Structure Database (LMSD)
Common Name
GP1c(d18:1/24:1(15Z))
Systematic Name
NeuAcα2-8NeuAcα2-3Galβ1-3GalNAcβ1-4(NeuAcα2-8NeuAcα2-8NeuAcα2-3)Galβ1-4Glcβ-Cer(d18:1/24:1(15Z))
Synonyms
LM ID
LMSP0601AX07
Formula
Exact Mass
Calculate m/z
2792.336588
Sum Composition
Status
Active (generated by computational methods)
No other lipid differing only in stereochemistry/bond geometry found
3D model of GP1c(d18:1/24:1(15Z))
Please note: Where there are chiral atoms but the stereochemistry is undefined, the 3D model takes an arbitrary conformation
Main
Classification
Category
Main Class
Sub Class
String Representations
InChiKey (Click to copy)
HVTHZVVCMKQFIH-LGTLYLQISA-N
InChi (Click to copy)
InChI=1S/C123H209N7O63/c1-9-11-13-15-17-19-21-23-24-25-26-27-28-29-30-32-34-36-38-40-42-44-82(154)130-66(67(146)43-41-39-37-35-33-31-22-20-18-16-14-12-10-2)59-176-111-97(163)96(162)100(80(57-138)179-111)181-113-99(165)109(193-123(118(174)175)49-72(151)87(128-64(7)144)107(191-123)95(161)79(56-137)186-121(116(170)171)47-70(149)85(126-62(5)142)105(189-121)93(159)77(54-135)184-119(114(166)167)45-68(147)83(124-60(3)140)103(187-119)89(155)73(152)50-131)101(81(58-139)180-113)182-110-88(129-65(8)145)102(91(157)75(52-133)177-110)183-112-98(164)108(92(158)76(53-134)178-112)192-122(117(172)173)48-71(150)86(127-63(6)143)106(190-122)94(160)78(55-136)185-120(115(168)169)46-69(148)84(125-61(4)141)104(188-120)90(156)74(153)51-132/h23-24,41,43,66-81,83-113,131-139,146-153,155-165H,9-22,25-40,42,44-59H2,1-8H3,(H,124,140)(H,125,141)(H,126,142)(H,127,143)(H,128,144)(H,129,145)(H,130,154)(H,166,167)(H,168,169)(H,170,171)(H,172,173)(H,174,175)/b24-23-,43-41+/t66-,67+,68-,69-,70-,71-,72-,73+,74+,75+,76+,77+,78+,79+,80+,81+,83+,84+,85+,86+,87+,88+,89+,90+,91-,92-,93+,94+,95+,96+,97+,98+,99+,100+,101-,102+,103+,104+,105+,106+,107+,108-,109+,110-,111+,112-,113-,119+,120+,121+,122-,123-/m0/s1
SMILES (Click to copy)
[C@](CO[C@@H]1O[C@H](CO)[C@@H](O[C@@H]2O[C@H](CO)[C@H](O[C@@H]3O[C@@H]([C@@H]([C@@H]([C@H]3NC(C)=O)O[C@H]3[C@H](O)[C@@H](O[C@]4(O[C@@]([H])([C@H](O)[C@H](O[C@]5(O[C@@]([H])([C@H](O)[C@H](O)CO)[C@H](NC(=O)C)[C@@H](O)C5)C(O)=O)CO)[C@H](NC(=O)C)[C@@H](O)C4)C(O)=O)[C@@H](O)[C@@H](CO)O3)O)CO)[C@H](O[C@]3(O[C@@]([H])([C@H](O)[C@H](O[C@]4(O[C@@]([H])([C@H](O)[C@H](O[C@]5(O[C@@]([H])([C@H](O)[C@H](O)CO)[C@H](NC(=O)C)[C@@H](O)C5)C(O)=O)CO)[C@H](NC(=O)C)[C@@H](O)C4)C(O)=O)CO)[C@H](NC(=O)C)[C@@H](O)C3)C(O)=O)[C@H]2O)[C@H](O)[C@H]1O)([H])(NC(CCCCCCCCCCCCC/C=C\CCCCCCCC)=O)[C@]([H])(O)/C=C/CCCCCCCCCCCCC
References
Calculated Physicochemical Properties
Heavy Atoms
193
Rings
9
Aromatic Rings
0
Rotatable Bonds
85
Van der Waals Molecular Volume
2619.03
Topological Polar Surface Area
1141.41
Hydrogen Bond Donors
40
Hydrogen Bond Acceptors
70
logP
7.51
Molar Refractivity
683.22
Reactions
Filter by species:
ⓘ
Reactions are shown if the E.C. number of the enzyme catalysing it is annotated in the UniProt database for a species belonging to the selected taxonomic class.
