Structure Database (LMSD)
Common Name
GD3(d18:1/24:1(15Z))
Systematic Name
NeuAcα2-8NeuAcα2-3Galβ1-4Glcβ-Cer(d18:1/24:1(15Z))
Synonyms
LM ID
LMSP0601AK07
Formula
Exact Mass
Calculate m/z
1553.918134
Sum Composition
Status
Computationally Generated
3D model of GD3(d18:1/24:1(15Z))
Please note: Where there are chiral atoms but the stereochemistry is undefined, the 3D model takes an arbitrary conformation
Classification
Category
Main Class
Sub Class
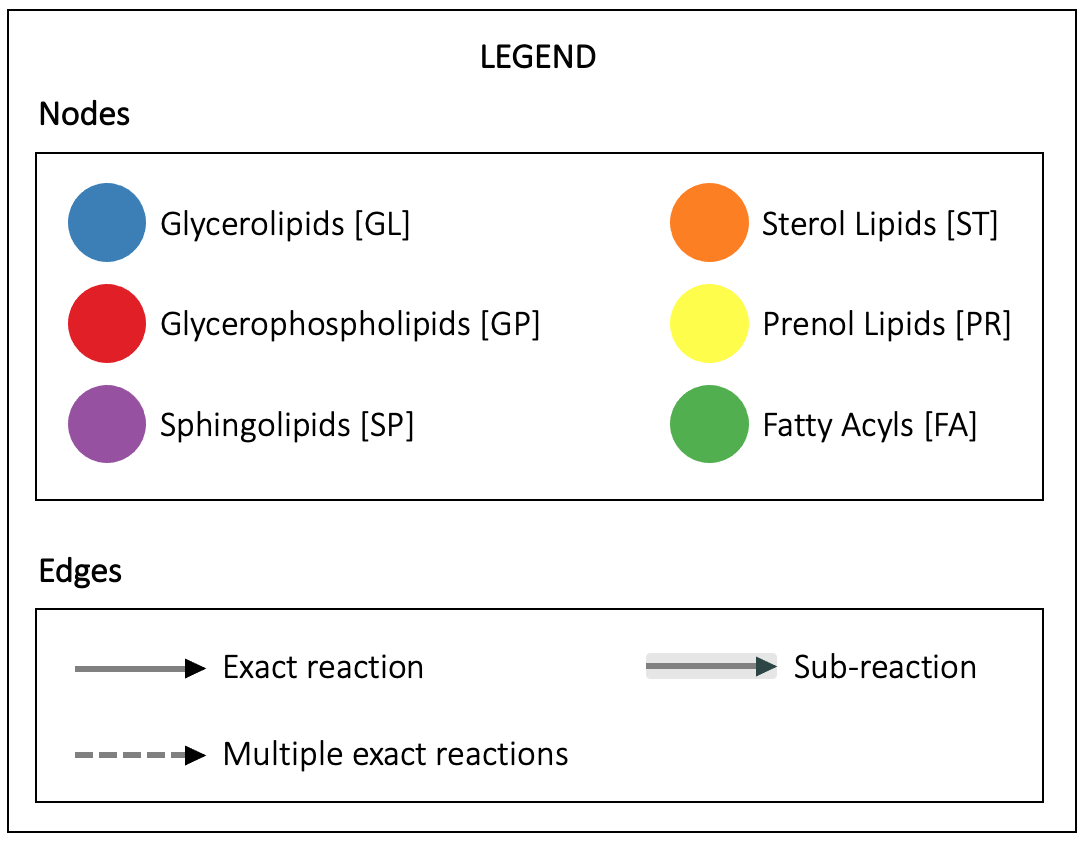
Reactions
Filter by species:
ⓘ
Reactions are shown if the E.C. number of the enzyme catalysing it is annotated in the UniProt database for a species belonging to the selected taxonomic class.
Click on an edge to display the reaction(s).
String Representations
InChiKey (Click to copy)
KTMUAIZWYIXNJC-WIJBFWOGSA-N
InChi (Click to copy)
InChI=1S/C76H135N3O29/c1-5-7-9-11-13-15-17-19-20-21-22-23-24-25-26-28-30-32-34-36-38-40-58(90)79-50(51(86)39-37-35-33-31-29-27-18-16-14-12-10-8-6-2)47-101-71-65(95)64(94)67(57(46-83)103-71)104-72-66(96)70(62(92)55(44-81)102-72)108-76(74(99)100)42-53(88)60(78-49(4)85)69(107-76)63(93)56(45-82)105-75(73(97)98)41-52(87)59(77-48(3)84)68(106-75)61(91)54(89)43-80/h19-20,37,39,50-57,59-72,80-83,86-89,91-96H,5-18,21-36,38,40-47H2,1-4H3,(H,77,84)(H,78,85)(H,79,90)(H,97,98)(H,99,100)/b20-19-,39-37+/t50-,51+,52-,53-,54+,55+,56+,57+,59+,60+,61+,62-,63+,64+,65+,66+,67+,68+,69+,70-,71+,72-,75+,76-/m0/s1
SMILES (Click to copy)
[C@](CO[C@@H]1O[C@H](CO)[C@@H](O[C@@H]2O[C@H](CO)[C@H](O)[C@H](O[C@]3(O[C@@]([H])([C@H](O)[C@H](O[C@]4(O[C@@]([H])([C@H](O)[C@H](O)CO)[C@H](NC(=O)C)[C@@H](O)C4)C(O)=O)CO)[C@H](NC(=O)C)[C@@H](O)C3)C(O)=O)[C@H]2O)[C@H](O)[C@H]1O)([H])(NC(CCCCCCCCCCCCC/C=C\CCCCCCCC)=O)[C@]([H])(O)/C=C/CCCCCCCCCCCCC
Other Databases
Calculated Physicochemical Properties
Heavy Atoms
108
Rings
4
Aromatic Rings
0
Rotatable Bonds
57
Van der Waals Molecular Volume
1543.35
Topological Polar Surface Area
527.24
Hydrogen Bond Donors
19
Hydrogen Bond Acceptors
32
logP
10.75
Molar Refractivity
405.27
Admin
Created at
-
Updated at
24th Aug 2021