP | .89 |
.80 |
.69 |
1.23 |
5.97 |
3.99 |
2.73 |
| KLA/ATP | .64 |
.53 |
.43 |
.77 |
3.48 |
3.78 |
2.37 |
|
| Pdgfb | 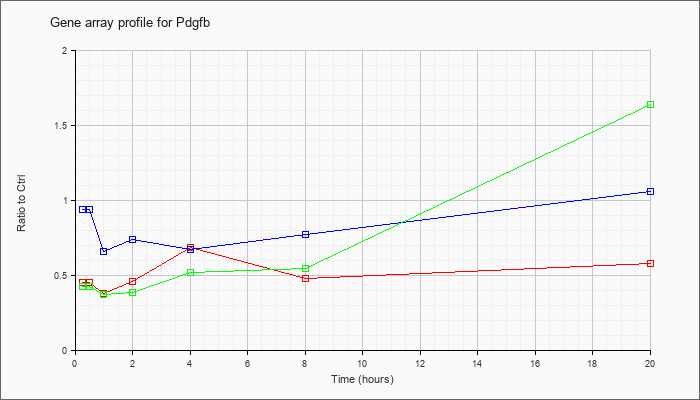 |
NM_011057 |
platelet derived growth factor, B polypeptide (Pdgfb), mRNA [NM_011057] |
KLA | .45 |
.43 |
.38 |
.46 |
.69 |
.48 |
.58 |
| ATP | .94 |
.95 |
.66 |
.74 |
.67 |
.77 |
1.06 |
| KLA/ATP | .43 |
.43 |
.37 |
.39 |
.52 |
.55 |
1.64 |
|
| Pdgfra | 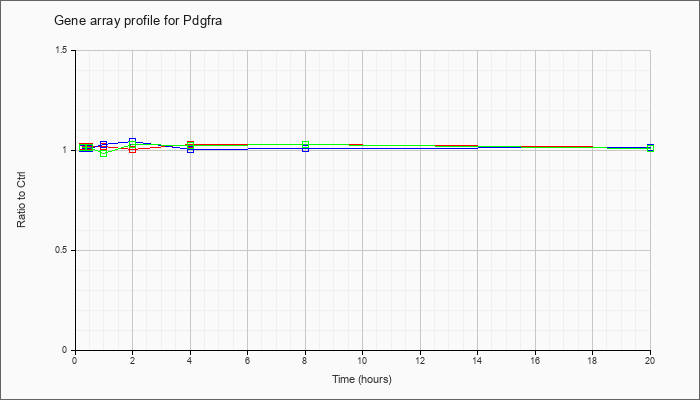 |
NM_011058 |
platelet derived growth factor receptor, alpha polypeptide (Pdgfra), transcript variant 1, mRNA [NM_011058] |
KLA | 1.02 |
1.05 |
1.02 |
1.01 |
1.03 |
1.03 |
1.01 |
| ATP | 1.01 |
1.02 |
1.03 |
1.04 |
1.00 |
1.01 |
1.01 |
| KLA/ATP | 1.01 |
1.01 |
.99 |
1.03 |
1.02 |
1.03 |
1.01 |
|
| Pdgfrb | 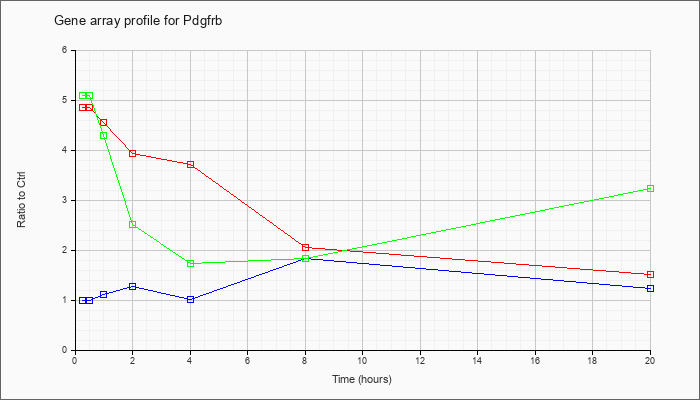 |
NM_008809 |
platelet derived growth factor receptor, beta polypeptide (Pdgfrb), mRNA [NM_008809] |
KLA | 4.85 |
5.49 |
4.56 |
3.94 |
3.71 |
2.05 |
1.51 |
| ATP | .99 |
1.00 |
1.11 |
1.27 |
1.02 |
1.83 |
1.23 |
| KLA/ATP | 5.09 |
5.86 |
4.30 |
2.51 |
1.74 |
1.84 |
3.23 |
|
| Pik3ca | 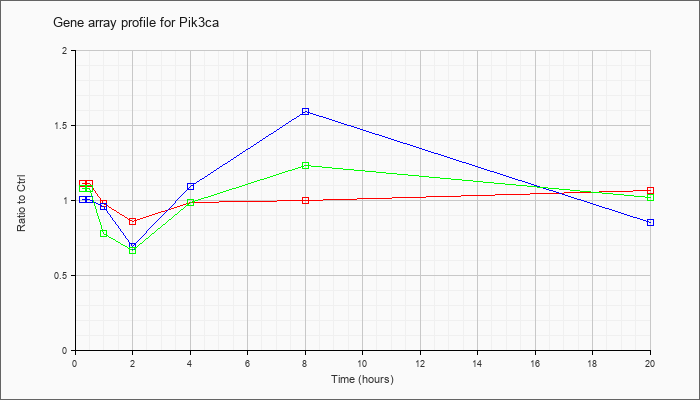 |
ENSMUST00000108243 |
ens|Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha isoform (EC 2.7.1.153) (PI3-kinase p110 subunit alpha) (PtdIns-3- kinase p110) (PI3K). [Source:Uniprot/SWISSPROT;Acc:P42337] [ENSMUST00000108243] |
KLA | 1.07 |
1.01 |
.91 |
.90 |
.98 |
.96 |
.86 |
| ATP | 1.03 |
.87 |
1.12 |
.63 |
1.22 |
1.86 |
.60 |
| KLA/ATP | 1.05 |
.82 |
.91 |
.61 |
1.17 |
1.26 |
.63 |
|
| Pik3ca |  |
NM_008839 |
phosphatidylinositol 3-kinase, catalytic, alpha polypeptide (Pik3ca), mRNA [NM_008839] |
KLA | 1.15 |
1.11 |
1.05 |
.82 |
.99 |
1.03 |
1.27 |
| ATP | .98 |
1.00 |
.79 |
.75 |
.96 |
1.32 |
1.10 |
| KLA/ATP | 1.10 |
.98 |
.64 |
.72 |
.80 |
1.20 |
1.41 |
|
| Pik3cb | 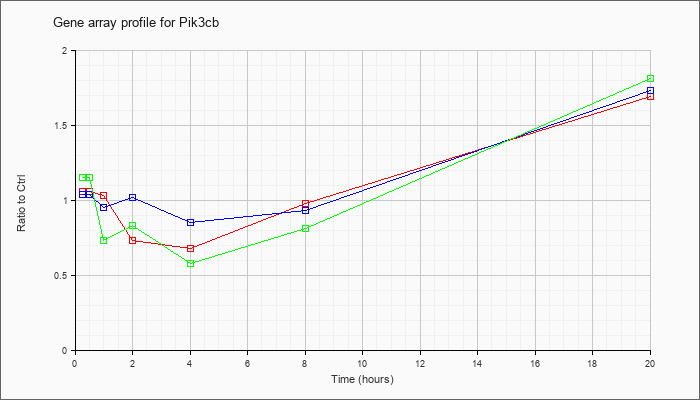 |
NM_029094 |
phosphatidylinositol 3-kinase, catalytic, beta polypeptide (Pik3cb), mRNA [NM_029094] |
KLA | 1.06 |
1.06 |
1.03 |
.73 |
.68 |
.98 |
1.69 |
| ATP | 1.04 |
1.21 |
.95 |
1.02 |
.85 |
.93 |
1.73 |
| KLA/ATP | 1.15 |
1.27 |
.73 |
.83 |
.58 |
.81 |
1.81 |
|
| Pik3cd | 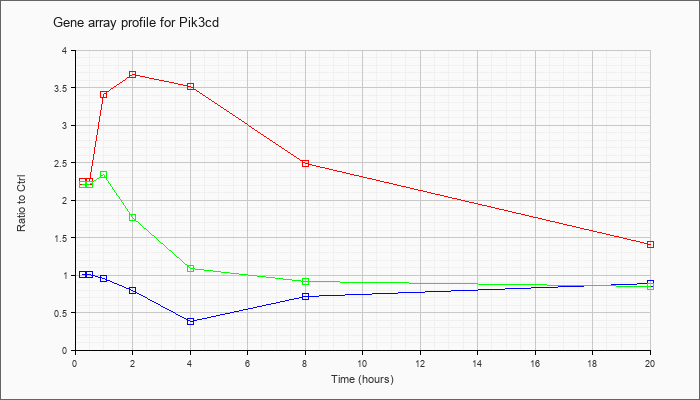 |
NM_008840 |
phosphatidylinositol 3-kinase catalytic delta polypeptide (Pik3cd), transcript variant 1, mRNA [NM_008840] |
KLA | 2.28 |
2.42 |
3.10 |
4.17 |
3.35 |
2.31 |
1.34 |
| ATP | 1.02 |
.91 |
.99 |
.63 |
.30 |
.67 |
.80 |
| KLA/ATP | 2.14 |
2.23 |
2.70 |
1.51 |
1.17 |
.85 |
.72 |
|
| Pik3cd |  |
U86587 |
phosphatidylinositol 3-kinase catalytic subunit p110 delta mRNA, complete cds. [U86587] |
KLA | 2.21 |
2.24 |
3.71 |
3.17 |
3.67 |
2.66 |
1.48 |
| ATP | 1.00 |
1.03 |
.93 |
.96 |
.46 |
.75 |
.97 |
| KLA/ATP | 2.27 |
2.32 |
1.97 |
2.02 |
1.01 |
.99 |
.98 |
|
| Pik3cg | 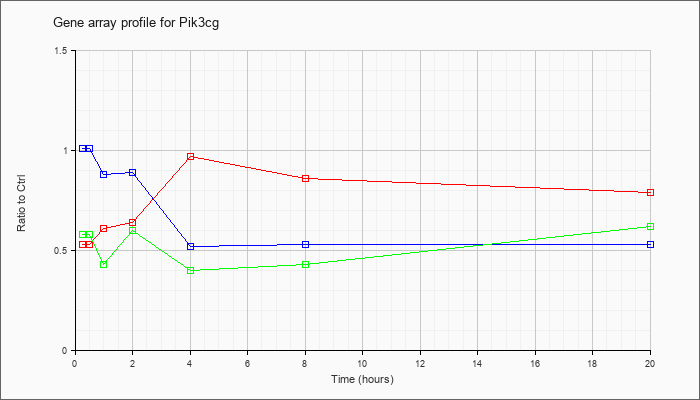 |
NM_020272 |
phosphoinositide-3-kinase, catalytic, gamma polypeptide (Pik3cg), mRNA [NM_020272] |
KLA | .53 |
.53 |
.61 |
.64 |
.97 |
.86 |
.79 |
| ATP | 1.01 |
1.13 |
.88 |
.89 |
.52 |
.53 |
.53 |
| KLA/ATP | .58 |
.53 |
.43 |
.60 |
.40 |
.43 |
.62 |
|
| Pik3r1 | 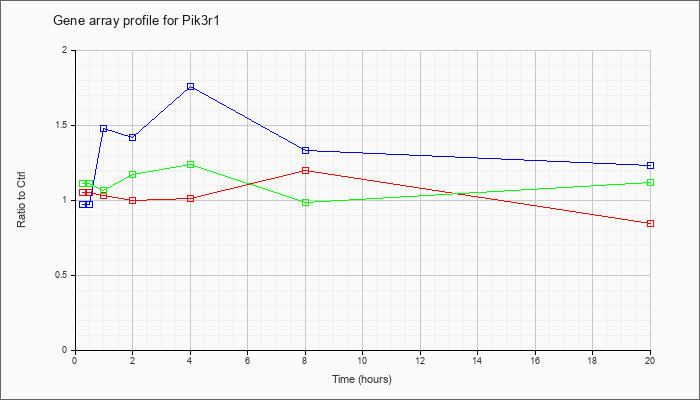 |
NM_001077495 |
phosphatidylinositol 3-kinase, regulatory subunit, polypeptide 1 (p85 alpha) (Pik3r1), transcript variant 2, mRNA [NM_001077495] |
KLA | 1.05 |
1.07 |
1.03 |
1.00 |
1.01 |
1.20 |
.85 |
| ATP | .97 |
1.02 |
1.48 |
1.42 |
1.76 |
1.33 |
1.23 |
| KLA/ATP | 1.11 |
1.12 |
1.06 |
1.17 |
1.24 |
.99 |
1.12 |
|
| Pik3r2 | 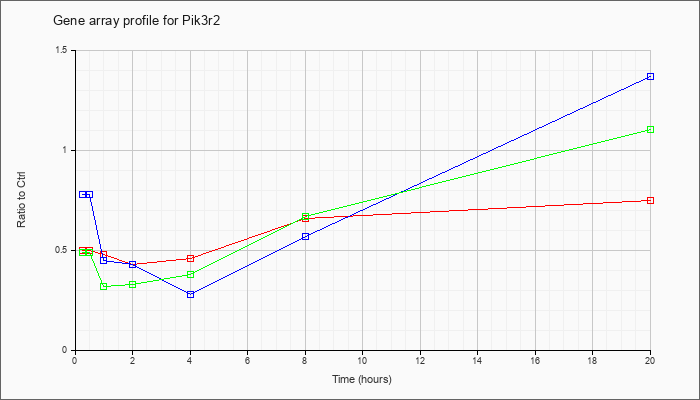 |
NM_008841 |
phosphatidylinositol 3-kinase, regulatory subunit, polypeptide 2 (p85 beta) (Pik3r2), mRNA [NM_008841] |
KLA | .50 |
.50 |
.48 |
.43 |
.46 |
.66 |
.75 |
| ATP | .78 |
.68 |
.45 |
.43 |
.28 |
.57 |
1.37 |
| KLA/ATP | .49 |
.38 |
.32 |
.33 |
.38 |
.67 |
1.10 |
|
| Pik3r3 | 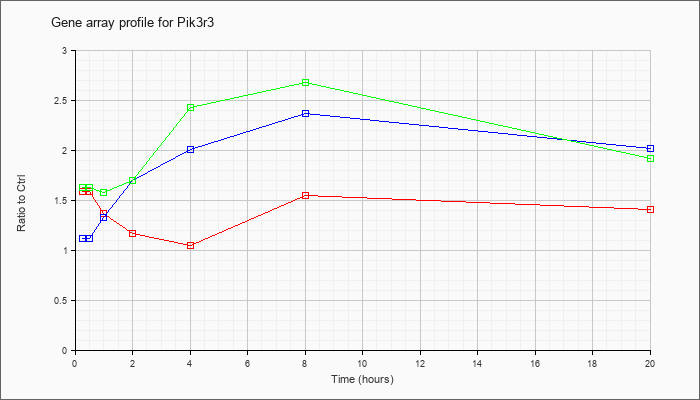 |
NM_181585 |
phosphatidylinositol 3 kinase, regulatory subunit, polypeptide 3 (p55) (Pik3r3), mRNA [NM_181585] |
KLA | 1.59 |
1.49 |
1.37 |
1.17 |
1.05 |
1.55 |
1.41 |
| ATP | 1.12 |
1.00 |
1.33 |
1.70 |
2.01 |
2.37 |
2.02 |
| KLA/ATP | 1.63 |
1.54 |
1.58 |
1.70 |
2.43 |
2.68 |
1.92 |
|
| Pik3r5 | 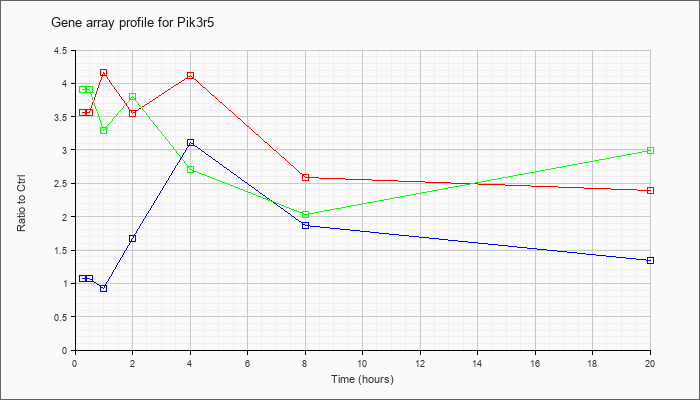 |
NM_177320 |
phosphoinositide-3-kinase, regulatory subunit 5, p101 (Pik3r5), mRNA [NM_177320] |