Click on an edge to display the reaction(s).
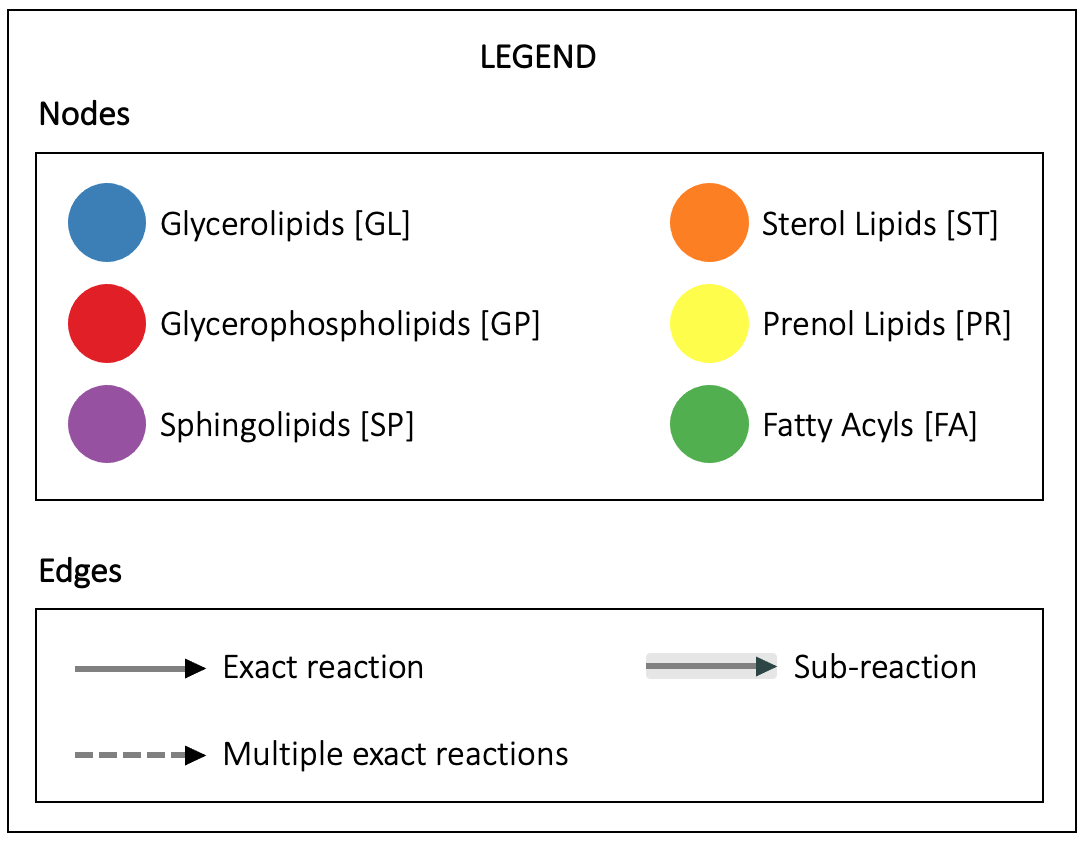
Admin
Created at
-
Updated at
26th Aug 2021