KLA | 3.57 |
3.58 |
4.16 |
3.54 |
4.11 |
2.59 |
2.39 |
| ATP | 1.08 |
1.08 |
.93 |
1.67 |
3.11 |
1.88 |
1.34 |
| KLA/ATP | 3.91 |
3.70 |
3.30 |
3.80 |
2.70 |
2.04 |
3.00 |
|
| Polb | 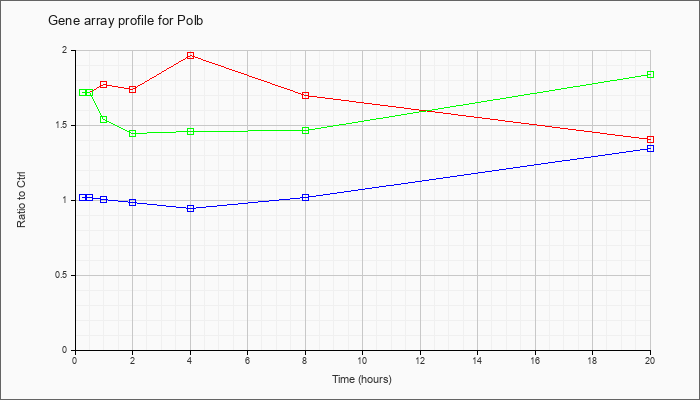 |
BC006681 |
polymerase (DNA directed), beta, mRNA (cDNA clone IMAGE:3581916), complete cds [BC006681] |
KLA | 1.62 |
1.31 |
1.52 |
1.37 |
1.57 |
1.35 |
1.17 |
| ATP | 1.04 |
1.03 |
.82 |
.63 |
.73 |
1.00 |
1.07 |
| KLA/ATP | 1.63 |
1.37 |
1.02 |
.72 |
.89 |
1.20 |
1.19 |
|
| Polb |  |
NM_011130 |
polymerase (DNA directed), beta (Polb), mRNA [NM_011130] |
KLA | 1.81 |
1.86 |
2.02 |
2.10 |
2.36 |
2.05 |
1.64 |
| ATP | .99 |
.99 |
1.19 |
1.34 |
1.16 |
1.04 |
1.62 |
| KLA/ATP | 1.80 |
1.83 |
2.05 |
2.17 |
2.03 |
1.73 |
2.49 |
|
| Pold1 | 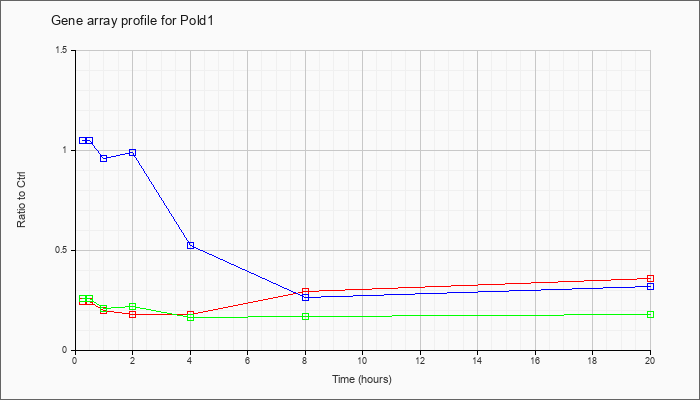 |
NM_011131 |
polymerase (DNA directed), delta 1, catalytic subunit (Pold1), mRNA [NM_011131] |
KLA | .24 |
.21 |
.20 |
.18 |
.18 |
.29 |
.36 |
| ATP | 1.05 |
1.05 |
.96 |
.99 |
.52 |
.26 |
.32 |
| KLA/ATP | .26 |
.22 |
.21 |
.22 |
.16 |
.17 |
.18 |
|
| Pold2 | 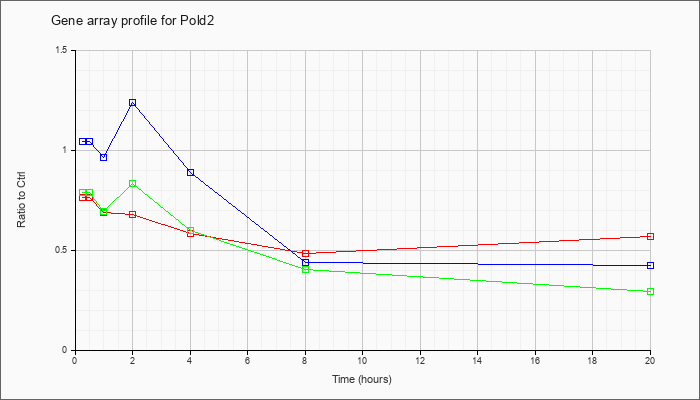 |
NM_008894 |
polymerase (DNA directed), delta 2, regulatory subunit (Pold2), mRNA [NM_008894] |
KLA | .77 |
.83 |
.69 |
.68 |
.59 |
.49 |
.57 |
| ATP | 1.05 |
1.09 |
.97 |
1.24 |
.89 |
.44 |
.43 |
| KLA/ATP | .79 |
.78 |
.70 |
.84 |
.60 |
.41 |
.30 |
|
| Pold3 | 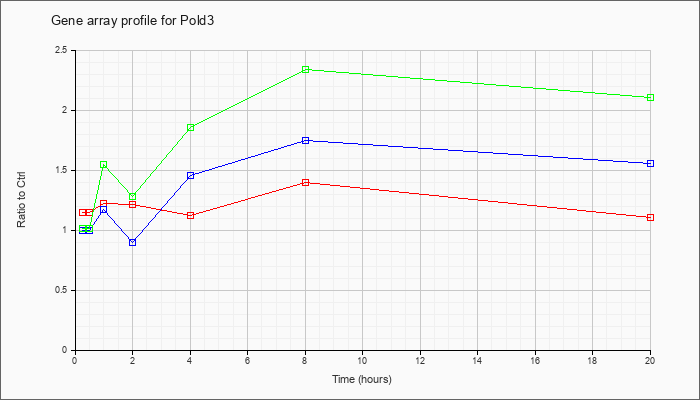 |
NM_133692 |
polymerase (DNA-directed), delta 3, accessory subunit (Pold3), mRNA [NM_133692] |
KLA | 1.14 |
1.07 |
1.22 |
1.22 |
1.12 |
1.40 |
1.10 |
| ATP | .99 |
.95 |
1.17 |
.89 |
1.46 |
1.74 |
1.55 |
| KLA/ATP | 1.01 |
1.08 |
1.54 |
1.28 |
1.86 |
2.34 |
2.11 |
|
| Pold4 | 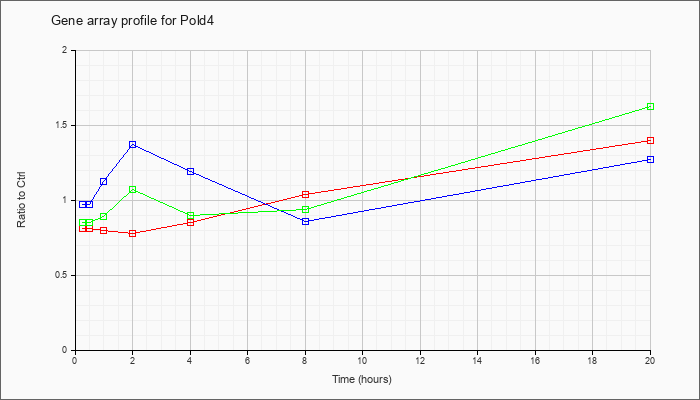 |
NM_027196 |
polymerase (DNA-directed), delta 4 (Pold4), mRNA [NM_027196] |
KLA | .81 |
.88 |
.80 |
.78 |
.85 |
1.04 |
1.40 |
| ATP | .97 |
1.07 |
1.12 |
1.37 |
1.19 |
.86 |
1.27 |
| KLA/ATP | .85 |
.90 |
.89 |
1.07 |
.90 |
.94 |
1.62 |
|
| Pole | 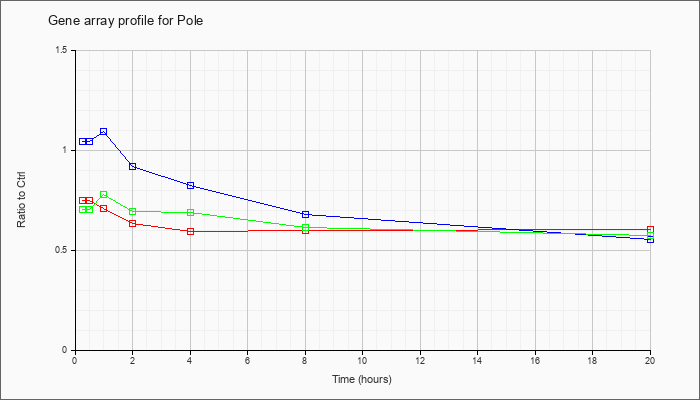 |
AK086490 |
15 days embryo head cDNA, RIKEN full-length enriched library, clone:D930032B21 product:DNA polymerase epsilon, full insert sequence. [AK086490] |
KLA | 1.00 |
1.00 |
1.04 |
.96 |
.94 |
.99 |
1.04 |
| ATP | .97 |
.95 |
.91 |
.95 |
.97 |
1.06 |
.95 |
| KLA/ATP | .96 |
.93 |
1.00 |
1.01 |
1.08 |
1.02 |
1.01 |
|
| Pole |  |
NM_011132 |
polymerase (DNA directed), epsilon (Pole), mRNA [NM_011132] |
KLA | .50 |
.44 |
.38 |
.31 |
.25 |
.21 |
.17 |
| ATP | 1.12 |
1.03 |
1.28 |
.89 |
.68 |
.30 |
.16 |
| KLA/ATP | .45 |
.44 |
.56 |
.38 |
.30 |
.21 |
.14 |
|
| Pole2 | 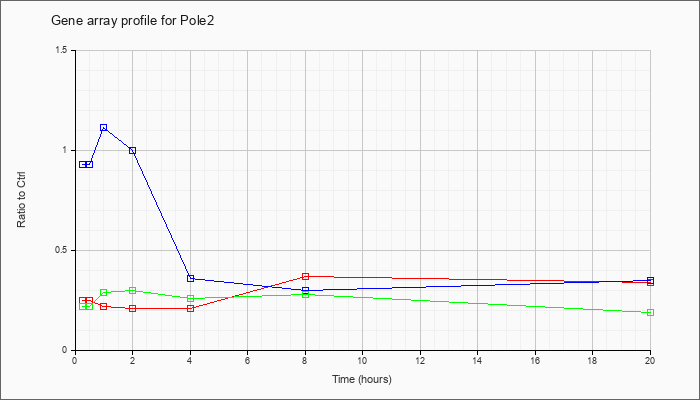 |
NM_011133 |
polymerase (DNA directed), epsilon 2 (p59 subunit) (Pole2), mRNA [NM_011133] |
KLA | .25 |
.24 |
.22 |
.21 |
.21 |
.37 |
.34 |
| ATP | .93 |
.98 |
1.11 |
1.00 |
.36 |
.30 |
.35 |
| KLA/ATP | .22 |
.22 |
.29 |
.30 |
.26 |
.28 |
.19 |
|
| Pole3 | 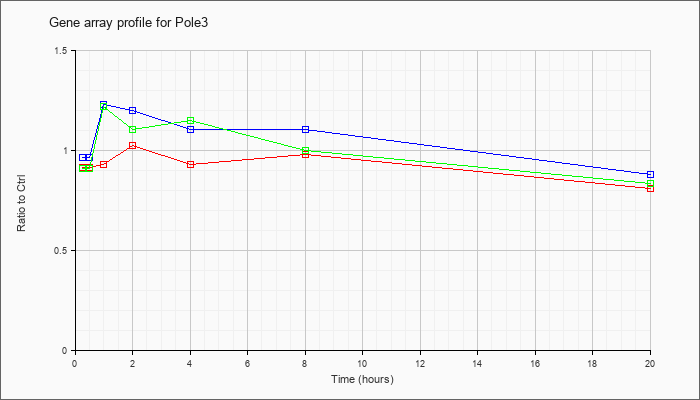 |
BC034004 |
polymerase (DNA directed), epsilon 3 (p17 subunit), mRNA (cDNA clone IMAGE:5117478) [BC034004] |
KLA | .69 |
.72 |
.63 |
.69 |
.68 |
.81 |
.61 |
| ATP | .91 |
.86 |
1.29 |
.96 |
.81 |
.94 |
.51 |
| KLA/ATP | .62 |
.54 |
.85 |
.64 |
.70 |
.75 |
.44 |
|
| Pole3 |  |
NM_021498 |
polymerase (DNA directed), epsilon 3 (p17 subunit) (Pole3), mRNA [NM_021498] |
KLA | 1.14 |
1.14 |
1.23 |
1.36 |
1.18 |
1.15 |
1.01 |
| ATP | 1.02 |
.98 |
1.17 |
1.44 |
1.40 |
1.27 |
1.25 |
| KLA/ATP | 1.20 |
1.24 |
1.58 |
1.57 |
1.60 |
1.25 |
1.23 |
|
| Pole4 | 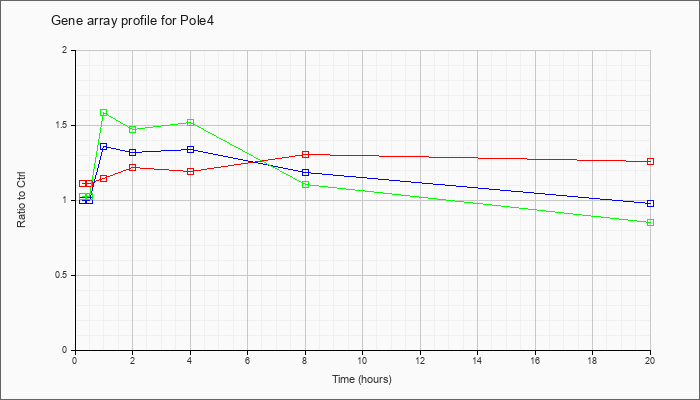 |
NM_025882 |
polymerase (DNA-directed), epsilon 4 (p12 subunit) (Pole4), mRNA [NM_025882] |
KLA | 1.11 |
1.15 |
1.15 |
1.22 |
1.19 |
1.31 |
1.26 |
| ATP | 1.00 |
.99 |
1.36 |
1.32 |
1.34 |
1.19 |
.98 |
| KLA/ATP | 1.02 |
1.15 |
1.58 |
1.47 |
1.52 |
1.10 |
.85 |
|
| Ppp3ca | 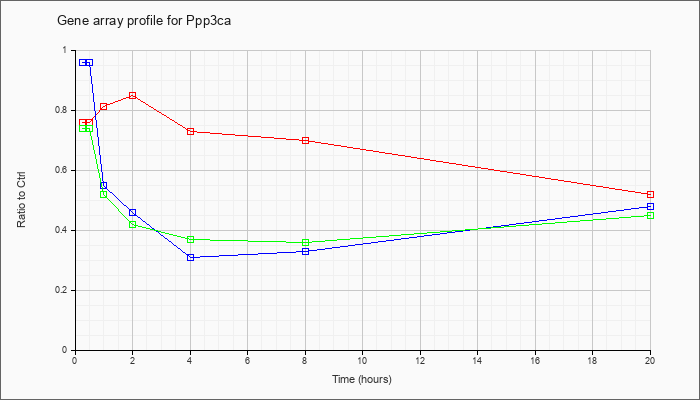 |
NM_008913 |
protein phosphatase 3, catalytic subunit, alpha isoform (Ppp3ca), mRNA [NM_008913] |
KLA | .76 |
.85 |
.81 |
.85 |
.73 |
.70 |
.52 |
| ATP | .96 |
.78 |
.55 |
.46 |
.31 |
.33 |
.48 |
| KLA/ATP | .74 |
.64 |
.52 |
.42 |
.37 |
.36 |
.45 |
|
| Ppp3cb | 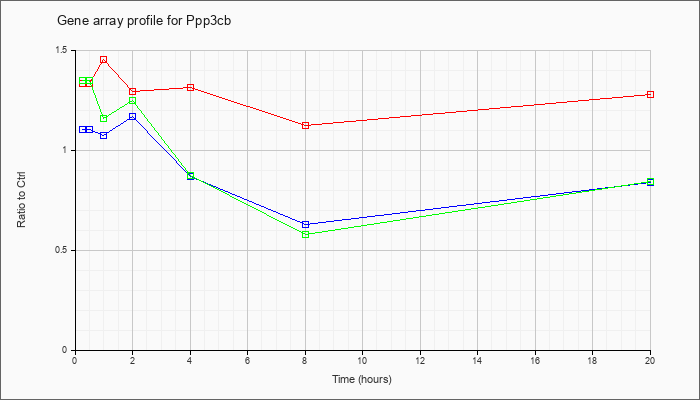 |
AK135203 |
12 days embryo male wolffian duct includes surrounding region cDNA, RIKEN full-length enriched library, clone:6720421K10 product:protein phosphatase 3, catalytic subunit, beta isoform, full insert sequence. [AK135203] |
KLA | 1.74 |
1.57 |
1.91 |
1.59 |
1.47 |
1.26 |
1.40 |
| ATP | 1.17 |
1.03 |
1.30 |
1.00 |
.81 |
.62 |
1.37 |
| KLA/ATP | 1.68 |
1.70 |
1.91 |
1.37 |
1.09 |
.47 |
1.13 |
|
| Ppp3cb |  |
AK171904 |
activated spleen cDNA, RIKEN full-length enriched library, clone:F830018L06 product:Protein phosphatase 3 catalytic subunit beta3 (EC 3.1.3.16) (Serine [AK171904] |
KLA | 1.29 |
1.29 |
1.21 |
1.15 |
1.22 |
1.13 |
1.28 |
| ATP | .90 |
.86 |
.80 |
.98 |
.94 |
.73 |
.92 |
| KLA/ATP | 1.06 |
1.03 |
.77 |
1.04 |
.92 |
.68 |
.90 |
|
| Ppp3cb |  |
NM_008914 |
protein phosphatase 3, catalytic subunit, beta isoform (Ppp3cb), mRNA [NM_008914] |
KLA | 1.21 |
1.29 |
1.38 |
1.24 |
1.29 |
1.08 |
1.23 |
| ATP | 1.15 |
1.31 |
1.09 |
1.29 |
.86 |
.60 |
.64 |
| KLA/ATP | 1.33 |
1.40 |
1.04 |
1.28 |
.79 |
.58 |
.73 |
|
| Ppp3cc | 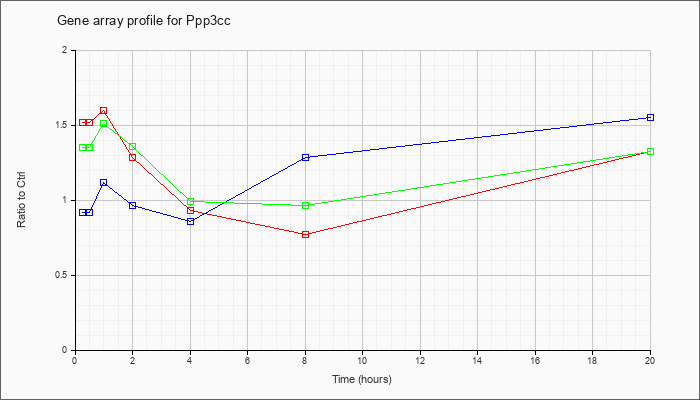 |
AK037563 |
16 days neonate thymus cDNA, RIKEN full-length enriched library, clone:A130026K22 product:unclassifiable, full insert sequence [AK037563] |
KLA | 1.27 |
1.18 |
1.21 |
1.15 |
1.01 |
.92 |
1.36 |
| ATP | .96 |
1.09 |
1.10 |
1.03 |
.85 |
1.06 |
1.47 |
| KLA/ATP | 1.17 |
1.24 |
1.07 |
1.13 |
.91 |
.88 |
1.36 |
|
| Ppp3cc |  |
NM_008915 |
protein phosphatase 3, catalytic subunit, gamma isoform (Ppp3cc), mRNA [NM_008915] |
KLA | 1.64 |
1.81 |
1.79 |
1.35 |
.90 |
.70 |
1.31 |
| ATP | .90 |
.90 |
1.13 |
.93 |
.86 |
1.40 |
1.60 |
| KLA/ATP | 1.45 |
1.62 |
1.74 |
1.47 |
1.04 |
1.01 |
1.31 |
|
| Ppp3r1 | 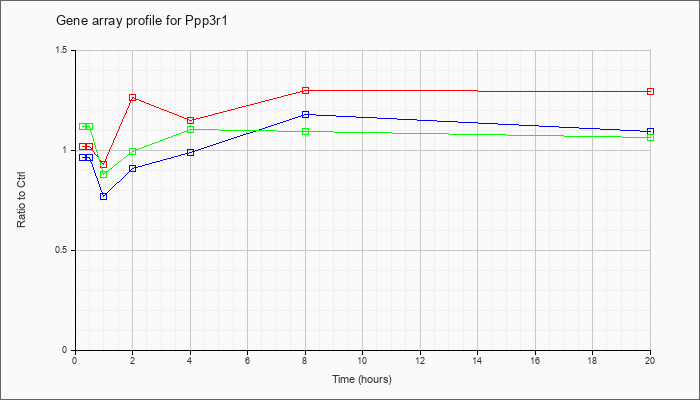 |
NM_024459 |
protein phosphatase 3, regulatory subunit B, alpha isoform (calcineurin B, type I) (Ppp3r1), mRNA [NM_024459] |
KLA | 1.02 |
1.17 |
.93 |
1.27 |
1.15 |
1.30 |
1.30 |
| ATP | .97 |
.95 |
.77 |
.91 |
.99 |
1.18 |
1.10 |
| KLA/ATP | 1.12 |
1.04 |
.88 |
1.00 |
1.10 |
1.10 |
1.07 |
|
| Ppp3r2 | 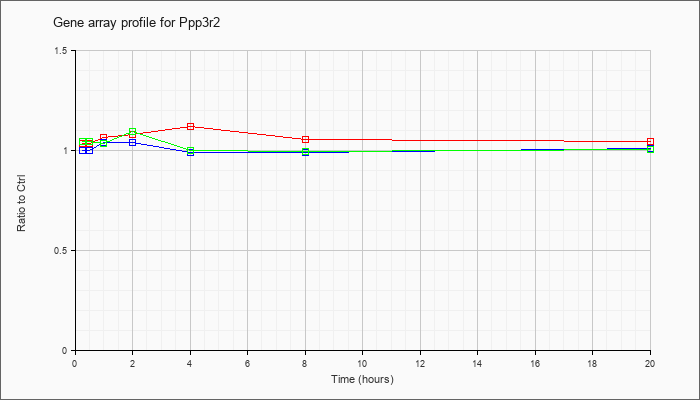 |
NM_001004025 |
protein phosphatase 3, regulatory subunit B, alpha isoform (calcineurin B, type II) (Ppp3r2), mRNA [NM_001004025] |
KLA | 1.03 |
1.05 |
1.06 |
1.08 |
1.12 |
1.05 |
1.04 |
| ATP | 1.00 |
1.02 |
1.04 |
1.04 |
.99 |
.99 |
1.01 |
| KLA/ATP | 1.04 |
1.12 |
1.03 |
1.09 |
1.00 |
.99 |
1.00 |
|
| Prkaca | 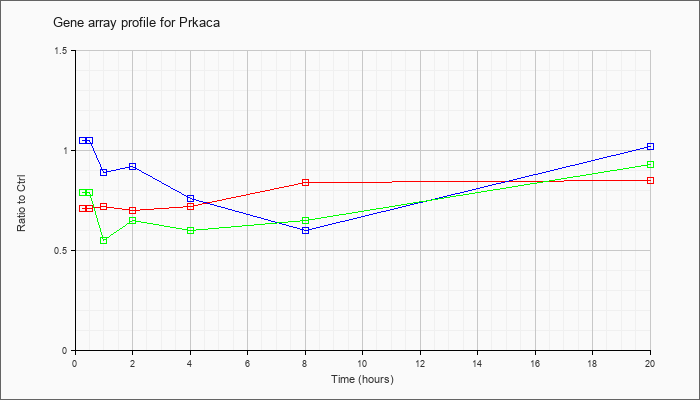 |
NM_008854 |
protein kinase, cAMP dependent, catalytic, alpha (Prkaca), mRNA [NM_008854] |
KLA | .71 |
.74 |
.72 |
.70 |
.72 |
.84 |
.85 |
| ATP | 1.05 |
.98 |
.89 |
.92 |
.76 |
.60 |
1.02 |
| KLA/ATP | .79 |
.74 |
.55 |
.65 |
.60 |
.65 |
.93 |
|
| Prkacb | 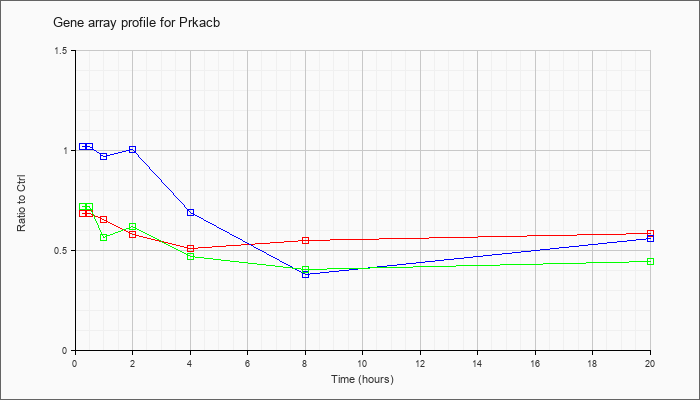 |
NM_011100 |
protein kinase, cAMP dependent, catalytic, beta (Prkacb), mRNA [NM_011100] |
KLA | .69 |
.70 |
.66 |
.58 |
.51 |
.55 |
.59 |
| ATP | 1.02 |
1.19 |
.97 |
1.01 |
.69 |
.38 |
.56 |
| KLA/ATP | .72 |
.73 |
.57 |
.62 |
.47 |
.41 |
.45 |
|
| Prkx | 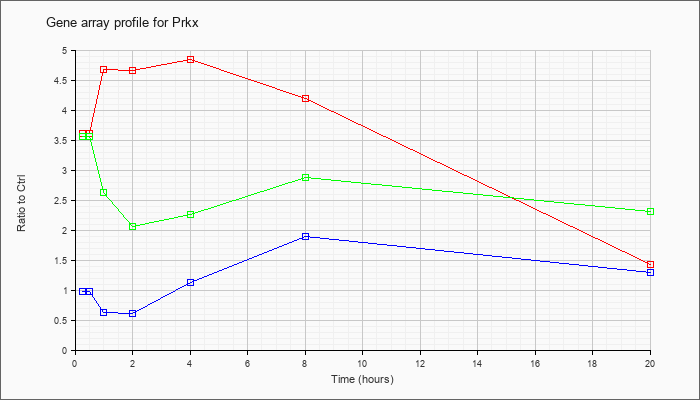 |
NM_016979 |
protein kinase, X-linked (Prkx), mRNA [NM_016979] |
KLA | 3.61 |
3.79 |
4.67 |
4.66 |
4.85 |
4.19 |
1.43 |
| ATP | .98 |
.90 |
.63 |
.61 |
1.12 |
1.90 |
1.29 |
| KLA/ATP | 3.56 |
3.36 |
2.62 |
2.06 |
2.26 |
2.87 |
2.31 |
|
| Pttg1 | 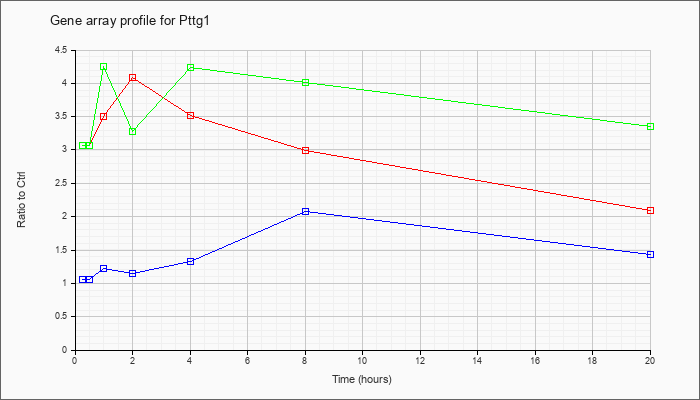 |
NM_013917 |
pituitary tumor-transforming 1 (Pttg1), mRNA [NM_013917] |
KLA | 3.06 |
2.73 |
3.51 |
4.09 |
3.52 |
2.99 |
2.09 |
| ATP | 1.06 |
.94 |
1.23 |
1.15 |
1.33 |
2.08 |
1.43 |
| KLA/ATP | 3.07 |
2.93 |
4.25 |
3.27 |
4.24 |
4.01 |
3.35 |
|
| Ran |  |
NM_009391 |
RAN, member RAS oncogene family (Ran), mRNA [NM_009391] |
KLA | .96 |
1.03 |
.96 |
.88 |
.78 |
.61 |
.75 |
| ATP | .96 |
.96 |
1.04 |
.87 |
1.10 |
1.18 |
.64 |
| KLA/ATP | .96 |
.97 |
.99 |
.84 |
1.03 |
.91 |
.61 |
|
| Ranbp1 | 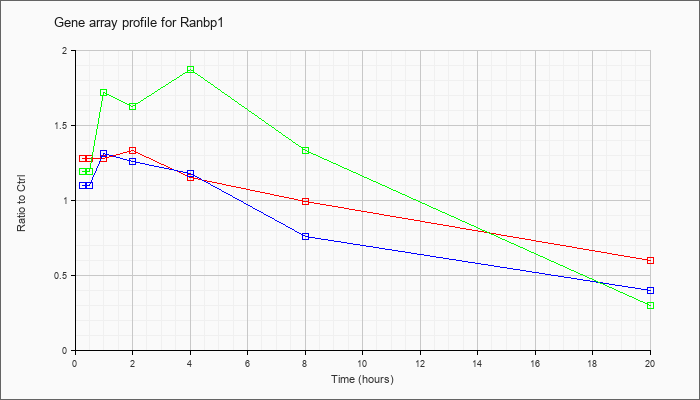 |
X56045 |
gb|Mouse mRNA (clone lambda-19) for hypothetical protein A. [X56045] |
KLA | 1.28 |
1.17 |
1.28 |
1.33 |
1.15 |
.99 |
.60 |
| ATP | 1.10 |
1.02 |
1.31 |
1.26 |
1.18 |
.76 |
.40 |
| KLA/ATP | 1.19 |
1.29 |
1.72 |
1.62 |
1.87 |
1.33 |
.30 |
|
| Ranbp3 | 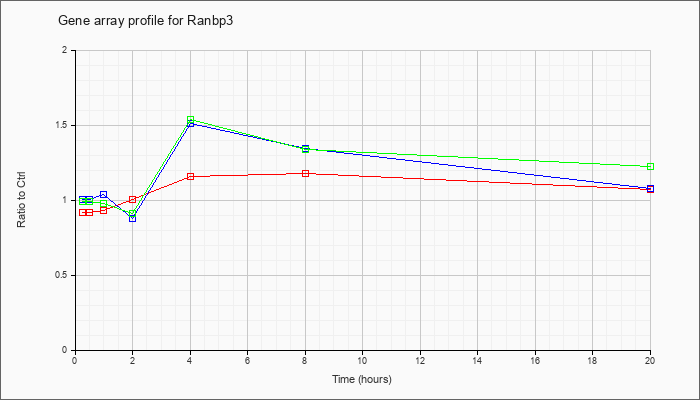 |
AK154201 |
NOD-derived CD11c +ve dendritic cells cDNA, RIKEN full-length enriched library, clone:F630008P20 product:unclassifiable, full insert sequence [AK154201] |
KLA | .86 |
.86 |
.87 |
.98 |
1.20 |
1.11 |
1.00 |
| ATP | 1.10 |
1.32 |
1.02 |
.77 |
1.84 |
1.33 |
1.04 |
| KLA/ATP | .99 |
1.04 |
.87 |
.82 |
1.80 |
1.19 |
1.06 |
|
| Ranbp3 |  |
NM_027933 |
RAN binding protein 3 (Ranbp3), mRNA [NM_027933] |
KLA | .97 |
1.01 |
.99 |
1.03 |
1.11 |
1.25 |
1.14 |
| ATP | .91 |
.98 |
1.05 |
.99 |
1.18 |
1.36 |
1.12 |
| KLA/ATP | .99 |
1.01 |
1.08 |
1.00 |
1.27 |
1.48 |
1.39 |
|
| Rasl2-9 |  |
NM_009028 |
RAS-like, family 2, locus 9 (Rasl2-9), mRNA [NM_009028] |
KLA | .94 |
1.01 |
1.05 |
.89 |
.81 |
.62 |
.70 |
| ATP | 1.05 |
1.05 |
1.13 |
.99 |
1.25 |
1.24 |
.68 |
| KLA/ATP | 1.00 |
.98 |
1.12 |
.94 |
1.10 |
.90 |
.67 |
|
| Rb1 | 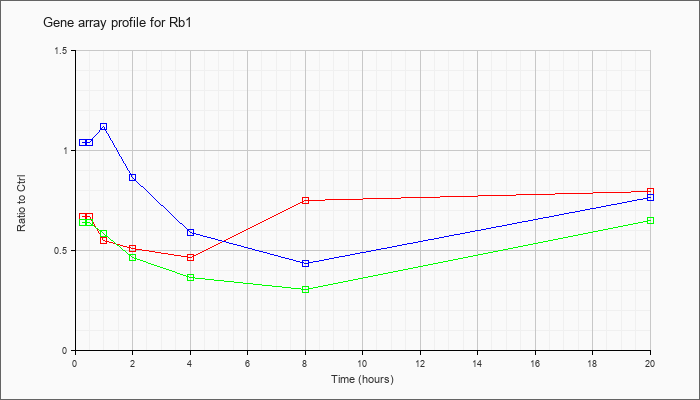 |
NM_009029 |
retinoblastoma 1 (Rb1), mRNA [NM_009029] |
KLA | .67 |
.60 |
.55 |
.51 |
.46 |
.75 |
.79 |
| ATP | 1.04 |
1.01 |
1.12 |
.86 |
.59 |
.43 |
.76 |
| KLA/ATP | .64 |
.59 |
.58 |
.46 |
.36 |
.30 |
.65 |
|
| Rela | 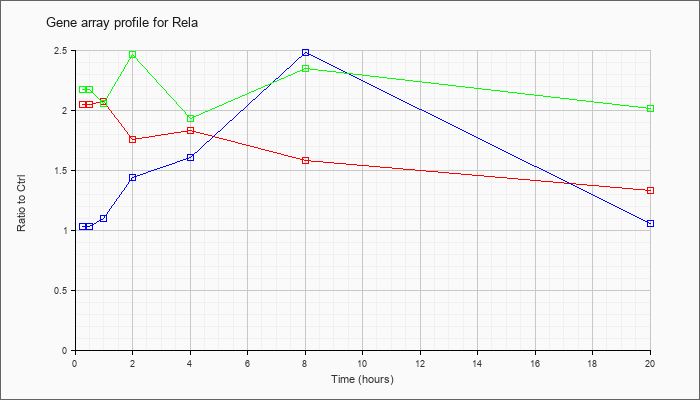 |
NM_009045 |
v-rel reticuloendotheliosis viral oncogene homolog A (avian) (Rela), mRNA [NM_009045] |
KLA | 2.05 |
2.05 |
2.07 |
1.75 |
1.83 |
1.58 |
1.33 |
| ATP | 1.03 |
1.08 |
1.10 |
1.44 |
1.61 |
2.48 |
1.05 |
| KLA/ATP | 2.17 |
2.27 |
2.05 |
2.47 |
1.93 |
2.35 |
2.02 |
|
| Relb |  |
NM_009046 |
avian reticuloendotheliosis viral (v-rel) oncogene related B (Relb), mRNA [NM_009046] |
KLA | 4.39 |
4.46 |
3.54 |
3.47 |
4.36 |
3.18 |
2.72 |
| ATP | 1.01 |
1.07 |
1.23 |
2.66 |
3.23 |
3.50 |
1.24 |
| KLA/ATP | 4.43 |
4.52 |
3.56 |
4.10 |
2.85 |
2.92 |
3.30 |
|
| Rras | 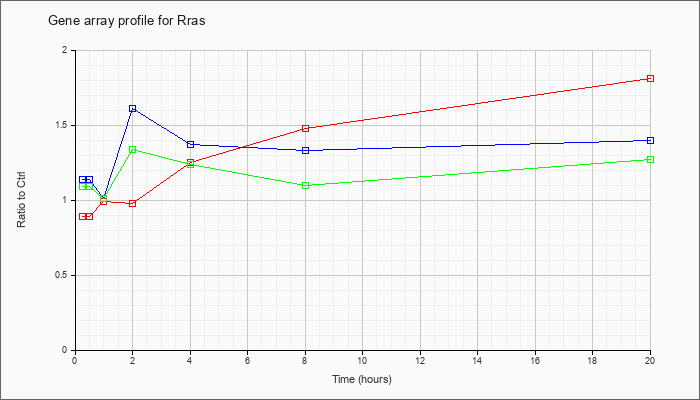 |
NM_009101 |
Harvey rat sarcoma oncogene, subgroup R (Rras), mRNA [NM_009101] |
KLA | .89 |
.89 |
.99 |
.98 |
1.25 |
1.48 |
1.81 |
| ATP | 1.14 |
1.17 |
1.01 |
1.61 |
1.37 |
1.33 |
1.40 |
| KLA/ATP | 1.09 |
1.05 |
1.01 |
1.34 |
1.24 |
1.10 |
1.27 |
|
| Rras2 | 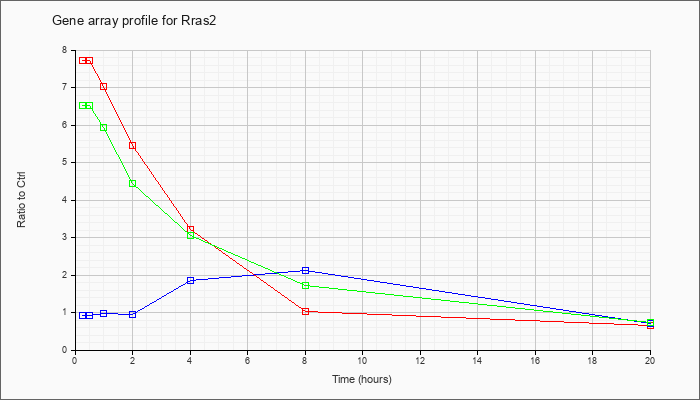 |
NM_025846 |
related RAS viral (r-ras) oncogene homolog 2 (Rras2), mRNA [NM_025846] |
KLA | 7.72 |
7.35 |
7.03 |
5.47 |
3.21 |
1.03 |
.65 |
| ATP | .92 |
.86 |
.99 |
.95 |
1.86 |
2.13 |
.72 |
| KLA/ATP | 6.51 |
6.50 |
5.94 |
4.45 |
3.05 |
1.72 |
.75 |
|
| Sfpi1 | 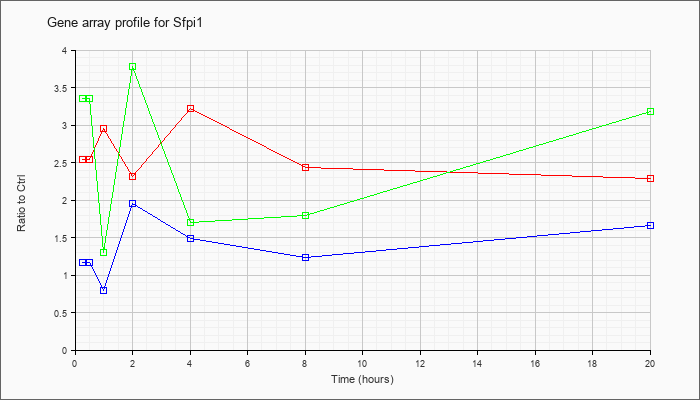 |
NM_011355 |
SFFV proviral integration 1 (Sfpi1), mRNA [NM_011355] |
KLA | 2.54 |
2.77 |
2.96 |
2.31 |
3.22 |
2.44 |
2.29 |
| ATP | 1.17 |
1.55 |
.79 |
1.96 |
1.49 |
1.23 |
1.66 |
| KLA/ATP | 3.35 |
3.40 |
1.30 |
3.78 |
1.70 |
1.80 |
3.18 |
|
Slc25a31
|  |
NM_178386 |
solute carrier family 25 (mitochondrial carrier; adenine nucleotide translocator), member 31 (Slc25a31), nuclear gene encoding mitochondrial protein, mRNA [NM_178386] |
KLA | .99 |
.96 |
.88 |
.97 |
.96 |
.91 |
.75 |
| ATP | 1.09 |
.96 |
.96 |
1.01 |
.92 |
.90 |
.77 |
| KLA/ATP | 1.02 |
.96 |
.89 |
.90 |
.96 |
.93 |
.74 |
|
| Slc25a4 | 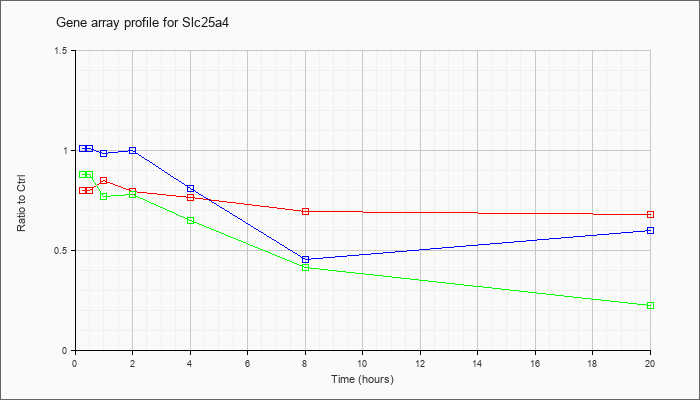 |
NM_007450 |
solute carrier family 25 (mitochondrial carrier, adenine nucleotide translocator), member 4 (Slc25a4), nuclear gene encoding mitochondrial protein, mRNA [NM_007450] |
KLA | .80 |
.84 |
.85 |
.80 |
.77 |
.70 |
.68 |
| ATP | 1.01 |
1.04 |
.99 |
1.00 |
.81 |
.46 |
.60 |
| KLA/ATP | .88 |
.85 |
.77 |
.78 |
.65 |
.42 |
.23 |
|
| Slc25a5 | 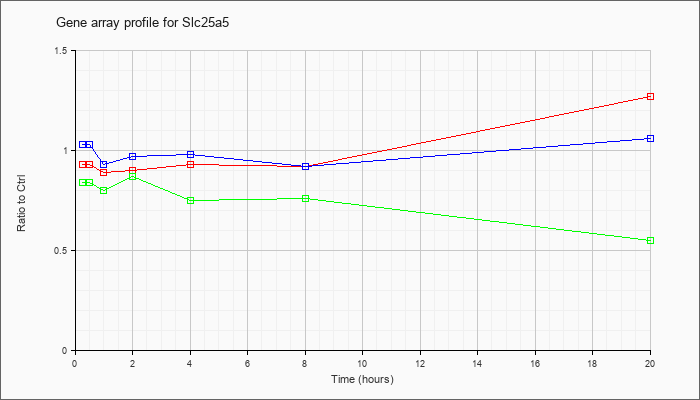 |
NM_007451 |
solute carrier family 25 (mitochondrial carrier, adenine nucleotide translocator), member 5 (Slc25a5), nuclear gene encoding mitochondrial protein, mRNA [NM_007451] |
KLA | .93 |
.93 |
.89 |
.90 |
.93 |
.92 |
1.27 |
| ATP | 1.03 |
1.08 |
.93 |
.97 |
.98 |
.92 |
1.06 |
| KLA/ATP | .84 |
.91 |
.80 |
.87 |
.75 |
.76 |
.55 |
|
| Slc2a1 | 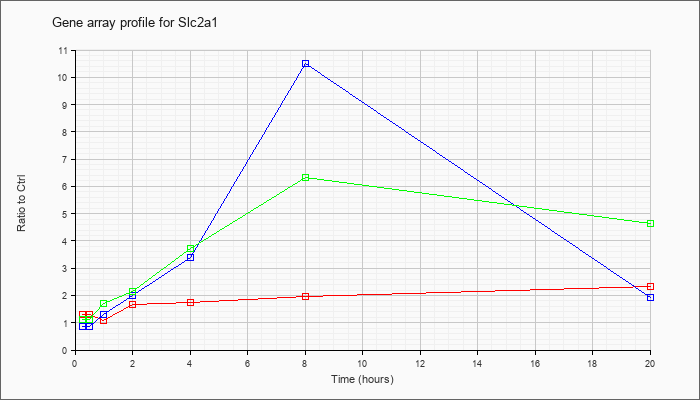 |
NM_011400 |
solute carrier family 2 (facilitated glucose transporter), member 1 (Slc2a1), mRNA [NM_011400] |
KLA | 1.31 |
1.46 |
1.10 |
1.68 |
1.75 |
1.95 |
2.34 |
| ATP | .88 |
.90 |
1.32 |
1.99 |
3.40 |
10.49 |
1.93 |
| KLA/ATP | 1.13 |
1.25 |
1.71 |
2.14 |
3.71 |
6.31 |
4.63 |
|
| Smad2 | 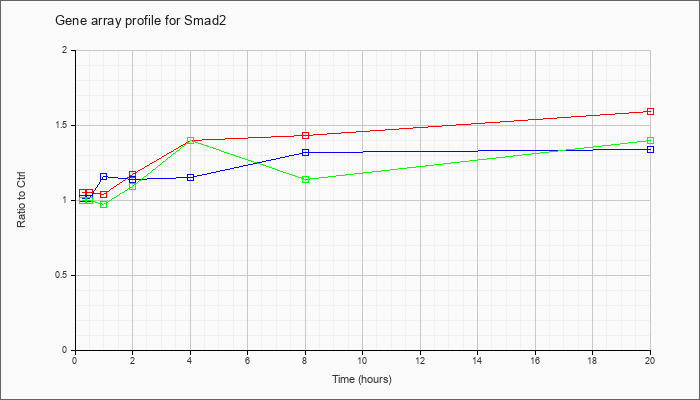 |
NM_010754 |
MAD homolog 2 (Drosophila) (Smad2), mRNA [NM_010754] |
KLA | 1.05 |
1.08 |
1.04 |
1.17 |
1.40 |
1.43 |
1.59 |
| ATP | 1.01 |
1.02 |
1.16 |
1.14 |
1.15 |
1.32 |
1.34 |
| KLA/ATP | 1.00 |
.99 |
.97 |
1.09 |
1.40 |
1.14 |
1.40 |
|
| Smad3 |  |
NM_016769 |
MAD homolog 3 (Drosophila) (Smad3), mRNA [NM_016769] |
KLA | .64 |
.60 |
.57 |
.64 |
.70 |
1.25 |
1.52 |
| ATP | 1.02 |
1.02 |
.84 |
1.21 |
2.42 |
.90 |
1.19 |
| KLA/ATP | .64 |
.60 |
.65 |
.66 |
1.29 |
1.26 |
2.26 |
|
| Smad4 | 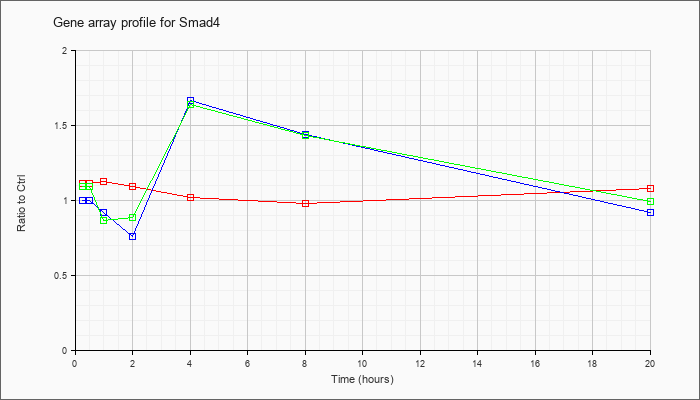 |
NM_008540 |
MAD homolog 4 (Drosophila) (Smad4), mRNA [NM_008540] |
KLA | 1.11 |
1.11 |
1.13 |
1.09 |
1.02 |
.98 |
1.08 |
| ATP | 1.00 |
1.04 |
.92 |
.76 |
1.67 |
1.44 |
.92 |
| KLA/ATP | 1.09 |
1.01 |
.87 |
.89 |
1.64 |
1.43 |
.99 |
|
| Srf | 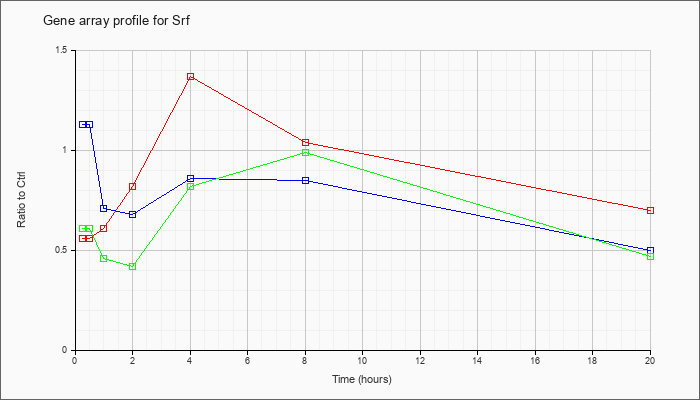 |
NM_020493 |
serum response factor (Srf), mRNA [NM_020493] |
KLA | .56 |
.53 |
.61 |
.82 |
1.37 |
1.04 |
.70 |
| ATP | 1.13 |
.88 |
.71 |
.68 |
.86 |
.85 |
.50 |
| KLA/ATP | .61 |
.51 |
.46 |
.42 |
.82 |
.99 |
.47 |
|
| Stat5a | 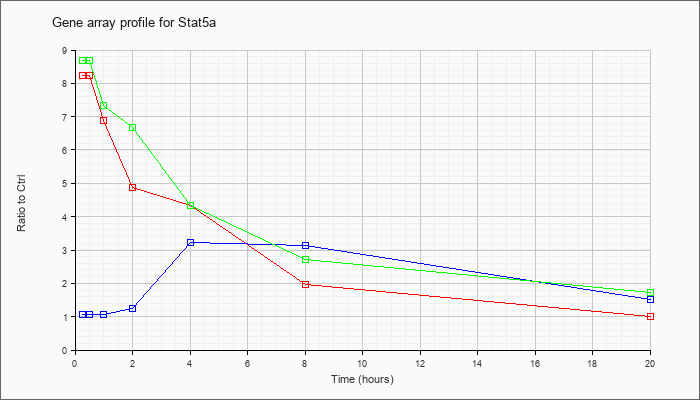 |
NM_011488 |
signal transducer and activator of transcription 5A (Stat5a), mRNA [NM_011488] |
KLA | 8.23 |
8.24 |
6.89 |
4.88 |
4.35 |
1.98 |
1.01 |
| ATP | 1.07 |
1.14 |
1.07 |
1.26 |
3.23 |
3.14 |
1.52 |
| KLA/ATP | 8.70 |
9.53 |
7.33 |
6.68 |
4.32 |
2.73 |
1.72 |
|
| Stat5b | 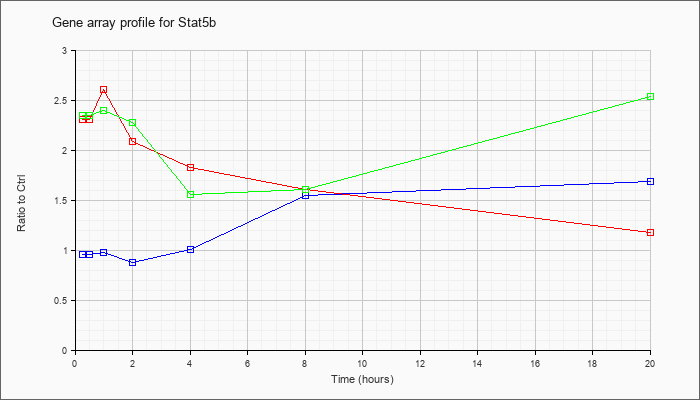 |
AK137889 |
16 days neonate thymus cDNA, RIKEN full-length enriched library, clone:A130035C01 product:unclassifiable, full insert sequence. [AK137889] |
KLA | 1.10 |
1.01 |
.98 |
.90 |
1.03 |
.93 |
.88 |
| ATP | .90 |
.87 |
.61 |
.69 |
1.01 |
.97 |
.89 |
| KLA/ATP | 1.12 |
1.00 |
.87 |
.85 |
.95 |
1.07 |
.83 |
|
| Stat5b |  |
NM_011489 |
signal transducer and activator of transcription 5B (Stat5b), transcript variant 1, mRNA [NM_011489] |
KLA | 2.91 |
2.81 |
3.42 |
2.69 |
2.23 |
1.95 |
1.32 |
| ATP | .99 |
1.04 |
1.16 |
.97 |
1.00 |
1.84 |
2.09 |
| KLA/ATP | 2.96 |
3.05 |
3.17 |
2.99 |
1.86 |
1.87 |
3.40 |
|
| Tbp | 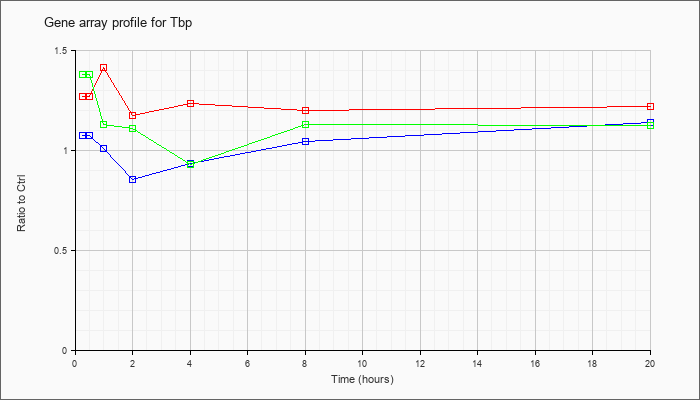 |
NM_013684 |
TATA box binding protein (Tbp), mRNA [NM_013684] |
KLA | 1.27 |
1.33 |
1.42 |
1.17 |
1.23 |
1.20 |
1.22 |
| ATP | 1.07 |
1.21 |
1.01 |
.85 |
.93 |
1.04 |
1.14 |
| KLA/ATP | 1.38 |
1.40 |
1.13 |
1.11 |
.93 |
1.13 |
1.12 |
|
| Tbpl1 | 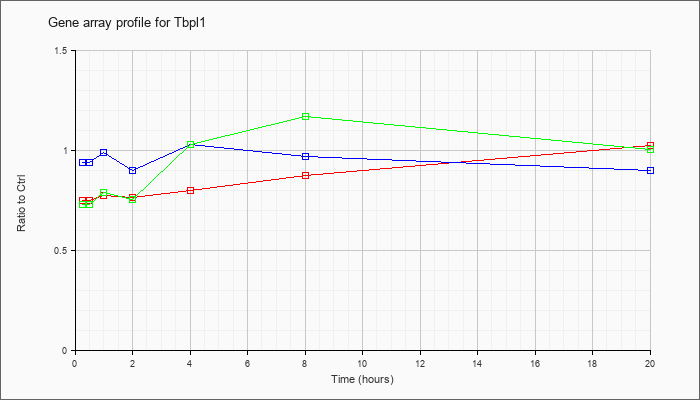 |
AK028621 |
10 days neonate skin cDNA, RIKEN full-length enriched library, clone:4732418P20 product:TATA box binding protein-like protein, full insert sequence [AK028621] |
KLA | 1.03 |
1.11 |
.99 |
.98 |
.97 |
.88 |
.92 |
| ATP | .95 |
1.03 |
1.05 |
.94 |
.94 |
.96 |
1.09 |
| KLA/ATP | .98 |
.95 |
.99 |
.98 |
.98 |
1.03 |
1.01 |
|
| Tbpl1 |  |
NM_011603 |
TATA box binding protein-like 1 (Tbpl1), mRNA [NM_011603] |
KLA | .68 |
.72 |
.72 |
.71 |
.76 |
.87 |
1.05 |
| ATP | .93 |
.97 |
.98 |
.89 |
1.05 |
.97 |
.85 |
| KLA/ATP | .67 |
.66 |
.74 |
.70 |
1.04 |
1.20 |
1.00 |
|
| Tbpl2 | 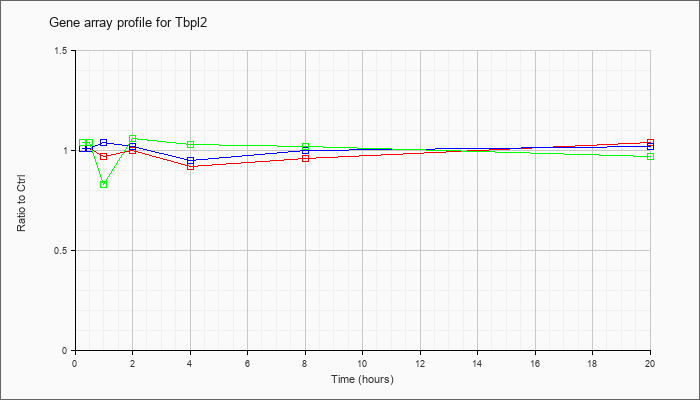 |
NM_199059 |
TATA box binding protein like 2 (Tbpl2), mRNA [NM_199059] |
KLA | 1.01 |
1.10 |
.97 |
1.00 |
.92 |
.96 |
1.04 |
| ATP | 1.01 |
1.03 |
1.04 |
1.02 |
.95 |
1.00 |
1.02 |
| KLA/ATP | 1.04 |
.92 |
.83 |
1.06 |
1.03 |
1.02 |
.97 |
|
| Tcfe2a | 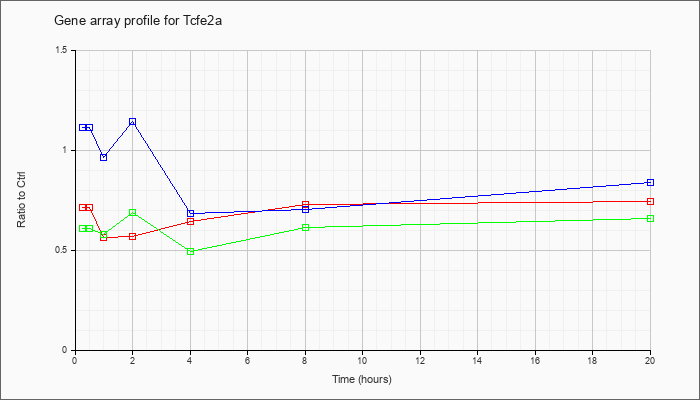 |
NM_011548 |
transcription factor E2a (Tcfe2a), mRNA [NM_011548] |
KLA | .72 |
.66 |
.57 |
.57 |
.65 |
.73 |
.75 |
| ATP | 1.11 |
1.26 |
.97 |
1.15 |
.69 |
.71 |
.84 |
| KLA/ATP | .61 |
.70 |
.58 |
.69 |
.50 |
.62 |
.66 |
|
| Tert | 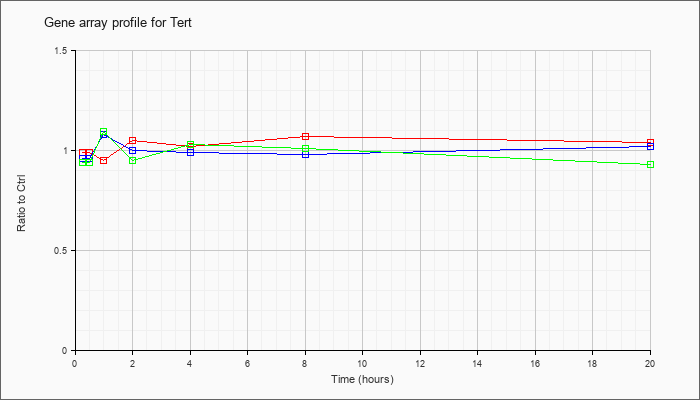 |
NM_009354 |
telomerase reverse transcriptase (Tert), mRNA [NM_009354] |
KLA | .99 |
1.08 |
.95 |
1.05 |
1.02 |
1.07 |
1.04 |
| ATP | .96 |
.99 |
1.08 |
1.00 |
.99 |
.98 |
1.02 |
| KLA/ATP | .94 |
1.06 |
1.09 |
.95 |
1.03 |
1.01 |
.93 |
|
| Tgfb1 | 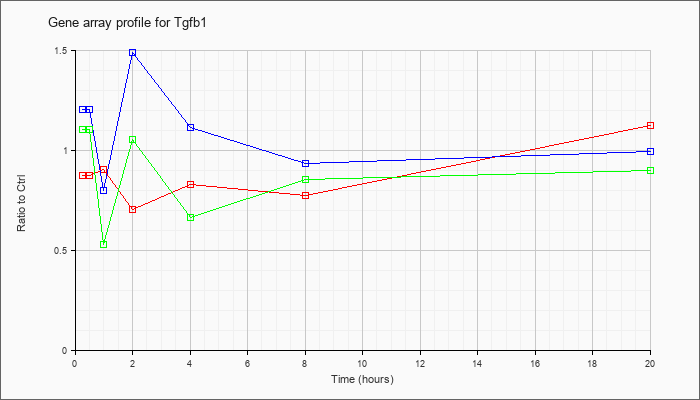 |
NM_011577 |
transforming growth factor, beta 1 (Tgfb1), mRNA [NM_011577] |
KLA | .87 |
.89 |
.90 |
.70 |
.83 |
.77 |
1.12 |
| ATP | 1.20 |
1.41 |
.80 |
1.49 |
1.11 |
.93 |
.99 |
| KLA/ATP | 1.10 |
1.20 |
.53 |
1.05 |
.66 |
.85 |
.90 |
|
| Tgfb2 | 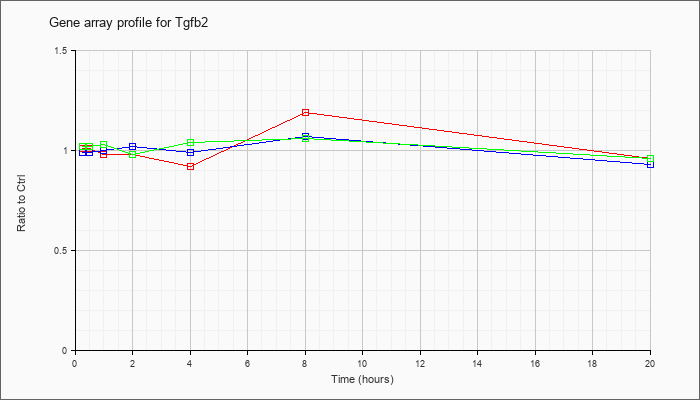 |
NM_009367 |
transforming growth factor, beta 2 (Tgfb2), mRNA [NM_009367] |
KLA | 1.01 |
.91 |
.98 |
.98 |
.92 |
1.19 |
.96 |
| ATP | .99 |
1.03 |
1.00 |
1.02 |
.99 |
1.07 |
.93 |
| KLA/ATP | 1.02 |
.99 |
1.03 |
.98 |
1.04 |
1.06 |
.96 |
|
| Tgfb3 | 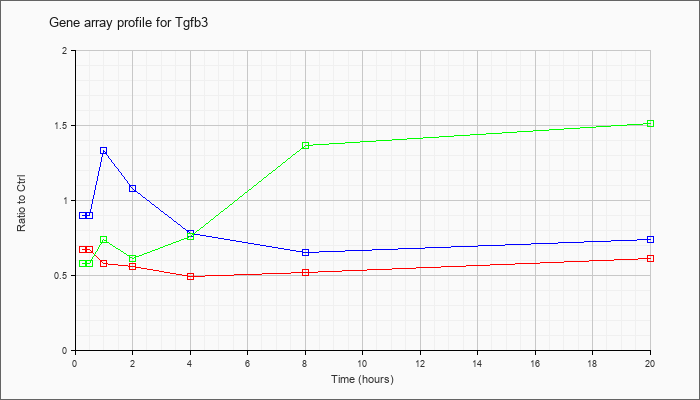 |
NM_009368 |
transforming growth factor, beta 3 (Tgfb3), mRNA [NM_009368] |
KLA | .67 |
.64 |
.58 |
.56 |
.49 |
.52 |
.61 |
| ATP | .90 |
.92 |
1.33 |
1.08 |
.78 |
.65 |
.74 |
| KLA/ATP | .58 |
.60 |
.74 |
.61 |
.76 |
1.36 |
1.51 |
|
| Tgfbr1 | 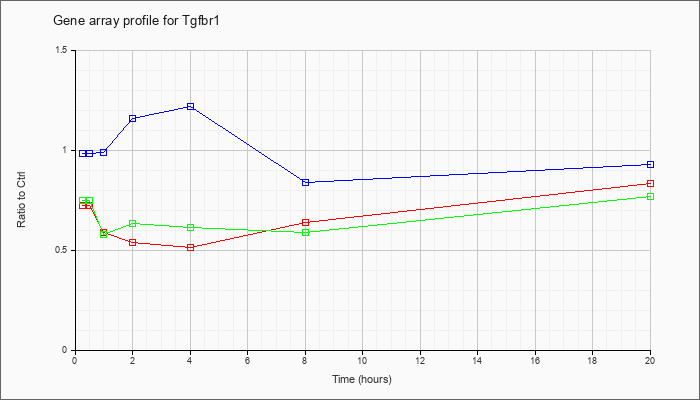 |
NM_009370 |
transforming growth factor, beta receptor I (Tgfbr1), mRNA [NM_009370] |
KLA | .73 |
.74 |
.59 |
.54 |
.52 |
.64 |
.84 |
| ATP | .99 |
1.12 |
.99 |
1.16 |
1.22 |
.84 |
.93 |
| KLA/ATP | .75 |
.78 |
.58 |
.64 |
.62 |
.59 |
.77 |
|
| Tgfbr2 | 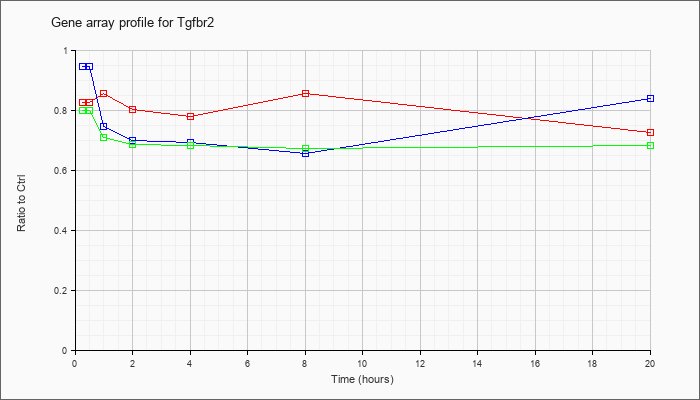 |
AK090393 |
10 days pregnant adult female ovary and uterus cDNA, RIKEN full-length enriched library, clone:G630098D03 product:transforming growth factor, beta receptor II, full insert sequence. [AK090393] |
KLA | .98 |
.89 |
1.00 |
.96 |
.94 |
1.00 |
.92 |
| ATP | .99 |
.99 |
.90 |
.94 |
.93 |
.96 |
1.02 |
| KLA/ATP | .92 |
.96 |
.98 |
.98 |
.98 |
.95 |
1.01 |
|
| Tgfbr2 |  |
NM_009371 |
transforming growth factor, beta receptor II (Tgfbr2), transcript variant 1, mRNA [NM_009371] |
KLA | .55 |
.50 |
.58 |
.50 |
.52 |
.55 |
.44 |
| ATP | .98 |
.75 |
.50 |
.32 |
.41 |
.29 |
.50 |
| KLA/ATP | .59 |
.45 |
.32 |
.23 |
.31 |
.27 |
.25 |
|
| Tgfbr2 |  |
NM_029575 |
transforming growth factor, beta receptor II (Tgfbr2), transcript variant 2, mRNA [NM_029575] |
KLA | .95 |
.93 |
.99 |
.95 |
.88 |
1.02 |
.82 |
| ATP | .87 |
.93 |
.84 |
.84 |
.74 |
.72 |
1.00 |
| KLA/ATP | .89 |
.85 |
.83 |
.85 |
.76 |
.80 |
.79 |
|
| Tln1 | 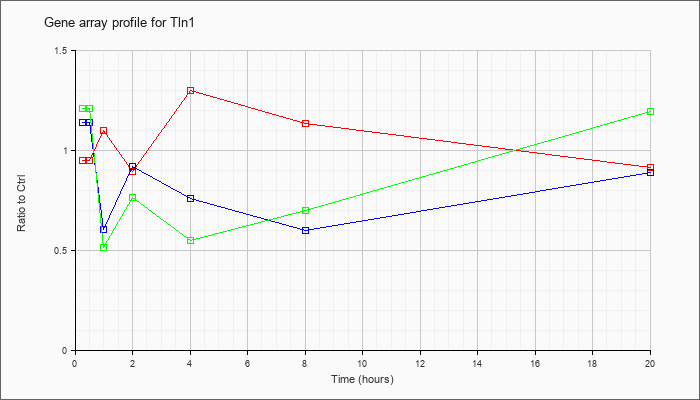 |
NM_011602 |
talin 1 (Tln1), mRNA [NM_011602] |
KLA | .95 |
1.02 |
1.10 |
.90 |
1.30 |
1.13 |
.92 |
| ATP | 1.14 |
1.35 |
.60 |
.92 |
.76 |
.60 |
.89 |
| KLA/ATP | 1.21 |
1.18 |
.51 |
.77 |
.55 |
.70 |
1.20 |
|
| Tln2 | 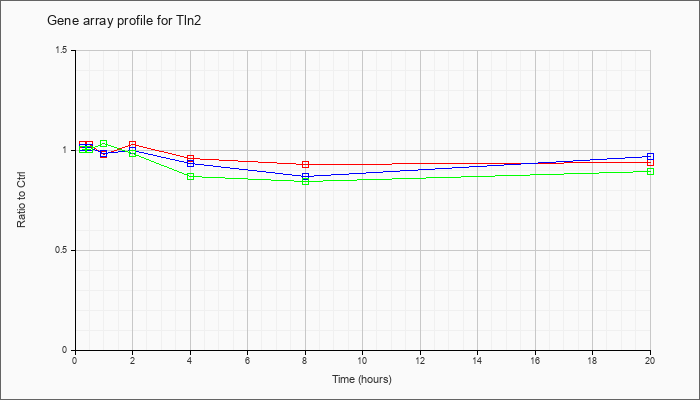 |
NM_001081242 |
talin 2 (Tln2), mRNA [NM_001081242] |
KLA | 1.03 |
1.01 |
.98 |
1.03 |
.96 |
.93 |
.94 |
| ATP | 1.01 |
.97 |
.98 |
1.00 |
.93 |
.87 |
.97 |
| KLA/ATP | 1.00 |
.99 |
1.03 |
.98 |
.87 |
.84 |
.89 |
|
| Tnf | 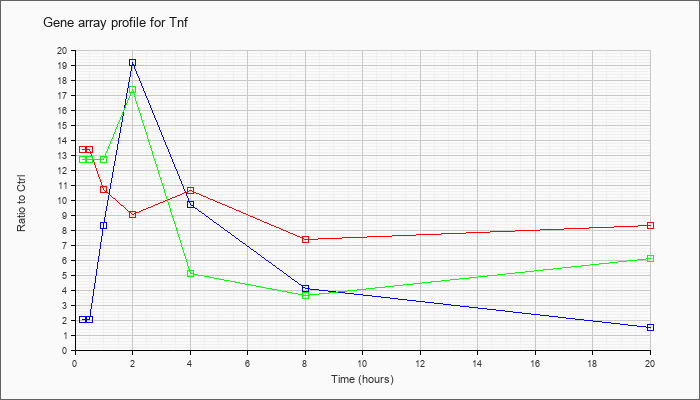 |
NM_013693 |
tumor necrosis factor (Tnf), mRNA [NM_013693] |
KLA | 13.35 |
12.76 |
10.71 |
9.03 |
10.64 |
7.39 |
8.32 |
| ATP | 2.01 |
2.48 |
8.27 |
19.20 |
9.72 |
4.09 |
1.49 |
| KLA/ATP | 12.72 |
15.76 |
12.73 |
17.34 |
5.10 |
3.61 |
6.07 |
|
Tnfrsf13
c | 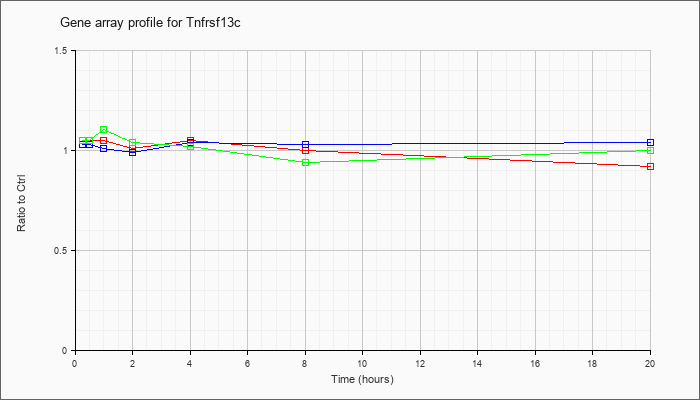 |
NM_028075 |
tumor necrosis factor receptor superfamily, member 13c (Tnfrsf13c), mRNA [NM_028075] |
KLA | 1.05 |
1.04 |
1.05 |
1.01 |
1.05 |
1.00 |
.92 |
| ATP | 1.03 |
1.01 |
1.01 |
.99 |
1.04 |
1.03 |
1.04 |
| KLA/ATP | 1.05 |
1.07 |
1.10 |
1.04 |
1.02 |
.94 |
1.00 |
|
Tnfrsf1a
| 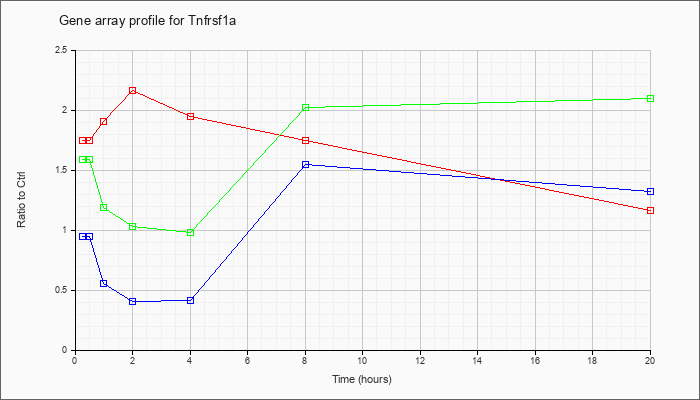 |
NM_011609 |
tumor necrosis factor receptor superfamily, member 1a (Tnfrsf1a), mRNA [NM_011609] |
KLA | 1.74 |
1.71 |
1.90 |
2.16 |
1.95 |
1.74 |
1.16 |
| ATP | .95 |
.83 |
.55 |
.41 |
.41 |
1.55 |
1.32 |
| KLA/ATP | 1.58 |
1.49 |
1.19 |
1.03 |
.98 |
2.02 |
2.10 |
|
| Trp53 | 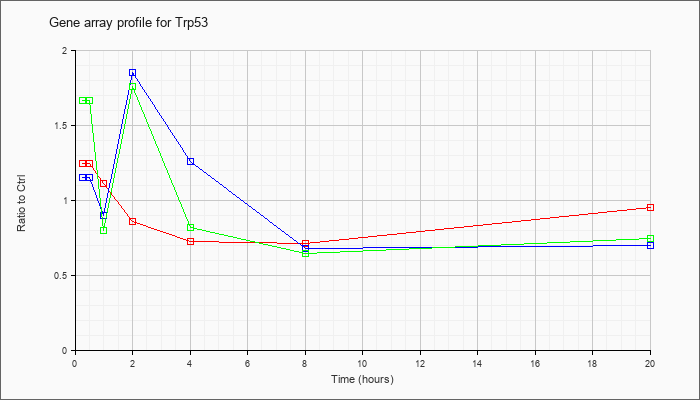 |
NM_011640 |
transformation related protein 53 (Trp53), mRNA [NM_011640] |
KLA | 1.25 |
1.37 |
1.11 |
.86 |
.72 |
.71 |
.95 |
| ATP | 1.15 |
1.32 |
.90 |
1.85 |
1.26 |
.68 |
.70 |
| KLA/ATP | 1.66 |
1.49 |
.80 |
1.76 |
.81 |
.65 |
.74 |
|
Trp53inp
1 | 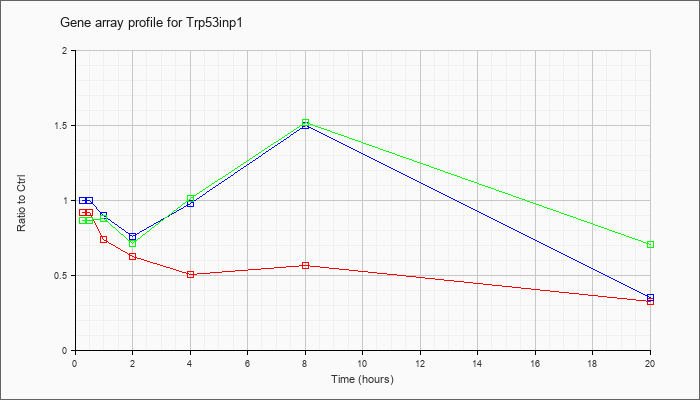 |
NM_021897 |
transformation related protein 53 inducible nuclear protein 1 (Trp53inp1), mRNA [NM_021897] |
KLA | .92 |
.81 |
.74 |
.63 |
.51 |
.57 |
.33 |
| ATP | 1.00 |
1.01 |
.90 |
.76 |
.98 |
1.50 |
.35 |
| KLA/ATP | .87 |
.85 |
.88 |
.71 |
1.01 |
1.52 |
.71 |
|
| Trrap | 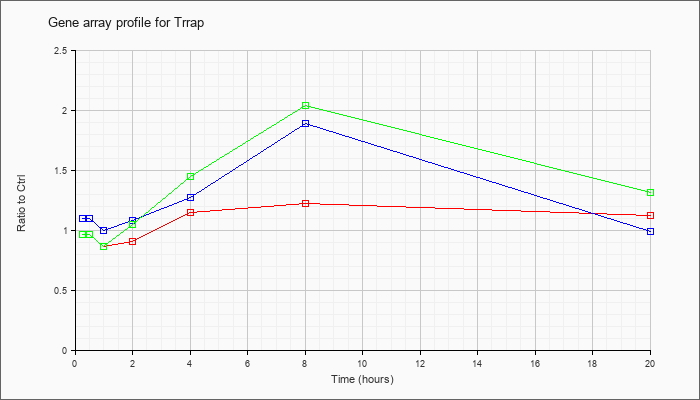 |
NM_001081362 |
transformation/transcription domain-associated protein (Trrap), mRNA [NM_001081362] |
KLA | .97 |
.88 |
.86 |
.91 |
1.15 |
1.23 |
1.13 |
| ATP | 1.10 |
1.16 |
1.00 |
1.08 |
1.27 |
1.89 |
.99 |
| KLA/ATP | .97 |
1.03 |
.86 |
1.05 |
1.45 |
2.04 |
1.32 |
|
| Tspo | 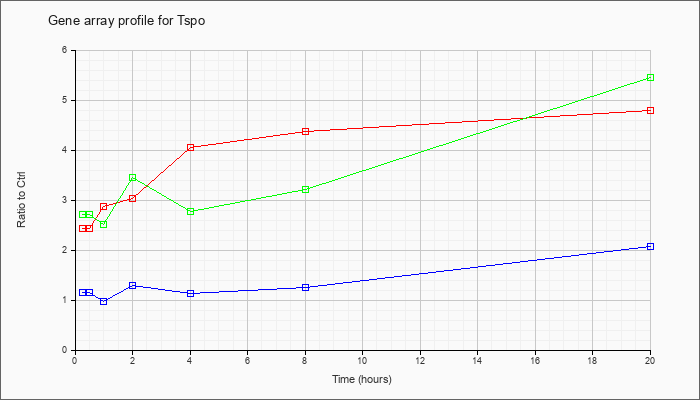 |
NM_009775 |
translocator protein (Tspo), mRNA [NM_009775] |
KLA | 2.43 |
2.59 |
2.88 |
3.03 |
4.05 |
4.37 |
4.80 |
| ATP | 1.15 |
1.16 |
.97 |
1.30 |
1.13 |
1.25 |
2.08 |
| KLA/ATP | 2.72 |
2.79 |
2.51 |
3.45 |
2.78 |
3.22 |
5.46 |
|
| Vac14 | 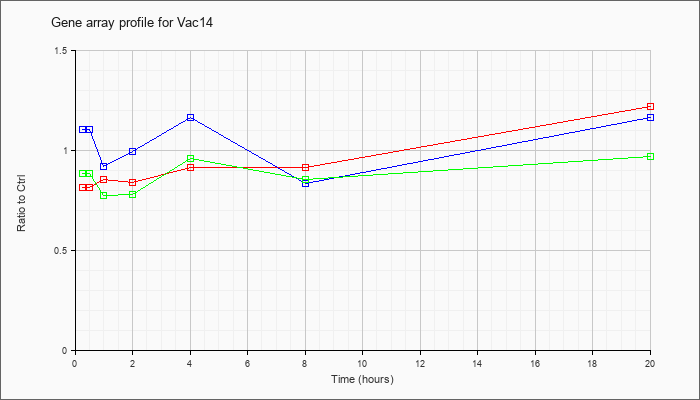 |
AK011396 |
10 days embryo whole body cDNA, RIKEN full-length enriched library, clone:2610014M01 product:unclassifiable, full insert sequence. [AK011396] |
KLA | .83 |
.81 |
.80 |
.90 |
1.01 |
.87 |
1.17 |
| ATP | 1.24 |
1.12 |
1.03 |
.92 |
1.45 |
.95 |
1.23 |
| KLA/ATP | .88 |
.99 |
.96 |
.79 |
1.28 |
.93 |
1.07 |
|
| Vac14 |  |
NM_146216 |
Vac14 homolog (S. cerevisiae) (Vac14), mRNA [NM_146216] |
KLA | .80 |
.86 |
.91 |
.78 |
.82 |
.96 |
1.26 |
| ATP | .97 |
1.07 |
.81 |
1.07 |
.88 |
.72 |
1.10 |
| KLA/ATP | .89 |
.80 |
.59 |
.77 |
.64 |
.78 |
.87 |
|
| Vcam1 | 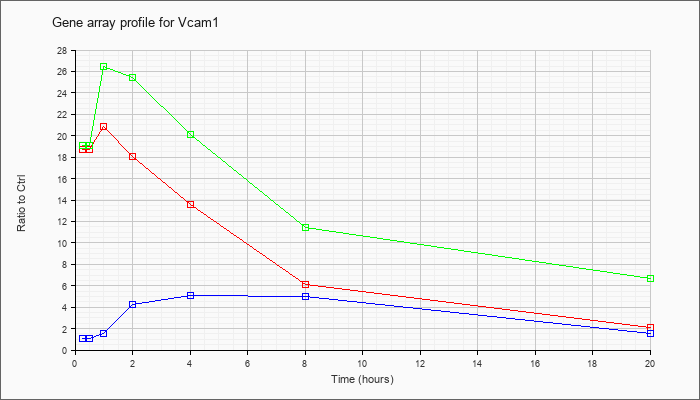 |
NM_011693 |
vascular cell adhesion molecule 1 (Vcam1), mRNA [NM_011693] |
KLA | 18.71 |
22.30 |
20.87 |
18.09 |
13.61 |
6.09 |
2.09 |
| ATP | 1.08 |
1.00 |
1.56 |
4.23 |
5.09 |
5.04 |
1.50 |
| KLA/ATP | 19.09 |
25.02 |
26.43 |
25.42 |
20.08 |
11.45 |
6.67 |
|
| Vdac1 | 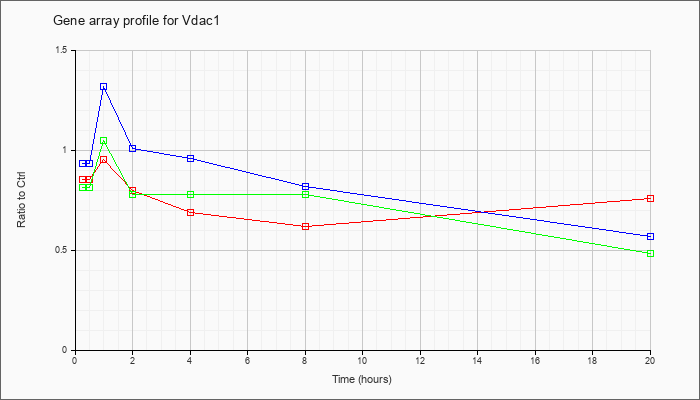 |
NM_011694 |
voltage-dependent anion channel 1 (Vdac1), mRNA [NM_011694] |
KLA | .85 |
.86 |
.95 |
.80 |
.69 |
.62 |
.76 |
| ATP | .93 |
1.01 |
1.32 |
1.01 |
.96 |
.82 |
.57 |
| KLA/ATP | .81 |
.85 |
1.05 |
.78 |
.78 |
.78 |
.48 |
|
| Vdac2 | 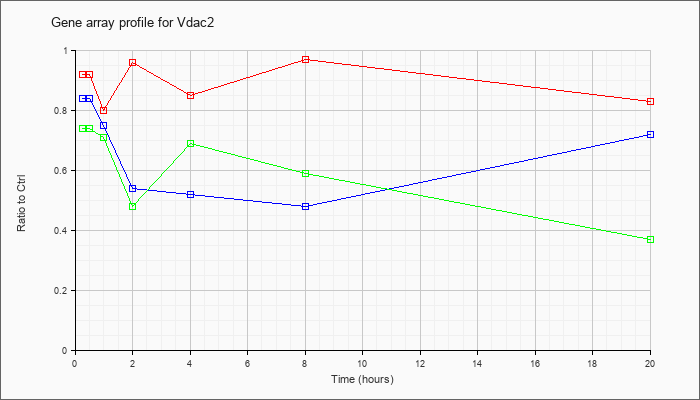 |
NM_011695 |
voltage-dependent anion channel 2 (Vdac2), mRNA [NM_011695] |
KLA | .92 |
.89 |
.80 |
.96 |
.85 |
.97 |
.83 |
| ATP | .84 |
.72 |
.75 |
.54 |
.52 |
.48 |
.72 |
| KLA/ATP | .74 |
.68 |
.71 |
.48 |
.69 |
.59 |
.37 |
|
| Vdac3 | 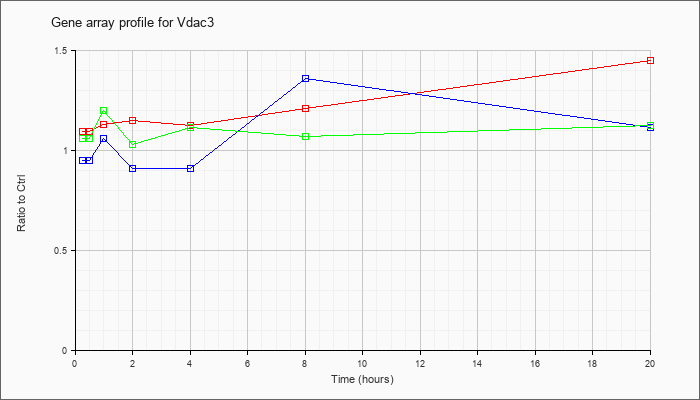 |
NM_011696 |
voltage-dependent anion channel 3 (Vdac3), mRNA [NM_011696] |
KLA | 1.09 |
1.11 |
1.13 |
1.15 |
1.12 |
1.21 |
1.45 |
| ATP | .95 |
.96 |
1.06 |
.91 |
.91 |
1.36 |
1.11 |
| KLA/ATP | 1.06 |
1.06 |
1.20 |
1.03 |
1.11 |
1.07 |
1.12 |
|
| Wnt1 | 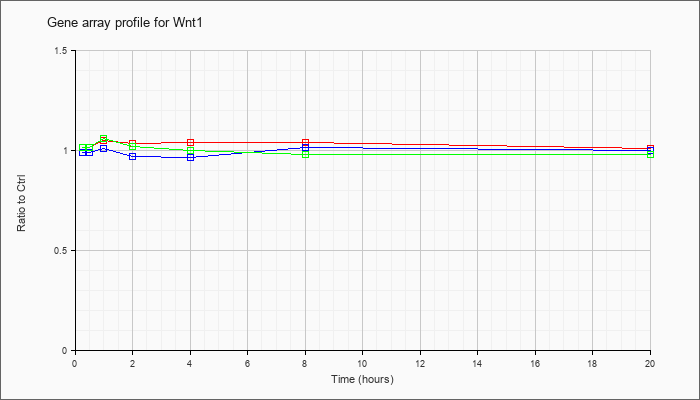 |
NM_021279 |
wingless-related MMTV integration site 1 (Wnt1), mRNA [NM_021279] |
KLA | 1.01 |
1.01 |
1.05 |
1.03 |
1.04 |
1.04 |
1.01 |
| ATP | .99 |
.99 |
1.01 |
.97 |
.96 |
1.01 |
1.00 |
| KLA/ATP | 1.01 |
1.04 |
1.06 |
1.02 |
1.00 |
.98 |
.98 |
|
| Wnt10a | 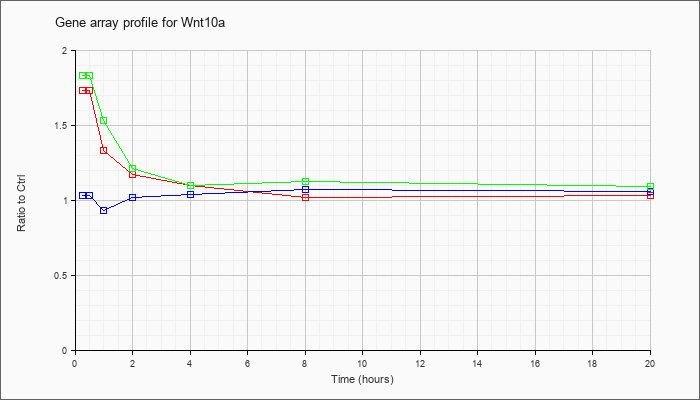 |
NM_009518 |
wingless related MMTV integration site 10a (Wnt10a), mRNA [NM_009518] |
KLA | 1.73 |
1.66 |
1.33 |
1.17 |
1.10 |
1.02 |
1.03 |
| ATP | 1.03 |
1.04 |
.93 |
1.02 |
1.04 |
1.07 |
1.06 |
| KLA/ATP | 1.83 |
1.67 |
1.53 |
1.21 |
1.10 |
1.12 |
1.09 |
|
| Wnt10b | 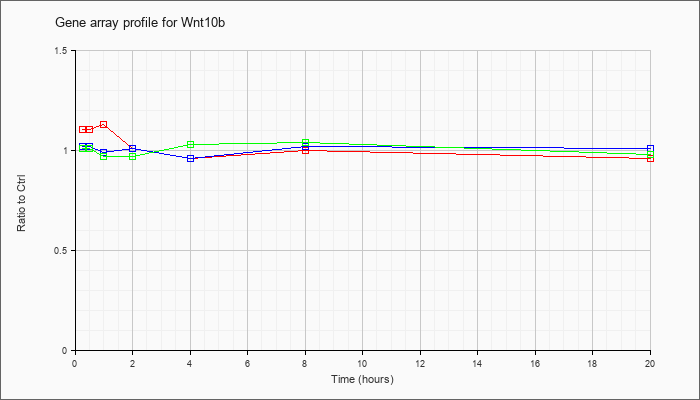 |
NM_011718 |
wingless related MMTV integration site 10b (Wnt10b), mRNA [NM_011718] |
KLA | 1.10 |
.98 |
1.13 |
1.01 |
.96 |
1.00 |
.96 |
| ATP | 1.02 |
1.02 |
.99 |
1.01 |
.96 |
1.02 |
1.01 |
| KLA/ATP | 1.01 |
1.18 |
.97 |
.97 |
1.03 |
1.04 |
.98 |
|
| Wnt11 | 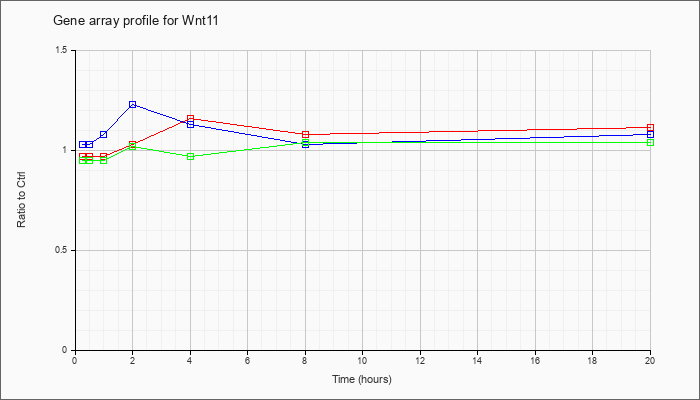 |
NM_009519 |
wingless-related MMTV integration site 11 (Wnt11), mRNA [NM_009519] |
KLA | .97 |
1.00 |
.97 |
1.03 |
1.16 |
1.08 |
1.11 |
| ATP | 1.03 |
1.06 |
1.08 |
1.23 |
1.13 |
1.03 |
1.08 |
| KLA/ATP | .95 |
1.00 |
.95 |
1.02 |
.97 |
1.04 |
1.04 |
|
| Wnt16 |  |
NM_053116 |
wingless-related MMTV integration site 16 (Wnt16), mRNA [NM_053116] |
KLA | 1.00 |
1.02 |
.97 |
1.02 |
.96 |
1.02 |
1.05 |
| ATP | 1.01 |
1.00 |
.97 |
.98 |
.97 |
.95 |
1.02 |
| KLA/ATP | .96 |
1.01 |
.98 |
1.03 |
.99 |
1.00 |
.97 |
|
| Wnt2 |  |
NM_023653 |
wingless-related MMTV integration site 2 (Wnt2), mRNA [NM_023653] |
KLA | 1.01 |
1.00 |
1.02 |
.96 |
1.06 |
1.03 |
.94 |
| ATP | .97 |
1.01 |
1.02 |
.98 |
1.00 |
1.05 |
1.01 |
| KLA/ATP | .97 |
.94 |
1.03 |
1.02 |
1.00 |
.99 |
1.05 |
|
| Wnt2b | 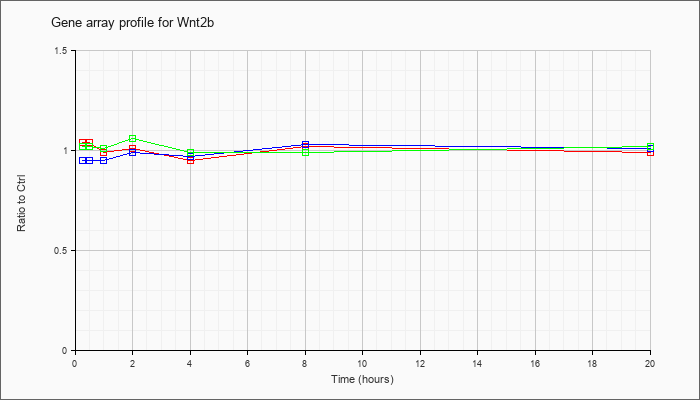 |
NM_009520 |
wingless related MMTV integration site 2b (Wnt2b), mRNA [NM_009520] |
KLA | 1.04 |
1.03 |
.99 |
1.01 |
.95 |
1.02 |
.99 |
| ATP | .95 |
1.00 |
.95 |
.99 |
.97 |
1.03 |
1.01 |
| KLA/ATP | 1.02 |
.99 |
1.01 |
1.06 |
.99 |
.99 |
1.02 |
|
| Wnt3 | 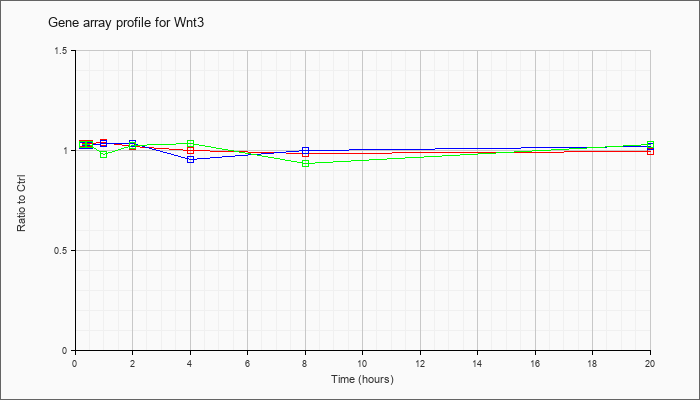 |
NM_009521 |
wingless-related MMTV integration site 3 (Wnt3), mRNA [NM_009521] |
KLA | 1.04 |
1.02 |
1.04 |
1.02 |
1.00 |
.99 |
1.00 |
| ATP | 1.03 |
.98 |
1.04 |
1.04 |
.96 |
1.00 |
1.02 |
| KLA/ATP | 1.03 |
.99 |
.98 |
1.03 |
1.04 |
.94 |
1.03 |
|
| Wnt3a |  |
NM_009522 |
wingless-related MMTV integration site 3A (Wnt3a), mRNA [NM_009522] |
KLA | 1.01 |
1.00 |
1.06 |
1.02 |
.99 |
1.02 |
.99 |
| ATP | .96 |
.97 |
1.05 |
1.04 |
1.02 |
1.02 |
1.04 |
| KLA/ATP | 1.02 |
1.03 |
.96 |
1.01 |
.99 |
1.06 |
1.00 |
|
| Wnt4 |  |
NM_009523 |
wingless-related MMTV integration site 4 (Wnt4), mRNA [NM_009523] |
KLA | 1.01 |
1.01 |
.97 |
1.06 |
1.12 |
1.02 |
.93 |
| ATP | 1.02 |
1.06 |
1.03 |
1.02 |
1.00 |
1.02 |
.98 |
| KLA/ATP | .94 |
1.01 |
1.06 |
1.05 |
.96 |
.96 |
1.00 |
|
| Wnt5a |  |
NM_009524 |
wingless-related MMTV integration site 5A (Wnt5a), mRNA [NM_009524] |
KLA | .98 |
1.06 |
1.01 |
.96 |
1.03 |
1.02 |
1.07 |
| ATP | 1.01 |
.96 |
1.06 |
1.11 |
1.02 |
.97 |
1.03 |
| KLA/ATP | 1.05 |
.96 |
.96 |
1.04 |
1.12 |
1.06 |
.96 |
|
| Wnt5b |  |
NM_009525 |
wingless-related MMTV integration site 5B (Wnt5b), mRNA [NM_009525] |
KLA | 1.04 |
1.04 |
1.19 |
1.11 |
1.10 |
1.11 |
1.05 |
| ATP | 1.04 |
1.01 |
.95 |
1.01 |
.99 |
1.04 |
1.05 |
| KLA/ATP | 1.05 |
1.06 |
1.11 |
1.07 |
1.04 |
1.08 |
1.05 |
|
| Wnt6 |  |
M89800 |
gb|Mouse Wnt-6 mRNA, complete cds. [M89800] |
KLA | 1.92 |
1.97 |
1.93 |
2.04 |
2.01 |
2.62 |
5.22 |
| ATP | .96 |
.97 |
1.07 |
1.03 |
.76 |
.94 |
1.18 |
| KLA/ATP | 2.04 |
1.91 |
1.96 |
1.72 |
1.40 |
1.31 |
2.45 |
|
| Wnt6 |  |
NM_009526 |
wingless-related MMTV integration site 6 (Wnt6), mRNA [NM_009526] |
KLA | 2.83 |
2.51 |
2.19 |
2.74 |
2.92 |
3.01 |
5.82 |
| ATP | 1.01 |
.87 |
1.07 |
.77 |
.59 |
1.11 |
1.06 |
| KLA/ATP | 2.31 |
2.49 |
2.69 |
1.79 |
1.63 |
1.26 |
2.64 |
|
| Wnt7a | 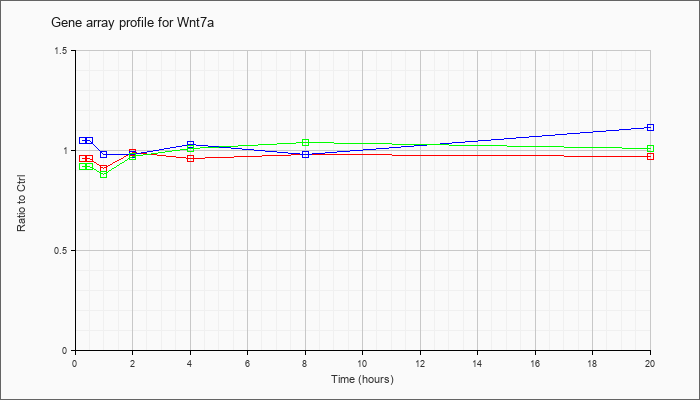 |
NM_009527 |
wingless-related MMTV integration site 7A (Wnt7a), mRNA [NM_009527] |
KLA | .96 |
.92 |
.91 |
.99 |
.96 |
.98 |
.97 |
| ATP | 1.05 |
.95 |
.98 |
.98 |
1.03 |
.98 |
1.11 |
| KLA/ATP | .92 |
.89 |
.88 |
.97 |
1.01 |
1.04 |
1.01 |
|
| Wnt7b | 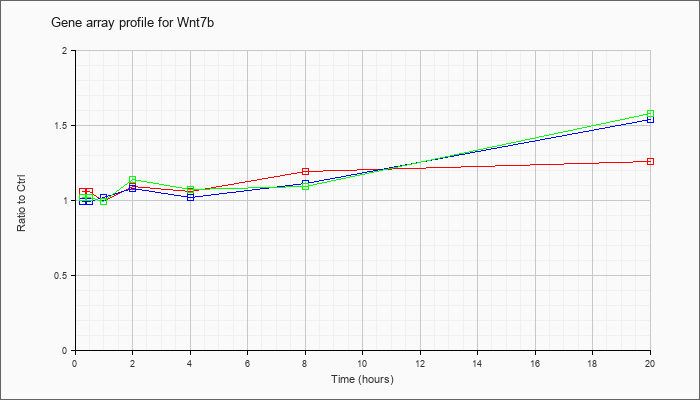 |
NM_009528 |
wingless-related MMTV integration site 7B (Wnt7b), mRNA [NM_009528] |
KLA | 1.06 |
1.09 |
.99 |
1.09 |
1.06 |
1.19 |
1.26 |
| ATP | .99 |
.98 |
1.02 |
1.08 |
1.02 |
1.11 |
1.54 |
| KLA/ATP | 1.02 |
1.01 |
.99 |
1.14 |
1.07 |
1.09 |
1.58 |
|
| Wnt8a |  |
NM_009290 |
wingless-related MMTV integration site 8A (Wnt8a), mRNA [NM_009290] |
KLA | .92 |
1.01 |
.99 |
1.00 |
1.06 |
1.01 |
1.04 |
| ATP | 1.00 |
1.01 |
1.01 |
1.04 |
1.00 |
1.06 |
1.05 |
| KLA/ATP | 1.07 |
1.05 |
.99 |
1.08 |
1.07 |
1.09 |
1.07 |
|
| Wnt8b | 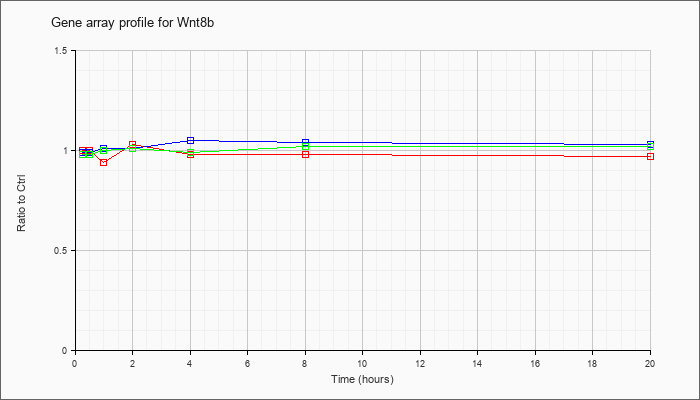 |
NM_011720 |
wingless related MMTV integration site 8b (Wnt8b), mRNA [NM_011720] |
KLA | 1.00 |
1.00 |
.94 |
1.03 |
.98 |
.98 |
.97 |
| ATP | .99 |
1.01 |
1.01 |
1.01 |
1.05 |
1.04 |
1.03 |
| KLA/ATP | .98 |
1.03 |
1.00 |
1.01 |
.99 |
1.02 |
1.02 |
|
| Wnt9a | 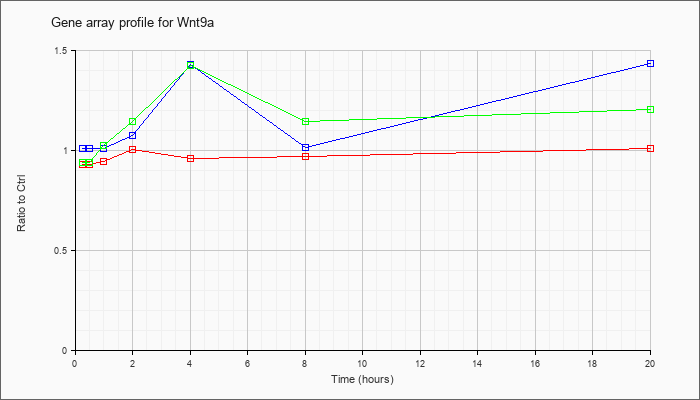 |
NM_139298 |
wingless-type MMTV integration site 9A (Wnt9a), mRNA [NM_139298] |
KLA | .93 |
.95 |
.95 |
1.01 |
.96 |
.97 |
1.01 |
| ATP | 1.01 |
.98 |
1.01 |
1.08 |
1.43 |
1.02 |
1.44 |
| KLA/ATP | .94 |
.96 |
1.03 |
1.15 |
1.43 |
1.15 |
1.21 |
|
| Wnt9b | 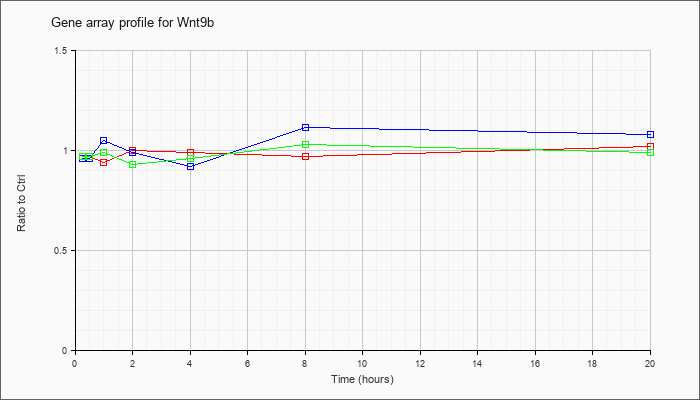 |
NM_011719 |
wingless-type MMTV integration site 9B (Wnt9b), mRNA [NM_011719] |
KLA | .97 |
1.00 |
.94 |
1.00 |
.99 |
.97 |
1.02 |
| ATP | .96 |
1.02 |
1.05 |
.99 |
.92 |
1.11 |
1.08 |
| KLA/ATP | .97 |
.96 |
.99 |
.93 |
.96 |
1.03 |
.99 |
|
| Xbp1 | 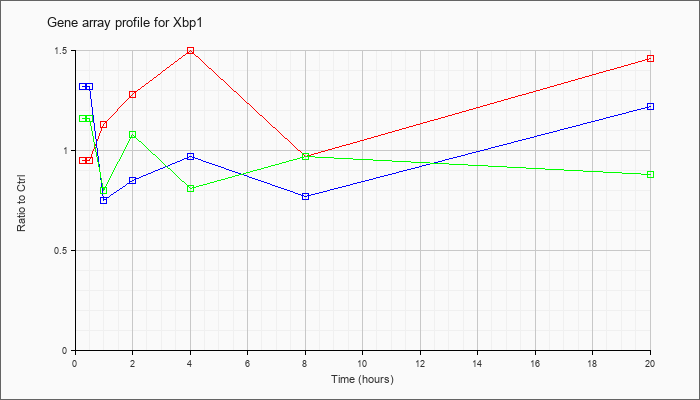 |
NM_013842 |
X-box binding protein 1 (Xbp1), mRNA [NM_013842] |
KLA | .95 |
.98 |
1.13 |
1.28 |
1.50 |
.97 |
1.46 |
| ATP | 1.32 |
1.37 |
.75 |
.85 |
.97 |
.77 |
1.22 |
| KLA/ATP | 1.16 |
1.29 |
.80 |
1.08 |
.81 |
.97 |
.88 |
|
| Xiap | 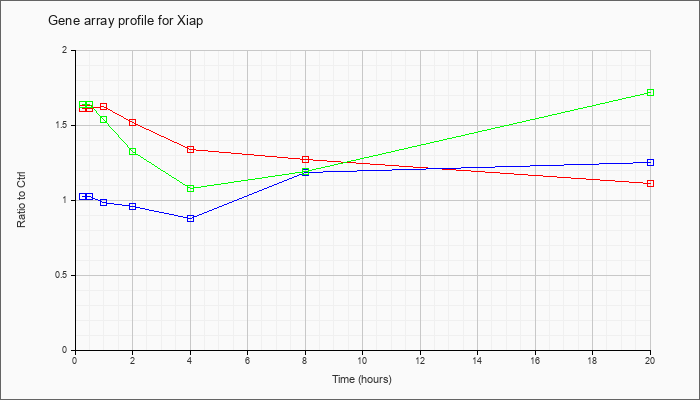 |
NM_009688 |
X-linked inhibitor of apoptosis (Xiap), mRNA [NM_009688] |
KLA | 1.61 |
1.63 |
1.62 |
1.52 |
1.34 |
1.27 |
1.11 |
| ATP | 1.03 |
1.01 |
.99 |
.96 |
.88 |
1.19 |
1.25 |
| KLA/ATP | 1.64 |
1.63 |
1.54 |
1.33 |
1.08 |
1.19 |
1.72 |
|
| Xpo1 |  |
NM_134014 |
exportin 1, CRM1 homolog (yeast) (Xpo1), transcript variant 1, mRNA [NM_134014] |
KLA | .85 |
.84 |
.92 |
.94 |
1.07 |
1.07 |
.94 |
| ATP | 1.00 |
.97 |
.86 |
.74 |
.61 |
.56 |
.77 |
| KLA/ATP | .87 |
.81 |
.71 |
.66 |
.70 |
.74 |
.77 |
|
| Zfp36 | 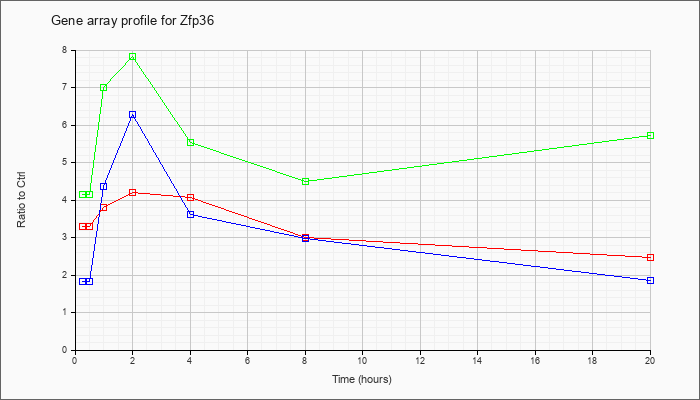 |
NM_011756 |
zinc finger protein 36 (Zfp36), mRNA [NM_011756] |
KLA | 3.30 |
3.27 |
3.79 |
4.20 |
4.08 |
3.01 |
2.48 |
| ATP | 1.84 |
2.37 |
4.35 |
6.29 |
3.62 |
2.97 |
1.85 |
| KLA/ATP | 4.16 |
5.16 |
7.00 |
7.83 |
5.54 |
4.48 |
5.73 |
|